Abstract
Purpose
Prognostically favorable IDH-mutant gliomas are known to produce oncometabolite D-2-hydroxyglutarate (2HG). In this study, we investigated metabolite-based features of patients with grade 2 and 3 glioma using 2HG-specific in vivo MR spectroscopy, to determine their relationship with image-guided tissue pathology and predictive role in progression-free survival (PFS).
Methods
Forty-five patients received pre-operative MRIs that included 3-D spectroscopy optimized for 2HG detection. Spectral data were reconstructed and quantified to compare metabolite levels according to molecular pathology (IDH1R132H, 1p/19q, and p53); glioma grade; histological subtype; and T2 lesion versus normal-appearing white matter (NAWM) ROIs. Levels of 2HG were correlated with other metabolites and pathological parameters (cellularity, MIB-1) from image-guided tissue samples using Pearson’s correlation test. Metabolites predictive of PFS were evaluated with Cox proportional hazards models.
Results
Quantifiable levels of 2HG in 39/42 (93%) IDH+ and 1/3 (33%) IDH– patients indicated a 91.1% apparent detection accuracy. Myo-inositol/total choline (tCho) showed reduced values in astrocytic (1p/19q-wildtype), p53-mutant, and grade 3 (vs. 2) IDH-mutant gliomas (p < 0.05), all of which exhibited higher proportions of astrocytomas. Compared to NAWM, T2 lesions displayed elevated 2HG+ γ-aminobutyric acid (GABA)/total creatine (tCr) (p < 0.001); reduced glutamate/tCr (p < 0.001); increased myo-inositol/tCr (p < 0.001); and higher tCho/tCr (p < 0.001). Levels of 2HG at sampled tissue locations were significantly associated with tCho (R = 0.62; p = 0.002), total NAA (R = − 0.61; p = 0.002) and cellularity (R = 0.37; p = 0.04) but not MIB-1. Increasing levels of 2HG/tCr (p = 0.0007, HR 5.594) and thresholding (≥ 0.905, median value; p = 0.02) predicted adverse PFS.
Conclusion
In vivo 2HG detection can reasonably be achieved on clinical scanners and increased levels may signal adverse PFS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diffuse infiltrating gliomas present as a family of diseases that demonstrate heterogenous malignancy and survival characteristics. Following the discovery that 70–90% of lower-grade (2/3) gliomas (LrGG) [1, 2] carry prognostically-favorable mutations in isocitrate dehydrogenase 1 & 2 (IDH 1/2), molecular classification has become increasingly emphasized under the precision medicine paradigm [1,2,3,4,5,6]. According to WHO 2021 classification guidelines [7], IDH mutations are now treated as exclusive features of LrGG, with 1p/19q-codeletion being used to further separate oligodendroglial versus astrocytic subtypes. Besides their prognostic significance and biologic implications [8], IDH mutations hold considerable importance based on the resultant enzymatic production of oncometabolite D-2-hydroxyglutarate (2HG) [9], which induces broad metabolic reprogramming [10] and functions as a marker of tumor that can confirm mutational status.
Non-invasive detection of 2HG was developed through a variety of 1H magnetic resonance spectroscopy (MRS) techniques [11,12,13,14]. These include the short echo time (TE = 30 ms) with spectral fitting [13]; long TE (97–110 ms) that exploits the unique signal evolution of 2HG’s strongly coupled spin system [14, 15]; and spectral editing methods (TE = 68 ms) designed to isolate 2HG from background resonances via coupling patterns [11, 14]. Clinical applications of 2HG MRS have shown diagnostic utility in identifying gliomas that express a range of IDH1/2 mutations [16, 17] and reliably quantifying concentrations in excess of 1 mM [18]. Moreover, levels of 2HG quantified from spectral data were demonstrated to increase with glioma progression [18] and decrease in response to treatment [18,19,20], while also displaying correlations with tumor cellularity [12, 13, 18, 20]. Patient lesions presenting with IDH2, versus IDH1, mutations have notably been characterized spectroscopically by more prodigious formation of 2HG [21, 22].
In this study, we investigated metabolomic characteristics of patients with IDH-mutant LrGG prior to surgery using a 2HG-targeted, long-TE [14] implementation of 3-D MR spectroscopic imaging (MRSI) that enabled whole-brain coverage. The goals were to evaluate metabolite features of molecular-based classifications of LrGG, correlate them with image-guided tissue pathology, and assess their predictive role on progression-free survival (PFS).
