Abstract
Prediction and control of emerging pathogens is a fundamental challenge for public health. To meet this challenge, new analytic tools are needed to characterize the underlying dynamics of the geographical spread of pathogens, identify predictable changes in their dynamics, and support strategic planning for disease elimination and control. Nonparametric and model-independent tools are particularly needed. Here, we propose a multivariate method that uses similarity in cross-spectral density between measured spatial time series of disease prevalence as a feature measuring the proximity of a tipping point, i.e., emergence or elimination. In particular, we show that the increase in the average value of spectral similarity in measured epidemiological time series contains crucial information about the underlying dynamics and proximity to critical points in infectious disease systems. Theoretical analysis of a standard metapopulation SIR model and empirical analysis of case reports of pertussis in the continental USA demonstrate that this increase is observed when the disease approaches elimination. Therefore, this nonparametric indicator provides insight into the fundamental underlying state of the epidemiological system, which is key in developing appropriate strategies to more quickly achieve elimination goals.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Emerging and re-emerging pathogens are among the hardest to predict threats to public health and global security [1]. In recent years, outbreaks of novel pathogens such as SARS-CoV-2 (the virus that causes COVID-19) have led to significant morbidity, mortality, and the global reduction in expected life span [2, 3], prompting substantial investment in emergency planning and preparedness across the globe. Similarly, endemic diseases such as tuberculosis, malaria, and dengue fever continue to tax the health and economic welfare of a large fraction of the global population [4,5,6,7,8,9]. To control such diseases and evaluate the effectiveness of interventions, e.g., vaccination, new methods are needed to anticipate emergence and elimination and, in general, characterize the associated critical phenomena. Despite recent advances in infectious disease analysis and forecasting [10,11,12], methods for anticipating disease emergence and elimination remain under-developed [13].
Predicting the emergence and elimination of infectious diseases is possible by fitting complex parametric mathematical models of disease transmission to incidence data [14,15,16,17,18,19]. While these models provide an in-depth picture of disease dynamics, their success relies on a detailed understanding of the underlying epidemiology, immunology and pathogenesis of the disease in addition to long-term data [20]. Such efforts are labor-intensive, technically demanding, and require detailed, fine-scale data that are often not readily available, especially during an outbreak of an emerging pathogen [21]. In addition, the underlying disease drivers and key epidemiological determinants including transmission route(s), the infectious period, infectiousness and rate of spillover are typically unknown or poorly quantified [22]. As a result, model-independent methods are for anticipating disease emergence and elimination that may be especially valuable in a wide range of scenarios. Such a need has resulted in a growing interest in model-free approaches to analyze the spatial and temporal pattern of simulated and real-world epidemics and extract features corresponding to the emergence of new disease [13, 23,24,25,26].
Recently, the availability of data and computational resources has inspired the use of data-driven techniques to address challenges in predicting the dynamics of epidemiological systems [27, 28]. However, the performance of these methods is inversely related to deviations identified from historically observed data [29]. In addition, emergence and elimination of infectious diseases are rare events and involve qualitative changes in dynamics [30] not just quantitative, for which empirical data for training algorithms are limited. Therefore, the development of metrics to forecast epidemic transitions that can provide reliable results without requiring substantial amounts of data is crucial. The development of such metrics can be achieved through systematic analysis of epidemiological dynamical models. By identifying metrics that are supported by the underlying principles of the system dynamics for predicting epidemic transitions, more reliable predictions can be made with a reduced need for extensive data sets.
In particular, it is known that emergence and elimination of infectious diseases are nonlinear phenomena and involve a critical transition [22, 31, 32], often reflected in deterministic models by a bifurcation. Early warning signals of critical transitions that are stochastic metrics extracted from online time-series measurements of system dynamics are examples of such approaches that indicate when systems approach instabilities [13, 33]. Existing early warning signals introduced as indicators of critical phenomena in disease dynamics mostly focus on temporal or spatial patterns observed in the dynamics of the disease. Studies on the dynamics of the disease spread, however, show that the spread of infectious disease is a spectro-spatio-temporal dynamical process [23, 24, 34]. As a result, analyzing epidemiological time series separately in the time, space or frequency domain might conceal essential characteristics of the dynamics. We hypothesized that analyzing spatial patterns in the time–frequency domain would identify more robust and informative features for data-driven algorithms to evaluate compared to other traditional patterns in natural frames of reference, particularly in infectious disease dynamics where a spatio-temporal travelling wave induces instability.
