Abstract
Background
The Eastern Tropical Pacific (ETP) harbors a great diversity of Porifera. In particular, the Aplysina genus has acquired biotechnological and pharmacological importance. Nevertheless, the ecological aspects of their species and populations have been poorly studied. Aplysina gerardogreeni is the most conspicuous verongid sponge from the ETP, where it is usually found on rocky-coralline ecosystems. We evaluated the polymorphism levels of 18 microsatellites obtained from next-generation sequencing technologies. Furthermore, we tested the null hypothesis of panmixia in A. gerardogreeni population from two Mexican-Pacific localities.
Methods and results
A total of 6,128,000 paired reads were processed of which primer sets of 18 microsatellites were designed. The loci were tested in 64 specimens from Mazatlan, Sinaloa (N = 32) and Isabel Island, Nayarit (N = 32). The microsatellites developed were moderately polymorphic with a range of alleles between 2 and 11, and Ho between 0.069 and 0.785. Fifteen loci displayed significant deviation from the Hardy–Weinberg equilibrium. No linkage disequilibrium was detected. A strong genetic structure was confirmed between localities using hierarchical Bayesian analyses, principal coordinates analyses, and fixation indices (FST = 0.108*). All the samples were assigned to their locality; however, there was a small sign of mixing between localities.
Conclusions
Despite the moderate values of diversity in microsatellites, they showed a strong signal of genetic structure between populations. We suggest that these molecular markers can be a relevant tool to evaluate all populations across the ETP. In addition, 17 of these microsatellites were successfully amplified in the species A. fistularis and A. lacunosa, meaning they could also be applied in congeneric sponges from the Caribbean Sea. The use of these molecular markers in population genetic studies will allow assessment of the connectivity patterns in species of the Aplysina genus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sponges (Phylum Porifera) have become an interesting model for studying ecological and evolutionary processes in marine environments; they are mainly characterized by limited larval dispersal capacity, sexual and asexual reproduction, and a sessile lifestyle [1,2,3,4,5]. Due to their high abundance and wide diversity, they have a pivotal ecological role in most aquatic ecosystems, filtering the water column and providing substrate and shelter for a wide variety of organisms [3, 6, 7]. Despite its relevance, biological aspects such as reproduction and population genetics have generally been little studied e.g., [8,9,10,11,12].
Studies of the population genetics in sponge species have been conducted mainly on traditional nuclear and mitochondrial genes (e.g.: ITS’s, 28 S, 18 S, COI, among others), which showed a low polymorphism level in most groups. Therefore, the approaches addressed with these markers have been mostly in taxonomy and systematics, limiting the knowledge of historical and contemporary demography, as well as the phylogeographic patterns in this group e.g., [13, 14]. Nevertheless, the use of Next Generation Sequencing (NGS) platforms in the development of hypervariable markers, such as microsatellites, has increased significantly in population genetic studies of Porifera e.g., [15, 16]. These markers have shown to be powerful tools for population studies providing valuable information about their population dynamics e.g., [9, 11, 12].
Genetic population studies on Porifera have evaluated the degree of structure and connectivity genetic among populations. These patterns have allowed to identification of marine areas that function as genetic reservoirs [10]. The identification of these reservoirs provides fundamental information for designing management plans and protection of marine areas [10]. In addition, the patterns of structure and connectivity genetic have been used to estimate and evaluate the invasive potential of some sponge species [17], as well as the effects of mass mortality [9], the effects of hydrodynamics on the distribution of populations e.g., [18, 19], and the assessment of endangered species [20].
