Abstract
Background
Ovarian cancer is a lethal gynecological cancer and no reliable minimally invasive early diagnosis tools exist. High grade serous ovarian carcinoma (HGSOC) is often diagnosed at advanced stages, resulting in poorer outcome than those diagnosed in early stage. Circulating microRNAs have been investigated for their biomarker potential. However, due to lack of standardization methods for microRNA detection, there is no consensus, which microRNAs should be used as stable endogenous controls. We aimed to identify microRNAs that are stably expressed in plasma of HGSOC and benign ovarian tumor patients.
Methods and results
We isolated RNA from plasma samples of 60 HGSOC and 48 benign patients. RT-qPCR was accomplished with a custom panel covering 40 microRNAs and 8 controls. Stability analysis was performed using five algorithms: Normfinder, geNorm, Delta-Ct, BestKeeper and RefFinder using an R-package; RefSeeker developed by our study group [1]. Among 41 analyzed RNAs, 13 were present in all samples and eligible for stability analysis. Differences between stability rankings were observed across algorithms. In HGSOC samples, hsa-miR-126-3p and hsa-miR-23a-3p were identified as the two most stable miRNAs. In benign samples, hsa-miR-191-5p and hsa-miR-27a-3p were most stable. In the combined HGSOC and benign group, hsa-miR-23a-3p and hsa-miR-27a-3p were identified by both the RefFinder and Normfinder analysis as the most stable miRNAs.
Conclusions
Consensus regarding normalization approaches in microRNA studies is needed. The choice of endogenous microRNAs used for normalization depends on the histological content of the cohort. Furthermore, normalization also depends on the algorithms used for stability analysis.
Similar content being viewed by others
Introduction
Ovarian cancer (OC) is the most lethal gynecological cancer with an estimated 313,959 new cases and 207,252 deaths worldwide in 2020 [2]. OC is subdivided into four main stages by the International Federation of Gynecology and Obstetrics (FIGO). Epithelial OC (EOC) which accounts for about 90–95% of OCs consists of four main histological subtypes: 75% Serous Carcinoma (SC) of which 70% is High grade (HGSOC) and 5% is Low Grade (LGSC), 10% are Clear Cell Carcinomas, 10% are Endometroid Carcinomas, and about 5% are Mucinous Carcinomas. Most HGSOCs are diagnosed in the late stages (FIGO III and IV) due to subtle symptoms of disease [3]. This is the most predominant and aggressive type of OC and due to high incidence and low survival rates caused by late-stage discovery, HGSOC is the most lethal of the EOCs [4], accounting for 70–80% of death related to ovarian cancer [5]. These cancers are characterized by frequent DNA gains and losses and chromosomal instability causing gene breakage and loss of hetero- and homozygosity [5].
The late discovery of HGSOC has a significant negative impact on overall survival and hence, there is an urgent clinical need for diagnostic and prognostic biomarkers. MicroRNAs (miRNAs) are small, noncoding RNAs and due to their involvement in regulation of mRNA and protein expression have gained increasing attention as biomarkers for various diseases [6,7,8]. However, the implementation of miRNA biomarkers in a clinical setting has been progressing slowly, in part due to missing standardization of methods for measuring, detecting and normalizing miRNA expression [9].
An important part of this standardization also pertains to the preprocessing of the data. Melt curve analysis is performed to identify poorly amplified products, the existence of primer dimer or genomic contamination. Besides being quite time consuming, melt curve evaluation is often very subjective. Spike-in controls is used by many to monitor efficiency and quality of RNA extraction, cDNA synthesis and final amplification. Many claim to perform these analyses but do not report methodology or list exclusion criteria and in our experience, it is difficult to locate specific guidelines and general acceptable cut-off and threshold values for variation in spike-in controls or differentiating between an insignificant shoulder and a second peak in a melt curve.
Ideally, normalization should be performed using stable endogenous references of the same type of RNA that is being quantified [10, 11]. However, identification of suitable references for normalization is not a trivial exercise. There is no universal endogenous control suitable for every tissue type, as the expression of most miRNAs varies with cell type and condition [12, 13]. An example of this is U6 (RNU-6-1) that have commonly been utilized as an endogenous control for miRNA expression in OC [14, 15] even though it has been shown to be differentially expressed in cancers and plasma [16]. Instead, references should be validated on a per study basis and several algorithms, assessing stability from RT-qPCR data are freely available. Normfinder [17], and geNorm [18] seem to be the most used but also delta-Ct [19] and Bestkeeper [20] have been used extensively. Normfinder ranks the stability of target RNAs based on a weighted geometric mean of the inter- and intra-group variations. GeNorm calculates a stability value M based roughly on the standard deviation of the linearized pairwise ratios between each target for each sample. Delta-Ct uses candidate pairwise ΔCt values, estimating a mean standard deviation for each candidate. BestKeeper stability is based on the mean absolute deviation (MAD) of each candidate. These different approaches may lead to different ranking of candidate references and discrepancy between research groups, comparing results from several algorithms may increase confidence in the selected reference [21]. Various combinations of the four have been employed. In the case with RefFinder [22], all these four algorithms are taken into account when calculating a geometric mean of the candidate’s stability rankings from the different algorithms. Ultimately, to advance the discovery and clinical utility of miRNA expression as biomarkers, consensus regarding these methodologies need to be build. There are no published studies on identification of stably expressed endogenous miRNAs in OC. In this study we aimed to provide a starting framework for discussing these methodological deficiencies while investigating stability of selected miRNAs in plasma samples from HGSOC patients and patients with benign gynecological tumors.
