Abstract
Pseudogymnoascus destructans (= Geomyces destructans) is a psychrophilic filamentous fungus that causes White-Nose Disease (WND; the disease associated with White-Nose Syndrome, WNS) in hibernating bats. The disease has caused considerable reductions in bat populations in the USA and Canada since 2006. Identification and detection of the pathogen in pure cultures and environmental samples is routinely based on qPCR or PCR after DNA isolation and purification. Rapid and specific direct detection of the fungus in the field would strongly improve prompt surveillance, and support control measures. Based on the genes coding for ATP citrate lyase1 (acl1) and the 28S-18S ribosomal RNA intergenic spacer (IGS) in P. destructans, two independent LAMP assays were developed for the rapid and sensitive diagnosis of the fungus. Both assays could discriminate P. destructans from 159 tested species of filamentous fungi and yeasts. Sensitivity of the assays was 2.1 picogram per reaction (pg/rxn) and 21 femtogram per reaction (fg/rxn) for the acl1 and IGS based assays, respectively. Moreover, both assays also work with spores and mycelia of P. destructans that are directly added to the master mix without prior DNA extraction. For field-diagnostics, we developed and tested a field-applicable version of the IGS-based LAMP assay. Lastly, we also developed a protocol for preparation of fungal spores and mycelia from swabs and tape liftings of contaminated surfaces or infected bats. This protocol in combination with the highly sensitive IGS-based LAMP-assay enabled sensitive detection of P. destructans from various sources.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During February 2006, the first evidence of North-American hibernating bats displaying visible signs associated with white-nose disease (WND) was documented in a cave near Albany, NY [1]. During subsequent years, the disease spread from New York to adjacent U.S. states and is now found in 38 U.S. states and seven Canadian provinces (see https://www.whitenosesyndrome.org/ for a current map) and recent studies showed that several bat species in North America show a population level response with a significant decrease in presence and abundance post-WND [2,3,4]. The causative agent of WND has been identified as the psychrophilic fungus Pseudogymnoascus destructans Minnis & D.L. Lindner (= Geomyces destructans Blehert & Gargas) [1, 5, 6]. Aside from North America, the fungus is prevalent in the Western Palearctic and Palearctic Asia [7,8,9,10,11,12]. However, in the Palearctic, observed mortality in bat colonies with confirmed P. destructans presence was similar to colonies from which P. destructans was not reported [11, 13]. The absence of mass mortality in the Palearctic combined with phylogenetic studies of European, North American and Asian P. destructans populations suggest that the fungus was introduced to North America from Europe rather than from Asia, even though the definitive source of the invading isolate has not yet been conclusively identified [14, 15].
For diagnostics, the detection and identification of P. destructans and its distinction from closely related species largely relies on molecular techniques such as quantitative PCR (qPCR). Single- or multi locus sequencing [6, 16] or microsatellite analysis [17,18,19,20,21] have also been used in research studies. Molecular techniques use genomic DNA that has to be isolated and purified from pure culture mycelia or sample materials before the analysis [22, 23]. However, due to its slow growth (8 mm in 16 d at 14 °C), cultivation of P. destructans is time consuming [5]. To aid diagnosis of the fungus, diagnostic PCR and qPCR assays were established for several genes: the 18S region of the rRNA gene [24, 25], the gene coding for alpha-L-rhamnosidase (ALR) [26], the rRNA intergenic spacer (IGS) region [27] and the rRNA internal transcribed spacer (ITS) region [28]. However, DNA also needs to be extracted and purified before running these assays.
Loop-mediated isothermal amplification (LAMP) [29] is an alternative technology for DNA-based diagnosis of microorganisms that runs under isothermal conditions at 65 °C. The specificity of the LAMP reaction is higher compared to qPCR because a set of four primers with six binding sites must hybridize correctly to their target sequence before autocycling DNA biosynthesis occurs. Amplification events can be detected visually by in-tube colour reaction of indicator dyes such as calcein in a fluorescence based assay [30] or neutral red in systems used under day light conditions [31]. Over the past 15 years, LAMP-based assays were established for a number of yeast-like and filamentous fungal species. Assays can be used to identify pure fungal cultures or to detect the target organisms in other sample types [32, 33]. Together with its lower sensitivity to typical polymerase inhibitors [34], the LAMP technology provides a versatile technology for rapid, user-friendly and inexpensive diagnostic assays that can be used in simple lab settings or even under field conditions. Such features make LAMP assays an ideal tool for the study of P. destructans.
We report the development and application of LAMP based assays for the rapid identification of P. destructans in pure cultures and other sample types as well as the development of innovative, rapid and simple protocols for the preparation of fungal material for testing with LAMP-based methods in the field.
Materials and Methods
Organisms and Culture Conditions
A total of 217 fungal cultures have been used during the current study (complete list in Table S1 of supplemental materials). The cultures originated from two sources. The large majority of non-Pseudogymnoascus sp. cultures were obtained from various official culture repositories or from collections of fungal cultures maintained at public research institutions (see column 3 and footnotes in Table S1, strains marked in green) while the majority of Pseudogymnoascus sp. cultures came from the authors reference collection (see further details in Table S1).The 53 cultures of P. destructans that were used in the current study were isolated from bat swabs, bat guano or bat hibernacula environments (sediments/walls) from various locations in Europe and North America [8, 15, 20, 35]. The self-isolated fungal strains shown in table S1 (marked in grey) were identified to species/clades level using micromorphology, FTIR spectroscopy or genetic methods (LAMP assay [36, 37], gene sequencing, e.g. 28S rRNA, ITS, beta tubulin, calmodulin, or multi locus sequencing [15]). Part of the pure cultures of P. destructans were identified using microsatellite analysis as described in [18] based on published markers from [17]. For analysis of fungi in bat guano from nine German roost sites (6 in Hesse and 3 in Baden Wuerttemberg, 2010 and 2011) as well as two samples of soil from Czech bat caves, bat guano pellets and soil particles were placed on Sabouraud´s dextrose agar (SDA, 0.5% (w/v) peptone from casein, 0.5% (w/v) meat peptone, 2% dextrose, 1.2% (w/v) agar) and incubated at 13 °C in the dark. Colonies were transferred to fresh SDA plates for pure culturing. For identification of strains, colonies that resembled Pseudogymnoascus spp. were maintained on SDA. Colonies of other species were re-inoculated to malt extract agar (MEA, 3% w/v malt extract (ApliChem, Darmstadt, Germany), 0.3% w/v soy peptone [Karl Roth, Karlsruhe, Germany), 2% w/v agar, pH 5.8) prior to microscopic identification using micromorphological characteristics and appropriate identification keys [38,39,40]. Cultures of environmental fungi that could not be readily identified microscopically were identified according to signature sequences, e.g. beta tubulin gene for Penicillium spp., calmodulin gene for Aspergillus spp., transcription elongation factor 1 alpha for Fusarium spp. and rRNA internal transcribed spacer (ITS) for all other genera.
For long-term storage, cultures were maintained as cryo-cultures at -80 °C in 80% glycerol as described previously [41]. Working cultures of non-P. destructans isolates were obtained from cryo-cultures by inoculation to MEA and incubation at ambient temperature under diffuse day light. Isolates of P. destructans and closely related taxa were cultivated on SDA at 13 °C in the dark. All Fusarium spp. were grown on Synthetic Nutrient Agar (SNA, [42]) amended with 0.4 g of glucose at ambient temperature under diffuse day light.