The genus Aplysina is the most conspicuous sponge group of the order Verongiida, with 47 valid species [21], many of which are widely recognized for their developed natural metabolites with cytotoxic and antimicrobial activity, and the use in bioengineering in regeneration tissue [22,23,24,25,26]. Despite its importance, some biological and ecological characteristics of this group of species are poorly understood. Although evolutionary aspects have been studied through mitochondrial and nuclear markers, they have exhibited low levels of polymorphism, even at the mitogenome level, which has limited their use in intraspecific studies [13, 27]. Species of this genus are characterized by larvae dispersal. The type of larvae has been described as clavablastula ciliated and swimming, with a period of settlement in the substrate after seven days [28]. Furthermore, this group presents asexual reproduction, which is carried out through fragmentation; where a part of the body of the sponge is detached and transported away several meters from the original parental site, where they settled and developed [29, 30]. These features allow hypothesized a low dispersal potential in Aplysina species.
The Eastern Tropical Pacific (ETP) is a region that extends from the Gulf of California to northern Peru, characterized by marine currents that provide unique oceanographic conditions promoting high levels of productivity and biodiversity [31, 32]. Aplysina gerardogreeni (Gómez and Bakus, 1992) is the most common verongid species from the ETP, usually found in rocky and coralline ecosystems [13, 33, 34]. Due to its high prevalence, the present study aims to develop microsatellite-type molecular markers to evaluate the genetic pattern of A. gerardogreeni in the ETP.
Materials and methods
Next-generation sequencing and microsatellite design
The procedure details for the NGS and specimen collections were previously described [27]. For microsatellite design, repetitive motifs of di-, tri-, and tetranucleotides were searched in the assembled contigs for a subsequent primer design using Msatcommander software [35]. All forward primers included the M13 primer sequence attached to their 5’ end following a protocol of dye-labeled universal primer [36].
Sample collection and DNA extraction
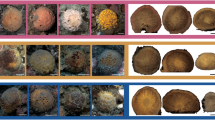
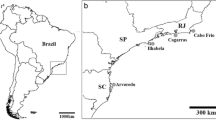
Sixty-four specimens of A. gerardogreeni were collected by SCUBA diving, 32 from Mazatlán Bay (23°15′29″N, 106°28′25″W) and 32 from Isabel Island (21°51′15″N, 105°53′33″W) Mexico (Fig. 1); samples were collected in different years (from 2010 to 2021) (Table S1). Genomic DNA was obtained using Promega’s Wizard® SV Genomic DNA Purification System protocol following the manufacturer’s instructions.
Location of the sampling stations in the Mexican Pacific and specimens’ images of Aplysina gerardogreeni. A Geographic distance between Mazatlan and Isabel Island. B Mazatlan. C Isabel Island. D A. gerardogreeni from Mazatlán. E A. gerardogreeni from Isabel Island (the yellow arrows indicate specimen collection sites for both sites). (Color figure online)
Amplification and genotyping of microsatellites
The PCR reaction mix contained 0.7 µl dNTPs (10mM) (Promega™), 2.0 µl 5x PCR Buffer (Promega™), 1.1 µl MgCl2 (25 mM) (Promega™), 0.5 µl unlabeled M13-tailed F-primer (10 mM), 0.5 µl R-Primer (10 mM), and 1.0 µl of fluorochrome-labeled F-primer (10 mM) (FAM, VIC, PET or NED), 1.0 µl BSA (20ng/µl) (Bovine Serum Albumin; SIGMA™), 0.1 µl Taq DNA polymerase (5u/µl) (Promega™), 1.5 µl of DNA (50 ng/µl), and fill out with H2O (styled Milli-Q Merck Millipore) to a final volume of 13 µl.
The thermocycling profile consisted of two stages: first 94 °C/4 min, followed by 30 cycles of 94 °C/30 s; 59 °C/30 s and 72 °C/60 s. Second, the fluorochrome-labeled forward primer was added to continue with 10 cycles of 94 °C/30 s; 53 °C/30 s and 72 °C/60 s; and a final elongation of 72 °C/20 min. Amplification products were visualized in 1.5% agarose gels stained with Gel Red™ Nucleic Acid Gel Stain (Biotium). To test the success of amplifying microsatellite loci on congeneric species, we randomly selected 3 specimens from each of the representative species of A. lacunosa and A. fistularis from the Mexican Caribbean and the Gulf of Mexico, respectively.
