Abstract
Introduction
A large subset of diffusely infiltrative gliomas contains a gain-of-function mutation in isocitrate dehydrogenase 1 or 2 (IDH1/2mut) which produces 2-hydroxglutarate, an inhibitor of α-ketoglutarate-dependent DNA demethylases, thereby inducing widespread DNA and histone methylation. Because histone deacetylase (HDAC) enzymes are localized to methylated chromatin via methyl-binding domain proteins, IDH1/2mut gliomas may be more dependent on HDAC activity, and therefore may be more sensitive to HDAC inhibitors.
Methods
Six cultured patient-derived glioma cell lines, IDH1wt (n = 3) and IDH1mut (n = 3), were treated with an FDA-approved HDAC inhibitor, panobinostat. Cellular cytotoxicity and proliferation assays were conducted by flow cytometry. Histone modifications and cell signaling pathways were assessed using immunoblot and/or ELISA.
Results
IDH1mut gliomas exhibited marked upregulation of genes associated with the HDAC activity. Glioma cell cultures bearing IDH1mut were significantly more sensitive to the cytotoxic and antiproliferative effects of panobinostat, compared to IDH1wt glioma cells. Panobinostat caused a greater increase in acetylation of the histone residues H3K14, H3K18, and H3K27 in IDH1mut glioma cells. Another HDAC inhibitor, valproic acid, was also more effective against IDH1mut glioma cells.
Conclusion
These data suggest that IDH1mut gliomas may be preferentially sensitive to HDAC inhibitors. Further, IDH1mut glioma cultures showed enhanced accumulation of acetylated histone residues in response to panobinostat treatment, suggesting a direct epigenetic mechanism for this sensitivity. This provides a rationale for further exploration of HDAC inhibitors against IDH1mut gliomas.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mutations in isocitrate dehydrogenase 1 and 2 (termed IDH1/2mut) are present in approximately 25–30% of all diffusely infiltrative gliomas in adults, and are now part of the diagnostic criteria for oligodendrogliomas [1]. Single point mutations at key arginine amino acid residues cause a dramatic change in enzyme activity, where α-ketoglutarate is converted into D-2-hydroxyglutarate (D2HG) [2]. D2HG, in turn, inhibits certain DNA- and histone-demethylating dioxygenases that require α-ketoglutarate as a co-substrate. This gradually leads to a global increase in DNA and histone methylation in IDH1/2mut gliomas [3,4,5]. While the exact mechanisms connecting this cellular activity to gliomagenesis are not yet completely understood, IDH1/2mut gliomas are generally much less aggressive than their IDH1/2 wild-type (IDHwt) counterparts. In fact, the distinction is so dramatic that the new World Health Organization (WHO) scheme has split adult-type infiltrative gliomas into three discrete subsets: “Astrocytoma, IDH1/2mut, WHO grades 2–4,” “Oligodendroglioma, IDH1/2mut and 1p/19q-codeleted, WHO grades 2–3” and “Glioblastoma, IDH1/2wt, WHO grade 4” [6].
Since IDH1/2mut occurs early in gliomagenesis, and the histone/DNA hypermethylation state of IDH1/2mut gliomas is largely retained even during disease progression [7,8,9,10], there is a possibility that epigenetic modifiers may be particularly effective against IDH1/2mut glioma. Histone deacetylases (HDACs) are a class of epigenetic “erasers” that promote chromatin compaction via removal of acetate residues from acetylated histones [11]. HDACs form complexes with methyl-binding domain (MBD) [12] and/or MeCP2 proteins [13], particularly the NuRD complex, which includes HDAC1/2 and MBD2/3 [14,15,16,17]. This interaction directs HDAC enzymes to methylated DNA to facilitate chromatin compaction and epigenetic repression of genes associated with these methylated DNA regions. We therefore sought to determine whether IDH1/2mut gliomas might be preferentially sensitive to HDAC inhibitors (HDACi), which are already being used to treat malignancies elsewhere in the body [18].
Results
Genes that promote HDAC function are upregulated in IDH1/2mut glioma
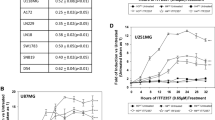
First, gene expression profiles from The Cancer Genome Atlas (TCGA) and the Chinese Glioma Genome Atlas (CGGA) were analyzed [19, 20]. IDH1/2wt and IDH1/2mut glioma samples were matched by grade (Grade 2/3 versus Grade 4) and subsequently subjected to differential gene expression analysis. Differentially expressed genes (DEGs) were further evaluated via gene ontology (GO) analysis. In both Grade 4 datasets from TCGA and CGGA, IDH1/2 status was significantly associated with upregulation of genes that modulate HDAC activity (Fig. 1b, d). In the low grade glioma cohort, this effect was only observed in the TCGA dataset, though this was not statistically significant based on adjusted p-value estimates (Fig. 1a, c). Nevertheless, 8/11 of the HDAC-related genes identified via GO analysis are significantly upregulated in the low grade TCGA and CGGA datasets (Fig. S1). HDAC-related genes upregulated in IDH1/2mut glioma include HDAC2/4/5, KDM1A, CHD4, MTA2/3, SIRT1/2, PHF21A, and BRMS1L. These data support the concept that IDH1/2mut gliomas have increased activity in HDAC pathways, and suggests targeting histone acetylation via HDAC inhibitors may exhibit profound effects in IDH1/2mut glioma.