Here, we propose a new spectro-spatio-temporal approach to analyzing geographical data of emerging and endemic pathogens and extract features associated with their transient and steady-state dynamics. From theoretical analysis of a dynamical disease model, we propose that monitoring the temporal evolution of spectral similarity in multivariate epidemiological time series may provide crucial information about the underlying dynamics and the proximity of the infectious disease system to a critical point. The method uses a multivariate wavelet analysis of the time series measured from the dynamics of the disease at different spatial locations to probe for significant changes in the pattern of similarity in wavelet cross-spectra. By analyzing a standard SIR model with vaccination as well as historical case reports of whooping cough (or pertussis) in the continental USA, we demonstrate that a measurable change in the average pairwise similarity in the reported time series is observed when the system approaches the critical point of disease elimination.
2 Metapopulation SIR model
Numerical simulations were performed with a standard SIR metapopulation model, a simple model of an immunizing disease among loosely coupled patches of a population [35, 36].
We consider a metapopulation consisting of \(n\) connected patches. Let \(S_{i} ,I_{i} , \) and \(R_{i}\) , respectively, denote the number of susceptible, infected, and removed individuals in patch \(i\). We model the dynamics of these variables in a population of \(n\) coupled patches as
for \(i \in 1,2, \ldots ,n\). Here \(r_{i}\) is the population birth rate, \(\nu\) is the probability of immunization at birth, \(\mu_{i} \) is the per capita death rate, \(\frac{1}{{\gamma_{i} }}\) is the mean infectious period and \(\beta_{ij}\) is the rate at which susceptibles in patch \( i\) are infected by infecteds in patch \(j\). In the deterministic model, the population size \(N_{i} = S_{i} + I_{i} + R_{i}\) in each patch is determined by \(r_{i} /\mu_{i}\); thus, the number of removed individuals in patch \(i\) follows \(R_{i} = r_{i} /\mu_{i} - S_{i} - I_{i}\).
For biologically meaningful parameters (i.e., all parameters are positive) and the assumption that the matrix of transmission rates is irreducible, the model always has a globally stable endemic equilibrium when \(R_{e} > 1\), where \(R_{e}\) is defined as the effective reproductive number [36]. Re is obtained via the spectral radius of the \(n \times n\) matrix \({\varvec{M}}\) with elements \(m_{ij} = \beta_{ij} S_{i}^{0} /[(\mu_{i} + \gamma_{i} )]\), where \(S_{i}^{0}\) denotes the population in patch \(i\) in the absence of disease (\(I_{1} = I_{2} = \cdots = I_{n} = 0\)) [37]. The endemic equilibrium reflects the presence of the disease in all patches. When \(R_{e} \le 1\) the disease-free equilibrium in which the disease is absent in all patches is globally stable.
To account for the finite size of the population and random fluctuations in disease transmission, we incorporated demographic stochasticity by adding an intrinsic noise vector, \(w_{2n \times 1} \left( t \right)\), to the equations [38]. The noise vector \(w_{2n \times 1} \left( t \right)\) contains zero mean Gaussian white noise sources with covariance calculated from the system size expansion [39]. The resulting stochastic differential equations are simulated to study the proposed method using simulated data before applying to real data.
3 Spatiotemporal features associated with the stability of the endemic equilibrium
In this study, we propose an approach to measure two main characteristics of the disease dynamics, the degree of interrelation/similarity between system components and the frequency band at which the interrelation is dominant. Here, we briefly discuss how these features of the disease process may vary over time as the disease prevalence drops, resulting in a measurable pattern in the dynamics of the disease.
Spectral similarity. Let \(\vec{S}_{i}^{*}\) and \(\vec{I}_{i}^{*}\) denote the stable endemic equilibrium of the SIR model of Eq. (1). Let \(\vec{z}_{{S_{i} }}\) and \(\vec{z}_{{I_{i} }}\) , respectively, denote deviations from the equilibrium of the susceptible and infected variables. In the neighborhood of the endemic equilibrium, the dynamics of \(\vec{z}_{{S_{i} }}\) and \(\vec{z}_{{I_{i} }}\) may be approximated with the linearization of the vector field of Eq. (1) at the endemic equilibrium, which we denote \({\varvec{J}}\). This linear system may be written as:
where the vectors \(\vec{z}_{S}\) and \(\vec{z}_{I}\) , respectively, collect the variables \(z_{{S_{i} }}\) and \(z_{{I_{i} }}\), and \(\vec{z}\) is a column vector containing \(\vec{z}_{S}\) stacked on top of \(\vec{z}_{I}\), i.e., \(\vec{z} = \left[ {\begin{array}{*{20}c} {\vec{z}_{s} } \\ {\vec{z}_{I} } \\ \end{array} } \right]\). The matrix \({\varvec{J}}\) may be partitioned into four \(n \times n\) blocks depending on whether its row and column number correspond to the index of susceptible or an infected deviation in \(\vec{z}\), that is
Each of these blocks has a unique structure based on a few common elements. Below is the expression of each block:
where \({\varvec{B}}\) denotes the matrix containing the transmission rates among individuals across network patches (i.e., \(\left( {\varvec{B}} \right)_{ij} = \beta_{ij}\)), \(\vec{S}^{*}\) and \(\vec{I}^{*}\) are column vectors containing \(S_{i}^{*}\) and \(I_{i}^{*}\) , respectively, and \(\vec{\mu }\) and \(\vec{\gamma }\) are column vectors containing \(\mu_{i}\) and \(\gamma_{i}\), respectively.