The final products were genotyped with an ABI3730 DNA analyzer (Applied Biosystems™). The genotypes were scored with Genemarker v3.0.1 software with GeneScan™ 500 LIZ (Soft Genetics, State College, PA, USA) and to convert and determine allelic size. The dataset to generate input-files was handled using a macro-Excel, Flexibin [37]. Finally, the presence of null alleles, large allele dropout and genotyping errors were assessed with Micro-Checker v.2.2.3 [38]. In addition, following the same methodology, we tested the cross-amplification in specimens of congeneric species: A. fistularis and A. lacunosa.
Data analyses
Polymorphism levels were estimated by the observed (HO) and expected heterozygosity (HE) indices, the number of alleles per locus and the polymorphic content index (PIC) using a macro excel Mstools v 3 [39]. Further, we evaluated the linkage disequilibrium (LD) between pairs of loci using Mstools. Tests of Hardy–Weinberg Equilibrium (HWE) for each locus were assessed using a probability test with a level of significance determined by Markov chain parameters of 1,000 dememorization steps, 100 batches and 1,000 iterations per batch using GENEPOP Web v 4.2 [40], we used the Weir and Cockerham for F-statistics [41]. The p-values of multiple comparison analyses (HWE and LD) were adjusted using the classical one-stage method of the False Discovery Rate (FDR) procedure [42].
For the statistical analyses of genetic structure, we only used those microsatellite loci in HWE. First, we used Structure v 2.3.4 software [43] with parameters set to 10 iterations discarded as a burn-in, and 100,000 Markov Chains Monte Carlo (MCMC) were run with a burn-in of 10,000 iterations. Moreover, Structure was run using an admixture ancestral model with independence of allele frequencies, and prior information of sample location. Ten replicates were run for each K value (K = 1 to 2). The K number was estimated with Structure Harvester Web v 0.6.94 [44]. According to plots of log probability LnP(K) of the data (Supplementary Material Fig. S1), the ten replicates for the best K were merged in Clumpp [45] and visualized by Distruct [46]. Second, Principal Coordinates Analysis (PCoA) was constructed using a pairwise codominant genotypic distance matrix using GenAlEx v 6.5 [47]. Third, population differentiation was assessed using pairwise FST in GenAlEx v 6.5 [47].
Results
Genetic diversity
Forty-one microsatellite loci were isolated, of which 18 presented at least two alleles per locus (Table 1). Genetic diversity indices were performed within loci with at least three alleles in each location (Table 2). Large allele dropout and genotyping errors were not detected, but seven loci exhibited the presence of possible null alleles. Six loci were in HWE and neither locus showed linkage disequilibrium (Table 2).
Almost all loci presented low levels of genetic diversity. The highest and the lowest numbers of alleles per locus were detected in the AGMX-6292 (11 alleles) and AGMX-180680 (2 alleles) loci. The PIC average in the data set was 0.587; the values for each locus were between low and moderate (0.255–0.765). The highest values of HO were detected in AGMX-37595 and AGMX-734 (0.614 and 0.817) loci. The FIS showed high values in loci AGMX-82588 (0.692) and AGMX-182674 (0.783), and AGMX-734 loci with exogamy (− 0.186; Table 2). Isabel Island presented higher values of genetic diversity than Mazatlan, except the FIS average was higher in Mazatlan.