Methods
Patient population
Forty-five patients diagnosed with new or recurrent LrGG and presenting for surgical resection were recruited to this IRB-approved study following informed consent. A summary of patient population characteristics is provided in Table 1. The median age at recruitment was 34 years, with a range of 19–72 (60% male; 40% female).
Pre-surgical MRI and MRSI
Pre-surgical MR imaging was acquired using a 3 Tesla scanner (GE Healthcare Technologies, Waukesha, WI) equipped with an 8-channel phased-array headcoil. The anatomical imaging protocol included T2-weighted fluid-attenuated inversion recovery (FLAIR), T1-weighted inversion recovery spoiled gradient echo (IR-SPGR) pre/post-gadolinium images, diffusion-weighted images acquired in the axial plane with six gradient directions (TR/TE = 1000/108 ms, voxel size = 1.7 × 1.7 × 3 mm3, b = 1000 s/mM2), and perfusion-weighted images acquired using a T2*-weighted echo-planar imaging sequence (TR/TE = 1250–1500/35–54 ms, 30–35° flip angle, 128 × 128 matrix, 3–5 mM slice thickness, 7–15 slices; 3 mL/s bolus injection of 0.1 mmol/kg gadolinium diethyltriamine pentaacetic acid). The 2HG-specific 3-D MRSI data were acquired using point-resolved spectroscopic selection (PRESS) (TR/TE1/TE2 = 1500/32/65 ms, matrix size = 18 × 18 × 16, nominal voxel size = 1 × 1 × 1 cm3) and flyback echo-planar readout, with a total acquisition time of 8.2 min [14, 23].
MRSI post-processing
MRSI data were reconstructed for the full natively acquired 3D volumes, as previously described [24], and for correlative analysis of spectra versus tissue assays, spectral voxels were recentered at the locations of image-guided tissue. The choline-to-N-acetylaspartate index (CNI) was computed automatically for each voxel using in-house software [25]. Quantification of metabolite data was performed using LCModel spectral fitting [26], with Cramer-Rao lower bounds (CRLBs) ≤ 10% for total choline (tCho; phosphocholine, glycerophosphocholine, choline), total creatine (tCr: creatine, phosphocreatine), total NAA (tNAA; N-acetylaspartate, N-acetylaspartylglutamate); 20% for glutamate (Glu), glutamine (Gln), myo-inositol (Ml), and glycine (Gly); and 25% for 2HG and GABA. Metabolite levels were expressed relative to tCr.
Anatomical segmentation and ROIs
The T2 lesion (T2L) and contrast-enhancing lesion (CEL) were segmented using 3D Slicer [27]. Normal-appearing white matter (NAWM) ROIs were generated by segmenting pre-contrast T1-weighted images using the FSL FAST algorithm [28] and subtracting the T2L. Volumes of each ROI were determined along with the spectroscopically-defined region of CNIT2L > 2 in the T2L. MRSI voxels containing ≥ 50% T2L or ≥ 75% NAWM volumes were classified according to these respective ROIs.
Image-guided tissue sampling
Based on pre-surgical MRI parameters, suspected tumor within the T2 lesion was designated for intra-operative tissue sampling. Regions with low apparent diffusion coefficients (ADC), high CNI (> 2; metabolic lesion), and/or elevated cerebral blood volume (CBV) were defined as 5-mm spherical targets using BrainLab (BrainLab Inc., Munich, Germany) surgical navigation software [29]. Additionally, targets had been chosen according to surgical accessibility and heterogeneity criteria that stipulated spatially distributed sampling. Surgeons located and removed these image-guided targets during surgery at their discretion, and the LPS-coordinates for the actual site of tissue removal were recorded. Tissue was acquired early in the surgery to minimize potential tissue shift that would impact registration with imaging. Upon excision, samples were immediately fixed in 10% zinc formalin, dehydrated by graded ethanols, and embedded in Paraplast Plus wax (McCormick Scientific LLC, St. Louis, MO) using standardized immunohistochemistry protocols.