Materials and methods
Study design
EDTA plasma samples were collected prior to primary surgery from 60 patients diagnosed with ovarian high grade serous carcinoma (HGSOC) and 48 patients with benign gynecological tumors relevant to our clinical setting. Samples were obtained through the Bio- and Genome Bank Denmark from two Danish projects: the Pelvic Mass study (2004–2014) and the GOVEC (Gynecological Ovarian Vulva Endometrial Cervix cancer) study (2015 – ongoing). Each patient has provided a written informed consent and the Danish National Committee for Research Ethics, Capital Region has approved the study (approval codes KF01-227/03 and KF01-143/04). During this study the Declaration of Helsinki guidelines was followed.
miRNA extraction of plasma samples
RNA was extracted using the miRNeasy Serum/Plasma Kit (Qiagen, Copenhagen, Denmark, cat. no. 217,184) as previously described [21]. Briefly, 200 µl plasma was lysed using 1 ml QIAzol lysis reagent, followed by addition of 1 µl RNA isolation control spike-ins mix consisting of UniSp2, UniSp4 and UniSp5, each at a different concentration with 100-fold increments: UniSp2 > UniSp4 > UniSp5. Afterwards, the samples were purified according to the manufacturer’s recommendations. RNA was eluted in 14 µl RNase free water and stored at -80 °C until further use. A no sample control extraction was also performed. This sample was subjected to the same procedure, only no plasma was added to the lysis buffer.
cDNA synthesis
cDNA synthesis was performed using miRCURY LNA RT Kit (Qiagen, Copenhagen, Denmark, cat. no. 339,340) following the manufacturer’s protocol. Briefly, for each 10 µl reaction, 2 µl 5X reaction buffer, 1 µl 10X miRCURY RT enzyme mix as well as 5.4 µl nuclease free water, 1.1 µl RNA and 0.5 µl RNA cDNA synthesis spike-in control mix containing UniSp6 and cel-miR-39-3p.
RT-qPCR
For RT-qPCR, on blood-plasma samples we decided to use a custom miRCURY LNA miRNA PCR Panels (Qiagen, Copenhagen, Denmark) previously designed for investigation miRNA expression in tissue samples from ovarian cancer patients [23]. See table S1. The PCR panel contained 40 miRNAs selected for their reported stability or potential as biomarkers in OC. Also, U6 (RNU6-1) which has previously been used as a reference for normalization was included. Assays for the spike-ins controls UniSp2, UniSp4 and UniSp5 added during RNA extraction as well as cel-miR-39-3p and UniSp6 that were added during cDNA synthesis were also selected. Additionally, the interplate calibrator UniSp3 was included to correct for inter plate variations as well as a blank spot to control for possible contaminations. A list of analyzed miRNAs can be found in Table 1 and further details regarding selection of these can be found in supplementary Table S1.
RT-qPCR reactions were performed as previously described [21] using the custom miRCURY LNA miRNA PCR Panels, in a 384-well plate format (Qiagen, Copenhagen, Denmark) and a LightCycler 480 384-well Block (Roche, Hvidovre, Denmark).
For each sample, a solution containing 264 µl 2X miRCURY SYBR Green master mix, 257 µl nuclease free water and 7 µl cDNA was prepared. Eight samples were prepared for each 384-well plate containing primers and polymerase enzymes and 10 µl reaction mix was aliquoted to each well. The PCR plates were then sealed, centrifuged for 1 min at 1500 x g, and subjected to real-time PCR amplification in a Roche LightCycler 480 according to the protocol, including 2 min heat activation at 95 °C, 45 amplification cycles of 95 °C for 10 s and 56 °C for 60 s ending with a melt curve analysis.
Crossing points (Cps) of the amplification curves were calculated by the LightCycler®480 software version 1.5 (Roche) using absolute quantification analysis/2nd derivative maximum method with high confidence setting. Melting temperature analysis (Tm calling) and calculating melting curve peaks were performed through the LightCycler®480 software. Cp and Tm tables as well as raw melt data were exported as txt-files for melt curve and data analysis.
Data analysis
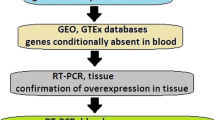
All data analyses were performed using R Statistical Software (version 4.1.1; R Foundation for Statistical Computing, Vienna, Austria) and R-studio IDE (version 1.4.1717, RStudio, Boston, United states, rstudio.com) and [24]. An overview of the analysis workflow can be seen in Fig. 1.
Workflow diagram explaining the quality control filtering process. Value in the first bubble represents number of samples (109 samples). All n values represent the number of Cp value data points arriving at the node. Note that when combining complete cases from the HGSOC and Benign group for stability analysis, only miRNAs that are in common in the two datasets are merged, meaning that, four miRNAs and 220 Cp values were removed from the HGSOC group
Melt curve analysis
Melt curve analyses were performed to prevent using data from poorly amplified products caused by primer dimer, non-specific targets, or genomic contamination. Prior to analysis, an adjusted Cp value was calculated based on the inter plate calibrator; UniSp3 according to manufacturer’s instructions. A Cp cutoff value of 35 was chosen and all Cp or adjusted Cp values above this threshold were set to not available (NA). Four of the 41 analyzed RNAs were removed due to all Cp values being above this cut-off value.