Genes involved in promoting HDAC activity are upregulated in IDH1mut glioma. a–d Gene ontology (GO) analysis of TCGA and CGGA datasets showing that genes which facilitate HDAC function are upregulated in IDH1/2mut glioma. The top 15 GO terms are ranked based on Combined Score, and the GO term relating to HDAC activity is highlighted in yellow
Characterization of IDH expression and activity in glioma cell lines
To assess the effect of HDAC inhibition on glioma based on IDH1/2 status, we employed six patient-derived glioma cell lines: three IDH1/2wt (0827, 0923, 0211) and three IDH1mut (0905, BT142, TB096). No patient-derived IDH2mut glioma cell lines were used as they are extremely rare (only 2.3% of all glioma patients bear an IDH2mut) [21]. Of the three IDH1mut glioma cell lines, two contain the canonical heterozygous R132H mutation (0905, TB096) while BT142 has a hemizygous R132H mutation, with the wild-type allele having been lost. Information on clinical pathology and other relevant genetic alterations, are provided in Table S1. IDH1 R132H expression was confirmed using western blot and reflected the genomic status for each line, with BT142 exhibiting enhanced production of the mutant R132H IDH1 enzyme (Fig. S2a). D2HG quantitation via ELISA also showed that 0905 and TB096 cell lines produced higher levels of D2HG, whereas BT142 more closely resembled IDH1/2wt glioma cells (Fig. S2b), consistent with prior studies indicating that both IDH1wt and IDH1mut alleles are required for prominent D2HG production [8, 22,23,24].
IDH1mut glioma cells display increased cytotoxicity with HDAC inhibition
Viability of IDH1/2wt and IDH1mut glioma cells was determined using DNA chromophore cell viability and Annexin V apoptosis assays. In cell viability dose–response assays, IDH1mut glioma cells were much more sensitive to an FDA-approved pan-HDACi, panobinostat, compared to their IDH1/2wt counterparts, with a 4.1 fold difference in IC50 values (Fig. 2a, b; Fig. S3; Table S2). This response was not dependent on culture conditions, as IDH1mut cells cultured in serum-free glioma stem cells (0905 and BT142) and in serum-containing media (TB096) showed the same heightened sensitivity. Annexin V apoptosis dose–response assays showed an even more pronounced sensitivity to panobinostat in IDH1mut cells, with a tenfold difference in average EC50 values compared to IDH1/2wt (Fig. 2c, d; Fig. S4; Table S2). (The disparity of magnitude between the apoptosis and cell viability data is likely due to the apoptosis assay detecting cells that were still viable, but were in early phases of apoptosis.) Western blot analysis showed selective induction of cleaved caspase-3 (CC3) exclusively in IDH1mut glioma cells cultured with 10 nM panobinostat (Fig. 2e), confirming that panobinostat triggers apoptosis in these cells.
IDH1mut glioma is particularly sensitive to the cytotoxic effects of panobinostat. a Normalized cell viability of IDH1/2wt and mutant glioma cells treated with the HDAC inhibitor panobinostat for 5 days (n = 3). Nonlinear regression statistical analysis was utilized to determine IC50 values. b Consolidation of cell viability IC50 values derived from a stratified based on IDH status (n = 9). Statistical analysis was performed via Student’s t-test. c Induction of apoptosis in IDH1/2wt and IDH1mut glioma cells treated with panobinostat for 5 days (n = 3). Nonlinear regression statistical analysis was utilized to determine IC50 values. d Consolidation of apoptosis EC50 values derived from c stratified based on IDH status (n = 9). Statistical analysis was performed via Student’s t-test. e Western blot of cleaved caspase 3 in whole cell lysates extracted from IDH1/2wt and IDH1mut glioma cells treated with 10 nM panobinostat for 5 days. f Phase-contrast microscopy of IDH1/2wt and IDH1mut glioma cells treated with 10 nM panobinostat for 5 days
Furthermore, phase-contrast microscopy images showed that 0827 and 0923 IDH1/2wt cells retained sphere size and number in culture when treated with 10 nM panobinostat for 5 days, whereas IDH1mut cells exhibited visible signs of cytotoxicity, including reduction in sphere size, sphere number, and a pronounced increase in single cells that were phase-dark (Fig. 2f). To examine whether these data simply reflect a general chemosensitivity in IDH1mut cells, both IDH1/2wt and IDH1mut cells were subjected to puromycin dose–response assays, which showed no difference in sensitivity based on IDH status (Fig. S5; Table S3). Together, these data suggest that IDH1mut glioma cells are preferentially sensitive to the cytotoxic effects of the HDACi panobinostat.
Panobinostat preferentially inhibits cell proliferation in IDH1mut glioma cells
To determine the effect of panobinostat on IDH1/2wt versus IDH1mut glioma cell proliferation, cells were treated with panobinostat, then subjected to BrdU proliferation assays. Whereas 10 nM panobinostat suppressed IDH1/2wt cell proliferation by 33% on average, it suppressed IDH1mut proliferation by 89.5% (Fig. 3a, b; Fig. S6a, b). The difference was much smaller, though still statistically significant, at 100 nM panobinostat. Cell cycle analysis further supported the notion that IDH1mut cells are more sensitive to panobinostat, with a statistically significant induction of G1 arrest/reduction in G2-M phase only observed in IDH1mut cells (Fig. 3c; Fig. S7). Finally, 10 nM panobinostat suppressed the pro-proliferation MAPK and AKT signaling pathways in IDH1mut cells, not IDH1/2wt cells (Fig. 3d). Further, the cell cycle suppressor, p21, was variably induced by panobinostat in both IDH1mut cell lines tested, and in only one of the IDH1/2wt cell lines (Fig. 3e). No effects were seen on the related cell cycle inhibitor p16 (Fig. 3e). Together, these data suggest that, in addition to increased cytotoxicity, IDH1mut glioma cells are also preferentially sensitive to the antiproliferative effects of HDACi.
Panobinostat preferentially reduces proliferation in IDH1mut glioma. a BrdU incorporation assays of IDH1/2wt and IDH1mut glioma treated with 10 nM and 100 nM panobinostat for 48 h (n = 3). b Consolidation of data from a stratified based on IDH status (n = 6 for IDH1/2wt and n = 9 for IDH1mut). Statistical analysis was performed via Student’s t-test. c Cell cycle analysis of IDH1/2wt and IDH1mut glioma treated with 20 nM and 100 nM panobinostat for 48 h (n = 3). d Western blot of ERK and AKT in whole cell lysates extracted from IDH1/2wt and IDH1mut glioma cells treated with 10 nM panobinostat for 2 days. e Western blot of p21 and p16 in whole cell lysates extracted from IDH1/2wt and IDH1mut glioma cells treated with 10 nM panobinostat for 2 days
HDACi elicits a greater increase in histone H3 acetylation in IDH1mut glioma cells
Both IDH1/2wt and IDH1mut glioma cells responded to panobinostat with increased H3K14/18/27Ac histone marks (Fig. 4a). However, IDH1mut cells showed a three to sixfold induction in those marks, compared to a twofold increase in IDH1/2wt cells (Fig. 4b, c; Fig. S8). To evaluate global changes in histone acetylation, an ELISA against total H3KAc was used, which showed that total H3KAc content increased by 20% at 10 nM panobinostat, and 35% at 50 nM panobinostat, with IDH1mut cells showing an approximate twofold increase in H3KAc compared to IDH1/2wt cells (Fig. 4d, e). These data indicate that the heightened cytotoxic and antiproliferative effects by panobinostat in IDH1mut glioma cells are associated with a preferential increase in acetylated chromatin.