Twenty-one field samples were collected from bats (n = 10; 8 swabs and 2 tape liftings) and bat hibernacula walls (n = 11 swabs) towards the end of the hibernation season in March/April [20]. Sampled bats were found either with (n = 7) or without (n = 3) visual fungal growth indicating P. destructans infection [66]. Swabs were stored in 1.5 mL tubes and tape liftings were stored on microscope slides, at ~ 5 °C until usage.
DNA Preparation from Pure Cultures
PCR grade genomic DNA was prepared from 30 mL liquid cultures of the 217 isolates grown under conditions that were optimum for the respective species. Mycelia were vacuum filtered and washed twice with sterile tap water. Fifty milligram of mycelia were treated for 1 min in a Fastprep® 24 system with addition of sterile glass beads (0.4 g of 0.5 mm diameter and 0.22 g of 1.25–1.65 mm diameter) at an amplitude of 6 m/s in lysis buffer PL2 provided with the peqGold Fungal DNA Mini Kit (VWR International, Erlangen, Germany). DNA purification from treated mycelia was performed according to the manufacturer´s protocol. DNA was maintained frozen at − 20 °C until used and further stored at 4 °C after initial thawing.
DNA Sequencing
A 766 bp partial sequence of the multicopy 28S-18S ribosomal RNA intergenic spacer (IGS) of P. destructans isolate 01NH07 (GenBank accession number JX270192; [43]) was downloaded from GenBank for primer design. To design an alternative primer set, from a single copy gene, the coding sequence of the P. destructans ATP citrate lyase subunit 1 gene (acl1) was retrieved from GenBank (isolate 20631-21, GenBank accession number XM_012884837). Since the P. destructans type 20631-21 was unavailable to our study, we used isolate P. destructans OT-38-2010 (isolated from Myotis mystacinus, Finnentrop, Germany; culture ‘Mmys-DE-2’ in [11]) as reference throughout the study. Presence and correctness of the acl1 sequence in the reference isolate P. destructans OT-38-2010 was checked by sequencing three overlapping portions of the gene using primers Geo-acl1-f to Geo-acl1-r2 (see Table 1 for primer sequences. See method descriptions in supplemental materials for a detailed description of acl1 sequencing).
Identity of LAMP products was verified by sequencing of the smallest amplified fragments. LAMP reactions were performed with primer sets IGS-ID10 and acl1-ID30, respectively and separated on 1.5% agarose gels. The smallest DNA fragment was cut from each of the two gels and purified using the QIAquick Gel Extraction Kit (QIAGEN, Hilden, Germany) according to the manufacturer's recommendations. Each of the purified LAMP fragments was re-amplified by PCR using primers F2-Pd-acl1/B2-Pd-acl1 and F2-Pd-IGS/B2-Pd-IGS, respectively (see Table 1) according to the following cycling protocol: 1 cycle 5 min initial melting at 95 °C, 40 cycles consisting of 1 min initial melting at 95 °C, 1 min annealing at 54 °C (primers Pd-IGS) or 58 °C (primers Pd-acl1), 1 min elongation at 72 °C, and 1 cycle of 5 min final elongation at 72 °C. Amplification products were purified using the QIAquick PCR Purification Kit (QIAGEN, Hilden, Germany) and sent off for bi-directional Sanger sequencing using PCR primers as sequencing primers. Consensus sequences were created and aligned using the BioEdit Software package [44]. The resulting consensus sequence was aligned to the acl1 and IGS gene sequences of P. destructans 20631-21, respectively. For verification, BLASTn [45] was used to search the NCBI nucleotide database for homologies with the consensus sequences from each of the two target genes (IGS and acl1).
Gene copy numbers were calculated from gDNA concentrations using the calculator available at the home page of the University of Rhode Island CELS Genomics and Sequencing Center (http://cels.uri.edu/gsc/cndna.html).
LAMP Primer Design
Primers for the LAMP acl1-specific assay were designed manually while those for the IGS-specific assay were designed using the PrimerExplorer software available on the Eiken GENOME website (see http://primerexplorer.jp/e/) (see Table 1 for primer sequences). Primer sets were selected to have a dimer(minimum)dG-value close to − 2.0, respectively. Default settings were applied for all other parameters adjustable in the primer design software. In silico analysis of primer specificity was done by testing each primer in a BLASTn [45] search against the NCBI sequence database. Target gene sequences of several Pseudogymnoascus spp. and from other fungal species for which > 90% similarities were found in at least three of the tested primers were extracted from the respective genome assemblies. Each primer was aligned with a multiple alignment (CustallW, BioEdit v7.2.5 [44]) of the respective target gene in the selected species for specificity analysis.
Vacuum Drying of LAMP Master Mix (‘dry-LAMP’)
A master mix consisting, per reaction, of 8 mM MgCl2, 1.4 mM each deoxynucleotide (dG, dA, dT, dC), 1.6 µM primers FIP/BIP, 0.8 µM primers LF/LB, 0.2 µM primers F3/B3, 0.1 mM neutral red, 8 U Bst 2.0 Warmstart DNA-polymerase (New England Biolabs, Frankfurt/M., Germany) and 15.9 µL sterile demineralized water was prepared and 25 µL per reaction were distributed into 200 µL PCR tubes (8-Strip PCR Tube, individually attached flat cap, optically clear, Star Lab, Hamburg, Germany). The master mixes were vacuum dried in a desiccator with the tube lids opened at ambient temperature (22 °C) and 3 mbar pressure for 60 min. For quality control, two LAMP reactions were tested directly after drying by adding 25 µL of P. destructans OT-38-2010 gDNA (2.4 ng/µL) in 1 × amplification buffer (10 mM (NH4)2SO4, 10 mM KCI, pH 8.7) as positive control and 25 µL 1 × amplification buffer without gDNA as negative control. Dried LAMP reactions (‘dry-LAMP’) were sealed in clear plastic bags together with a silica desiccant pack and stored at 4 °C in the dark until use.
Specificity Testing of LAMP Assays
LAMP assays were run under the conditions shown in Table 2 using calcein as fluorescence indicator and purified genomic DNA of the fungal isolates listed in Table S1 as template. Five microliters of DNA at 20 ng/µL was added to the LAMP reactions to ensure that even in case of a weak cross-reaction, a LAMP signal would occur. Purified genomic DNA of 217 fungal isolates representing 159 different species, subspecies and varieties from 33 genera of filamentous fungi and yeasts was tested for cross reactivity with both LAMP assays. Among others, DNA of 53 isolates of P. destructans originating from 12 countries were tested as reference material.
Preparation of Tape Lifting Samples
The tape lifting technique was developed based on a previously described touch imprint method used to obtain samples from P. destructans infected bats for microscopic and PCR analysis [7]. We used two types of adhesive tape, a common household/office tape (tesafilm®, BNR 57284, tesa SE, Norderstedt, Germany), similar to the tape that has been used so far [7] and Water Soluble Wave Solder tape (Scotch™ Brand Mask Plus II 1″ × 36 yrd no. 5414, 3 M Industrial Tape Division, St. Paul, MN, USA). For both materials, strips of approximately 4 cm were cut from a roll and pressed to a sampling surface with its adhesive side. After removal from the sampling surface, the tape was either adhered to a microscope glass slide or folded with the adhesive sides of both ends touching and transferred to a sterile 2.0 mL round bottom reaction vessel after two more foldings (only for the water soluble tape).
For the tesafilm® tape lifting samples (field experiment), small pieces (~ 1 to 2 cm2) containing lifted sample material were directly transferred into 300 µL LAMP buffer and 25 µL of this template-buffer-solution was used in the dry-LAMP reaction (see field experiment section).