Genetic differentiation
The hierarchical Bayesian analysis revealed two genetic clusters among the organisms. The average LnP(K) value was maximal at 2, and the membership probabilities of the sample individuals reflected a clear geographical pattern of genetic differentiation (Fig. 2). In addition, the genetic distribution of the individuals was graphically represented under a vector plane in the PCoA, the results suggested one genetic group associated with each locality (Fig. 3). Nonetheless, there is an overlap of some Mazatlan individuals with Isabel Island cluster (Fig. 3). The initial two principal components (PCs) explain 45.09% of the observed variation: PC 1 explained 30.28%, whereas PC 2 explained 14.81%. The AMOVA showed a moderate genetic structure between localities (FST = 0.108; P-value < 0.05).
Discussion
Genetic diversity
The high number of loci out of HWE could be indicative of either technical issues, such as null alleles [48, 49] (Table S2), or biological features of this species, including the potential of inbreeding [28]. Although there is little information about the reproductive biology of A. gerardogreeni, one record in Isabel Island showed that 5.2% and 2.5% of samples developed oocytes and spermatic cysts, with a female-male sex proportion of 3:1 [50]. Those results indicate sexual reproduction in a small portion, leaving the possibility that most sponges of this species reproduce asexually. This would be consistent with low levels of genetic diversity and high values of inbreeding in both locations (Table 2).
In general, sponge species tend to present low levels of genetic diversity, which could be associated with the asexual reproduction seen in many species e.g., [51,52,53]. This type of reproduction is associated to the response of massive population reductions by meteorological phenomena such as storms and hurricanes, and hydrodynamic local events [8, 18, 19, 54]. Mazatlan and Isabel Island are localized in the mouth of Gulf of California, a region with high oceanographic dynamics, because converged oceanic currents such as the California Current near-surface and the Mexican Coastal Current at sub-surface [55]. In addition, this region is characterized by high activity of tropical cyclones and hurricanes [56]; therefore, it is probable that sponge species recover their populations after a drastic decline through asexual reproduction. Nevertheless, we did not detect identical genotypes across both localities. This finding could be due to (1) the sampling method possibly preventing collection of clones because there were from three to five meters of separation between samples, and (2) this species present both types of reproduction according to environmental conditions (stressful and non-stressful) [54]. To corroborate our findings, studies at smaller geographic scales must be conducted e.g., [12, 53].
Genetic differentiation
Sponges from Mazatlan and Isabel Island conform to two genetically isolated populations (FST = 0.108*; Figs. 2 and 3). Although there is little evidence of sexual reproduction in A. gerardogreeni, it is possible exchange organisms between populations through larval dispersal by currents or by hitchhiking invasive and floating buds [8, 11, 12, 54]. Under the premise that A. gerardogreeni could develop a type of clavablastula larva like its congeneric (A. aerophoba) [28], it is possible there is limited dispersal. In addition, the environmental conditions between locations play a relevant role in fixing different alleles [57, 58]. Both localities present contrasting environmental conditions. Mazatlan is a coastal region near many estuaries and river mouths where sediment entrainment is characteristic; sponge species are suspension feeders and changes in sediment levels can affect the abundance of populations [59]. In contrast, Isabel Island present low sediment deposition and is not affected by anthropogenic impacts, thus is considered a site with better environmental conditions for reef communities [60].
To conclude, we observed a high degree of genetic structure in A. gerardogreeni using only six microsatellite loci; we hope to increase the number of microsatellites by increasing the study area at the Mexican Pacific (investigation in progress). In addition, these markers can be amplified in two congeneric species from the Caribbean Sea (A. fistularis and A. lacunosa), therefore, it offers the possibility of evaluating the patterns of genetic structure population in these species.
Data availability
Information on the designated primers and microsatellite sequences will be available in GenBank once the manuscript is accepted.
