Abstract
Background
Bacterial leaf blight (BLB) is one of the major biotic stress in rice cultivation. Management techniques, such as the development of BLB-resistant cultivars, are required to lessen the severity of the disease attack and yield losses. Pratikshya was selected in the present investigation as the recipient parent, as it is one of the popular high-yielding rice varieties of Odisha, India, which is having excellent grain as well as cooking quality. However, Pratikshya is highly susceptible to BLB which is prevalent in Eastern Indian region.
Methods and results
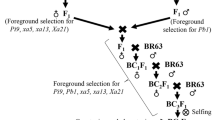
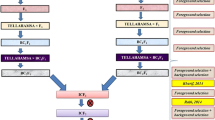
Three major BLB resistance genes xa5, xa13, and Xa21 from the donor source Swarna MAS (CR Dhan 800) were attempted to introduce into Pratikshya through a marker-assisted backcross breeding program. Those markers closely linked to the target genes were employed for foreground selection in the segregating generations till BC2F3. In each backcross generation, progenies containing all three targeted resistance genes and phenotypically more similar to the recipient parent, Pratikshya were selected and backcrossed. Screening of 1,598 plants of the BC2F2 population was conducted against BLB using Xoo inoculum and 35 resistant plants similar to Pratikshya were carried forward to the next generation. In the BC2F3 generation, 31 plants were found to possess all the three resistance genes. For background selection of plants carrying resistance genes 45 polymorphic SSR markers were employed. Evaluation of the pyramided lines at BC2F4 generation exhibited that, most pyramided lines were similar to Pratikshya in terms of morphological features and yield parameters, and some lines were superior to the recurrent parent in terms of morphological features and yield parameters.
Conclusion
The three-gene pyramided lines showed a high level of resistance to BLB infection and are anticipated to offer a significant yield advantage over the recipient parent Pratikshya. The pyramided lines can further be used for multi-location trial, so as to be released as a variety or can be used as a potential donor for BLB resistance genes.





Similar content being viewed by others
Data Availability
All data generated or analysed during this study are included in this published article.
References
Rahman H, Dakshinamurthi V, Ramasamy S, Manickam S, Ashok Kumar Kaliyaperumal AK, Raha S, Panneerselvam N, Ramanathan V, Nallathambi J, Sabarippan R, Raveendran M (2018) Introgression of Submergence Tolerance into CO 43, a Popular Rice Variety of India, through marker-assisted Backcross breeding. Czech J Genet Plant Breed. 54:101–108
Khush GS (2005) What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol. 59:1–6
Muduli L, Dash M, Das Mohapatra S, Mohapatra KK, Nayak HS, Bastia DN, Pradhan B, Tripathy SK, Jena RC, Pradhan SK (2023) Phenotypic and genotypic assessment of elite rice varieties for brown plant hopper (Nilaparvata lugens Stål.) Resistance. Cereal Res Commun. 1–13
Srinivasan B, Gnanamanickam S (2005) Identification of a new source of resistance in wild rice, Oryza rufipogon to bacterial blight of rice caused by indian strains of Xanthomonas oryzae pv. Oryzae. Curr Sci. 88:1229–1231
Mew TW (2003) Current status and future prospects of research on bacterial blight of rice. Annu Rev Phytopathol. 25:359–382
Lee K, Rasabandith S, Angeles E, Khush G (2003) Inheritance of resistance to bacterial blight in 21 cultivars of rice. Phytopathol. 93:147–152
McManus PS, Stockwell VO, Sundin GW, Jones AL (2002) Antibiotic use in plant agriculture. Annu Rev Phytopathol. 40:443–465. https://doi.org/10.1146/annurev.phyto.40.120301
Ahmad N, Joji RM, Shahid M (2023) Evolution and implementation of one health to control the dissemination of antibiotic-resistant bacteria and resistance genes: a review. Front Cell Infect Microbiol. 12:1065796. https://doi.org/10.3389/fcimb.2022.1065796
Khush GS, Mackill DJ, Sidhu GS (1989) Breeding rice for resistance to bacterial blight. Bacterial blight of rice. IRRI, pp 207–217
Xie LH, Lin QY, Wu ZJ, Zhou ZJ, Duan YP (1994) Diagnosis, detection and control of rice virus disease in China. J Fujian Agric Univ. 23:280–285