Since tesafilm® can very easily stick to the tube and thus LAMP buffer might not always reach the tape areas with spores, we developed an alternative method using water soluble tape. The water soluble tape was solubilized by addition of 2 mL sterile 8 M aqueous urea (for details see method descriptions in supplemental materials). Five microliters of the resulting solution were used as template in the LAMP reaction.
LAMP-Based Analysis of Pure Culture Material
We evaluated the potential of the IGS-based LAMP assay to detect P. destructans after direct addition of fungal material into the master mix. Minute amounts of P. destructans pure culture material including mycelium and spores were removed from SDA grown colonies with either a heat sterilized steel needle or a sterile swab from which spores were later obtained by centrifugation in demineralized water. Cell material (dry or in solution) was transferred directly to the LAMP master mix prior to incubation at 65 °C for 60 min. For testing the influence of culture age on direct detection of fungal material, samples were collected from colonies daily, starting at day three until day 31 of cultivation at 13 °C on SDA. Since colonies started developing guttation droplets from day 23 onward, these colonies were sampled both from the margin and centre and samples were tested in separate LAMP reactions.
Isothermal DNA Amplification
Loop-mediated isothermal amplification (LAMP) of P. destructans DNA was performed using essentially the protocol described by Tomita, Mori [30] with slight modifications. The master mix was prepared containing per 25 µL reaction: 2.5 µl 10 × LAMP buffer (200 mM 3-(N-morpholino)propanesulfonic acid (MOPS), 100 mM KCl, 100 mM (NH4)2SO4, 80 mM MgCl2, pH 8.8, all chemicals from Sigma-Aldrich (Taufkirchen, Germany) and Carl Roth (Karlsruhe,Germany), 3.5 µL dNTP mix (10 mM each, MP Biomedicals, Heidelberg, Germany), 2.6 µL primer mix (resulting in final concentrations of 1.6 µM each FIP and BIP, 0.8 µM each LF and LB, 0.2 µM each F3 and B3 in the master mix, see Table 1 for sequences, all primers by Eurofins, Ebersberg, Germany), 8 U (1.0 µl) Bst DNA polymerase, large fragment (8000 U/mL, New England Biolabs, Frankfurt, Germany), 1 µL calcein reagent (see Niessen and Vogel [46]), or V13-01184 dye (10 µg/mL working solution (Dyomics, Jena, Germany] [47], or neutral red reagent [31, 48]. Demineralized water was added to result in a 25 µL total reaction volume, including sample DNA. Regularly, 5 µL of template DNA solution or spore suspension were added per reaction. Visual detection under day light conditions was achieved using neutral red as indicator dye. In this case the 10 × LAMP buffer was composed of 100 mM (NH4)2SO4, 100 mM KCI, pH 8.7. Reaction components used for the different LAMP assays are listed in Table 2. Twenty microliters of the master mix per sample were distributed into 200 µL PCR tubes (8-Strip PCR Tube, individually attached flat cap, optically clear, Star Lab, Hamburg, Germany) and 5 µL sample DNA was added. DNA of P. destructans OT-38-2010 (1.0 ng/reaction) was used as positive control in both assays throughout the study. Water was added instead of DNA in no template controls (NTC). Sterile filter pipette tips were used for all liquid handling (Star Lab, Hamburg, Germany). Two different sets of pipettes were used for DNA-free operations and operations involving DNA, respectively. Preparation of master mixes and addition of sample DNA were performed in separate rooms. Agarose gel electrophoresis of LAMP products was performed in a room separate from master mix preparation and template addition. Reaction tubes were placed in the heating block of a Mastercycler® gradient thermal cycler (Eppendorf, Hamburg, Germany) operated at constant temperature according to the optimized protocol for respective LAMP reactions (Table 2). For inspection of calcein-based assays, reaction tubes were placed upon black photo cardboard under a 365 nm UV lamp (MinUVIS, Desaga, Heidelberg, Germany) and inspected for the presence of a bright green fluorescence. Neutral-red based assays were documented by placing tubes on a white-light transilluminator. Results were documented using a hand held digital camera.
Detection of P. destructans in Field Swab and Tape Samples Using dry-LAMP
We used dry-LAMP reactions to test two different field sample types: (1) nylon swabs (CLASSIQSwabs, COPAN, Brescia, Italy) collected from bats and hibernacula walls and (2) tesafilm® (tape liftings) collected from bats. One negative control was included every 7th sample. Swabs and pieces of the tesafilm® (~ 1 to 2 cm2) were transferred directly to 300 µL LAMP buffer in 1.5 mL tubes and vigorously shaken by hand for 1 min and 25 µL of the solution was transferred to the dry-LAMP tubes. After brief manual shaking until the dry-LAMP pellet was dissolved, tubes were incubated at 65 °C for 60 min using a Heating Thermomixer (HLC BioTech, Bovenden, Germany). All dry-LAMP procedures and reactions were run outside a laboratory on an office desk (to mimic field conditions). Surfaces and materials (pipettes, scalpel blades, forceps) were cleaned and disinfected with 70% aqueous 2-propanol solution.
Results
Sequence Determination of the acl1 Gene in P. destructans
BLASTn [45] was used to search the NCBI nucleotide database for the acl1 gene in the genomic sequence of P. destructans strain 20631-21. The search retrieved a 2067 nt sequence between positions 1,116,082 and 1,118,148 on the unplaced genomic scaffold_9 of the P. destructans whole genome shotgun sequence (locus_tag = VC83_04411) [49]. The coding sequence included a start and stop codon with one intron of 57 nt between nucleotide positions 481 and 537. The translated protein sequence (Expasy translate tool http://web.expasy.org/translate/) consisted of 667 amino acids. SignalP 3.0 Server analysis [50] predicted no signal peptide providing evidence that the protein is not secreted. Sequencing of the acl1 gene of the reference isolate OT-38-2010 (GenBank Accession Number JN242242) confirmed the presence of the acl1 gene. Both sequences showed 99.9% homology. Two nucleotide exchanges were found at positions 527 (A–T) and 540 (T–C), i.e. in the intron sequence of the gene and will therefore not alter the amino acid sequence of the resulting protein in comparison to the type strain.
Primer Design and LAMP Product Verification
Two sets of LAMP primers, including a pair of loop primers for each gene, were designed on the basis of the acl1 gene and IGS sequences (Table 1 and Fig. 1). Similarity analysis of acl1 and IGS sequences for each primer-binding site in P. destructans as well as other Pseudogymnoascus spp. and other fungal species that showed high in silico sequence similarities for some of the primers is shown in Figs. S1 and S2, respectively. The binding sites of all Pd-acl1-ID30 primers were in 100% homology with the corresponding sequences of two P. destructans strains for which acl1 sequences were available (see Fig. S1). Moreover, the binding sites of three out of eight Pd-acl1-ID30 primers shared 100% identity with the respective homologues sequences of eight Pseudogymnoascus spp. of yet unidentified taxonomic association to P. destructans. Moreover, four out of eight primers showed only 1–2 mismatches. Only primer B3-Pd-acl1-ID30 shared a higher degree of sequence dissimilarity with the analysed Pseudogymnoascus spp. isolates. Non-Pseudogymnoascus acl1 sequences showed between 0 and 10 mismatches with the Pd-acl1-ID30 primer sequences. No acl1 sequences were available for P. pannorum and P. verrucosus. Figure S2 shows that all IGS-specific primers shared 100% homology with the respective binding sequences in two P. destructans strains but showed a high percentage of mismatches with 10 Pseudogymnoascus spp. isolates as well as P. verrucosus. No IGS sequence was available from the genome of P. pannorum.