As Re approaches 1, the number of infectious individuals, \(\vec{I}^{*}\), gradually declines towards zero (or its minimum)—the tipping point at which the disease is eliminated. Looking at the terms in Eq. (4), such a change in disease prevalence results in a decrease in the effect of within-patch processes on the dynamics of the connected system and consequently an increase in the effect of cross-coupling terms that come to dominate transmission dynamics. It can be shown that such an increase in interrelation is reflected in a noticeable increase in similarity of the measured power spectra of system time series. Given two signals \(x_{1} \left( t \right)\) and \(x_{2} \left( t \right)\), the cross-spectra identifying the relationship between two time series as a function of frequency are defined as:
where \(R_{12} \left( \tau \right)\) is the cross-correlation function of signals \(x_{1} \left( t \right)\) and \(x_{2} \left( t \right)\). The matrix of cross-spectra can be constructed as \(\left( {{\varvec{S}}\left( \omega \right)} \right)_{uv}\), where element in row \(u\) and column \(v\) of the matrix represents \(S_{uv} \left( \omega \right)\).
Analyzing Eq. (4) shows that when \(\vec{I}^{*}\) gradually approaches zero, a pair of complex conjugate eigenvalues, namely \(\lambda_{1}\) and \(\lambda_{2}\) (\(\lambda_{2} = \overline{\lambda }_{1}\)) also approach zero. We can show (see supplementary information) that the elements of the matrix of cross-spectra \({\varvec{S}}\) for the system in Eq. (3) at an angular frequency \(\omega\) can be obtained as:
where \(\left( {{\varvec{S}}\left( \omega \right)} \right)_{uv}\) denotes element in row \(u\) and column \(v\) of the matrix \({\varvec{S}}\), \({\tilde{\mathbf{\Sigma }}}_{ }\) is the pseudo-covariance matrix of the model variables projected into the space spanned by the eigenvectors of \({\varvec{J}}\), and constant coefficients \(a_{ij} \left( {u,v} \right)\) are determined by the necessary change of basis matrix. It can be shown that the elements of covariance matrix \({\tilde{\mathbf{\Sigma }}}_{ }\) are inversely proportional to the system eigenvalues in the form of \(\left( {{\tilde{\mathbf{\Sigma }}}} \right)_{ij} \propto \frac{1}{{\lambda_{i} + \lambda_{j} }}\) (see supplementary information). As a result, when \({\text{Re}} \left( {\lambda_{1} } \right)\) (consequently \({\text{Re}} \left( {\lambda_{2} } \right)\)) becomes small relative to the other eigenvalues, \(\left( {{\tilde{\mathbf{\Sigma }}}} \right)_{12} = \left( {{\tilde{\mathbf{\Sigma }}}} \right)_{21}\) grows relatively large. Therefore, the corresponding term in Eq. (6) becomes increasingly dominant in all elements of \({\varvec{S}}\left( \omega \right)\). Additionally, decreases in the real parts of these eigenvalues lead to component terms in Eq. (6) that have relatively sharper and larger peaks near \({\text{Im}} \left( {\lambda_{1} } \right) = \omega\). This analysis suggests that when a disease approaches a tipping point (corresponding to \({\text{Re}} \left( {\lambda_{1} } \right) \to 0\)), i.e., elimination or emergence, an increased similarity in the frequency contents of the measured time series is expected. In the supplementary information, we provide a more rigorous and detailed argument for this assertion based on the linearized model. While the details of this method are demonstrated in Eq. (1), an increased spectral similarity is due to the existence of a tipping point in the system dynamics (i.e., when the dominant system eigenvalue approaching to zero) and is not specific to the set of equations studied in this paper. Hence, the results can be extended to other epidemiological models.