IHC parameters
A single pathologist performed immunohistochemical analysis of the image-guided tissue samples. While CNS WHO grading [7] characterized the overall tumor, each image-guided tissue sample was assessed according to tumor cellularity, measured by the mean number of cells within a 200 × field-of-view (0.776 mM2), and proliferation (MIB-1 index), as determined by immunostaining with Ki-67 (clone MIB-1; Ventana, 790-4286). IDH mutation (IDH+) was assessed primarily via immunostaining with an anti-IDH1R132H antibody (Dianova, H09), and in some instances by sequencing targeted to IDH1/2 mutational variants, allowing IDH-wildtype tumor to also be identified. Fluorescence in situ hybridization (FISH) analysis was used to assess 1p/19q co-deletion and immunostaining was performed for p53 mutation. Strong nuclear staining for p53 in ≥ 10% of tumor cells was used as a surrogate for TP53 mutation [30].
Statistical analysis
Wilcoxon rank sum tests in R were used to compare MRSI and volumetric parameters from IDH+ patients, according to ROIs, glioma grade, histological subtype, and status of pathological mutation (p53). Correlations were performed in R among continuous metabolite and pathology parameters (Pearson), and ordinal pathology parameters (Kendall-Tau). The influence of metabolite parameters on progression-free survival (PFS) was evaluated with Cox proportional hazards models, and PFS was compared between subgroups using the log-rank test. Due to the exploratory nature of this study, correction for multiple comparisons was not made.
Results
WHO 2021 tumor characterization
A total of 108 image-guided tissue samples were acquired from 45 patients undergoing tumor resection as part of standard of care at UCSF. Table 1 details pathological and molecular tumor characteristics within the patient population. Analysis of the tissue revealed 42/45(93%) patients with IDH-mutant diffuse glioma, determined on the basis of IHC staining for the IDH1R132H mutation (n = 39), IDH1 sequencing (n = 1) and IDH2 sequencing (n = 2). The remaining 3/45(7%) gliomas were verified as IDH1-negative by both IDH1R132H immunostaining and IDH1 sequencing; IDH2 sequencing showed no mutation in 2/3 of these cases, with the third case being unanalyzable but unlikely IDH2 mutated (p53-negative, 1p/19q intact). Among the 42 patients with IDH-mutant tumors (hereafter referred to as IDH+) there were 29(69%) grade 2 gliomas (13 oligodendrogliomas; 16 astrocytomas) and 13(31%) grade 3 gliomas (3 oligodendrogliomas; 10 astrocytomas). A majority of IDH+ patients (37/42; 88%) were newly diagnosed with LrGG and formed the focus of reported findings based on their treatment-naïve status, while some patients (5/42; 12%) had at least one recurrence. Pathology demonstrated p53 expression in 27/34 (79%) assayed newly diagnosed tumors.
With respect to radiological presentation, the IDH+ patients displayed mostly non-enhancing tumors (36/42; 86%) on anatomical MRI, with mean ± standard deviation T2L volumes of 48.3 ± 36.9 cm3 and CEL volumes (n = 6) ranging 0.14–2.58 cm3.
In vivo brain metabolism
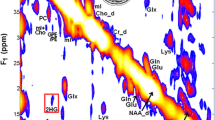
Example pre-surgical MRSI data from a patient presenting with newly diagnosed IDH+ grade 2 astrocytoma are shown in Fig. 1. A CNI map overlaid on a FLAIR image highlights the metabolic abnormality within the T2 lesion (Fig. 1A), as illustrated by the corresponding spectra (Fig. 1B). Data reconstructed at the location of surgical tissue sampling indicated by the black ring show the 2HG resonance alongside other common brain metabolites (Fig. 1C). Based on LCModel quantification, the fitted contributions of individual metabolites were determined in the region of 2HG, with the red-dotted line depicting the specific chemical shift of 2HG at 2.26 ppm (Fig. 1D). The map of 2HG/tCr (Fig. 1E) demonstrates the localization of the oncometabolite within the T2L; whereas MI/tCho (Fig. 1F) increases outside of the lesion.