Localization of LAMP primers within their target sequences. Binding sites of LAMP primer sets for specific detection of P. destructans are marked in grey. A Partial sequence of the acl1 gene (nucleotide positions 841-1130 of 2067, Genbank accession number JN242242) in P. destructans with the smallest LAMP fragment predicted to be 139 nt. B Partial sequence of the 18S-28S ribosomal RNA intergenic spacer IGS (nucleotide positions 361-659 of 766, partial sequence of Genbank accession number GQ489025) in P. destructans with the smallest LAMP fragment predicted to be 175 nt → indicates forward primers, ← indicates backward primers
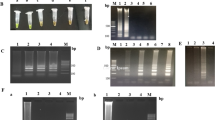
Presence of a ladder-like banding pattern that typically occurs in LAMP reactions was verified by agarose gel electrophoresis (AGE) of reactions run with a serial dilution of template DNA from P. destructans OT-38-2010. The ladder-like pattern of DNA fragments of sizes between 150 bp and several Kbp occurred with primers specific for the acl1 and IGS sequence, respectively (Fig. 2). Comparison of banding patterns and LAMP reaction tubes made it obvious that DNA was synthesized in all reactions where the calcein indicator showed a bright green fluorescence (Fig. 2A, B). The same concordance between DNA biosynthesis and visual signal detection under day light conditions was found when neutral red was used as indicator (Fig. 2C).
Confirmation of DNA biosynthesis in LAMP reactions and correlation with calcein fluorescence and neutral red colour change. The LAMP reactions shown in Figure S3 (A, B, C) were separated on 1.5% agarose gels and stained with dimidium bromide. LAMP reactions were electronically mounted onto respective agarose gels for comparison between the presence of DNA and calcein fluorescence/neutral red colour change. M1 = Gene ruler 100 bp DNA ladder; 1 = 213 ng per reaction (ng/rxn) of P. destructans OT-38–2010 gDNA; 2 = 21.3 ng/rxn; 3 = 2.1 ng/rxn; 4 = 210 pg/rxn; 5 = 21 pg/rxn; 6 = 2.1 pg/rxn; 7 = 210 fg/rxn; 8 = 21 fg/rxn; 9 = 2.1 fg/rxn; 10 = 210 ag/rxn; NTC = not template control, demineralized water instead of DNA-template figureM2 = Gene ruler 1 kb DNA ladder
Product verification was performed by comparing the nucleotide sequence of the smallest LAMP fragment with the respective template sequences. The partial sequences obtained for both LAMP products were 98.6% and 95.6% identical with the target sequences in acl1 and IGS, respectively (Fig. 3). Although sequence identity was not 100%, as expected when using polymerases with low fidelity and comparing different isolates, results nevertheless suggest that both primers specifically amplify their target gene.
DNA-sequence based verification of the LAMP products obtained in the acl1 (above) and IGS (below) based assays. Consensus sequences obtained from sequencing of the smallest DNA fragment in each assay were aligned to the respective reference sequences of target genes.-= nucleotide position not represented in consensus sequence, * = nucleotide position matching in consensus and reference sequences, ß = unmatched positions
Optimization of LAMP Protocols and Sensitivity Testing
To establish optimum incubation conditions for both LAMP assays, 12 LAMP reactions were prepared as previously described. Five microliters of purified genomic DNA from P. destructans OT-38-2010 were added as template. Reactions were incubated for 60 min in the heating block of a gradient thermal cycler, set to provide constant temperatures with 0.83 °C increments between 60.0 and 70.0 °C in individual wells. A clearly visible signal of bright green fluorescence occurred at temperatures between 62.8 and 65.5 °C for both primer sets. Incubation at lower and higher temperatures led to weak or no fluorescence, respectively. In subsequent experiments, acl1- and IGS-based reactions were incubated at 64.0 °C and 65.0 °C, respectively. Sensitivity of both assays was tested by setting up LAMP reactions using a tenfold serial dilution of OT-38-2010 genomic DNA as template. Reactions were incubated for 60 min. The acl1-based assay had a limit of detection (LOD) of 2.1 pg/rxn (Fig. S3 A). Based on a genome size of 35.818201 Mbp for P. destructans [51], this is equivalent to 54 theoretical genome copies per reaction. The same serial dilution of P. destructans OT-38-2010 DNA with the IGS specific primer set, using either calcein (Fig. S3 B) or neutral red (Fig. S3 C) as indicator, revealed an LOD of 21 fg/rxn for both indicators. This concentration was equivalent to 0.54 theoretical genomes per reaction [51]. Six repetitions of this experiment showed that the mentioned LOD was reached in about 50% of experiments suggesting an LOD of rather 1 genome equivalent per reaction. Table 3 gives an overview of the different LAMP experiments carried out in the laboratory as well as their detection limits.
Sensitivity of the IGS-based LAMP assay for detection of P. destructans spores after direct addition to the master mix was tested with untreated conidia of isolate OT-38-2010. Five microliters per reaction were added directly to the master mix from a tenfold serial dilution of a stock solution containing 2.1 × 108 conidia mL−1. In order to increase the sensitivity of direct spore analysis, 300 µl from each tenfold dilution was mixed with sterile glass beads and vortexed as described in materials and methods. Following the treatment, 5 µL from each dilution were transferred to a LAMP master mix and incubated at 65.0 °C for 60 min. The assay was sensitive to the detection of 1.05 × 105 conidia per reaction without any treatment and 1.05 × 103 conidia per reaction after simple glass beads vortex treatment with neutral red as indicator (Fig. 4). These results show that a simple mechanical treatment of conidia during sample preparation can increase the sensitivity of the assay by 100-fold.
Sensitivity of the IGS-based LAMP assay for the detection of P. destructans conidia after direct addition to the master mix. Reactions were run under the conditions described in Table 2 with neutral red as indicator and a serial dilution of conidia from P. destructans OT-38-2010 as template. A LAMP assay with untreated conidial suspensions. B LAMP assay with conidial suspensions after vortexing with sterile glass beads. 1 = 1.05 × 106 conidia /rxn; 2 = 1.05 × 105 conidia/rxn; 3 = 1.05 × 104 conidia/rxn; 4 = 1.05 × 103 conidia/rxn; 5 = 1.05 × 102 conidia/rxn; 6 = 1.05 × 101 conidia/rxn; 7 = 1.05 conidia/rxn; 8 = 0 conidia/rxn
Specificity Testing of LAMP Assays
Both LAMP assays successfully amplified for the diverse set of 53 P. destructans isolates tested. No positive reactions were obtained with DNA from any other of the 159 fungal species tested, including the 25 Pseudogymnoascus species/variants herein tested (see Table S1 in supplemental materials). This result suggests that both LAMP assays are highly specific for the detection of P. destructans and that there will be no cross-reactions with the various taxonomic variants of P. destructans that are phylogenetically closely related and may co-occur in bat environments.
Direct Testing of Pure Culture Mycelia
Signals were detected as early as 3 d post inoculation showing that the assay can be used on young cultures, obviating the need of waiting for the production of conidia. A signal was also detected, when material from older colonies was used. However, no signal was detected from colonies as soon as production of guttation droplets occurred indicating the presence of LAMP inhibitors. However, material taken from the droplet free margin of older colonies still gave a positive signal in the assay when colonies were at least 4 weeks old.