IDH1mut glioma preferentially upregulates Histone H3 acetylation in response to panobinostat treatment. a Western blot of H3K14Ac, H3K18Ac, and H3K27Ac in whole cell lysates extracted from IDH1/2wt and IDH1mut glioma cells treated with 10 nM panobinostat for 2 days. b Densitometry analysis of H3K14Ac, H3K18Ac, and H3K27Ac western blots normalized to GAPDH. Whole cell lysates were extracted from IDH1/2wt and IDH1mut glioma cells treated with 10 nM panobinostat for 2 days. Plots show total signal based on three independent experiments. c Consolidation of data from b stratified based on IDH status (n = 6) showing fold change in H3K14Ac, H3K18Ac, and H3K27Ac in response to 10 nM panobinostat for 2 days. Statistical analysis was performed via Student’s t-test. d Total H3KAc levels analyzed via ELISA in nuclear extracts from IDH1/2wt and IDH1mut glioma cells treated with 10 and 50 nM panobinostat for 2 days (n = 3). e Consolidation of data from d stratified based on IDH status (n = 6). Statistical analysis was performed via Student’s t-test
IDH1mut glioma cells are also more sensitive to the HDACi valproic acid
To investigate whether other HDACi are also preferentially active against IDH1mut glioma cells, IDH1/2wt and IDH1mut cells were treated with valproic acid (VPA), an antiepileptic with HDACi properties [25]. As with panobinostat, VPA was more toxic to IDH1mut cells than IDH1/2wt cells, with an IC50 of ~ 2 mM for IDH1mut cells versus 4.6 mM for IDH1/2wt cells (Fig. 5, Fig. S9).
Valproic acid recapitulates the preferential cytotoxicity in IDH1mut glioma. a Normalized cell viability of IDH1/2wt and IDH1mut glioma cells treated with the HDAC inhibitor valproic acid for 5 days (n = 3). Nonlinear regression statistical analysis was utilized to determine IC50 values. b Consolidation of cell viability IC50 values from a stratified based on IDH status (n = 9). Statistical analysis was performed via Student’s t-test
Discussion
Despite the comparatively longer survival of patients with IDH1mut gliomas versus IDH1/2wt glioblastomas, IDH1mut gliomas remain an ultimately fatal, incurable disease, and thus new treatment options are needed for these tumors. Targeting the canonical IDH1 R132H variant, both by small molecule inhibitors and by a vaccine, are in clinical trials [26,27,28]. Other therapeutic strategies exploiting potential metabolic vulnerabilities in IDH1mut gliomas, including inhibitors of poly (ADP-ribose) polymerase and nicotinamide phosphoribosyltransferase, are also being pursued [29,30,31]. Yet, to date, comparatively little has been done to target the unique effects of IDH1mut on the glioma epigenome and chromatin.
In this study, we showed that cultured IDH1mut glioma cells are more sensitive than IDH1/2wt cells to the FDA-approved pan-HDACi, panobinostat, and that this is mediated through both cytotoxic and cytostatic effects (Figs. 2 and 3). This contrasts with some other IDH1mut-specific cancer therapies, such as selective IDH1mut inhibitors that suppress proliferation but are not cytotoxic [20]. Panobinostat was effective against IDH1mut cells cultured in either serum free media (0905 and BT142) or serum-supplemented media (TB096), suggesting that the effect is not unique to a particular glioma cell type or culture condition. Even BT142 cells, which deleted the wild-type IDH1 allele and therefore no longer produce as much D2HG, were still more sensitive to panobinostat and therefore suggest a durable therapeutic response. This indicates that HDACi may still be a useful strategy against IDH1mut gliomas that have decreased D2HG production during disease progression. Additionally, the ribosomal inhibitor, puromycin, was not more active against IDH1mut glioma cells, demonstrating that the effects of panobinostat are unlikely to be the result of a more generalized chemosensitivity of IDH1mut cells.
One key finding in this data is the greater upregulation of total histone H3 acetylation in IDH1mut glioma cells treated with panobinostat, specifically in H3K14/18/27Ac active chromatin marks (Fig. 4). Such upregulation may be driving the increased sensitivity of IDH1mut cells to HDACi. Our data suggests that this sensitivity may be due to enhanced regulation of the acetylome in IDH1mut glioma due to increased HDAC activity (Fig. 1). Considering that panobinostat is a pan-HDACi, future experiments will investigate the role of specific HDAC isoforms in mediating increases in histone acetylation in response to panobinostat, as well as to determine the potential of HDACi against in vivo models of IDH1mut gliomas.
In addition to their proven role in treating multiple myeloma [18], HDACi are being explored in a number of other solid tumors, including glioblastomas and diffuse intrinsic pontine gliomas [32,33,34,35]. However, results thus far in IDH1/2wt glioblastoma patients have been disappointing [36,37,38]. One study that did include 10 IDH1mut astrocytoma patients was inconclusive, perhaps in part because of a reliance on historical controls, and an exclusive focus on recurrent cases that had already been treated with standard therapy [39]. A recent study by others suggested that IDH1mut gliomas were actually more resistant to HDACi [40]; however, that study used overexpression of IDH1 R132H in U87MG and U373MG glioblastoma cells. Our data demonstrate the importance of using patient-derived models, with endogenous IDH1mut, when studying therapeutics aimed at altering the epigenome and/or chromatin.