2HG-specific MRSI. Spectral data acquired prior to surgery from a patient with newly diagnosed IDH-mutant grade 2 astrocytoma. A CNI map overlaid on a FLAIR image (A) and corresponding spectra (B) illustrate elevated levels of choline species associated with cell turnover in the lesion; the black ring in the T2 lesion indicates where tissue was surgically sampled. Spectra reconstructed (C) and quantified (D) at the site of tissue sampling highlight the 2HG resonance among other common brain metabolites. The map of 2HG/tCr (E) shows the localization of 2HG within the T2L, while the map of MI/tCho (F) is conversely elevated in normal-appearing tissue
MRSI spectra from T2Ls displayed apparent quantifiable levels of 2HG in 39/42(93%) IDH+ and 1/3(33%) IDH− patients, thus providing an overall detection accuracy of 91.1% for the entire LrGG population. Table 2 presents a summary of significant features describing newly diagnosed IDH+ lesions. Among newly diagnosed IDH+ patients (n = 42), levels of 2HG + GABA/tCr in the T2L were significantly elevated compared to NAWM (p < 1.0 × 10–5); whereas levels of Glu/tCr (p < 1.0 × 10–3) showed a corresponding reduction (Table 2; Fig. 2). Relative to NAWM, the IDH + T2L was also characterized by elevations in Gln/tCr (p = 0.001), MI/tCr (p < 1.0 × 10–4), MI + Gly (mIG)/tCr (p = 1.0 × 10–9), MI/tCho (p = 0.001), mIG/tCho (p < 1.0 × 10–4), tCho/tCr (p < 1.0 × 10–5) and tCho/tNAA (p < 1.0 × 10–6); and a reduction in tNAA/tCr (p < 1.0 × 10–10) (Table 2; Fig. 2).
MRSI in IDH+ patients. 3D MRSI data quantified by LCModel showing the ratios of 2HG+ GABA/tCr (A), Glu/tCr (B), Gln/tCr (C) and MI/tCho (D) in patients with newly diagnosed IDH-mutant gliomas: T2L versus NAWM. The ratio of MI/tCho is also shown for the comparison of grade 2 versus 3 (E) and oligodendroglioma versus astrocytoma (F). See Table 2
Among newly diagnosed IDH+ LrGG, the grade 2 gliomas demonstrated an elevated ratio of MI/tCho (p = 0.020) compared to grade 3 counterparts (Table 2; Fig. 2). When evaluated according to histological subtype, oligodendrogliomas (81% grade 2; 19% grade 3) showed higher MI/tCho (p = 0.037) and mIG/tCho (p = 0.023) relative to astrocytomas (52% grade 2; 48% grade 3). Grade-based differences in histological subtypes were noted as a source of bias in this comparison. Additionally, astrocytic tumors, which were characterized by a greater proportion of grade 3 versus 2 pathology, consistently demonstrated the largest volumes of CNIT2L > 2 (p < 0.01) and T2L (p < 0.05) (Table 2; Fig. 2).
Available p53 expression data among newly diagnosed IDH+ LrGG [18(69%) astrocytoma; 8(31%) oligodendroglioma] showed higher levels of tCho/tCr (p = 0.017), together with lower levels of MI/tCho (p = 0.021) and mIG/tCho (p = 0.018), when compared against p53-wildtype tumor [2(22%) astrocytoma; 7(78%) oligodendroglioma] (Table 2; Fig. 2). The p53-mutant tumors also presented larger volumes of CNIT2L > 2 (p = 0.008) and T2L (p = 0.009) relative to non-mutant counterparts (Table 2).
Spectroscopic and pathologic correlations
Tumor was verified by histopathology in 75/108 image-guided samples, 28 of which displayed reconstructed 2HG spectra at the site of sampling with CRLB < 25 (17 patients). Figure 3 shows 2HG levels in sampled regions correlated with tCho/tCr (R = 0.62, p = 0.002), tNAA/tCr (R = − 0.61, p = 0.002), and cellularity (R = 0.37, p = 0.04; CRLB < 30, n = 32); no correlation between 2HG and MIB-1 was observed. Evaluation of metabolite-pathology correlations across all tumor-containing samples, revealed tNAA was inversely correlated with cellularity (R = − 0.35, p = 0.002; n = 75); and CNI was correlated with MIB-1 (R = 0.27, p = 0.041, n = 58). MRSI metabolite correlations within the T2Ls of newly diagnosed IDH+ glioma are presented in Supplementary Fig. 1. CNI-related parameters were inversely correlated with levels of tNAA/tCr and MI/tCho; 2HG and tCho/tCr were weakly correlated.