References
Maldonado M (2006) The ecology of the sponge larva. Can J Zool 84:2. https://doi.org/10.1139/z05-177
Ereskovsky A (2010). The comparative embryology of sponges. https://doi.org/10.1007/978-90-481-8575-7
Carballo JL, Bell JJ (2017). Climate change, ocean acidification and sponges: impacts across multiple levels of organization. https://doi.org/10.1007/978-3-319-59008-0
Pires AC, Marinoni L (2010) DNA barcoding and traditional taxonomy unified through integrative taxonomy: a view that challenges the debate questioning both methodologies. Biota Neotrop 10:2. https://doi.org/10.1590/S1676-06032010000200035
Morrow C, Cárdenas P (2015) Proposal for a revised classification of the Demospongiae (Porifera). Front Zool 12:1. https://doi.org/10.1186/s12983-015-0099-8
van Soest RWM, Boury-Esnault N, Vacelet J, Dohrmann M, Erpenbeck D, de Voogd NJ, Santodomingo N, Vanhoorne B, Kelly M, Hooper JNA (2012) Global diversity of sponges (Porifera). PLoS ONE 7:4. https://doi.org/10.1371/journal.pone.0035105
Schönberg CHL (2021) No taxonomy needed: sponge functional morphologies inform about environmental conditions. Ecol Indic. https://doi.org/10.1016/j.ecolind.2021.107806
DeBiasse MB, Richards VP, Shivji MS (2010) Genetic assessment of connectivity in the common reef sponge, Callyspongia vaginalis (Demospongiae: Haplosclerida) reveals high population structure along the Florida reef tract. Coral Reefs 29:1. https://doi.org/10.1007/s00338-009-0554-0
Riesgo A, Pérez-Portela R, Pita L, Blasco G, Erwin PM, López-Legentil S (2016) Population structure and connectivity in the Mediterranean sponge Ircinia fasciculata are affected by mass mortalities and hybridization. Heredity 117:6. https://doi.org/10.1038/hdy.2016.41
Taboada S, Kenny NJ, Riesgo A, Wiklund H, Paterson GLJ, Dahlgren TG, Glover AG (2018) Mitochondrial genome and polymorphic microsatellite markers from the abyssal sponge Plenaster craigi Lim & Wiklund, 2017: tools for understanding the impact of deep-sea mining. Marine Biodivers 48:1. https://doi.org/10.1007/s12526-017-0786-0
Cavalcanti FF, Padua A, Cunha H, Halasz M, Nikolić V, Barreto S, Klautau M (2020) Population differentiation supports multiple human-mediated introductions of the transatlantic exotic sponge Paraleucilla magna (Porifera, Calcarea). Hydrobiologia 847:17. https://doi.org/10.1007/s10750-020-04368-x
Shaffer MR, Davy SK, Bell JJ (2021) Reproductive isolation between two cryptic sponge species in New Zealand: high levels of connectivity and clonality shape Tethya species boundaries. Mar Biol 168:5. https://doi.org/10.1007/s00227-021-03880-5
Cruz-Barraza JA, Carballo JL, Rocha-Olivares A, Ehrlich H, Hog M (2012) Integrative taxonomy and molecular phylogeny of genus Aplysina (Demospongiae: Verongida) from Mexican Pacific. PLoS ONE 7:8. https://doi.org/10.1371/journal.pone.0042049
Gómez P, González-Acosta B, Sánchez-Ortíz C, Hoffman Z, Hernández-Guerrero CJ (2018) Amended definitions for Aplysinidae and Aplysina (Porifera, Demospongiae, Verongiida): on three new species from a remarkable population in the gulf of California. Zootaxa 4455:2. https://doi.org/10.11646/zootaxa.4455.2.4
Riesgo A, Blasco G, Erwin PM, Pérez-Portela R, López-Legentil S (2014) Optimization of 14 microsatellite loci in a Mediterranean demosponge subjected to population decimation, Ircinia fasciculata. Conserv Genet Res 6:2. https://doi.org/10.1007/s12686-013-0137-9