In this study, another HDACi, VPA, was also more effective against IDH1mut glioma cells (Fig. 5). VPA has already been tested in clinical trials against IDH1/2wt pediatric and adult gliomas, with varying results [41,42,43]. To date, however, it has not been tested specifically in patients with IDH1mut gliomas. Considering that VPA is a powerful antiepileptic often used to control seizures in glioma patients, and that IDH1mut gliomas are highly epileptogenic [44], VPA might have a dual purpose in IDH1mut glioma patients.
In summary, we identified that cultured glioma cells with endogenous IDH1mut are preferentially sensitive to the cytotoxic and antiproliferative effects of the FDA-approved HDACi, including panobinostat and valproic acid. HDACi increases the amount of acetylated chromatin in IDH1mut, providing a potential molecular rationale for this increased sensitivity to HDACi. In sum, this study suggests the potential of HDACi inhibition as a therapeutic strategy specifically for the treatment of IDH1mut glioma.
Materials and methods
Cell lines
Glioma stem cells 0827, 0923, 0211 and 0905 were previously isolated at the Neuro-Oncology Branch, NIH and have been previously characterized for glioma stem-like properties and genomic alterations [22,23,24,25]. BT142 [26] was purchased from ATCC, while TB096 [11] was a kind gift from Dr. Hai Yan at Duke University. All cell lines were grown in medium that includes Neurobasal-A base medium, B-27 and N2 supplements along with EGF and FGF growth factors (NBE). Two cell lines, 0905 and BT142, required 100 ng/mL PDGF-AA in the medium. TB096 was cultured in half NBE and half DMEM with 10% FBS. All cell cultures were cultured at 37C and 5% CO2 with 1 × penstrep and were frequently monitored for mycoplasma contamination. Additionally, cells were cultured in suspension as neurospheres except for TB096 which exhibited adherent growth. Cell culture medium was changed 3x/week. Panobinostat (Fischer Scientific; NC0021665) was dissolved in 100% DMSO, sterile filtered, and diluted to 1000 × in DMSO for drug treatments. For all treatments, vehicle concentrations were 0.1% or less.
Analysis of TCGA and CGGA Data
Gene expression data from TCGA and CGGA datasets was downloaded as raw counts and processed using the online tool Appyter [19, 20, 45]. DEGs between IDHwt/mut gliomas were generated using the Limma package with a p-value cutoff of 0.05 and a fold change cutoff of 0.2. GO analysis employed the use of the online tool Enrichr using the top 1000 DEGs [45]. Gene expression plots were created by uploading data to the UCSC Xena Browser, organizing data by IDH status and tumor grade, and then generating grade- and IDH-matched gene expression boxplots with corresponding t-test statistics [46].
Cell viability and apoptosis assays
Cell viability and apoptosis assays were conducted using a Luminex Guava® Muse® Cell Analyzer in conjunction with a Muse® Cell Count and Viability Kit or a Muse® Annexin V and Dead Cell Kit. Single-cell suspensions were analyzed on the Muse® Cell Analyzer according to the manufacturer’s protocols. Briefly, cells were trypsinized (0.05%) and then mixed with Cell Count and Viability reagent or 1:1 NBE to Annexin V and Dead Cell reagent, incubated for the specified amount of time on manufacturer’s protocol, and then inserted into the Muse® Cell Analyzer for data acquisition.
Phase-contrast microscopy
Phase-contrast microscopy was performed using an Olympus CK40 inverted microscope with a Canon EOS 700D camera mounted via an OM-Mount Photomicro Adapter L. Briefly, cells were placed on the microscope stage, allowed to settle, and 2–3 images were taken of different fields of view. Images were captured using the EOS Utility software.
Immunoblotting
Cell samples were washed with 1X DPBS, pelleted via centrifugation, flash-frozen in liquid nitrogen, and finally stored at − 80 °C until cell lysis. Total cell extracts were prepared by resuspending cell pellets in 1 × RIPA buffer containing 1 × protease and phosphatase inhibitor and then subjecting pellets to sonication for 3 × 20 s. Insoluble fractions were pelleted and discarded, and then subsequent protein quantitation was performed on clarified cell lysates via Bradford assay. SDS-PAGE and transfer steps were conducted via the Novex immunoblot system. Blots were visualized using a ProteinSimple Fluorchem® E Imager. Bands on immunoblots were quantified via AlphaView SA v3.5. Quantified immunoblots all experienced the same transfer conditions, primary/secondary antibody concentrations, and exposure times. Antibody information can be found in Supplementary Methods.
Enzyme-linked immunosorbent assay (ELISA)
Detection of intracellular 2-HG levels was performed using the Abcam D-2-Hydroxyglutarate Assay kit (ab211070) according to the manufacturer’s protocol. Briefly, cells were trypsinized (0.05%), pelleted via centrifugation, and flash frozen in liquid nitrogen. Once ready for further analysis, cells were lysed according to manufacturer’s protocol, stably spotted on strip-wells, and then processed for final colorimetric analysis at 450 nm via a Molecular Devices SpectraMax® 340 microplate reader.
Detection of total histone H3 acetylation was performed via Epigentek’s EpiQuik® Global Histone H3 Acetylation Assay kit according to the manufacturer’s protocol. Briefly, histones were acid extracted, stably spotted on strip-wells, and antibodies bound to antigen. Finally, samples were analyzed colorimetrically at 450 nm and quantified via standard curve.
Cell cycle analysis
Cells were analyzed for changes in cell cycle distribution via Muse® Cell Cycle kit according to manufacturer’s protocol. Briefly, cells were trypsinized (0.05%) to form a single cell suspension, washed with 1 × DPBS, and fixed in 70% ethanol at − 20 °C overnight. Fixed cells were then incubated with Muse® Cell Cycle reagent for 30 min at room temp away from light and then injected onto the Muse Cell Analyzer.