In addition to an increased cross-coupling, investigation of the natural period of oscillations in disease incidence is of fundamental importance in mathematical epidemiology [38, 40, 41]. Empirical data of disease incidence show identifiable cyclic patterns in many common diseases, including diseases for which environmental influences do not appear to play an important role, such as measles, pertussis, chicken pox, and mumps [42, 43]. In an epidemiological system with vaccination rate as a control parameter, a progressive temporal change in the dominant period of the epidemic is reported when vaccine uptake increases and the disease approaches elimination [24, 44]. Such a progressive change is also a sign of approaching epidemic to a tipping point [7].
For the metapopulation model of Eq. (1), we can show that as the disease prevalence gradually decreases over time, lower frequency components of the power spectra become more dominant (see supplementary information). As a result, a change in the dominant period of the measurements can be used as an indicator of approaching to the tipping point in the dynamics of the disease.
3.1 Multivariate time-varying spectral similarity analysis
Based on the identified features associated with approaching an epidemic transition, i.e., spectral similarity and frequency shift, we propose a multivariate spectro-spatio-temporal method to monitor the status of the endemic equilibrium. In particular, we define a spectral similarity coefficient \(c\) for a multivariate system as a function that measures the spectral similarity among measured spatial signals. Consider \(n\) time series, namely \(x_{1} \left( t \right), x_{2} \left( t \right), \ldots , x_{n} \left( t \right)\), where \(n\) denotes the variable index. In addition, consider \({\varvec{S}}_{n \times n} \left( \omega \right)\) to denote the cross-spectral matrix of the time series. The spectral similarity for the measured spatial time series is defined as follows
where \(N_{uv} = \mathop \smallint \limits_{ - \infty }^{\infty } \left| {\left( {{\varvec{S}}\left( \omega \right)} \right)_{uv} } \right|{\text{d}}\omega\) is a normalization constant. Theoretical analysis of the SIR equations shows that the similarity coefficient in a metapopulation SIR model increases exponentially as the disease approaches an elimination (supplementary information).
The approximated similarity essentially depends on the power spectra of the time series. However, many real time series including the epidemiological time series are nonstationary. In such cases, classical spectral analysis approaches (e.g., the Fourier transform) do not provide time-dependent spectrum of the time series. To address this issue, we use wavelet analysis, which disentangles nonstationary and periodic components of the spatiotemporal features of nonstationary signals. In this study, we measured the spectral similarity using a continuous wavelet transform, and Eq. (7) is updated as
where \(N_{uv} \left( t \right) = \mathop \smallint \limits_{ - \infty }^{\infty } \left| {\left( {{\varvec{W}}\left( {t,\omega } \right)} \right)_{uv} } \right|{\text{d}}\omega\) is a normalization constant; \(c\left( {t,\omega } \right)\) measures the spectral similarity between two signals at both time and frequency domain. As a result, one can monitor how the spectral similarity patterns change in both time and frequency to evaluate the proximity to elimination threshold of infectious disease.
The proposed approach can be advantageous compared with existing methods, including temporal and spatial correlation and variance, which are used as warning signals in the literature. This method shows the covariation between two signals as a function of frequency and thus provides a set of coefficients (in contrast to a single coefficient), which better describe the underlying connections between system components. More importantly, spectral similarity is able to reveal the increased connectivity in systems with traveling waves in their dynamics. While time-domain methods like temporal correlation capture connectivity in a system with stationary oscillations, they do not capture changes in the connectivity in systems where a travelling wave induces instability. The proposed approach, in contrast, considers correlation in the frequency domain and can be an efficient method to be applied to systems exhibiting traveling wave in their dynamics. Since previous studies have revealed existence of traveling waves in the process of disease spread in spatial systems [23, 24], the proposed approach provides an advantage in studying disease dynamical systems.
4 Results and discussion
4.1 Simulation study
Two-patch SIR metapopulation model. To illustrate the application of the proposed indicator, we first evaluated the method with a simple, 2-patch metapopulation model. The parameters of this model are given in Table S2. In Fig. 1a, we show the equilibrium number of infected for the deterministic model in both patches across the range of transmission rates for which the endemic equilibrium is stable. As the vaccination rate is increased, \(R_{e}\) approaches one and the prevalence declines, approaching the disease-free steady state. To evaluate the validity of the proposed indicator in the absence of errors and approximations associated with numerical and data observation/reporting processes, we first analytically computed the spectral similarity coefficient \(c\left( t \right)\) as a function of vaccination rate using \(I_{1}\) and \(I_{2}\) variables and Eq. (6). Results of this analysis show that the theoretical value of \(c\) rises exponentially as \(R_{e}\) approaches its critical value (Fig. 1b).