Taboada S, Ríos P, Mitchell A, Cranston A, Busch K, Tonzo V, Cárdenas P, Sánchez F, Leiva C, Koutsouveli V, Cristobo J, Xavier JR, Hentschel U, Rapp HT, Morrow C, Drewery J, Romero PE, Arias MB, Whiting C, Riesgo A (2022) Genetic diversity, gene flow and hybridization in fan-shaped sponges (Phakellia spp.) in the North-East Atlantic deep sea. Deep-Sea Res Part I: Oceanogr Res Papers. https://doi.org/10.1016/j.dsr.2021.103685
Guardiola M, Frotscher J, Uriz MJ (2016) High genetic diversity, phenotypic plasticity, and invasive potential of a recently introduced calcareous sponge, fast spreading across the Atlanto-Mediterranean basin. Mar Biol 163:5. https://doi.org/10.1007/s00227-016-2862-6
Padua A, Cavalcanti FF, Cunha H, Klautau M (2013) Isolation and characterization of polymorphic microsatellite loci from Clathrina aurea (Porifera, Calcarea). Marine Biodivers 43:4. https://doi.org/10.1007/s12526-013-0167-2
Richards VP, Bernard AM, Feldheim KA, Shivji MS (2016) Patterns of population structure and dispersal in the long-lived redwood of the coral reef, the giant barrel sponge (Xestospongia muta). Coral Reefs 35:3. https://doi.org/10.1007/s00338-016-1435-y
Li R, Nitsche F, Arndt H (2018) Mesoscale investigations based on microsatellite analysis of the freshwater sponge Ephydatia fluviatilis in the River-Sieg system (Germany) reveal a genetic divergence. Conserv Genet 19:4. https://doi.org/10.1007/s10592-018-1069-4
de Voogd NJ, Alvarez B, Boury-Esnault N, Carballo JL, Cárdenas P, Díaz M-C, Dohrmann M, Downey R, Hajdu E, Hooper JNA, Kelly M, Klautau M, Manconi R, Morrow CC, Pisera AB, Ríos P, Rützler K, Schönberg C, Vacelet J, van Soest RWM (2023) World Porifera database. Accessed 16 September, 2023, from https://www.marinespecies.org/porifera
Gutleben J, Koehorst JJ, McPherson K, Pomponi S, Wijffels RH, Smidt H, Sipkema D (2019) Diversity of tryptophan halogenases in sponges of the genus Aplysina. FEMS Microbiol Ecol 95:8. https://doi.org/10.1093/femsec/fiz108
Lever J, Brkljača R, Rix C, Urban S (2021) Application of networking approaches to assess the chemical diversity, biogeography, and pharmaceutical potential of Verongiida natural products. Mar Drugs 19:10. https://doi.org/10.3390/md19100582
Ehrlich H, Steck E, Ilan M et al (2010) Three-dimensional chitin-based scaffolds from Verongida sponges (Demospongiae: Porifera). Part II: biomimetic potential and applications. Int J Biol Macromol 47:2. https://doi.org/10.1016/j.ijbiomac.2010.05.009
Binnewerg B, Schubert M, Voronkina A et al (2020) Marine biomaterials: biomimetic and pharmacological potential of cultivated Aplysina aerophoba marine demosponge. Mater Sci Eng C. https://doi.org/10.1016/j.msec.2019.110566
Mutsenko V, Bazhenov V, Rogulska O et al (2017) 3D chitinous scaffolds derived from cultivated marine demosponge Aplysina aerophoba for tissue engineering approaches based on human mesenchymal stromal cells. Int J Biol Macromol. https://doi.org/10.1016/j.ijbiomac.2017.03.116
Salas-Castañeda M, Castillo-Páez A, Rocha-Olivares A, Cruz-Barraza J (2019) The complete mitogenome of the Eastern Pacific sponge Aplysina gerardogreeni (Demospongiae, Verongida, Aplysinidae). Mitochondrial DNA Part B Res 4:2. https://doi.org/10.1080/23802359.2019.1643804
Maldonado M (2009) Embryonic development of verongid demosponges supports the Independent acquisition of spongin skeletons as an alternative to the siliceous skeleton of sponges. Biol J Linn Soc 97:2. https://doi.org/10.1111/j.1095-8312.2009.01202.x
Wulff JL (1991) Asexual fragmentation, genotype success, and population dynamics of erect branching sponges. J Exp Mar Biol Ecol 149:2. https://doi.org/10.1016/0022-0981(91)90047-Z