BrdU incorporation assays
Cells were analyzed for BrdU incorporation via BD Pharmingen® BrdU Flow kit according to the manufacturer’s protocol. Briefly, cells were treated for 48 h with drug and then pulsed with 10 µM BrdU (except for our No BrdU control) for 2.5 h before washing, fixing, and storing at − 80 °C in solution of 1:9 FBS to DMSO. When ready for analysis, cells were then thawed, permeabilized, and incubated with anti-BrdU antibody according to manufacturer’s protocol. Once stained cell samples and unstained controls were ready for analysis, data was acquired using a Beckman Coulter Cytomics® FC500 flow cytometer with FITC filter applied. Cells were gated using the No BrdU control and then treated samples were assessed for BrdU-positive cells within this gate (Supplementary Figure).
Statistical analyses
Student’s t test and nonlinear regression statistical analyses were performed via GraphPad Prism 8.0 software package. All replicate analyses are presented as mean ± standard deviation of three independent replicates at a significance level (α) of 0.05 unless otherwise indicated in the figure captions.
References
Horbinski C (2013) What do we know about IDH1/2 mutations so far, and how do we use it? Acta Neuropathol 125:621–636. https://doi.org/10.1007/s00401-013-1106-9
Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, Marks KM, Prins RM, Ward PS, Yen KE, Liau LM, Rabinowitz JD, Cantley LC, Thompson CB, Vander Heiden MG, Su SM (2009) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462:739–744
Turcan S, Rohle D, Goenka A, Walsh LA, Fang F, Yilmaz E, Campos C, Fabius AW, Lu C, Ward PS, Thompson CB, Kaufman A, Guryanova O, Levine R, Heguy A, Viale A, Morris LG, Huse JT, Mellinghoff IK, Chan TA (2012) IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483:479–483. https://doi.org/10.1038/nature10866
Xu W, Yang H, Liu Y, Yang Y, Wang P, Kim S-H, Ito S, Yang C, Wang P, Xiao M-T (2011) Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19:17–30
Duncan CG, Barwick BG, Jin G, Rago C, Kapoor-Vazirani P, Powell DR, Chi J-T, Bigner DD, Vertino PM, Yan H (2012) A heterozygous IDH1(R132H/WT) mutation induces genome-wide alterations in DNA methylation. Genome Res 22:2339–2355. https://doi.org/10.1101/gr.132738.111
Brat DJ, Aldape K, Colman H, Holland EC, Louis DN, Jenkins RB, Kleinschmidt-DeMasters BK, Perry A, Reifenberger G, Stupp R, von Deimling A, Weller M (2018) cIMPACT-NOW update 3: recommended diagnostic criteria for “Diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma, WHO grade IV.” Acta Neuropathol 136:805–810
Noushmehr H, Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RG, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H, Neuvial P, Cope LM, Buckley J, Herman JG, Baylin SB, Laird PW, Aldape K (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17:510–522. https://doi.org/10.1016/j.ccr.2010.03.017
Mazor T, Chesnelong C, Pankov A, Jalbert LE, Hong C, Hayes J, Smirnov IV, Marshall R, Souza CF, Shen Y, Viswanath P, Noushmehr H, Ronen SM, Jones SJM, Marra MA, Cairncross JG, Perry A, Nelson SJ, Chang SM, Bollen AW, Molinaro AM, Bengtsson H, Olshen AB, Weiss S, Phillips JJ, Luchman HA, Costello JF (2017) Clonal expansion and epigenetic reprogramming following deletion or amplification of mutant IDH1. Proc Natl Acad Sci USA 114:10743–10748. https://doi.org/10.1073/pnas.1708914114
Moure CJ, Diplas BH, Chen LH, Yang R, Pirozzi CJ, Wang Z, Spasojevic I, Waitkus MS, He Y, Yan H (2019) CRISPR editing of mutant IDH1 R132H induces a CpG methylation-low state in patient-derived glioma models of G-CIMP. Mol Cancer Res 17:2042–2050
Turcan S, Makarov V, Taranda J, Wang Y, Fabius AWM, Wu W, Zheng Y, El-Amine N, Haddock S, Nanjangud G (2018) Mutant-IDH1-dependent chromatin state reprogramming, reversibility, and persistence. Nat Genet 50:62–72
Brehm A, Miska EA, McCance DJ, Reid JL, Bannister AJ, Kouzarides T (1998) Retinoblastoma protein recruits histone deacetylase to repress transcription. Nature 391:597–601
Feng Q, Zhang Y (2001) The MeCP1 complex represses transcription through preferential binding, remodeling, and deacetylating methylated nucleosomes. Genes Dev 15:827–832
Jones PL, Veenstra GJ, Wade PA, Vermaak D, Kass SU, Landsberger N, Strouboulis J, Wolffe AP (1998) Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat Genet 19:187–191
Zhang Y, Ng H-H, Erdjument-Bromage H, Tempst P, Bird A, Reinberg D (1999) Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation. Genes Dev 13:1924–1935
Desai MA, Webb HD, Sinanan LM, Scarsdale JN, Walavalkar NM, Ginder GD, Williams DC Jr (2015) An intrinsically disordered region of methyl-CpG binding domain protein 2 (MBD2) recruits the histone deacetylase core of the NuRD complex. Nucleic Acids Res 43:3100–3113
Basta J, Rauchman M (2015) The nucleosome remodeling and deacetylase complex in development and disease. Transl Res 165:36–47