PFS analysis
The median PFS among all IDH+ patients was 3.8 years (95% CI 2.6–4.9 years; 17 censored), and 4.0 years (95% CI 2.7–5.3 years) in the newly diagnosed cohort (Supplementary Fig. 2A; 15 censored). Tumor grade, histological subtype and recurrence status were not significantly correlated with PFS. Cox proportional hazard models indicated that higher levels of 2HG/tCr (p = 0.0007, HR 5.594) and Glu/tCr (p = 0.006, HR 32.567), along with larger volumes of T2L (p = 0.02, HR 1.014) and non-enhancing lesion (p = 0.03, HR 1.014), were predictive of worse PFS in newly diagnosed patients (Supplementary Table 1). Higher levels of 2HG/tCr (≥ 0.905, median value; p = 0.02; Fig. S2B) and higher levels of Glu/tCr (≥ 0.945, median value; log-rank test, p = 0.02; Fig. S2C) were associated with a significant reduction in PFS (Table S1). Based on Cox regression analysis, there was no significant difference between populations empirically thresholded by 2HG/tCr or Glu/tCr levels with respect to extent of resection or treatment post surgery (Supplementary Tables 2 and 3).
Discussion
This MRSI study demonstrated the detection and quantification of 2HG in patients with LrGG, who underwent image-guided tissue sampling during surgery to assess regional histopathology. Among IDH+ LrGG patients, the full complement of metabolite data derived from 3-D MRSI helped to noninvasively differentiate tumor with respect to pathological grade, histological subtype (1p/19q codeletion), p53 expression, and NAWM. Additionally, levels of 2HG quantified from data that was reconstructed at tissue sampling locations correlated with other metabolites and cellularity. Based on empirical thresholding of lesion-wide 2HG levels, it was possible to distinguish a subpopulation of IDH+ patients that manifested adverse PFS associated with elevations of the oncometabolite.
Using a clinically feasible MRS strategy for detecting 2HG with enhanced 3-D coverage and 1-cm3 resolution (tacq = 8.2 min.), this study achieved 91% apparent accuracy in predicting IDH mutational status, which is in line with prior literature detailing technical reliability [16, 17]. However, this assessment of diagnostic accuracy must be contextualized by the considerable class imbalance created by the 3 IDH− patients relative to the larger overall population and the absence of a validation cohort to enrich the number of IDH- patients.
While a majority (93%) of patients expressing mutant IDH displayed 2HG-positive quantification, 1/3(33%) patients who lacked immunohistochemical and sequencing evidence of IDH mutation nevertheless indicated 2HG presence. This discordant case was evaluated by IHC for the most common IDH1R132H-mutant enzyme, which accounts for approximately 90% of identified IDH1 mutations [1], and also sequenced for the range of IDH1 R132/R100 and rarer IDH2 R172/R140 mutational variants [31]. As a matter of technical limitation, such false identification of 2HG can arise with LCModel quantification of background metabolites GABA, Glu and Gln, as well as lipids from necrotic lesions [32, 33], resonating in the H4,H4’-proton region of 2HG around 2.26 ppm. Other challenges to absolute detection can stem from partial volume effects in small lesions [20] and relatedly low 2HG concentrations (< 1 mM) [18] (here indicated by relatively higher tNAA levels), coupled with the relatively high spectroscopic spatial resolution that limited 2HG signal most noticeably from data reconstructed at tissue sampling sites (28/75).
Because the study population was overwhelmingly comprised of patients with IDH+ LrGG, it was difficult to assess the broader metabolic implications of IDH mutations [10] relative to IDH-lesions. Earlier investigations have demonstrated glutamate reduction in IDH-mutagenized cell lines [34], surgically-derived IDH+ tumor samples [35, 36], and high-grade IDH+ lesions (7 T MRS) [37] compared to non-mutated counterparts, perhaps as a consequence of conversion to 2HG-precursor alpha-ketoglutarate via glutamate dehydrogenase [36, 38]. Some of the same studies additionally reported increased glutamine levels in IDH+ versus IDH- lesions, which may represent heightened metabolic dependency in the mutated phenotype [34, 36, 37]. Although primarily treating IDH+ lesions here, the characteristically reduced glutamate and elevated glutamine levels in tumor versus NAWM follow similar trends.
Besides 2HG production and apparent alterations to glutamine/glutamate metabolism, the IDH+ T2Ls displayed a variety of features distinguishing them from NAWM. The elevation in choline species, together with reduced levels of tNAA, was consistent with established literature characterizing increased cell turnover and reduced neuronal function in glioma, respectively [39, 40]. Several studies have suggested that increased MI within the T2L versus NAWM can be understood as the result of heightened glial cellularity, particularly in astrocytomas [41]. As a known marker of malignant tumor, the elevation in Gly, which was fit in combination with MI due to spectral overlap, represents an expected finding [42]. The ratio of MI/tCho, previously evaluated as a marker of gliosis versus recurrent tumor [43], was shown to be reduced in the lesion owing to the excess choline content.