The Cp table, the Tm table, and the melt curve data provided by the LightCycler®480 software was used to evaluate initial sample quality. Using the color and status column present in the exported Cp data, each reaction was characterized as either; approved, uncertain, absent, or late. For the Tm calling data, three categories were identified: approved, inconclusive, or absent. In this way, each reaction was categorized according to supplementary Table S2. Manual melt curve assessment was performed on amplifications where a Cp value was reported, but not auto-approved (approved by both the Cp and Tm calling produced by the LightCycler®480 software) (supplementary Table S2, grey shaded cells).
The manual evaluation was performed by comparing the melt curve for the reaction in question with all auto-approved melt curves from the same miRNA. In general, reactions were rejected manually if no clear peak was seen, a peak was seen but did not reach a threshold of 0.5 fluorescent units, a melt curve contained two peaks and one peak was at least half the size of the other or if more than two peaks were seen. A total of 468 melt curves were analyzed manually out of which 357 datapoints were approved, the rest were rejected and removed from the dataset Fig. 1.
Spike-in analysis
To control for variation in extraction efficiencies, three spike-in RNAs (UniSp4, UniSp4 and UniSp5) were added to the lysis buffer during RNA extraction. Samples were split in two groups: HGSOC and benign. Based on these groups, samples were removed if their Cp value for UniSp2 was an outlier as defined by the ± 1.5xIQR rule [25] or if the difference in Cp values of UniSp2 and UniSp4 were not between five and eight cycles. UniSp5 was not considered since the Cp values for this spike-in were above Cp = 35 cut-off. Hemolysis was checked on 20 random samples using the miRCURY QC Panel according to the protocol and as described in [21, 26]. No hemolysis was observed and was therefore considered negligible.
A summary of the spike-in analysis can be seen in Supplementary Table S3, and an overview of the workflow can be seen in Fig. 1. No HGSOC sample was identified as a UniSp2 outlier, six samples that had a difference in UniSp2 and UniSp4 Cp values of less than five. Two samples had a difference in UniSp2 and UniSp4 Cp values of more than eight. In total, eight samples were excluded from the HGSOC group resulting in the removal of one miRNA and a total of 52 remaining. Of the benign samples, five had a difference in Cp values of UniSp2 and UniSp4 of less than five, one of which was also identified as a UniSp2 outlier. No samples had a difference in UniSp2 and UniSp4 Cp values of more than eight. In total, five samples were excluded from further analyses, leaving 43 samples for further analyses.
Stability analysis and normalization
To investigate the stabilities of miRNAs, we developed the R package RefSeeker [1] based on the web tool RefFinder [22] found at https://www.heartcure.com.au/reffinder/. RefFinder utilizes four widely used algorithms for stability analysis; BestKeeper [20], Normfinder [17], GeNorm [18] and the comparative delta-Ct method [19]. Using the ranking of stability values from the four other algorithms RefFinder calculates a geometric mean, representing an overall stability ranking value for each miRNA. These algorithms do not perform equally well on all datasets. The geNorm algorithm tends to favor highly correlated genes excluding potential stable genes with bad correlation to regulated genes [27]. Compared to geNorm, Normfinder has been described as less robust with smaller sample sizes [28] and the delta-Ct method was found to work better with heterogeneously correlated sets of genes [27]. The full BestKeeper analysis calculates two main measures of stability, one of which is the raw MAD that is incorporated into the RefFinder method. The MAD, however, is best suited for samples with a very fixed amount of input material [28].
The RefFinder rank was calculated separately for each of the HGSOC and benign groups as well as the HGSOC and benign combined on miRNAs that were detected in all samples. Stabilities of candidate pairs were also checked using the Normfinder analysis of expression data grouped in benign and HGSOC samples.
Results
Patients
Clinical and pathologic characteristics of the patients are summarized in Table 2. For the control group, patients with various benign gynecological tumors relevant in the clinical setting was selected. A group of 60 HGSOC patients were selected as test group. Of these, one patient was categorized as FIGO stage II, 38 as FIGO stage III and 21 as FIGO stage IV. Outcome of the patients was checked on 2 November 2021.
Identification of stable miRNA
After quality control of the data, 17 miRNAs presented complete cases in the HGSOC group, 19 miRNAs had missing values and five miRNAs were excluded since all Cp values were eliminated (Table 3). In the benign group, 13 miRNAs were present in all remaining samples, 24 miRNAs had missing data and four miRNAs were eliminated due to removal of all Cp values (Table 4). Remarkably, U6 were missing in 56% of the HGSOC samples and 77% of the benign samples and were never considered as a candidate reference RNA.
Stability assessment was performed on the complete cases using the RefFinder method through the R package RefSeeker [1]. This method provides results from the delta-Ct, BestKeeper, Normfinder and geNorm algorithms as well as an overall agreement score. Complete case datasets can be found in the supplemental material Table S4, Table S5 and Table S6. In HGSOC samples, stability of 17 miRNAs were evaluated (Table 5). In these samples the delta-Ct method ranks hsa-miR-23a-3p as most stable and hsa-miR-126-3p as second most stable miRNA. This is in line with Normfinder that ranks these as third and second most stable respectively, with hsa-miR-191-5p as most stable. GeNorm also ranks these two as some of the most stable: with hsa-miR-126-3p and hsa-miR-223-3p as most stable and hsa-miR-23a-3p as the fourth most stable. This is contrasted by BestKeeper that ranks hsa-miR-126-3p as the eighth most stable and hsa-miR-23a-3p as 12th most stable corresponding to the sixth least stable miRNA. Instead, BestKeeper finds hsa-miR-193a-5p as the most stable and hsa-miR-486-5p as second. Interestingly, the four other algorithms find hsa-miR-193a-5p to be the least stable of all miRNAs. Using these different rankings, RefFinder identify hsa-miR-126-3p as most stable and hsa-miR-23a-3p as second most stable followed by hsa-miR-191-5p and hsa-miR-223-3p.