Zhan X, Guo S, Li Y, Ran H, Huang H, Mi L, Wu J, Wang X, Xiao D, Chen L (2020) Glioma stem-like cells evade interferon suppression through MBD3/NuRD complex-mediated STAT1 downregulation. J Exp Med. https://doi.org/10.1084/jem.20191340
San-Miguel JF, Hungria VT, Yoon SS, Beksac M, Dimopoulos MA, Elghandour A, Jedrzejczak WW, Günther A, Nakorn TN, Siritanaratkul N, Corradini P, Chuncharunee S, Lee JJ, Schlossman RL, Shelekhova T, Yong K, Tan D, Numbenjapon T, Cavenagh JD, Hou J, LeBlanc R, Nahi H, Qiu L, Salwender H, Pulini S, Moreau P, Warzocha K, White D, Bladé J, Chen W, de la Rubia J, Gimsing P, Lonial S, Kaufman JL, Ocio EM, Veskovski L, Sohn SK, Wang MC, Lee JH, Einsele H, Sopala M, Corrado C, Bengoudifa BR, Binlich F, Richardson PG (2014) Panobinostat plus bortezomib and dexamethasone versus placebo plus bortezomib and dexamethasone in patients with relapsed or relapsed and refractory multiple myeloma: a multicentre, randomised, double-blind phase 3 trial. Lancet Oncol 15:1195–1206. https://doi.org/10.1016/s1470-2045(14)70440-1
Zhao Z, Meng F, Wang W, Wang Z, Zhang C, Jiang T (2017) Comprehensive RNA-seq transcriptomic profiling in the malignant progression of gliomas. Sci Data. https://doi.org/10.1038/sdata.2017.24
Tomczak K, Czerwińska P, Wiznerowicz M (2015) The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol 19:A68
AACR Project GENIE Consortium (2017) AACR Project GENIE: powering precision medicine through an International Consortium. Cancer Discov 7:818–831. https://doi.org/10.1158/2159-8290.Cd-17-0151
Pietrak B, Zhao H, Qi H, Quinn C, Gao E, Boyer JG, Concha N, Brown K, Duraiswami C, Wooster R, Sweitzer S, Schwartz B (2011) A tale of two subunits: how the neomorphic R132H IDH1 mutation enhances production of alphaHG. Biochemistry 50:4804–4812. https://doi.org/10.1021/bi200499m
Jin G, Reitman ZJ, Duncan CG, Spasojevic I, Gooden DM, Rasheed BA, Yang R, Lopez GY, He Y, McLendon RE, Bigner DD, Yan H (2013) Disruption of wild-type IDH1 suppresses D-2-hydroxyglutarate production in IDH1-mutated gliomas. Cancer Res 73:496–501. https://doi.org/10.1158/0008-5472.CAN-12-2852
Ward PS, Lu C, Cross JR, Abdel-Wahab O, Levine RL, Schwartz GK, Thompson CB (2013) The potential for isocitrate dehydrogenase mutations to produce 2-hydroxyglutarate depends on allele specificity and subcellular compartmentalization. J Biol Chem 288:3804–3815. https://doi.org/10.1074/jbc.M112.435495
Göttlicher M (2004) Valproic acid: an old drug newly discovered as inhibitor of histone deacetylases. Ann Hematol 83(Suppl 1):S91-92. https://doi.org/10.1007/s00277-004-0850-2
Mellinghoff IK, Ellingson BM, Touat M, Maher E, De La Fuente MI, Holdhoff M, Cote GM, Burris H, Janku F, Young RJ, Huang R, Jiang L, Choe S, Fan B, Yen K, Lu M, Bowden C, Steelman L, Pandya SS, Cloughesy TF, Wen PY (2020) Ivosidenib in isocitrate dehydrogenase 1-mutated advanced glioma. J Clin Oncol 38:3398–3406
Andronesi OC, Loebel F, Bogner W, Marjanska M, Vander Heiden MG, Iafrate AJ, Dietrich J, Batchelor TT, Gerstner ER, Kaelin WG, Chi AS, Rosen BR, Cahill DP (2016) Treatment response assessment in IDH-mutant glioma patients by noninvasive 3D functional spectroscopic mapping of 2-hydroxyglutarate. Clin Cancer Res 22:1632–1641. https://doi.org/10.1158/1078-0432.ccr-15-0656
Platten M, Bunse L, Wick A, Bunse T, Le Cornet L, Harting I, Sahm F, Sanghvi K, Tan CL, Poschke I, Green E, Justesen S, Behrens GA, Breckwoldt MO, Freitag A, Rother LM, Schmitt A, Schnell O, Hense J, Misch M, Krex D, Stevanovic S, Tabatabai G, Steinbach JP, Bendszus M, von Deimling A, Schmitt M, Wick W (2021) A vaccine targeting mutant IDH1 in newly diagnosed glioma. Nature 592:463–468
Sulkowski PL, Corso CD, Robinson ND, Scanlon SE, Purshouse KR, Bai H, Liu Y, Sundaram RK, Hegan DC, Fons NR, Breuer GA, Song Y, Mishra-Gorur K, De Feyter HM, de Graaf RA, Surovtseva YV, Kachman M, Halene S, Gunel M, Glazer PM, Bindra RS (2017) 2-Hydroxyglutarate produced by neomorphic IDH mutations suppresses homologous recombination and induces PARP inhibitor sensitivity. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aal2463
Tateishi K, Wakimoto H, Iafrate AJ, Tanaka S, Loebel F, Lelic N, Wiederschain D, Bedel O, Deng G, Zhang B, He T, Shi X, Gerszten RE, Zhang Y, Yeh JR, Curry WT, Zhao D, Sundaram S, Nigim F, Koerner MV, Ho Q, Fisher DE, Roider EM, Kemeny LV, Samuels Y, Flaherty KT, Batchelor TT, Chi AS, Cahill DP (2015) Extreme vulnerability of IDH1 mutant cancers to NAD+ depletion. Cancer Cell 28:773–784. https://doi.org/10.1016/j.ccell.2015.11.006
Wang Y, Wild AT, Turcan S, Wu WH, Sigel C, Klimstra DS, Ma X, Gong Y, Holland EC, Huse JT, Chan TA (2020) Targeting therapeutic vulnerabilities with PARP inhibition and radiation in IDH-mutant gliomas and cholangiocarcinomas. Sci Adv. https://doi.org/10.1126/sciadv.aaz3221