Next, to analyze the trend of the spectral similarity coefficient using observations of infected individuals in each patch as in real-world scenarios, we performed numerical stochastic simulations of the metapopulation model with the same vaccination rates as the deterministic study. At each vaccination rate, the model is simulated for 10 years and the dynamics is sampled at increments of 0.1 year. The similarity coefficients are then approximated numerically using the power spectra of sampled data of infected individuals in each patch (Eq. (6)). Results of this numerical analysis are plotted on the top of the analytical results in Fig. 1b. Results show that the value of spectral similarity coefficient calculated from stochastic simulations grows as the vaccination rate increases but falls in the immediate neighborhood of the critical value. Analysis of the simulations shows that numerically approximated power spectra agree with Eq. (6) when the number infected at equilibrium is large and the linear noise approximation performs well (see Fig. S5). In the vicinity of the critical point, however, the linear noise approximation fails and the approximated similarity coefficient does not follow the analytically expected path. As a result, it is expected that in the practical application of this approach, the peak in the similarity coefficient is observed ahead but in a close neighborhood of the critical point. Such an observation is consistent with what we observe in the empirical study presented in Sect. 4.2.
Multi-patch SIR metapopulation model. Here, we consider a 8-patch metapopulation model as described in Sect. 2. The parameters and interactions were selected to generate a spatially heterogeneous epidemic. In particular: \(r_{i} \in \left[ {5e4, 5e5} \right]\), \(\mu_{i} \in \left[ {0.02, 0.04} \right] \;{\text{year}}^{ - 1} \), \(\gamma_{i} = 14\;{\text{year}}^{ - 1}\), where \({\text{year}}^{ - 1}\) is the unit and indicates the frequency of the event per individual per year. The parameters \(r_{i}\), \(\mu_{i}\), and \(\gamma_{i}\) were randomly selected from a uniform distribution within the specified ranges. We define interaction terms \((\beta_{ij}\)) in the model in the form of an interaction matrix \({\varvec{B}}\), where \(\left( {\varvec{B}} \right)_{ij} = \beta_{ij}\). We assume that \({\varvec{B}}\) is symmetric and that the within patch transmissions are similar among all patches, i.e., \(\beta_{0} = \beta_{ii} , i \in 1, \ldots ,n\). In addition, we set \(\beta_{ij} = \epsilon_{i} \beta_{0}\), where \(\epsilon_{i} \ll 1\) and \(\epsilon_{i} \ne \epsilon_{j }\), i.e., we defined a nonhomogeneous coupling in the system. Figure 2a shows the stable endemic equilibria of the system obtained by varying the vaccine uptake rate. For the set of parameters selected, the critical immunization level for disease elimination is \(\nu = 0.95\).
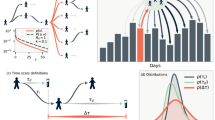
a Variation of endemic equilibrium (disease prevalence) in the simulated metapopulation model as a function of immunization level, b dynamics of the number infected in a slow approach to elimination in one of the patches, c time evolution of system level spectral similarity \(c(t,\omega )\) in the simulated metapopulation model as the immunization level gradually increases, d time evolution of spectral similarity coefficient \(c(t)\) obtained from Eq. (6) as the immunization level gradually increases
Unlike the previous example (Fig. 1), here we chose the vaccination rate to be time dependent and to vary continuously over time, resulting in a nonstationary time series. We chose initial conditions close to the endemic equilibrium when the immunization rate is \(\nu = 0\) and the model dynamics resulted from demographically driven stochastic oscillations in the absence of seasonal forcing [38]. The vaccine uptake increased gradually from \(\nu = 0\) to \(\nu = 0.95\) over 80 years, and the number of infected individuals was recorded at weekly intervals and used as an input to the multivariate similarity analysis (Fig. 2b).
Using the recorded monthly incidences at each of ten patches, the spectral similarity coefficients \(c\left( {t,\omega } \right)\) were estimated and are shown in Fig. 2. We observed that as the vaccination rate increased, there is an increase in the estimated similarity coefficient \(c\left( {t,\omega } \right)\), where the maximum value is observed in the vicinity but ahead of the elimination point. In addition to the change in the system-wide spectral similarity, we observed a progressive change in the period at which the similarity is maximum (Fig. 2c). This observation implies that the period at which the dynamics of the patches are dominantly coupled changes over time as a function of the vaccination rate, consistent with our analysis in the supplementary information.