In benign samples, stability of 13 miRNAs were evaluated (Table 5). Here, hsa-miR-191-5p and hsa-miR-27a-3p is found to be the most and second most stable respectively, by delta-Ct. The results of the Normfinder ranking are, except for a single switch in the order of hsa-miR-199a-3p and hsa-miR-103a-3p, identical to delta-Ct. The most and second most stable miRNAs are therefore also here hsa-miR-191-5p and hsa-miR-27a-3p. GeNorm ranks these in reverse order with hsa-miR-27a-3p and hsa-miR-126-3p as most stable and hsa-miR-191-5p as fourth most stable with hsa-miR-23a-3p as third most stable. BestKeeper ranks hsa-miR-191-5p as second most stable and hsa-miR-27a-3p as seventh most stable with hsa-miR-101-3p. Combining these rankings, RefFinder orders hsa-miR-191-5p, hsa-miR-27a-3p, hsa-miR-126-3p, and hsa-miR-23a-3p as most, second most, third most and fourth most stable respectively.
When combining the two groups (HGSOC and benign), only the miRNAs in common between these groups were evaluated. Opportunely, the 13 miRNAs evaluated for the benign group were all represented in the HGSOC group as well, thus stability of 13 miRNAs were assessed for all samples (Table 5). Here, hsa-miR-23a-3p, hsa-miR-27a-3p, hsa-miR-126-3p and hsa-miR-191-5p ranked as most, second most, third most and fourth most stable respectively. Normfinder mostly agrees on this, however, with a slightly different order with hsa-miR-23a-3p, hsa-miR-191-5p, hsa-miR-27a-3p and hsa-miR-126-3p ordered from most stable to least stable. GeNorm largely agrees with the two others, however it finds hsa-miR-191-5p to be the fifth most stable with hsa-miR-27a-3p and hsa-miR-126-3p as the most stable miRNAs and hsa-miR-23a-3p and hsa-miR-223-3p as the third and fourth most stable, respectively. Interestingly, BestKeeper finds quite different rankings compared to the three other algorithms. Though hsa-miR-191-5p also here is found to be quite stable as the third most stable miRNA, hsa-miR-23a-3p ranks as the ninth most stable, hsa-miR-27a-3p as the 10th most stable and hsa-miR-126-3p as the seventh most stable miRNA. The most stable, second most stable and fourth most stable miRNA are instead found by BestKeeper to be hsa-miR-486-5p, hsa-miR-101-3p and hsa-miR-199a-3p, respectively.
To further validate our findings that hsa-miR-23a-3p and hsa-miR-27a-3p were the most stable miRNAs across HGSOC and benign samples, we performed grouped Normfinder analysis on the combined dataset assessing the most stable pair of miRNAs across the two sample groups. This resulted in hsa-miR-23a-3p and hsa-miR-27a-3p being the most stable pair with a stability value of 0.042 (supplementary Table S7).
Discussion
Discovery and detection of blood-based miRNA biomarkers largely depends upon the accurate and robust measurement of the presence of miRNAs in an analyzed sample. RT-qPCR is a proven and reliable method for the detection and quantification of miRNA and other nucleotide sequences. However, the accuracy of RT-qPCR quantification is dependent on the selection of a stable reference for normalization [29, 30].
Missing consensus on how to perform quality control and preprocessing of data to identify and assess the stability of reference miRNA candidates also makes direct comparisons challenging. Different algorithms are being used and selection of an algorithm often seems arbitrary [30,31,32,33]. Since these algorithms often provide very different recommendations and the outcome of a given study is dependent on the selected references, results could end up appearing random and inconsistent. Moreover, in our experience, the algorithms are quite sensitive to small changes in the datasets, hence the data preprocessing has a major impact on results.
We used the RefSeeker package to perform the RefFinder analysis which provide results from four widely used algorithms (delta-Ct, BestKeeper, Normfinder and geNorm) as well as an overall ranking.
In this study, we found hsa-miR-23a-3p and hsa-miR-126-3p to be the best candidates to use for reference if only HGSOC samples were to be compared (Table 5). For the benign samples hsa-miR-191-5p and hsa-miR-27a-3p were the most stable (Table 5). This indicates that the choice of references and the subsequent results largely depend upon the specific cohort composition and the available reference candidates.
To accommodate this discrepancy between the two sample groups, a combined set was analyzed and hsa-miR-23a-3p and hsa-miR-27a-3p were found to the most stable candidates across both sample types (Table 5). Specific recommendations for good stability value thresholds of most of the algorithms are not clearly communicated and different cut-offs are being used for the different algorithms [30, 34, 35].
For Normfinder a used cut-off stability value of references is 0.5-1. For geNorm 1-1.5 is used. For delta-Ct and BestKeeper, about 1 is common [30, 34, 35]. We found that stabilities for the four miRNAs (hsa-miR-23a-3p, hsa-miR-27a-3p, hsa-miR-126-3p and hsa-miR-191-5p) highlighted in this study all performed below these thresholds (Table 5). Curiously, BestKeeper seem to systematically contradict the three other algorithms. Only one miRNA (hsa-miR-486-5p) in the dataset barely pass a BestKeeper stability threshold of 1.5 (MAD = 1.432) (Table 5).