Analysis of Tape Liftings
A protocol was developed in which water-soluble adhesive tape was used to lift fungal material from the surface of agar cultures and microscope glass slides as a model surface. First experiments showed that the tape material readily dissolved in demineralized water at 65.0 °C. However, using this method, a pellet consisting of globular microscopic structures remained after centrifugation and washing which inhibited the LAMP reaction (result not shown). We therefore tested different chaotropic compounds to enhance solubility of the material. Tests showed that 8 M urea resulted in complete disintegration of the tape material with no inhibition of the LAMP assays after three washings with demineralized water. Analysis of solution behaviour at different temperatures showed that the material dissolved completely even at ambient temperature of 23 °C. Figure 5A, B show the efficiency with which spores were lifted from a glass surface using the water soluble Mark Plus II tape. It was noteworthy that the tape removed spores with an efficiency close to 100% while leaving mycelial fragments on the surface. Lifting of spores from a tenfold serial dilution before preparation of spores for the LAMP assay using glass beads as described previously showed that a minimum of 2.3 104 conidia per lifting were detected (see Fig. 6).
Tape lifting of P. destructans spores from a glass surface using 1″ Mask Plus II water soluble wave solder tape (spores per spot = number of spores dried on surface from 100 µL of spore suspension). A Suspension of 2.3 × 106 spores per spot of P. destructans OT-38–2010 air-dried onto surface of a microscope glass slide, 400× magnification. B The same section of the surface after removal of spores with water soluble adhesive tape, 400× magnification. C Suspension of 2.3× 105 spores per spot of P. destructans OT-38-2010 air-dried onto surface of a microscope glass slide, 400× magnification. D The same section of the surface after removal of spores with water soluble adhesive tape, 400× magnification
LAMP reaction with direct addition of: A fungal material prepared according to the new protocol from tape liftings of spores dried previously onto a glass surface, B Conidia added directly from a tenfold serial dilution of P. destructans OT-38-2010 conidial suspension. 1 = 2.3 × 106 conidia per spot (A)/2.3 × 107 conidia per mL (B); 2 = 2.3 × 105 per spot/2.3 × 106 per mL, 3 = 2.3 × 104 conidia per spot/2.3 × 105 per mL; 4 = 2.3 × 103 per spot/2.3 × 104 per mL; 5 = 2.3 × 102 per spot/2.3 × 103 per mL; 6 = 23 per spot/230 per mL, 7 = 0 per spot/0 per ml, 8 = positive control with 1.0 pg/rxn gDNA of P. destructans OT-38-2010, NTC = no template control with water instead of DNA. Pink color = positive reaction, yellow color = negative reaction
Field Experiment: Testing Different Sample Types in Dry-LAMP Reactions
Four out of five samples from bats that have been visually classified as P. destructans-infected gave positive results in the dry-LAMP assay; the one ‘false’ negative was a sample that contained yellowish fluid (suspected inhibitor) which was collected from the bat together with the fungal material. One out of three samples from visually non-infected bats was positive in the LAMP assay. Two out of 11 swabs from hibernacula walls, where P. destructans-infected bats had been previously recorded, were positive in the LAMP assay. Two negative controls (one swab transferred in the tube within the hibernaculum and one during the LAMP preparation) were negative in the LAMP assay. Two tape liftings (tesafilm®) from P. destructans-infected bats were positive in the LAMP assay, the negative control (tape collected on a microscope slide without previously touching a bat) was negative. An overview of the results obtained with field samples is given in supplementary Table S2.
Discussion
Specificity of the LAMP Assays
We herein developed two LAMP assays for the detection of P. destructans. The acl1-based assay detects a typical single copy fungal housekeeping gene. The same gene was used by Niessen, Gräfenhan [52] as target sequence in a Fusarium tricinctum specific LAMP assay. Authors made use of the acl1 sequence because it readily separated F. tricinctum from the closely related F. avenaceum which was not possible when other typical genes such as ß-tubulin, EF1α or calmodulin were targeted. The IGS-based assay developed here for P. destructans uses primers specific for a region that separates eukaryotic cistrons coding for different units of ribosomal RNA. The non-transcribed sequences usually display a high degree of variation between species since mutations are less subject to selection and can therefore be used to separate genetically closely related taxa such as the pathogenic P. destructans and the various close relatives that may occur in the same environments [6]. The specificity of our LAMP-assays was tested against gDNA of a wide variety of fungal species (Table S1), many of which (or their conspecifics) have been found in underground sites used by bats (e.g. [53,54,55,56]). The absence of cross-species reaction demonstrates the high specificity of our LAMP assays. However, the in silico analysis of primer binding revealed that the IGS primer set can be expected to have a superior selectivity for P. destructans since the number of mismatches with closely related taxa was much higher when compared with the acl1 primer set. The higher the mismatch rate of a primer set with closely related non-target species, the higher the selectivity for the target species.
Sensitivity of the LAMP Assays
As fungal cells usually contain several up to several hundred copies of rRNA coding genes [57], an assay based on one of those units, i.e. the IGS sequence, is supposed to provide a higher number of primer binding sites per DNA molecule and should thus have a higher sensitivity for the detection of its target organism. Our results fully confirmed this expectation because the IGS-based assay was about 100-fold more sensitive compared to the single-copy gene acl1 assay. Depending on the detection system applied, the IGS-based LAMP assay had an LOD of 21–30 fg per reaction which is equivalent to 0.54–0.85 theoretical genome copies, respectively. Comparison to LOD of PCR or qPCR assays for P. destructans shows that the new assay has a similarly high sensitivity as the multicopy ITS- and IGS-based qPCR assays described previously [27, 28]. The acl1-based singlecopy LAMP assay had an LOD of 2.1 pg per reaction (equivalent to 54 genome copies per reaction) and was slightly less sensitive than the alr-gene based qPCR published by Chaturvedi, Rudd [26] with an LOD of 0.8 pg (equivalent to 20.7 genome copies per reaction). The IGS-based LAMP assay had a sufficiently high sensitivity to detect P. destructans even in samples with a low concentration of the pathogen. IGS-based LAMP assays were also developed for other fungi such as Metharizium anisopliae [58], Fusarium oxysporum f. sp. cubense tropical race 4 [59] or Trichosporon asahii and T. mucoides [60]. All these assays were also highly sensitive for their respective target organisms. Because of its higher sensitivity, the IGS-based LAMP assay should be given priority over the acl1-based assay when used in practical applications such as screening for the fungus in bat environments (e.g. hibernacula walls [20, 61, 62]).
Advantages and Disadvantages of the LAMP Assay
In contrast to qPCR-based assays, LAMP is a useful tool for presence/absence assessment rather than for precise quantification of target organisms. Even though there is a correlation between time to result and the initial concentration of target DNA, the reaction runs with a dynamic that is too high and with moderate replicability to allow for precise DNA quantification. Few attempts were made in the past to apply LAMP in quantitative assays, but the results were generally less accurate as compared to qPCR [63, 64]. It was demonstrated that LAMP is rather insensitive to matrix inhibition in qualitative assays but shows a strong influence on the quantitative results [65].