Arunakumari K, Durgarani CV, Satturu V, Sarikonda KR, Chittoor PD, Vutukuri B, Laha GS, Nelli AP, Gattu S, Jamal M, Prasadbabu A (2016) Marker-assisted pyramiding of genes conferring resistance against bacterial blight and blast diseases into indian rice variety MTU1010. Rice Sci. 23:306–316
He D, Zhan J, Xie L (2016) Problems, challenges and future of plant disease management from an ecological point of view. J Integr Agric. 15:705–715
Pradhan M, Bastia DN (2022) Validation of linked markers of bacterial leaf blight resistance genes in rice variety of Odisha (Oryza sativa). J Pharm Innov. 11:446–448
Das G, Rao G (2015) Molecular marker assisted gene stacking for biotic and abiotic stress resistance genes in an elite rice cultivar. Front Plant Sci. 6
Chen S, Wang C, Yang J, Chen B, Wang W, Su J (2020) Identification of the novel bacterial blight resistance gene Xa46(t) by mapping and expression analysis of rice mutant H120. Sci Rep. 10:14642
Chen L, Yin F, Zhang D, Xiao S, Zhong Q, Wang B, Cheng Z (2022) Unveiling a novel source of resistance to bacterial blight in medicinal wild rice, Oryza officinalis. Life. 12:827. https://doi.org/10.3390/life12060827
Jiang N, Yan J, Liang Y (2020) Resistance genes and their interactions with bacterial Blight/Leaf Streak Pathogens (Xanthomonas oryzae) in Rice (Oryza sativa L.) - an updated review. Rice. 13:3. https://doi.org/10.1186/s12284-019-0358-y
Singh S, Sidhu J, Huang N, Vikal Y, Li Z, Brar D, Dhaliwal H, Khush G (2001) Pyramiding three bacterial blight resistance genes (xa5, xa13 and Xa21) using marker-assisted selection into indica rice cultivar PR106. Theor Appl Genet. 102:1011–1015
Rajpurohit D, Kumar R, Kumar M, Paul P, Awasthi A, Basha PO, Puri A, Jhang T, Singh K, Dhaliwal HS (2010) Pyramiding of two bacterial blight resistance and a semi dwarfing gene in type 3 basmati using marker-assisted selection. Euphytica. 178:111–126
Collard BC, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos Trans R Soc Lond B Biol Sci. 363(1491):557–572. https://doi.org/10.1098/rstb.2007.2170
Balachiranjeevi CH, Bhaskar NS, Abhilash V, Akanksha S, Viraktamath BC, Madhav MS, Hariprasad AS, Laha GS, Prasad MS, Balachandran SM, Neeraja CN, Kumar MS, Senguttvel P, Kemparaju KB, Bhadana VP, Ram T, Harika G, Swamy HKM, Hajira SK, Yugandhar A, Pranathi K, Anila M, Rekha G, Kousik BVN, Kumar TD, Swapnil RK, Archana G, Sundaram RM (2015) Marker-assisted introgression of bacterial blight and blast resistance into DRR17B, an elite, fine-grain type maintainer line of rice. Mol Breed. 35:151
Mohapatra S, Panda AK, Bastia AK, Mukherjee AK, Sanghamitra P, Meher J, Mohanty SP, Pradhan SK (2021) Development of Submergence-Tolerant, bacterial Blight-Resistant, and high-yielding Near Isogenic Lines of Popular Variety, ‘Swarna’ through marker-assisted breeding Approach. Front Plant Sci. 12:672618
Kauffman HE, Reddy APK, Hsien SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Disease Rep. 57:537–554
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus. 12:13–15
Jena PP, Bharathkumar S, Reddy JN, Mohapatra T (2015) Introgression of Sub1 locus into highly Preferred Rice Cultivars (Pooja and Pratikshya) in Eastern Region of India for Submergence Tolerance through marker assisted Backcrossing. Adv Biores. 6:45–53
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: Software for association mapping of complex traits in diverse samples. J Bioinform 23:2633–2635
IRRI (2013) Standard evaluation system for rice. International Rice Research Institute, Philippines
Gopinath PP, Parsad R, Joseph B, Adarsh VS (2020) GRAPES: General Rshiny Based Analysis Platform Empowered by Statistics. Available at: https://www.kaugrapes.com/home. version 1.0.0. Accessed on: 1st May 2022
Sarangi, S. K., Maji, B., Mahanta, K. K., Digar, S., Burman, D., Mandal, S., … Bell,R. W. (2019). Alternate kharif rice crop establishment methods and medium duration varieties to enable cropping system intensification in coastal saline region. Journal of Indian Society of Coastal Agricultural Research. 37(2), 115–122. Available at: https://krishi.icar.gov.in/jspui/handle/123456789/49625