Tsurumi M, Reiswig HM (1997) Sexual versus asexual reproduction in an oviparous rope-form sponge, Aplysina cauliformis (Porifera; Verongida). Invertebr Reprod Dev. https://doi.org/10.1080/07924259.1997.9672598
Enright SR, Meneses-Orellana R, Keith I (2021) The eastern tropical pacific marine corridor (CMAR): the emergence of a voluntary regional cooperation mechanism for the conservation and sustainable use of marine biodiversity within a fragmented regional ocean governance landscape. Front Mar Sci. https://doi.org/10.3389/fmars.2021.674825
Fiedler PC, Lavín MF (2017) Oceanographic conditions of the eastern tropical pacific. Coral Reefs East Trop Pacific. https://doi.org/10.1007/978-94-017-7499-4_3
Gómez, Bakus GJ (1992) Aplysina gerardogreeni and Aplysina aztecus (Porifera: Demospongiae), new species from the Mexican Pacific. Anales Del Instituto de Ciencias Del Mar y Limnología 19:2
Caballero-George C, Bolaños J, Ochoa E, Carballo JL, Cruz JA, Elizabeth AA (2010) Protocol to isolate sponge-associated fungi from tropical waters and an examination of their cardioprotective potential. Curr Trends Biotechnol Pharm 4:4
Faircloth B (2008) Msatcommander: detection of microsatellite repeat arrays & automated, locus-specific primer design. Mol Ecol Res 8:92–94. https://doi.org/10.1111/j.1471-8286.2007.01884.x
de Arruda MP, Gonçalves EC, Schneider MPC, da Costa da Silva AL, Morielle-Versute E (2010) An alternative genotyping method using dye-labeled universal primer to reduce unspecific amplifications. Mol Biol Rep 37:4. https://doi.org/10.1007/s11033-009-9655-7
Amos W, Hoffman J, Frodsham A, Zhang L, Best S, Hill A (2007) Automated binning of microsatellite alleles: problems and solutions. Mol Ecol Notes 7:1. https://doi.org/10.1111/j.1471-8286.2006.01560.x
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:3. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Park S (2001) MStools v 3 (Excel spreadsheet toolkit for data conversion) Smurfit Institute of Genetics. Trinity College. Dublin 2
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:3. https://doi.org/10.1093/oxfordjournals.jhered.a111573
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:6. https://doi.org/10.1111/j.1558-5646.1984.tb05657.x
Pike N (2011) Using false discovery rates for multiple comparisons in ecology and evolution. Methods Ecol Evol 2:3. https://doi.org/10.1111/j.2041-210X.2010.00061.x
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:2. https://doi.org/10.1093/genetics/155.2.945
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Res 4:2. https://doi.org/10.1007/s12686-011-9548-7
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:14. https://doi.org/10.1093/bioinformatics/btm233
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:1. https://doi.org/10.1046/j.1471-8286.2003.00566.x
Peakall R, Smouse PE (2012) GenALEx 6.5: genetic analysis in excel. Population genetic software for teaching and research—an update. Bioinformatics 28:9. https://doi.org/10.1093/bioinformatics/bts460
Chapuis MP, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24:3. https://doi.org/10.1093/molbev/msl191
Ahmed L, Al-Najjar Y, Cramer E, Thareja G, Suhre K, Chen K-C (2022) Development and characterization of microsatellite primers for Triops granarius (Branchiopoda: Notostraca) using MiSeq technology. Mol Biol Rep 49:10. https://doi.org/10.1007/s11033-022-07804-4
Chávez-Castro (2014) Ciclo reproductivo de dos esponjas del género Aplysina (esponja: Demospongiae) en la Isla Isabel, Nayarit. MSc thesis. Faculty of Marine Sciences. Autonomous University of Sinaloa, pp54. http://posgradofacimar.maz.uasnet.mx/tesis/