Interestingly neither hsa-miR-191-5p nor hsa-miR-126-3p was part of the top performing pair in the combined sample group, even though they both performed better in the HGSOC group compared to hsa-miR-27a-3p and better then hsa-miR-23a-3p in the benign group. Nonetheless, both hsa-miR-191-5p and hsa-miR-126-3p were determined to be stable and using these as references could probably also be a viable option.
As previously seen, BestKeeper evaluation of stability is largely in direct opposition to the other algorithms [21]. For example, when looking at stabilities of the HGSOC and benign groups combined, hsa-miR-23a-3p is determined to be the most stable miRNA by delta-Ct and Normfinder, and third most stable by geNorm. However, BestKeeper ranks this miRNA as number 9 out of 13 (Table 5). BestKeeper uses mean absolute deviation of the Cp values for each candidate and in doing so do not consider comparing with other miRNAs taking experimental and technical variations into account. Since it has not yet been determined which algorithms are superior and in which contexts, we implemented RefFinder to provide an overview and compare results. Though this approach has been used in a number of studies it has to our knowledge never been properly validated, emphasizing the need for more precise standardization of methods for miRNA quantifications [28, 36]. It is therefore imperative, that the results from the RefFinder Rank is validated.
In this study we validated the findings using two methods. We used the grouped Normfinder analysis which uses a weighted mean of two parameters inter- and intragroup variations. Adding the intergroup variation into the calculation considers that a given stable miRNA should be stable in both within and between the analyzed sample groups.
To our knowledge, there are no publicly available datasets that contain suitable miRNA RT-qPCR data, to validate our findings in external OC cohorts. Shapira et al. published a study based on miRNA profiles obtained by RT-qPCR on presurgical plasma samples from 42 women with serous EOC, 36 women diagnosed with a benign neoplasm, and 23 comparably age-matched women with no known pelvic mass, however no raw data is provided. Hsa-miR-126-3p was reported downregulated in plasma from 42 serous EOC patients compared with 23 controls [37].
Interestingly, Resnick et al. found hsa-miR-126-3p to be overexpressed in serum from 19 EOC patients compared to that of 11 healthy controls. These discrepancies might be explained by relatively low number of examined patients and differences in the cohort heterogeneity, as exact subtypes of OC being included are not precisely described. Moreover, the normalization strategies are different: Shapira et al. normalized the data by subtracting the mean of complete cases miRNAs expression values in each sample from all miRNAs in that sample, whereas Resnick et al. used two miRNAs (hsa-miR-142-3p and hsa-miR-16) as endogenous stable controls. However, the details on how these normalizers have been chosen were not provided [38].
Hsa-miR-23a-3p has previously been reported to be stable in blood and plasma as well as being unaffected by hemolysis [26]. Indeed, it is used in combination with miR-451, present in high amounts in red blood cells [39], to assess the degree of hemolysis in blood-based samples [26].
Interestingly, U6-snRNA that is often used as a stable endogenous reference for normalization [14, 15] was excluded from the stability analysis in our study. After quality control and filtering U6-snRNA was only represented in 44% HGSOC samples and only 23% of benign tumor samples. Of the 60 HGSOC samples, only 26 U6-snRNA Cp values were below 35 cycles and in the benign samples only 10 out of 48 were below 35. Furthermore, none of these values were below 31 cycles, making the use of U6-snRNA as an endogenous control problematic. It should be noted that in this study RT-qPCR were performed on a limited number of selected targets miRNAs and U6-snRNA.
In this study we found 13 miRNA candidates expressed in all samples. To validate ubiquitous expression of the most stable candidates we compared the results with a previous study performed in our research group (Table S8). In that study, miRNAs were investigated in plasma from two different cohorts each consisting of 95 malignant and 95 benign pelvic mass patients. Here, only hsa-miR-191-5p were present in all samples in both cohorts (n = 365), hsa-miR-126-3p was missing in 1 sample (0.27%), hsa-miR-27a-3p was missing in two samples (0.55%) and hsa-miR-23a-3p was missing in 8 samples (2.19%). Due to the ratio of missing values being very low we assess these to be negligeable.
Additional research needs to be performed to further validate our findings and to explore other suitable reference miRNAs.
As discussed previously, RT-qPCR results are heavily affected by the data composition, robust and trustworthy data require prober data cleaning and quality control. Few papers report their methodology for data evaluation and data point exclusion criteria and if quality control like Spike-in or melt curves are reported, often very little information is provided on the specific use of these [40,41,42]. Here, we found that using a cut-off for Cp values of 35 cycles, 19.8% (1065) of our dataset Cp values was removed. Additionally, 8.9% (481) of remaining Cp values needed manual evaluation of melt curves and 24.7% of these (119) were excluded from the further analysis. In summary, 22% of the data was excluded in this initial validation. Many uses a Cp cut-off, and 35 cycles is a very common number to use in this respect (Fig. 1). However, at least regarding the LightCycler software, not many describe their approach to handling Cp values that is marked by the software to be of uncertain quality [41].
In this project we used the LightCycler 480 (Roche) in combination with the official LightCycler®480 software for RT-qPCR. This software allowed us to evaluate melt-curves for Cp values that were marked by either the Cp table or the Tm table as of uncertain quality.
It is worth noting that this study is limited by the miRNA targets selected for the RT-qPCR panel used. We are here presenting a workflow for data preprocessing and stability analysis and support and emphasize the notion that these or similar steps must be performed for each individual study [13]. We propose to include a set of candidate references in every RT-qPCR panel and to determine which of these are most stable and ubiquitously expressed in the given sample set. In this respect, the four miRNAs mentioned in here could be included in such a candidate list designed for blood plasma analysis as they in our study are proven stable.