Despite lacking practical utility in discerning tumor lesions from NAWM, the ratio of MI/tCho was shown to function as a broad indicator of inter-tumoral malignancy when performing pairwise comparisons of molecular/pathological glioma classifications. A major caveat to this finding, however, was that aggressive tumors, which demonstrated an inverse relationship with MI/tCho, tended to be astrocytic (1p/19q-wildtype) and of higher grade. Higher values of MI/tCho, for example, provided the best means of discriminating grade 2 versus 3 IDH+ glioma, with prior research showing effective stratification of grades 2–4 [44]. This ratio was likewise found to be higher in the least aggressive forms of IDH+ glioma across histological and molecular sub-types: e.g., oligodendroglial (1p/19q co-deleted) and p53-wildtype. Volumetric trends showing smaller T2L and CNIT2L > 2 regions in p53-wildtype and oligodendroglial tumors also distinguished less aggressive gliomas.
2HG correlations within surgically sampled regions supported a conventional understanding of the tumor microenvironment. While showing threshold-level significance, the relationship observed between 2HG and cellularity comports with the notion that tumor cell density influences 2HG productive capacity, as evidenced by other pre-surgical studies [12, 13, 18, 20] and those employing ADC as a surrogate for cellularity [20, 45]. Image-guided samples of tumors displayed a correlation between pathological measurement of cellular proliferation via MIB-1 and the spectroscopic marker of CNI, consistent with prior research [46]. Although 2HG was not found to be associated with the MIB-1 index, metabolite correlations with 2HG highlighted the abovementioned elevation of total choline and concomitant reduction in tNAA, which have historically been used as non-invasive measures of tumor malignancy. Despite the challenges that image-guided studies pose in terms of tissue shift and image-tissue misregistration, the technical implementation of 3-D MRSI coverage demonstrated in this study increased the opportunity for evaluating in vivo correlates of sampled tissue.
Of particular importance in this study were the findings that adverse PFS could most reliably be predicted by increasing levels of 2HG and glutamate, the latter having previously been associated with malignant progression [35]. These results suggest that spectroscopic measures of 2HG and other metabolites provide information that extends beyond the mere identification of IDH mutation by IHC and genomic analyses. Moreover, empirically thresholding levels of 2HG in IDH+ patients enabled a practical means of discriminating subpopulations with disparate PFS, showing the potential utility of 2HG as a prognostic marker. With the current challenges in monitoring response to treatment across a range of standard-of-care and experimental therapies, several promising studies have already sought to leverage this utility by capturing longitudinal changes in 2HG associated with disease progression and radiochemotherapy using MRS [18,19,20]. The collective advances being made in 2HG imaging lend support to the emergent clinical value of in vivo spectroscopy and its potential to enhance existing MR protocols for patients with LrGG.
Conclusion
In vivo 2HG detection can be achieved on clinical scanners and may have prognostic value based on the relationship between quantified levels and PFS. Spectroscopic strategies for resolving 2HG benefit from the technical implementation of 3-D MRSI that extends volumetric coverage.
Data availability
Data available upon request.