Nguyen TTT, Zhang Y, Shang E, Shu C, Torrini C, Zhao J, Bianchetti E, Mela A, Humala N, Mahajan A, Harmanci AO, Lei Z, Maienschein-Cline M, Quinzii CM, Westhoff MA, Karpel-Massler G, Bruce JN, Canoll P, Siegelin MD (2020) HDAC inhibitors elicit metabolic reprogramming by targeting super-enhancers in glioblastoma models. J Clin Invest 130:3699–3716
Radoul M, Najac C, Viswanath P, Mukherjee J, Kelly M, Gillespie AM, Chaumeil MM, Eriksson P, Delos Santos R, Pieper RO, Ronen SM (2019) HDAC inhibition in glioblastoma monitored by hyperpolarized (13) C MRSI. NMR Biomed. https://doi.org/10.1002/nbm.4044
Grasso CS, Tang Y, Truffaux N, Berlow NE, Liu L, Debily MA, Quist MJ, Davis LE, Huang EC, Woo PJ, Ponnuswami A, Chen S, Johung TB, Sun W, Kogiso M, Du Y, Qi L, Huang Y, Hütt-Cabezas M, Warren KE, Le Dret L, Meltzer PS, Mao H, Quezado M, van Vuurden DG, Abraham J, Fouladi M, Svalina MN, Wang N, Hawkins C, Nazarian J, Alonso MM, Raabe EH, Hulleman E, Spellman PT, Li XN, Keller C, Pal R, Grill J, Monje M (2015) Functionally defined therapeutic targets in diffuse intrinsic pontine glioma. Nat Med 21:555–559
Hennika T, Hu G, Olaciregui NG, Barton KL, Ehteda A, Chitranjan A, Chang C, Gifford AJ, Tsoli M, Ziegler DS, Carcaboso AM, Becher OJ (2017) Pre-clinical study of panobinostat in xenograft and genetically engineered murine diffuse intrinsic pontine glioma models. PLoS ONE. https://doi.org/10.1371/journal.pone.0169485
Puduvalli VK, Wu J, Yuan Y, Armstrong TS, Vera E, Wu J, Xu J, Giglio P, Colman H, Walbert T, Raizer J, Groves MD, Tran D, Iwamoto F, Avgeropoulos N, Paleologos N, Fink K, Peereboom D, Chamberlain M, Merrell R, Penas Prado M, Yung WKA, Gilbert MR (2020) A Bayesian adaptive randomized phase II multicenter trial of bevacizumab with or without vorinostat in adults with recurrent glioblastoma. Neuro Oncol 22:1505–1515
Galanis E, Anderson SK, Miller CR, Sarkaria JN, Jaeckle K, Buckner JC, Ligon KL, Ballman KV, Moore DF Jr, Nebozhyn M, Loboda A, Schiff D, Ahluwalia MS, Lee EQ, Gerstner ER, Lesser GJ, Prados M, Grossman SA, Cerhan J, Giannini C, Wen PY (2018) Phase I/II trial of vorinostat combined with temozolomide and radiation therapy for newly diagnosed glioblastoma: results of Alliance N0874/ABTC 02. Neuro Oncol 20:546–556
Friday BB, Anderson SK, Buckner J, Yu C, Giannini C, Geoffroy F, Schwerkoske J, Mazurczak M, Gross H, Pajon E, Jaeckle K, Galanis E (2012) Phase II trial of vorinostat in combination with bortezomib in recurrent glioblastoma: a north central cancer treatment group study. Neuro Oncol 14:215–221
Lee EQ, Reardon DA, Schiff D, Drappatz J, Muzikansky A, Grimm SA, Norden AD, Nayak L, Beroukhim R, Rinne ML, Chi AS, Batchelor TT, Hempfling K, McCluskey C, Smith KH, Gaffey SC, Wrigley B, Ligon KL, Raizer JJ, Wen PY (2015) Phase II study of panobinostat in combination with bevacizumab for recurrent glioblastoma and anaplastic glioma. Neuro Oncol 17:862–867
Kim GH, Choi SY, Oh TI, Kan SY, Kang H, Lee S, Oh T, Ko HM, Lim JH (2019) IDH1(R132H) causes resistance to HDAC inhibitors by increasing NANOG in glioblastoma cells. Int J Mol Sci. https://doi.org/10.3390/ijms20112679
Su JM, Murray JC, McNall-Knapp RY, Bowers DC, Shah S, Adesina AM, Paulino AC, Jo E, Mo Q, Baxter PA, Blaney SM (2020) A phase 2 study of valproic acid and radiation, followed by maintenance valproic acid and bevacizumab in children with newly diagnosed diffuse intrinsic pontine glioma or high-grade glioma. Pediatr Blood Cancer. https://doi.org/10.1002/pbc.28283
Weller M, Gorlia T, Cairncross JG, van den Bent MJ, Mason W, Belanger K, Brandes AA, Bogdahn U, Macdonald DR, Forsyth P, Rossetti AO, Lacombe D, Mirimanoff RO, Vecht CJ, Stupp R (2011) Prolonged survival with valproic acid use in the EORTC/NCIC temozolomide trial for glioblastoma. Neurology 77:1156–1164
Krauze AV, Myrehaug SD, Chang MG, Holdford DJ, Smith S, Shih J, Tofilon PJ, Fine HA, Camphausen K (2015) A phase 2 study of concurrent radiation therapy, temozolomide, and the histone deacetylase inhibitor valproic acid for patients with glioblastoma. Int J Radiat Oncol Biol Phys 92:986–992
Chen H, Judkins J, Thomas C, Wu M, Khoury L, Benjamin CG, Pacione D, Golfinos JG, Kumthekar P, Ghamsari F, Chen L, Lein P, Chetkovich DM, Snuderl M, Horbinski C (2017) Mutant IDH1 and seizures in patients with glioma. Neurology 88:1805–1813
Clarke DJB, Jeon M, Stein DJ, Moiseyev N, Kropiwnicki E, Dai C, Xie Z, Wojciechowicz ML, Litz S, Hom J, Evangelista JE, Goldman L, Zhang S, Yoon C, Ahamed T, Bhuiyan S, Cheng M, Karam J, Jagodnik KM, Shu I, Lachmann A, Ayling S, Jenkins SL, Ma’ayan A (2021) Appyters: Turning Jupyter Notebooks into data-driven web apps. Patterns. https://doi.org/10.1016/j.patter.2021.100213
Goldman M, Craft B, Swatloski T, Cline M, Morozova O, Diekhans M, Haussler D, Zhu J (2015) The UCSC cancer genomics browser: update 2015. Nucl Acids Res 43:D812–D817
Acknowledgements
This research was supported by the UC Davis Floyd and Mary Schwall Dissertation Fellowship, the UC Davis Murray B. Gardner Junior Faculty Research Fellowship, and the Elsa U. Pardee Foundation Research Grant.