Conclusion
In this study, we provide the basis for further investigations of stability analysis and exclusion principles when performing miRNA expression studies by RT-qPCR. We found that hsa-miR-126-3p and hsa-miR-23a-3p were most stable among tested miRNAs in HGSOC patients, whereas in the benign samples, hsa-miR-191-5p and hsa-miR-27a-3p were the most stable. When combining the two groups sets, hsa-miR-23a-3p and hsa-miR-27a-3p were found as most suitable endogenous references. The choices were based on the stability rankings provided by RefFinder and discrepancies between various algorithms were observed. U6 detection failed in many of the samples and were found to be unreliable as stable reference. Moreover, to achieve consensus between miRNAs studies across various research groups, access to raw data and more detailed reports regarding raw data handling and pre-processing in terms of inclusion should be provided to enable reliable validation across studies.
Data Availability
All relevant data are within the paper and its Supporting Information files.
References
Petersen PHD, Lopacinska-Joergensen J, Høgdall CK, Høgdall EV (2023) RefSeeker - RefFinder R Package. https://github.com/Hannibal83dk/RefSeeker
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Momenimovahed Z, Tiznobaik A, Taheri S, Salehiniya H (2019) Ovarian cancer in the world: epidemiology and risk factors. Int J Womens Health 11:287–299. https://doi.org/10.2147/IJWH.S197604
Torre LA, Trabert B, DeSantis CE, Miller KD, Samimi G, Runowicz CD, Gaudet MM, Jemal A, Siegel RL (2018) Ovarian cancer statistics, 2018. CA Cancer J Clin 68:284–296. https://doi.org/10.3322/caac.21456
Bowtell DD, Böhm S, Ahmed AA, Aspuria P-J, Bast RC, Beral V, Berek JS, Birrer MJ, Blagden S, Bookman MA, Brenton JD, Chiappinelli KB, Martins FC, Coukos G, Drapkin R, Edmondson R, Fotopoulou C, Gabra H, Galon J, Gourley C, Heong V, Huntsman DG, Iwanicki M, Karlan BY, Kaye A, Lengyel E, Levine DA, Lu KH, McNeish IA, Menon U, Narod SA, Nelson BH, Nephew KP, Pharoah P, Powell DJ, Ramos P, Romero IL, Scott CL, Sood AK, Stronach EA, Balkwill FR (2015) Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat Rev Cancer 15:668–679. https://doi.org/10.1038/nrc4019
Saral MA, Tuncer SB, Odemis DA, Erdogan OS, Erciyas SK, Saip P, Ozel S, Yazici H (2022) New biomarkers in peripheral blood of patients with ovarian cancer: high expression levels of miR-16-5p, miR-17-5p, and miR-638. Arch Gynecol Obstet 305:193–201. https://doi.org/10.1007/s00404-021-06138-z
Prahm KP, Høgdall C, Karlsen MA, Christensen IJ, Novotny GW, Høgdall E (2018) Identification and validation of potential prognostic and predictive miRNAs of epithelial ovarian cancer. PLoS ONE 13:e0207319. https://doi.org/10.1371/journal.pone.0207319
Sanders I, Holdenrieder S, Walgenbach-Brünagel G, von Ruecker A, Kristiansen G, Müller SC, Ellinger J (2012) Evaluation of reference genes for the analysis of serum miRNA in patients with prostate cancer, bladder cancer and renal cell carcinoma. Int J Urol 19:1017–1025. https://doi.org/10.1111/j.1442-2042.2012.03082.x
Tiberio P, Callari M, Angeloni V, Daidone MG, Appierto V (2015) Challenges in using circulating miRNAs as cancer biomarkers. Biomed Res Int 2015. https://doi.org/10.1155/2015/731479
Huggett J, Dheda K, Bustin S, Zumla A (2005) Real-time RT-PCR normalisation; strategies and considerations. Genes Immun 6:279–284. https://doi.org/10.1038/sj.gene.6364190
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Das MK, Andreassen R, Haugen TB, Furu K (2016) Identification of endogenous controls for use in miRNA quantification in human cancer cell lines. Cancer Genomics and Proteomics 13:63–68
Kaur J, Saul D, Doolittle ML, Rowsey JL, Vos SJ, Farr JN, Khosla S, Monroe DG (2022) Identification of a suitable endogenous control miRNA in bone aging and senescence. Gene 835:146642. https://doi.org/10.1016/j.gene.2022.146642
Zhu Z, Chen Z, Wang M, Zhang M, Chen Y, Yang X, Zhou C, Liu Y, Hong L, Zhang L (2022) Detection of plasma exosomal miRNA-205 as a biomarker for early diagnosis and an adjuvant indicator of ovarian cancer staging. J Ovarian Res 15:1–10. https://doi.org/10.1186/s13048-022-00961-x
Talaat A, Helmy MA, Saadawy SF (2022) Evaluation of miRNA-21 and CA-125 as a promising diagnostic biomarker in patients with ovarian cancer. Egypt J Med Hum Genet 23. https://doi.org/10.1186/s43042-022-00342-5
Xiang M, Zeng Y, Yang R, Xu H, Chen Z, Zhong J, Xie H, Xu Y, Zeng X (2014) U6 is not a suitable endogenous control for the quantification of circulating microRNAs. Biochem Biophys Res Commun 454:210–214. https://doi.org/10.1016/j.bbrc.2014.10.064