4.2 Empirical study
To demonstrate the use of this method in practice, we analyzed monthly whooping cough incidence in the continental USA. Whooping cough is a highly contagious respiratory disease caused by the bacterium Bordetella pertussis. Despite concerted and long-standing pediatric immunization programs dating to the 1950s, pertussis control in many developed nations remains a significant public health challenge [3]. These data include monthly cases of pertussis in 49 continental US states (including the District of Columbia) from 1938 to 1980, which reflect temporal and spatial patterns of the disease over many decades. In the 1950s, a national immunization program led to a drastic reduction in incidence over the subsequent 3 decades [45]. As a result, this time series provides a unique opportunity to study the dynamics of an infectious disease exhibiting elimination due to an increase in the immunization level.
Figure 3 shows the monthly incidence of pertussis aggregated over the 49 states from 1938 to 1980. In addition, monthly incidences in six selected states are also shown in Fig. 3. The spectral similarity coefficient \(c\left( {t,\omega } \right)\) was calculated over time and is plotted in Fig. 4. We observed that as the vaccination uptake rate increased in 1950s, there was a significant increase in the estimated similarity coefficient \(c\left( t \right)\). The time at which the similarity coefficient was maximized is around 1955, which is a few years before the disease was successfully controlled in some of the states. Figure 3 shows that between 1950 and 1960 the disease was controlled in some states. In addition, the time–frequency map of the similarity coefficient \(c\left( {t,\omega } \right)\) shows a change in the frequency at which the spectral similarity was at its maximum ranged from 2 years in 1940s to 4.5 years in 1950s, which is consistent with what observed in theoretical and numerical analysis [23]. Results of this analysis show potential practical applicability of the spectral similarity coefficient in empirical epidemiological systems to reveal the proximity of the disease dynamics to a critical point.
a Time evolution of the system level spectral similarity \(c(t,\omega )\) between states of the United States using historical time series of pertussis from 1938 to 1980, b time evolution spectral similarity coefficient \(c(t)\) (i.e., \(c(t,\omega )\) averaged over all frequency ranges) for the continental USA
5 Conclusion
The spatial dynamics of infectious disease are complex, hard to model and difficult to predict. To aid the creation of efficient strategies for disease elimination and control, new model-independent methods that characterize the underlying processes of spread may be extremely useful. In this study, we introduced spectral similarity analysis of multivariate epidemiological time series as a tool for estimating the proximity of an infectious disease system to a critical point. Theoretical analyses demonstrated that when the disease prevalence gradually decreases over time and approaches elimination, the power spectrum of small fluctuations of the infected populations in each patch around their equilibrium values becomes increasingly dominated by a common function, resulting in an increased system-wide power spectral similarity. This approach exploits information contained in the spatial pattern and time–frequency structure of the epidemiological data. Results of simulated disease spread on a metapopulation model and analysis of historical spatiotemporal data of pertussis cases provide empirical validation of the method under both ideal and real-world data reporting.
The proposed approach is motivated by dynamical systems theory and represents a model-independent early warning signal of critical transitions in epidemiological systems. This early warning signal is supported by theory from nonlinear dynamics, and specifically the occurrence of critical slowing down in the proximity of certain bifurcations. This approach is distinct from traditional data analyses that extract feature from data and identify trends in data without considering that data results from nonlinear system dynamics. In contrast, using theoretical analysis of a well-stablished epidemiological system, this method suggests that one can identify whether the epidemiological system is approaching a critical event (emergence or elimination) in its dynamics by monitoring the multivariate spectral similarity in measurements. Since anticipating critical events in infectious disease is a formidable task, this approach combined with other methods in this active area of research may advance the field of data-driven prediction in infectious disease.
Data availability
The data and source codes used in this work can be made available upon reasonable request to the corresponding author.