Funding
This study was funded by the UC Davis Schwall Dissertation Fellowship and UC Davis La Pittus Fellowship. CH was supported by NIH grants R01NS118039, R01NS117104, R01NS102669, and P50CA221747.
Author information
Authors and Affiliations
Contributions
Conceptualization: TS, KW; methodology: TS, CH, KW; formal analysis and investigation: TS; writing—original draft preparation: TS; writing—review and editing: TS, CH, KW; funding acquisition: TS, KW; resources: TS, KW; supervision: KW.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11060_2021_3829_MOESM1_ESM.tif
Supplementary file 1 Fig. S1 Gene expression of HDAC-related genes identified as significantly upregulated in IDH1/2mut glioma based on GO analysis. Statistical analyses were performed via Welch’s T-test. a, b Expression of GO-identified genes that promote HDAC function in Grade 2/3 IDH1/2mut glioma using samples from TCGA and CGGA datasets. c, d Expression of GO-identified genes that promote HDAC function in Grade 4 IDH1/2mut glioma using samples from TCGA and CGGA datasets (TIF 17402 kb)
11060_2021_3829_MOESM3_ESM.tif
Supplementary file 3 Fig. S2 Glioma cell characterization. a Table showing known genetic alterations in glioma cells utilized in this study. Studies with analysis of genetic alterations are cited in the table. NIF = no information found. b Western blot of R132H IDH1 in whole cell lysates extracted from IDH1/2wt (0827, 0923, 0211) and IDH1mut (0905, BT142, TB096) glioma cells grown in NBE medium. c 2-HG levels assessed by ELISA in IDH1/2wt and IDH1mut glioma cells used in this study (TIF 24802 kb)
11060_2021_3829_MOESM4_ESM.tif
Supplementary file 4 Fig. S3 Cell viability dose-response assay with puromycin in IDH1/2wt and IDH1mut glioma cells. a Normalized cell viability of IDH1/2wt and IDH1mut glioma cells treated with puromycin for 3 days (n = 3). Nonlinear regression statistical analysis was utilized to determine IC50 values. b Consolidation of cell viability IC50 values derived from a stratified based on IDH status (n = 6). Statistical analysis was performed via Student’s t-test. c Representative Muse® cell analyzer flow cytometry plots from a using the Muse® Cell Count and Viability Kit. Y-axis represents uptake of a membrane permeable DNA dye by all cells, whereas the x-axis represents uptake of a membrane impermeable DNA dye e.g. 7-AAD in dead cells that have lost membrane integrity (TIF 44121 kb)
11060_2021_3829_MOESM5_ESM.tif
Supplementary file 5 Fig. S4 Representative Muse® cell analyzer flow cytometry plots from Fig. 2a, b using the Muse® Cell Count and Viability Kit. Y-axis represents uptake of a membrane permeable DNA dye by all cells, whereas the x-axis represents uptake of a membrane impermeable DNA dye e.g. 7-AAD in dead cells that have lost membrane integrity (TIF 44121 kb)
11060_2021_3829_MOESM6_ESM.tif
Supplementary file 6 Fig. S5 Representative Muse® cell analyzer flow cytometry plots from Fig. 2c, d using the Muse® Annexin V and Dead Cell Kit. Y-axis represents uptake of a membrane impermeable DNA dye e.g. 7-AAD in dead cells that have lost membrane integrity, whereas the x-axis represents binding of Annexin V to phosphatidyl-serine residues (TIF 19243 kb)
11060_2021_3829_MOESM7_ESM.tif
Supplementary file 7 Fig. S6 Flow cytometry plots from Fig. 3a, b BrdU incorporation assays. Plots were generated using the FCSalyzer v0.9.17 software program. BC = Background Control, Veh = Vehicle Control. a Representative bivariate forward- and side-scatter plots showing the gating method for our WT and mutant glioma cells. b Representative univariate plots (FITC) showing BrdU uptake in our IDH1/2wt and IDH1mut glioma cells. Included background stain controls did not receive BrdU treatment but were still stained with anti-BrdU antibody (TIF 44121 kb)
11060_2021_3829_MOESM9_ESM.tif
Supplementary file 9 Fig. S7 Representative Muse® cell analyzer flow cytometry plots from Fig. 3c using the Muse® Cell Cycle Kit. DNA content on the x-axis is used as an indicator for cell cycle phase (TIF 27083 kb)
11060_2021_3829_MOESM10_ESM.tif
Supplementary file 10 Fig. S8 Validation of linearity with our antibodies used for quantitative immunoblot analysis. H3K27Ac shows issues with linearity, but the H3K27Ac signal from our quantitative experiments does not extend past the linear range. Linearity was determined using linear regression analysis via Graphpad Prism 8.0.0 (TIF 18111 kb)
11060_2021_3829_MOESM11_ESM.tif
Supplementary file 11 Fig. S9 Representative Muse® cell analyzer flow cytometry plots from Fig. 5a, b using the Muse® Cell Count and Viability Kit. Y-axis represents uptake of a membrane permeable DNA dye by all cells, whereas the x-axis represents uptake of a membrane impermeable DNA dye e.g. 7-AAD in dead cells that have lost membrane integrity (TIF 59441 kb)
11060_2021_3829_MOESM12_ESM.png
Supplementary file 12 Table S1 Table showing known genetic and molecular alterations in the glioma cells used in this study with references to sources in the literature (PNG 105 kb)
11060_2021_3829_MOESM13_ESM.png
Supplementary file 13 Table S2 Table showing IC50 and EC50 values from our cell viability and apoptosis studies in Fig. 2 (PNG 76 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sears, T.K., Horbinski, C.M. & Woolard, K.D. IDH1 mutant glioma is preferentially sensitive to the HDAC inhibitor panobinostat. J Neurooncol 154, 159–170 (2021). https://doi.org/10.1007/s11060-021-03829-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-021-03829-0