Even though LAMP assays, similarly to other DNA- or RNA-based detection methods, do not provide direct information on the extent of wing damage or other symptoms nor information on the viability of their target organisms [66], the LAMP method presents several important advantages over qPCR. First, although extracted DNA can be used as template in the LAMP, as demonstrated herein, unextracted material can equally be used as template, providing a great advantage for applications in the field where limited laboratory equipment is available. Results presented herein show that a limit of 1.05 103 conidia can be detected per reaction after minimal mechanical treatment from a solution of pure culture conidia. This LOD should be sufficiently sensitive to detect fungal colonization during early infection stages. Since P. destructans is a strongly sporulating species, even a detection limit of 1.05 105 conidia per reaction that was obtained for pure culture conidia without mechanical treatment should be sufficient for the diagnosis of visible fungal infections. Second, a thermocycler is not needed to perform the LAMP as the amplification only requires a constant temperature, which can be obtained via a thermal block or even in a water bath (as demonstrated in our field trial, see below). Third, when using indicator dyes for colour change inside the reaction tube, the result is directly readable to the naked eye without the need for any further equipment. Therefore, these LAMP assays provide a greatly needed field-deployable molecular genetic tool for P. destructans detection that will considerably reduce the time necessary for accurate diagnosis. Given that the assays can directly use unextracted material as template, the LAMP assays also provide a quick and cheap method for genetically confirming species ID from culture material, adhesive tape samples or swab samples as classically used (e.g. [7,8,9, 11, 67]). These analyses can be performed in the field but also in the lab where they will provide considerable savings in terms of time and costs compared to qPCR. This new diagnostic LAMP assay and the recently published Visual-Pd scoring system [66] offer significant additions to the toolbox to better understand and monitor White-Nose Disease in bats. While the Visual-Pd score can be considered as non-invasive [66], the collection of samples from bats necessitates a soft contact between the collection item (swab, tape) and the animal skin. However, this can be achieved with minimal or even without disturbance when the procedure is carried out carefully and swiftly, in compliance with general arousal-avoidance measures during the hibernacula surveys [66].
Swabbing or Tape Lifting in Combination with LAMP Assay as a Field-Deployable Method
The current study includes the development of a new method for the sampling of fungal material from solid surfaces and the preparation of such samples for application in LAMP assays. Results showed that spores were detected from a model glass surface with an LOD of 2.3 × 104. Swabbing and tape lifting are common techniques in criminal forensics or veterinary medical mycology and are also used to sample bats for the presence of P. destructans [7, 22]. Tape lifting is particularly useful to obtain a rapid morphological identification of P. destructans by using light microscopy [7]. However, once the fungal material sticks to such tape, it appears difficult to recover it easily for further analyses. Thus, we tested both, common tesafilm® as well as tape lifting material which is water soluble, for direct amplification of the lifted material in the newly developed LAMP assay.
Our tests involved samples from pure cultures as well as swabs and tape liftings collected from bats and hibernacula walls. All sample types were rinsed directly in the LAMP buffer and an aliquot was directly used as LAMP target without further manipulation. Additionally, we developed and successfully tested a dried version of the LAMP reaction (‘dry-LAMP’) which is particularly convenient for shipment and storage prior to testing samples in the field. Although the sample sizes are low, bat swabs and tape samples were successfully tested under field conditions, providing a proof-of-concept for the applicability of the method in the field. Further investigations and optimization are nevertheless necessary before the dry-LAMP can be applied at large scale, especially for hibernacula wall swabs. Indeed, some negative results could be due to inhibition caused by impure sampling material [34] or might have been caused by the absence of high spore loads on the tested hibernacula walls.
The simplicity of swabbing or tape lifting and sample preparation, in combination with a rapid and sensitive LAMP assay as a field deployable detection tool, provides great practical potential to the newly developed methods.
Moreover, the sample preparation protocol is not restricted to applications involving LAMP but can also be used to prepare amplifiable DNA for PCR or qPCR applications. Specifically, the application of swabs or tape that can be tested in the field can be used much to the advantage of researchers, wildlife and conservation biologists involved in WND studies or even for the study of other dermatophytes in other animals or even in humans [35].
Data Availability
All data have been published, either as part of the manuscript or as supplemental materials.
Materials Availability
Fungal strains have been deposited in the culture collection of Chair of Microbiology at Technical University of Munich, Freising, Germany (all TMW clones) or at the culture collection of ISEM, University of Montpellier, France.
Code Availability
Not applicable.
References
Blehert DS, Hicks AC, Behr M, Meteyer CU, Berlowski-Zier BM, Buckles EL, et al. Bat white-nose syndrome: an emerging fungal pathogen? Science. 2009;323(5911):227.
Johnson C, Brown DJ, Sanders C, Stihler CW. Long-term changes in occurrence, relative abundance, and reproductive fitness of bat species in relation to arrival of White-nose Syndrome in West Virginia, USA. Ecol Evol. 2021;11(18):12453–67.
Frick WF, Puechmaille SJ, Hoyt JR, Nickel BA, Langwig KE, Foster JT, et al. Disease alters macroecological patterns of N orth A merican bats. Glob Ecol Biogeo. 2015;24(7):741–9.
Dzal Y, McGuire LP, Veselka N, Fenton MB. Going, going, gone: the impact of white-nose syndrome on the summer activity of the little brown bat (Myotis lucifugus). Biol Lett. 2011;7:392–4.
Gargas A, Trest M, Christensen M, Volk TJ, Blehert D. Geomyces destructans sp. Nov. associated with bat white-nose syndrome. Mycotaxon. 2009;108(1):147–54.
Minnis AM, Lindner DL. Phylogenetic evaluation of Geomyces and allies reveals no close relatives of Pseudogymnoascus destructans, comb. nov., in bat hibernacula of eastern North America. Fung Biol. 2013;117(9):638–49.
Wibbelt G, Kurth A, Hellmann D, Weishaar M, Barlow A, Veith M, et al. White-nose syndrome fungus (Geomyces destructans) in bats, Europe. Emerg Infect Dis. 2010;16(8):1237.
Puechmaille SJ, Verdeyroux P, Fuller H, Gouilh MA, Bekaert M, Teeling EC. White-nose syndrome fungus (Geomyces destructans) in bat, France. Emerg Infect Dis. 2010;16(2):290.
Martínková N, Bačkor P, Bartonička T, Blažková P, Červený J, Falteisek L, et al. Increasing incidence of Geomyces destructans fungus in bats from the Czech Republic and Slovakia. PLoS ONE. 2010;5(11):e13853.
Zukal J, Bandouchova H, Brichta J, Cmokova A, Jaron KS, Kolarik M, et al. White-nose syndrome without borders: Pseudogymnoascus destructans infection tolerated in Europe and Palearctic Asia but not in North America. Sci Rep. 2016;6:19829.
Puechmaille SJ, Wibbelt G, Korn V, Fuller H, Forget F, Mühldorfer K, et al. Pan-European distribution of white-nose syndrome fungus (Geomyces destructans) not associated with mass mortality. PLoS ONE. 2011;6:e19167.
Hoyt JR, Sun K, Parise KL, Lu G, Langwig KE, Jiang T, et al. Widespread bat white-nose syndrome fungus, northeastern China. Emerg Infect Dis. 2016;22:140.
Fritze M, Puechmaille SJ. Identifying unusual mortality events in bats: a baseline for bat hibernation monitoring and white-nose syndrome research. Mamm Rev. 2018;48(3):224–8.
Drees KP, Lorch JM, Puechmaille SJ, Parise KL, Wibbelt G, Hoyt JR, et al. Phylogenetics of a fungal invasion: origins and widespread dispersal of white-nose syndrome. MBio. 2017;8(6):e01941-e2017.
Leopardi S, Blake D, Puechmaille SJ. White-nose syndrome fungus introduced from Europe to North America. Curr Biol. 2015;25(6):R217–9.
Khankhet J, Vanderwolf KJ, McAlpine DF, McBurney S, Overy DP, Slavic D, et al. Clonal expansion of the Pseudogymnoascus destructans genotype in North America is accompanied by significant variation in phenotypic expression. PLoS ONE. 2014;9(8):e104684.
Drees KP, Parise KL, Rivas SM, Felton LL, Puechmaille SJ, Keim P, et al. Characterization of microsatellites in Pseudogymnoascus destructans for white-nose syndrome genetic analysis. J Wild Dis. 2017;53(4):869–74.