Code availability
SIVIC visualization software available at: https://github.com/SIVICLab/sivic
References
Yan H, Parsons DW, Jin G et al (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360:765–773
Balss J, Meyer J, Mueller W et al (2008) Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol 116:597–602
Houillier C, Wang X, Kaloshi G, K, et al (2010) IDH1 or IDH2 mutations predict longer survival and response to temozolomide in low-grade gliomas. Neurology 75:1560–1566
Brat DJ, Verhaak RG, Aldape KD et al (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372:2481–2498
Eckel-Passow JE, Lachance DH, Molinaro AM et al (2015) Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N Engl J Med 372:2499–2508
Olar A, Wani KM, Alfaro-Munoz KD et al (2015) IDH mutation status and role of WHO grade and mitotic index in overall survival in grade II-III diffuse gliomas. Acta Neuropathol 129:585–596
Louis DN, Perry A, Wesseling P et al (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol 23:1231–1251
Du X, Hu H (2021) The roles of 2-Hydroxyglutarate. Front Cell Dev Biol 9:651317
Dang L, White DW, Gross S et al (2009) Cancer- associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462:739–744
Ježek P (2020) 2-Hydroxyglutarate in cancer cells. Antioxid Redox Signal 33:903–926
Andronesi OC, Kim GS, Gerstner E et al (2012) Detection of 2-hydroxyglutarate in IDH-mutated glioma patients by in vivo spectral-editing and 2D correlation magnetic resonance spectroscopy. Sci Transl Med. https://doi.org/10.1126/scitranslmed.3002693
Elkhaled A, Jalbert LE, Phillips JJ et al (2012) Magnetic resonance of 2-hydroxyglutarate in IDH1-mutated low-grade gliomas. Sci Transl Med. https://doi.org/10.1126/scitranslmed.3002796
Pope WB, Prins RM, Thomas MA et al (2012) Non-invasive detection of 2-hydroxyglutarate and other metabolites in IDH1 mutant glioma patients using magnetic resonance spectroscopy. J Neurooncol 107:197–205
Choi C, Ganji SK, DeBerardinis RJ et al (2012) 2-hydroxyglutarate detection by magnetic resonance spectroscopy in IDH-mutated patients with gliomas. Nat Med 18:624–629
Shams A, van der Kemp WJM, Emir U et al (2021) Comparison of 2-hydroxyglutarate with sLASER and MEGA-sLASER. Front Neurol 7:718423
Suh CH, Kim HS, Jung SC et al (2018) 2-hydroxyglutarate MR spectroscopy for prediction of isocitrate dehydrogenase mutant glioma: a systemic review and metal-analysis using individual patient data. Neuro Oncol 20:1573–1583
Zhou M, Zhou Y, Liao H et al (2018) Diagnostic accuracy of 2-hydroxyglutarate magnetic resonance spectroscopy in newly diagnosed brain mass and suspected recurrent gliomas. Neuro Oncol 20:1262–1271
Choi CC, Raisanen JM, Ganji SK et al (2016) Prospective longitudinal analysis of 2-hydroxyglutarate magnetic resonance spectroscopy identifies broad clinical utility for the management of patients with IDH-mutant glioma. J Clin Oncol 34:4030–4039
Andronesi OC, Loebel F, Bogner W et al (2016) Treatment response assessment in IDH-mutant glioma patients by noninvasive 3D functional spectroscopic mapping of 2-hydroxyglutarate. Clin Cancer Res 22:1632–1641
De la Fuente MI, Young RJ, Rubel J et al (2016) (2016) Integration of 2-hydroxyglutarate- proton magnetic resonance spectroscopy into clinical practice for disease monittoring in isocitrate dehydrogenase-mutant glioma. Neuro Oncol 18(2):283–290
Emir UE, Larkin SJ, de Pennington N et al (2015) Noninvasive quantification of 2-hydroxyglutarate in human gliomas with IDH1 and IDH2 mutations. Cancer Res 76:43–49
Shen X, Voets NL, Larkin SJ et al (2019) A noninvasive comparison study between human gliomas with IDH1 and IDH2 mutations by MR spectroscopy. Metabolites 9:35
Park I, Chen AP, Zierhut ML et al (2011) Implementation of 3 T lactate-edited 3D 1H MR spectroscopic imaging with flyback echo-planar readout for gliomas patients. Ann Biomed Eng 39:193–204
Nelson SJ, Kadambi AK, Park I et al (2017) Association of early changes in H-1 MRSI parameters with survival for patients with newly diagnosed glioblastoma receiving a multimodality treatment regimen. Neuro Oncol 19:430–439
McKnight TR, von dem Bussche MH, Vigneron DB et al (2002) Histopathological validation of a three-dimensional magnetic resonance spectroscopy index as a predictor of tumor presence. J Neurosurg 97:794–802
Provencher SW (2001) Automatic quantitation of localized in vivo 1H spectra with LC Model. NMR Biomed 14:260–264