Andersen CL, Jensen JL, Ørntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64:5245–5250. https://doi.org/10.1158/0008-5472.CAN-04-0496
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. https://doi.org/10.1186/gb-2002-3-7-research0034. 3:
Silver N, Best S, Jiang J, Thein SL (2006) Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol Biol 7:1–9. https://doi.org/10.1186/1471-2199-7-33
Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP (2004) Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper - Excel-based tool using pair-wise correlations. Biotechnol Lett 26:509–515. https://doi.org/10.1023/B:BILE.0000019559.84305.47
Petersen PHD, Lopacinska-Jørgensen J, Oliveira DVNP, Høgdall CK, Høgdall EV (2022) miRNA expression in Ovarian Cancer in Fresh Frozen, Formalin-fixed paraffin-embedded and plasma samples. Vivo (Brooklyn) 36:1591–1602. https://doi.org/10.21873/invivo.12869
Xie F, Xiao P, Chen D, Xu L, Zhang B (2012) miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol Biol 80:75–84. https://doi.org/10.1007/s11103-012-9885-2
Lopacinska-Jørgensen J, Petersen PHD, Oliveira DVNP, Høgdall CK, Høgdall EV (2023) Strategies for data normalization and missing data imputation and consequences for potential diagnostic microRNA biomarkers in epithelial ovarian cancer. PLoS ONE 18:e0282576. https://doi.org/10.1371/journal.pone.0282576
R Core Team (2022) R: A language and environment for statistical computing. http://www.r-project.org
Tukey JW (1977) Exploratory data analysis. Pearson Education
Blondal T, Jensby Nielsen S, Baker A, Andreasen D, Mouritzen P, Wrang Teilum M, Dahlsveen IK (2013) Assessing sample and miRNA profile quality in serum and plasma or other biofluids. Methods 59:S1–S6. https://doi.org/10.1016/j.ymeth.2012.09.015
Sundaram VK, Sampathkumar NK, Massaad C, Grenier J (2019) Optimal use of statistical methods to validate reference gene stability in longitudinal studies. PLoS ONE 14:1–18. https://doi.org/10.1371/journal.pone.0219440
De Spiegelaere W, Dern-Wieloch J, Weigel R, Schumacher V, Schorle H, Nettersheim D, Bergmann M, Brehm R, Kliesch S, Vandekerckhove L, Fink C (2015) Reference gene validation for RT-qPCR, a note on different available software packages. PLoS ONE 10:1–13. https://doi.org/10.1371/journal.pone.0122515
Grätz C, Bui MLU, Thaqi G, Kirchner B, Loewe RP, Pfaffl MW (2022) Obtaining Reliable RT-qPCR results in Molecular Diagnostics—MIQE Goals and Pitfalls for Transcriptional Biomarker Discovery. https://doi.org/10.3390/life12030386. Life 12:
Faraldi M, Gomarasca M, Sansoni V, Perego S, Banfi G, Lombardi G (2019) Normalization strategies differently affect circulating miRNA profile associated with the training status. Sci Rep 9:1–13. https://doi.org/10.1038/s41598-019-38505-x
Morata-Tarifa C, Picon-Ruiz M, Griñan-Lison C, Boulaiz H, Perán M, Garcia MA, Marchal JA (2017) Validation of suitable normalizers for miR expression patterns analysis covering tumour heterogeneity. Sci Rep 7:39782. https://doi.org/10.1038/srep39782
ÖZCAN K, PEKTAŞ AN, BERK Ş (2022) Identification and validation of reference genes for RT-qPCR normalization in Nauphoeta cinerea (Olivier, 1789) (Blattodea, Blaberidae). Bilge Int J Sci Technol Res 6:62–72. https://doi.org/10.30516/bilgesci.1067570
Bignotti E, Calza S, Tassi RA, Zanotti L, Bandiera E, Sartori E, Odicino FE, Ravaggi A, Todeschini P, Romani C (2016) Identification of stably expressed reference small non-coding RNAs for microRNA quantification in high-grade serous ovarian carcinoma tissues. J Cell Mol Med 20:2341–2348. https://doi.org/10.1111/jcmm.12927
Linardić M, Braybrook SA (2021) Identification and selection of optimal reference genes for qPCR-based gene expression analysis in Fucus distichus under various abiotic stresses. PLoS ONE 16:1–19. https://doi.org/10.1371/journal.pone.0233249
Wang H, Zhang X, Liu Q, Liu X, Ding S (2017) Selection and evaluation of new reference genes for RT-qPCR analysis in Epinephelus akaara based on transcriptome data. PLoS ONE 12:1–18. https://doi.org/10.1371/journal.pone.0171646
Rice J, Roberts H, Rai SN, Galandiuk S (2015) Housekeeping genes for studies of plasma microRNA: a need for more precise standardization. Surg (United States) 158:1345–1351. https://doi.org/10.1016/j.surg.2015.04.025
Shapira I, Oswald M, Lovecchio J, Khalili H, Menzin A, Whyte J, Dos Santos L, Liang S, Bhuiya T, Keogh M, Mason C, Sultan K, Budman D, Gregersen PK, Lee AT (2014) Circulating biomarkers for detection of ovarian cancer and predicting cancer outcomes. Br J Cancer 110:976–983. https://doi.org/10.1038/bjc.2013.795