Blanquer A, Uriz MJ (2010) Population genetics at three spatial scales of a rare sponge living in fragmented habitats. BMC Evol Biol 10:1. https://doi.org/10.1186/1471-2148-10-13
Kelly M (1986) Systematics and ecology of the sponges of Motupore Island, Papua New Guinea. MSc thesis. University of Auckland, p 101. https://researchspace.auckland.ac.nz/handle/2292/5966
Duran S, Pascual M, Estoup A, Turon X (2004) Strong population structure in the marine sponge Crambe crambe (Poecilosclerida) as revealed by microsatellite markers. Mol Ecol 13:3. https://doi.org/10.1046/j.1365-294X.2004.2080.x
Uriz M, Turon X (2012) Sponge ecology in the molecular era. Adv Mar Biol. https://doi.org/10.1016/B978-0-12-387787-1.00006-4
Gómez-Valdivia F, Parés-Sierra A, Flores-Morales AL (2015) The Mexican coastal current: a subsurface seasonal bridge that connects the tropical and subtropical Northeastern Pacific. Cont Shelf Res. https://doi.org/10.1016/j.csr.2015.10.010
Kimberlain TB (2014) The 2013 Eastern North Pacific Hurricane season: Mexico takes the brunt. Weatherwise 67:3. https://doi.org/10.1080/00431672.2014.899802
Leiva C, Taboada S, Kenny NJ, Combosch D, Giribet G, Jombart T, Riesgo A (2019) Population substructure and signals of divergent adaptive selection despite admixture in the sponge Dendrilla antarctica from shallow waters surrounding the Antarctic Peninsula. Mol Ecol 28:13. https://doi.org/10.1111/mec.15135
Shaffer M, Luter H, Webster N, Abdul W, Bell J (2020) Evidence for genetic structuring and limited dispersal ability in the great barrier reef sponge Carteriospongia foliascens. Coral Reefs 39:1. https://doi.org/10.1007/s00338-019-01876-8
Bell JJ, McGrath E, Biggerstaff A, Bates T, Bennett H, Marlow J, Shaffer M (2015) Sediment impacts on marine sponges. Mar Pollut Bull 94:1–2. https://doi.org/10.1016/j.marpolbul.2015.03.030
Nava H, Carballo JL (2013) Environmental factors shaping boring sponge assemblages at Mexican Pacific coral reefs. Mar Ecol 34:3. https://doi.org/10.1111/maec.12012
Acknowledgements
The first author received a CONACYT scholarship No. 782102. Thanks to Misha Yazmín Hernández-Lozano for assistance in the laboratory and Francisco Ponce Núñez for his help in the design of geographic maps.
Funding
This work has been funded by the Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica PAPIIT (DGAPA): PAPIIT-UNAM IN210018.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation, all authors performed data collection and analysis. The first draft of the manuscript was written by [MRS-C] and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
No animal testing was performed during this study. Aplysina gerardogreeni is not a protected or endangered species. Sampling activities were not performed at locations where specific permission is required.
Consent to participate
Not applicable.
Research involving human and animal participants
No other studies with other animals or human participants were performed by any of the authors.
Additional information
Publisher’s Note
Springer nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salas-Castañeda, M.R., Saavedra-Sotelo, N.C., Cruz-Barraza, J.A. et al. Novel microsatellite markers suggest significant genetic isolation in the Eastern Pacific sponge Aplysina gerardogreeni. Mol Biol Rep 51, 87 (2024). https://doi.org/10.1007/s11033-023-09043-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-023-09043-7