Menze BH, Jakab A, Bauer S et al (2015) The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans Med Imaging 34:1993–2024
Zhang Y, Brady M, Smith S (2001) Segmentation of brain MR images through a hidden random field model and the expectation maximization algorithm. IEEE Trans Med Imaging 20:45–57
Luks TL, McKnight TR, Jalbert LE et al (2018) Relationship of in vivo MR parameters to histopathological and molecular characteristics of newly diagnosed, nonenhancing lower-grade gliomas. Transl Oncol 11:941–949
Takami H, Yoshida A, Fukushima S et al (2015) Revisiting TP53 mutations and immunohistochemistry: a comparative study in 157 diffuse gliomas. Brain Pathol 25:256–265
Parsons DW, Jones S, Zhang X et al (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321:1807–1812
Suh CH, Kim HS, Paik W et al (2019) False positive measurement of 2-hydroxyglutarate MR spectroscopy in isocitrate dehydrogenase wild-type glioblastoma: a multifactorial analysis. Radiology 291:752–762
Tietze A, Choi C, Mickey B et al (2018) Noninvasive assessment of isocitrate dehydroge- nase mutation status in cerebral gliomas by magnetic resonance spectroscopy in a clinical setting. J Neurosurg 128:391–398
Reitman ZJ, Jin G, Karoly ED (2011) Profiling the effects of isocitrate dehydrogenase 1 and 2 mutations on the cellular metabolome. Proc Natl Acad Sci USA 108:3270–3275
Jalbert LE, Elkhaled A, Phillips JJ et al (2017) Metabolic profiling of IDH mutation and malignant progression in infiltrating glioma. Sci Rep 7:44792
Nagashima H, Tanaka K, Sasayama T et al (2016) Diagnostic value of glutamate with 2-hydroxyglutarate in magnetic resonance spectroscopy for IDH1 mutant glioma. Neuro Oncol 18:1559–1568
Hangel G, Cadrien C, Lazen P et al (2020) High-resolution metabolic imaging of high-grade gliomas using 7T-CRT-FID-MRSI. Neuroimage Clin 28:102433
Yoo HC, Yu YC, Sung Y et al (2020) Glutamine reliance in cell metabolism. Exp Mol Med 52:1496–1516
Saraswathy S, Crawford FW, Lamborn KR et al (2009) Evaluation of MR markers that predict survival in patients with newly diagnosed GBM prior to adjuvant therapy. J Neurooncol 91:69–81
Usenius JP, Vaino P, Hernesniemi J et al (1994) Choline-containing compounds in human astrocytomas studied by 1H NMR spectroscopy in vivo and in vitro. J Neurochem 63:1538–1543
Candiota AP, Majos C, Julia-Sape M et al (2011) Non-invasive grading of astrocytic tumours from the relative contents of myo-inositol and glycine measured by in vivo MRS. JBR-BTR 94(6):319–329
Righi V, Andronesi OC, Mintzopoulos D et al (2010) High-resolution magic angle spinning magnetic resonance spectroscopy detects glycine as a biomarker in brain tumors. Int J Oncol 36:301–306
Srinivasan R, Phillips JJ, Vandenberg SR et al (2010) Ex vivo MR spectroscopic measure differentiates tumor from treatment effects in GBM. Neuro Oncol 12:1152–1161
Elkhaled A, Jalbert L, Constantin A et al (2014) Characterization of metabolites in infiltrating gliomas using ex vivo 1H high-resolution magic angle spinning spectroscopy. NMR Biomed 27:578–593
Andronesi OC, Arrillaga-Romany IC, Ly KI et al (2018) Pharmacodynamics of mutant-IDH1 inhibitors in glioma patients probed by in vivo 3D MRS imaging of 2-hydroxyglutarate. Nat Commun 9:1474
McKnight T, Lamborn KR, Love TD et al (2007) Correlation of magnetic resonance spectroscopic and growth characteristic within grades 2 and 3 gliomas. J Neurosurg 106:660–666
Acknowledgements
This study was funded by NCI R01 CA127612, P01 CA118816 and NCI T32 CA151022 (AA).
Funding
NCI R01 CA127612, P01 CA118816; NCI T32 CA151022 (AA).
Author information
Authors and Affiliations
Contributions
MB and SMC recruited patients. ML, LJ, and AA collected MR data, SMC and JV interpreted MR data, and ML, LJ, EP, and YL processed MR data. MB provided guidance for the image-guided tissue sampling, and JJ performed tissue analysis. AA, ML, and YL carried out data analysis, and AA wrote the first draft of the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial conflicts to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Autry, A.W., Lafontaine, M., Jalbert, L. et al. Spectroscopic imaging of D-2-hydroxyglutarate and other metabolites in pre-surgical patients with IDH-mutant lower-grade gliomas. J Neurooncol 159, 43–52 (2022). https://doi.org/10.1007/s11060-022-04042-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-022-04042-3