References
Morens, D.M., Folkers, G.K., Fauci, A.S.: The challenge of emerging and re-emerging infectious diseases. Nature 430, 242–249 (2004)
Dayan, G.H., Quinlisk, M.P., Parker, A.A., Barskey, A.E., Harris, M.L., Schwartz, J.M.H., Hunt, K., Finley, C.G., Leschinsky, D.P., O’Keefe, A.L.: Recent resurgence of mumps in the United States. N. Engl. J. Med. 358, 1580–1589 (2008)
Rohani, P., Drake, J.M.: The decline and resurgence of pertussis in the US. Epidemics 3, 183–188 (2011)
World Health Organization: Global Tuberculosis Report 2013. World Health Organization, Geneva (2013)
Reid, M.J.A., Arinaminpathy, N., Bloom, A., Bloom, B.R., Boehme, C., Chaisson, R., Chin, D.P., Churchyard, G., Cox, H., Ditiu, L.: Building a tuberculosis-free world: The Lancet Commission on tuberculosis. Lancet 393, 1331–1384 (2019)
World Health Organization: World Malaria Report 2021. World Health Organization, Geneva (2021)
P. PAHO: Epidemiological Alert: Increase of Malaria in the Americas (2018)
Cohen, J.M., Smith, D.L., Cotter, C., Ward, A., Yamey, G., Sabot, O.J., Moonen, B.: Malaria resurgence: a systematic review and assessment of its causes. Malar. J. 11, 1–17 (2012)
Nedjadi, T., El-Kafrawy, S., Sohrab, S.S., Desprès, P., Damanhouri, G., Azhar, E.: Tackling dengue fever: current status and challenges. Virol. J. 12, 1–11 (2015)
Ertem, Z., Raymond, D., Meyers, L.A.: Optimal multi-source forecasting of seasonal influenza. PLoS Comput. Biol. 14, e1006236 (2018)
Han, B.A., Drake, J.M.: Future directions in analytics for infectious disease intelligence: toward an integrated warning system for emerging pathogens. EMBO Rep. 17, 785–789 (2016)
Yang, W., Olson, D.R., Shaman, J.: Forecasting influenza outbreaks in boroughs and neighborhoods of New York City. PLoS Comput. Biol. 12, e1005201 (2016)
Drake, J.M., Brett, T.S., Chen, S., Epureanu, B.I., Ferrari, M.J., Marty, É., Miller, P.B., O’Dea, E.B., O’Regan, S.M., Park, A.W.: The statistics of epidemic transitions. PLoS Comput. Biol. 15, e1006917 (2019)
Ferrari, M.J., Grais, R.F., Bharti, N., Conlan, A.J.K., Bjørnstad, O.N., Wolfson, L.J., Guerin, P.J., Djibo, A., Grenfell, B.T.: The dynamics of measles in sub-Saharan Africa. Nature 451, 679–684 (2008)
Bharti, N., Djibo, A., Ferrari, M.J., Grais, R.F., Tatem, A.J., McCabe, C.A., Bjornstad, O.N., Grenfell, B.T.: Measles hotspots and epidemiological connectivity. Epidemiol. Infect. 138, 1308–1316 (2010)
Finkenstädt, B., Grenfell, B.: Empirical determinants of measles metapopulation dynamics in England and Wales. Proc. R. Soc. Lond. Ser. B Biol. Sci. 265, 211–220 (1998)
De Celles, M.D., Magpantay, F.M.G., King, A.A., Rohani, P.: The impact of past vaccination coverage and immunity on pertussis resurgence. Sci. Transl. Med. 10, eaaj1748 (2018)
Magpantay, F.M.G., King, A.A., Rohani, P.: Age-structure and transient dynamics in epidemiological systems. J. R. Soc. Interface 16, 20190151 (2019)
Blackwood, J.C., Cummings, D.A.T., Broutin, H., Iamsirithaworn, S., Rohani, P.: Deciphering the impacts of vaccination and immunity on pertussis epidemiology in Thailand. Proc. Natl. Acad. Sci. 110, 9595–9600 (2013)
Metcalf, C.J.E., Lessler, J.: Opportunities and challenges in modeling emerging infectious diseases. Science (80-.) 357, 149–152 (2017)
Graham, M., Winter, A.K., Ferrari, M., Grenfell, B., Moss, W.J., Azman, A.S., Metcalf, C.J.E., Lessler, J.: Measles and the canonical path to elimination. Science (80-.) 364, 584–587 (2019)
Brett, T.S., Drake, J.M., Rohani, P.: Anticipating the emergence of infectious diseases. J. R. Soc. Interface 14, 20170115 (2017)
Choisy, M., Rohani, P.: Changing spatial epidemiology of pertussis in continental USA. Proc. R. Soc. B Biol. Sci. 279, 4574–4581 (2012)
Grenfell, B.T., Bjørnstad, O.N., Kappey, J.: Travelling waves and spatial hierarchies in measles epidemics. Nature 414, 716–723 (2001)
Brett, T.S., Rohani, P.: Dynamical footprints enable detection of disease emergence. PLoS Biol. 18, e3000697 (2020)