Dool S, Altewischer A, Fischer NM, Drees KP, Foster JT, Fritze M, et al. Mating type determination within a microsatellite multiplex for the fungal pathogen Pseudogymnoascus destructans, the causative agent of white-nose disease in bats. Cons Gen Res. 2020;12(1):45–8.
Fischer NM, Dool SE, Puechmaille SJ. Seasonal patterns of Pseudogymnoascus destructans germination indicate host–pathogen coevolution. Biol Lett. 2020;16(6):20200177.
Fischer NM, Altewischer A, Ranpal S, Dool SE, Kerth G, Puechmaille SJ. Population genetics as a tool to elucidate pathogen reservoirs: Lessons from Pseudogymnoascus destructans, the causative agent of white-nose disease in bats. Mol Ecol. 2022;31:675–90.
Zhelyazkova V, Hubancheva A, Radoslavov G, Toshkova N, Puechmaille SJ. Did you wash your caving suit? Cavers’ role in the potential spread of Pseudogymnoascus destructans, the causative agent of white-nose disease. Int J Speleol. 2020;49:145–54.
Verant ML, Bohuski EA, Lorch JM, Blehert DS. Optimized methods for total nucleic acid extraction and quantification of the bat white-nose syndrome fungus, Pseudogymnoascus destructans, from swab and environmental samples. J Vet Diag Invest. 2016;28:110–8.
Zhelyazkova VL, Toshkova NL, Dool SE, Bonaccorso FJ, Pinzari CA, Montoya-Aiona K, et al. Screening and biosecurity for white-nose fungus Pseudogymnoascus destructans (Ascomycota: Pseudeurotiaceae) in Hawai ‘i. Pacific Sci. 2019;73:357–65.
Lorch JM, Gargas A, Meteyer CU, Berlowski-Zier BM, Green DE, Shearn-Bochsler V, et al. Rapid polymerase chain reaction diagnosis of white-nose syndrome in bats. J Vet Diag Invest. 2010;22(2):224–30.
Lindner DL, Gargas A, Lorch JM, Banik MT, Glaeser J, Kunz TH, et al. DNA-based detection of the fungal pathogen Geomyces destructans in soils from bat hibernacula. Mycologia. 2011;103(2):241–6.
Chaturvedi S, Rudd RJ, Davis A, Victor TR, Li X, Appler KA, et al. Rapid real-time PCR assay for culture and tissue identification of Geomyces destructans: the etiologic agent of bat geomycosis (white nose syndrome). Mycopathologia. 2011;172(4):247–56.
Muller LK, Lorch JM, Lindner DL, O’Connor M, Gargas A, Blehert DS. Bat white-nose syndrome: a real-time TaqMan polymerase chain reaction test targeting the intergenic spacer region of Geomyces destructans. Mycologia. 2013;105(2):253–9.
Shuey MM, Drees KP, Lindner DL, Keim P, Foster JT. Highly sensitive quantitative PCR for the detection and differentiation of Pseudogymnoascus destructans and other Pseudogymnoascus species. Appl Env Microbiol. 2014;80(5):1726–31.
Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, Amino N, et al. Loop-mediated isothermal amplification of DNA. Nucl Acids Res. 2000;28(12):e63.
Tomita N, Mori Y, Kanda H, Notomi T. Loop-mediated isothermal amplification (LAMP) of gene sequences and simple visual detection of products. Nat Prot. 2008;3(5):877.
Tanner NA, Zhang Y, Evans TC Jr. Visual detection of isothermal nucleic acid amplification using pH-sensitive dyes. Biotechniques. 2015;58(2):59–68. https://doi.org/10.2144/000114253.
Niessen L. Current state and future perspectives of loop-mediated isothermal amplification (LAMP)-based diagnosis of filamentous fungi and yeasts. Appl Microbiol Biotechnol. 2015;99(2):553–74.
Niessen L. The application of loop-mediated isothermal amplification (LAMP) assays for the rapid diagnosis of food-borne mycotoxigenic fungi. Curr Op Food Sci. 2018;23:11–22.
Francois P, Tangomo M, Hibbs J, Bonetti EJ, Boehme CC, Notomi T, et al. Robustness of a loop-mediated isothermal amplification reaction for diagnostic applications. FEMS Immunol Med Microbiol. 2011;62(1):41–8. https://doi.org/10.1111/j.1574-695X.2011.00785.x.
Kubátová A, Koukol O, Nováková A. Geomyces destructans, phenotypic features of some Czech isolates. Czech Mycol. 2011;63:65–75.
Luo J, Taniwaki MH, Iamanaka BT, Vogel RF, Niessen L. Application of loop-mediated isothermal amplification assays for direct identification of pure cultures of Aspergillus flavus, A. nomius, and A. caelatus and for their rapid detection in shelled Brazil nuts. Int J Food Microbiol. 2013;172:5–12.
Köck J, Gottschalk C, Ulrich S, Schwaiger K, Gareis M, Niessen L. Rapid and selective detection of macrocyclic trichothecene producing Stachybotrys chartarum strains by loop-mediated isothermal amplification (LAMP). Anal Bioanal Chem. 2021. https://doi.org/10.1007/s00216-021-03436-y.
Klich M. Identification of common Aspergillus spcies. Utrecht, The Netherlands: Centraalbureau voor Schimmelcultures; 2002.
Domsch KH, Gams W, Anderson T-H. Compendium of Soil Fungi. London: Academic Press Ltd; 1980.
Nelson P, Tousson T, Marasas WO. Fusarium species: an illustrated manual for identification. Pennsylvania: The Pennsylvania State University Press; 1983.
Niessen ML, Vogel RF. Specific identification of Fusarium graminearum by PCR with gaoA targeted primers. System Appl Microbiol. 1997;20(1):111–23.
Nirenberg HI. A simplified method for identifying Fusarium spp. occurring on wheat. Can J Bot. 1981;59(9):1599–609.
Lorch JM, Lindner DL, Gargas A, Muller LK, Minnis AM, Blehert DS. A culture-based survey of fungi in soil from bat hibernacula in the eastern United States and its implications for detection of Geomyces destructans, the causal agent of bat white-nose syndrome. Mycologia. 2013;105(2):237–52.
Hall, T.A. (1999) BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res. 1997;25(17):3389–402.
Niessen L, Vogel RF. Detection of Fusarium graminearum DNA using a loop-mediated isothermal amplification (LAMP) assay. Int J Food Microbiol. 2010;140(2):183–91.
Denschlag C, Vogel RF, Niessen L. Hyd5 gene based analysis of cereals and malt for gushing-inducing Fusarium spp. by real-time LAMP using fluorescence and turbidity measurements. Int J Food Microbiol. 2013;162(3):245–51.
Niessen L, Bechtner J, Fodil S, Taniwaki MH, Vogel RF. LAMP-based group specific detection of aflatoxin producers within Aspergillus section Flavi in food raw materials, spices, and dried fruit using neutral red for visible-light signal detection. Int J Food Microbiol. 2018;266:241–50.
Chibucos MC, Crabtree J, Nagaraj S, Chaturvedi S, Chaturvedi V. Draft genome sequences of human pathogenic fungus Geomyces pannorum sensu lato and bat white nose syndrome pathogen Geomyces (Pseudogymnoascus) destructans. Gen Announc. 2013;1:e01045-e1113.
Bendtsen JD, Nielsen H, von Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol. 2004;340(4):783–95.
Drees KP, Palmer JM, Sebra R, Lorch JM, Chen C, Wu C-C, et al. Use of multiple sequencing technologies to produce a high-quality genome of the fungus Pseudogymnoascus destructans, the causative agent of bat white-nose syndrome. Gen Announc. 2016. https://doi.org/10.1128/genomeA.00445-16.