Duy PN, Lan DT, Pham Thu H, Thi Thu HP, Nguyen Thanh H, Pham NP, Auguy F, Bui Thi Thu H, Manh TB, Cunnac S, Pham XH (2021) Improved bacterial leaf blight disease resistance in the major elite vietnamese rice cultivar TBR225 via editing of the OsSWEET14 promoter. PLoS ONE. 16(9):e0255470. https://doi.org/10.1371/journal.pone.0255470. PMID: 34499670; PMCID: PMC8428762
Dokku P, Das KM, Rao GJN (2013) Pyramiding of four resistance genes of bacterial blight in Tapaswini, an elite rice cultivar, through marker-assisted selection. Euphytica. 192:87–96
Hajira SK, Yugander A, Balachiranjeevi CH, Pranathi K, Anila M, Mahadevaswamy HK (2014) Development of durable bacterial blight resistant lines of Samba Mahsuri possessing Xa33, Xa21, Xa13&Xa5. Progressive Res. 9:1224–1227
Babu R, Nair SK, Prasanna B, Gupta H (2004) Integrating marker-assisted selection in crop breeding-prospects and challenges. Curr Sci. 87:607–619
Hsu YC, Chiu CH, Yap R, Tseng YC, Wu YP (2020) Pyramiding bacterial blight resistance genes in Tainung82 for broad-spectrum resistance using marker-assisted selection. Int J Mol Sci. 21(4):1281. https://doi.org/10.3390/ijms21041281. PMID: 32074964; PMCID: PMC7072918
Pradhan SK, Nayak DK, Pandit E, Behera L, Anandan A, Lenka S, Barik DP (2016) Incorporation of bacterial leaf blight resistance genes into low-land rice cultivars through marker-assisted breeding. Phytopathol. 106:710–718
Wang GL, Song WY, Ruan DL, Sideris S, Ronald PC (1996) The cloned gene, Xa21, confers resistance to multiple Xanthomonas oryzae pv. Oryzae isolates in transgenic plants. Mol Plant-Microbe Interact. 9:850–855
Iyer AS, McCouch SR (2004) The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol Plant Microbe Interact. 17:1348–1354
Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL (2006) Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev. 20:1250–1255
Sanchez A, Brar D, Huang N, Li Z, Khush G (2000) Sequence tagged site marker-assisted selection for three bacterial blight resistance genes in rice. Crop Sci. 40:792–797
Gopalakrishnan S, Sharma RK, Rajkumar KA, Joseph M, Singh VP, Singh AK, Bhat KV, Singh NK, Mohapatra T (2008) Integrating marker-assisted background analysis with foreground selection for identification of superior bacterial blight resistant recombinants in Basmati rice. Plant Breed. 127:131–139
Pradhan SK, Pandit E, Pawar S, Baksh SY, Mukherjee AK, Mohanty SP (2019) Development of flash-flood tolerant and durable bacterial blight resistant versions of mega rice variety ‘swarna’ through marker-assisted backcross breeding. Sci Rep. 9:12810
Baliyan N, Malik R, Rani R, Mehta K, Vashisth U, Dhillon S, Boora KS (2018) Integrating marker-assisted background analysis with foreground selection for pyramiding bacterial blight resistance genes into Basmati rice. C R Biologies. 341:1–8
Ellur RK, Khanna A, Bhowmick PK, Vinod KK, Nagarajan M, Mondal KK, Singh NK, Singh K, Prabhu KV, Singh AK (2016) Marker-aided incorporation of Xa38, a Novel Bacterial Blight Resistance Gene, in PB1121 and comparison of its resistance spectrum with xa13 + Xa21. Sci Rep. 6:29188
Chen JM, Fu ZY, Quan BQ, Tian DG, Li G, Wang F (2009) Breeding hybrid rice restoring line with double resistance to rice blast and bacterial blight by marker-assisted selection. Mol Plant Breed. 7:465–470
Funding
The authors acknowledge the support of IRRI-OUAT project (Pyramiding of resistance genes for BLB and submergence tolerance into Pratikshya: A popular rice variety of Odisha).
Author information
Authors and Affiliations
Contributions
DB, KCS supervised the work. MP, DB developed the genetic materials. MP, DB, KCS, MD, and JPS performed and supported the molecular analyses. MP, DB performed the field evaluations. MP, DB, and JPS performed and supported the analyses of data. MP, DB, KCS, MD, and JPS contributed in writing and critically revised the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pradhan, M., Bastia, D., Samal, K.C. et al. Pyramiding resistance genes for bacterial leaf blight (Xanthomonas oryzae pv. Oryzae) into the popular rice variety, Pratikshya through marker assisted backcrossing. Mol Biol Rep 50, 9047–9060 (2023). https://doi.org/10.1007/s11033-023-08805-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08805-7