O’Regan, S.M., O’Dea, E.B., Rohani, P., Drake, J.M.: Transient indicators of tipping points in infectious diseases. J. R. Soc. Interface 17, 20200094 (2020)
Lucas, B., Vahedi, B., Karimzadeh, M.: A spatiotemporal machine learning approach to forecasting COVID-19 incidence at the county level in the USA. Int. J. Data Sci. Anal. 15, 1–20 (2022)
Santangelo, O.E., Gentile, V., Pizzo, S., Giordano, D., Cedrone, F.: Machine learning and prediction of infectious diseases: a systematic review. Mach. Learn. Knowl. Extr. 5, 175–198 (2023)
Wang, L., Adiga, A., Chen, J., Lewis, B., Sadilek, A., Venkatramanan, S., Marathe, M.: In: Knowledge-Guided Machine Learning, pp. 55–82. Chapman and Hall (2023)
Keeling, M.J., Rohani, P.: Modeling infectious diseases in humans and animals. Princeton University Press, Princeton (2011)
Brauer, F.: Backward bifurcations in simple vaccination models. J. Math. Anal. Appl. 298, 418–431 (2004)
Magpantay, F.M.G., Riolo, M.A., De Celles, M.D., King, A.A., Rohani, P.: Epidemiological consequences of imperfect vaccines for immunizing infections. SIAM J. Appl. Math. 74, 1810–1830 (2014)
O’Regan, S.M., Drake, J.M.: Theory of early warning signals of disease emergenceand leading indicators of elimination. Theor. Ecol. 6, 333–357 (2013)
Viboud, C., Bjørnstad, O.N., Smith, D.L., Simonsen, L., Miller, M.A., Grenfell, B.T.: Synchrony, waves, and spatial hierarchies in the spread of influenza. Science (80-.) 312, 447–451 (2006)
Anderson, R.M., Anderson, B., May, R.M.: Infectious Diseases of Humans: Dynamics and Control. Oxford University Press, Oxford (1992)
Keeling, M.J., Rohani, P.: Modeling Infectious Diseases in Humans and Animals, vol. 47. Princeton University Press, Princeton (2008). https://doi.org/10.1086/591197
Guo, H., Li, M.Y., Shuai, Z.: Global stability of the endemic equilibrium of multigroup SIR epidemic models. Can. Appl. Math. Q. 14, 259–284 (2006)
Alonso, D., McKane, A.J., Pascual, M.: Stochastic amplification in epidemics. J. R. Soc. Interface. 4, 575–582 (2007)
Van Kampen, N.G.: Stochastic Processes in Physics and Chemistry, vol. 1. Elsevier, Amsterdam (1992)
Rohani, P., Keeling, M.J., Grenfell, B.T.: The interplay between determinism and stochasticity in childhood diseases. Am. Nat. 159, 469–481 (2002)
Black, A.J., McKane, A.J.: Stochasticity in staged models of epidemics: quantifying the dynamics of whooping cough. J. R. Soc. Interface. 7, 1219–1227 (2010)
Li, M.Y., Liu, W., Shan, C., Yi, Y.: Turning points and relaxation oscillation cycles in simple epidemic models. SIAM J. Appl. Math. 76, 663–687 (2016)
London, W.P., Yorke, J.A.: Recurrent outbreaks of measles, chickenpox and mumps: I. Seasonal variation in contact rates. Am. J. Epidemiol. 98, 453–468 (1973)
Anderson, R.M., May, R.M.: Vaccination against rubella and measles: quantitative investigations of different policies. Epidemiol. Infect. 90, 259–325 (1983)
Simpson, D.M., Ezzati-Rice, T.M., Zell, E.R.: Forty years and four surveys: how does our measuring measure up? Am. J. Prev. Med. 20, 6–14 (2001)
Funding
Open access funding provided by SCELC, Statewide California Electronic Library Consortium. This research was supported by the National Institute of General Medical Sciences of the National Institutes of Health under Award No. U01GM110744. The content is solely the responsibility of the authors and does not necessarily reflect the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Amin Ghadami and Eamon B. O’Dea. The first draft of the manuscript was written by Amin Ghadami and Eamon B. O’Dea and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
We declare we have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ghadami, A., O’Dea, E.B., Drake, J.M. et al. Anticipating epidemic transitions in metapopulations with multivariate spectral similarity. Nonlinear Dyn 111, 17605–17615 (2023). https://doi.org/10.1007/s11071-023-08727-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11071-023-08727-w