Niessen L, Gräfenhan T, Vogel RF. ATP citrate lyase 1 (acl1) gene-based loop-mediated amplification assay for the detection of the Fusarium tricinctum species complex in pure cultures and in cereal samples. Int J Food Microbiol. 2012;158(3):171–85.
Vanderwolf KJ, Malloch D, McAlpine DF, Forbes GJ. A world review of fungi, yeasts, and slime molds in caves. Int J Speleol. 2013;42:9.
Zhang T, Victor TR, Rajkumar SS, Li X, Okoniewski JC, Hicks AC, et al. Mycobiome of the bat white nose syndrome affected caves and mines reveals diversity of fungi and local adaptation by the fungal pathogen Pseudogymnoascus (Geomyces) destructans. PLoS ONE. 2014;9:e108714.
Nováková A. Microscopic fungi isolated from the Domica Cave system (Slovak Karst National Park, Slovakia): a review. Int J Speleol. 2009;38:8.
Lorch JM, Lindner DL, Gargas A, Muller LK, Minnis AM, Blehert DS. A culture-based survey of fungi in soil from bat hibernacula in the eastern United States and its implications for detection of Geomyces destructans, the causal agent of bat white-nose syndrome. Mycologia. 2013;105:237–52.
Lofgren LA, Uehling JK, Branco S, Bruns TD, Martin F, Kennedy PG. Genome-based estimates of fungal rDNA copy number variation across phylogenetic scales and ecological lifestyles. Mol Ecol. 2019;28:721–30.
Li Y, Cai S-H. Sensitive and rapid detection of the insect pathogenic fungus Metarhizium anisopliae var. anisopliae by loop-mediated isothermal amplification. Curr Microbiol. 2011;62(5):1400–4.
Zhang X, Zhang H, Pu J, Qi Y, Yu Q, Xie Y, et al. Development of a real-time fluorescence loop-mediated isothermal amplification assay for rapid and quantitative detection of Fusarium oxysporum f. sp. cubense tropical race 4 in soil. PLoS ONE. 2013;8(12):e82841.
Kasahara K, Ishikawa H, Sato S, Shimakawa Y, Watanabe K. Development of multiplex loop-mediated isothermal amplification assays to detect medically important yeasts in dairy products. FEMS Microbiol Lett. 2014;357(2):208–16.
Vanderwolf KJ, Malloch D, Mcalpine DF. Detecting viable Pseudogymnoascus destructans (Ascomycota: Pseudeurotiaceae) from walls of bat hibernacula: effect of culture media. J Cave Karst Stud. 2016;78:158.
Vanderwolf KJ, Malloch D, McAlpine DF. No change detected in culturable fungal assemblages on cave walls in eastern Canada with the introduction of Pseudogymnoascus destructans. Diversity. 2019;11:222.
Li L, Zhang Sy, Zhang C-Q. Establishment of a rapid detection method for rice blast fungus based on one-step loop-mediated isothermal amplification (LAMP). Plant Dis. 2019. https://doi.org/10.1094/PDIS-11-18-1964-RE.
Peng J, Zhang H, Chen F, Zhang X, Xie Y, Hou X, et al. Rapid and quantitative detection of Fusarium oxysporum f. sp. cubense race 4 in soil by real-time fluorescence loop-mediated isothermal amplification. J Appl Microbiol. 2014;117(6):1740–9.
Nixon GJ, Svenstrup HF, Donald CE, Carder C, Stephenson JM, Morris-Jones S, et al. A novel approach for evaluating the performance of real time quantitative loop-mediated isothermal amplification-based methods. Biomol Det Quant. 2014;2:4–10. https://doi.org/10.1016/j.bdq.2014.11.001.
Fritze M, Puechmaille SJ, Fickel J, Czirják GÁ, Voigt CC. A rapid, in-situ minimally-invasive technique to assess infections with Pseudogymnoascus destructans in bats. Act Chiropterol. 2021;23:259–70.
Wibbelt G, Puechmaille SJ, Ohlendorf B, Mühldorfer K, Bosch T, Görföl T, et al. Skin lesions in European hibernating bats associated with Geomyces destructans, the etiologic agent of white-nose syndrome. PLoS ONE. 2013;8(9):e74105.
Acknowledgements
We thank Alena Kubátová, Charles University in Prague (CUNI), Department of Botany for providing cultures of P. destructans from the Culture Collection of Fungi (CCF), Jeff Lorch for providing cultures of Pseudogymnoascus spp. from hibernacula sediments, and all who contributed samples from which cultures were isolated and herein used.
Funding
This work was supported by Bat Conservation International (awarded to SJP); the Deutsche Forschungsgemeinschaft (PU 527/2-1, awarded to SJP); and the Institut Universitaire de France (IUF, awarded to SJP). Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
LN conceived and designed the study, performed the research, analyzed data and wrote the paper. MF performed research, contributed new methods, and edited the paper. GW analyzed data and edited the paper. SJP helped with the study design, performed research, analyzed data and edited the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that none of them has a conflict of interest or competing interests.
Ethical Approval
This research involved no studies on humans or animals.
Additional information
Handling Editor: Koichi MAKIMURA.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11046_2022_650_MOESM2_ESM.pdf
Figure S1 Binding site homologies between primer set Pd-acl1-ID30 and acl1 gene sequences of fungal species retrieved from GenBank. The used sequences shared >85 % total homology with the P. destructans acl1 gene sequence. Nucleotide positions with homology to the respective positions in the primer sequences are marked in grey, non-homologues positions are marked in white (PDF 16 KB)
11046_2022_650_MOESM3_ESM.pdf
Figure S2 Binding site homologies between primer set Pd-IGS-ID10 and the 28S-18S intergenic spacer (IGS) sequence of three P. destructans strains and 10 Pseudogymnoascus spp. and P. verrucosus. Sequences retrieved from GenBank. The used sequences shared >90 % total homology with the corresponding P. destructans sequence. Nucleotide positions with homology to the respective positions in the primer sequences marked in grey, non-homologues positions marked in white (PDF 531 KB)
11046_2022_650_MOESM4_ESM.pdf
Figure S3 Sensitivity of LAMP assays for the detection of P. destructans purified gDNA. Reactions were run under the conditions described in table 3 with calcein and neutral red as indicator dyes and a serial 10fold dilution of purified gDNA of strain OT-38-2010 as template. A LAMP assay using primer set Pd-acl1-ID30 with calcein indicator under UV light. Bright green fluorescence = positive reaction; no fluorescence = negative reaction. B LAMP assay using primer set Pd-IGS-ID10 with calcein indicator under UV light. C LAMP assay using primer set Pd-IGS-ID10 with neutral red indicator under day light conditions (pink = positive reaction; yellow = negative reaction). 1 = 213 ng per reaction (ng/rxn) of P. destructans OT-38-2010 gDNA; 2 = 21.3 ng/rxn; 3 = 2.1 ng/rxn; 4 = 210 pg/rxn; 5 = 21 pg/rxn; 6 = 2.1 pg/rxn; 7 = 210 fg/rxn; 8 = 21 fg/rxn; 9 = 2.1 fg/rxn; 10 = 210 ag/rxn; NTC = not template control, demineralized water instead of DNA-template (PDF 351 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Niessen, L., Fritze, M., Wibbelt, G. et al. Development and Application of Loop-Mediated Isothermal Amplification (LAMP) Assays for Rapid Diagnosis of the Bat White-Nose Disease Fungus Pseudogymnoascus destructans. Mycopathologia 187, 547–565 (2022). https://doi.org/10.1007/s11046-022-00650-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-022-00650-9