Resnick KE, Alder H, Hagan JP, Richardson DL, Croce CM, Cohn DE (2009) The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol Oncol 112:55–59. https://doi.org/10.1016/j.ygyno.2008.08.036
Rasmussen KD, Simmini S, Abreu-Goodger C, Bartonicek N, Di Giacomo M, Bilbao-Cortes D, Horos R, Von Lindern M, Enright AJ, O’Carroll D (2010) The miR-144/451 locus is required for erythroid homeostasis. J Exp Med 207:1351–1358. https://doi.org/10.1084/jem.20100458
Wu X, Yang J, Yu L, Long D (2018) Plasma miRNA-223 correlates with risk, inflammatory markers as well as prognosis in sepsis patients. Med (United States) 97:1–6. https://doi.org/10.1097/MD.0000000000011352
Géczi D, Nagy B, Szilágyi M, Penyige A, Klekner Á, Jenei A, Virga J, Birkó Z (2021) Analysis of circulating mirna profile in plasma samples of glioblastoma patients. Int J Mol Sci 22. https://doi.org/10.3390/ijms22105058
Zaporozhchenko IA, Morozkin ES, Ponomaryova AA, Rykova EY, Cherdyntseva NV, Zheravin AA, Pashkovskaya OA, Pokushalov EA, Vlassov VV, Laktionov PP (2018) Profiling of 179 miRNA expression in blood plasma of Lung Cancer Patients and Cancer-Free individuals. Sci Rep 8:1–13. https://doi.org/10.1038/s41598-018-24769-2
Lopacinska-Joergensen J, Oliveira DVNP, Hoegdall CK, Hoegdall EV (2022) Identification of stably expressed reference microRNAs in epithelial ovarian Cancer. In Vivo 36:1059–1066. https://doi.org/10.21873/invivo.12803
Yokoi A, Matsuzaki J, Yamamoto Y, Yoneoka Y, Takahashi K, Shimizu H, Uehara T, Ishikawa M, Ikeda S ichi, Sonoda T, Kawauchi J, Takizawa S, Aoki Y, Niida S, Sakamoto H, Kato K, Kato T, Ochiya T (2018) Integrated extracellular microRNA profiling for ovarian cancer screening. Nat Commun 9:2–6. https://doi.org/10.1038/s41467-018-06434-4
Jiang W, Pan JJ, Deng YH, Liang MR, Yao LH (2017) Down-regulated serum microRNA-101 is associated with aggressive progression and poor prognosis of cervical cancer. J Gynecol Oncol 28:1–10. https://doi.org/10.3802/jgo.2017.28.e75
Lopacinska-Jørgensen J, Oliveira DVNP, Novotny GW, Høgdall CK, Høgdall EV (2021) Integrated microRNA and mRNA signatures associated with overall survival in epithelial ovarian cancer. PLoS ONE 16:1–15. https://doi.org/10.1371/journal.pone.0255142
Prahm KP, Høgdall CK, Karlsen MA, Christensen IJ, Novotny GW, Høgdall E (2021) MicroRNA characteristics in epithelial ovarian cancer. PLoS ONE 16:1–18. https://doi.org/10.1371/journal.pone.0252401
Zhong C, Dong Y, Zhang Q, Yuan C, Duan S (2022) Aberrant expression of miR-1301 in Human Cancer. Front Oncol 11:1–13. https://doi.org/10.3389/fonc.2021.789626
Tang W, Jiang Y, Mu X, Xu L, Cheng W, Wang X (2014) MiR-135a functions as a tumor suppressor in epithelial ovarian cancer and regulates HOXA10 expression. Cell Signal 26:1420–1426. https://doi.org/10.1016/j.cellsig.2014.03.002
Fukagawa S, Miyata K, Yotsumoto F, Kiyoshima C, Nam SO, Anan H, Katsuda T, Miyahara D, Murata M, Yagi H, Shirota K, Yasunaga S, Kato K, Miyamoto S (2017) MicroRNA-135a-3p as a promising biomarker and nucleic acid therapeutic agent for ovarian cancer. Cancer Sci 108:886–896. https://doi.org/10.1111/cas.13210
Shi M, Mu Y, Zhang H, Liu M, Wan J, Qin X, Li C (2018) MicroRNA-200 and microRNA-30 family as prognostic molecular signatures in ovarian cancer: a meta-analysis. Med (United States) 97:1–9. https://doi.org/10.1097/MD.0000000000011505
Sharma PC, Gupta A (2020) MicroRNAs: potential biomarkers for diagnosis and prognosis of different cancers. Transl Cancer Res 9:5798–5818. https://doi.org/10.21037/tcr-20-1294
Tokumaru Y, Asaoka M, Oshi M, Katsuta E, Yan L, Narayanan S, Sugito N, Matsuhashi N, Futamura M, Akao Y, Yoshida K, Takabe K (2020) High expression of microRNA-143 is associated with favorable tumor immune microenvironment and better survival in estrogen receptor positive breast cancer. Int J Mol Sci 21:1–17. https://doi.org/10.3390/ijms21093213
Wilczyński M, Żytko E, Szymańska B, Dzieniecka M, Nowak M, Danielska J, Stachowiak G, Wilczyński JR (2017) Expression of miR-146a in patients with ovarian cancer and its clinical significance. Oncol Lett 14:3207–3214. https://doi.org/10.3892/ol.2017.6477
Oliveira DNP, Carlsen AL, Heegaard NHH, Prahm KP, Christensen IJ, Høgdall CK, Høgdall EV (2019) Diagnostic plasma miRNA-profiles for ovarian cancer in patients with pelvic mass. PLoS ONE 14:1–15. https://doi.org/10.1371/journal.pone.0225249
Wang Q, Ye B, Wang P, Yao F, Zhang C, Yu G (2019) Overview of microRNA-199a regulation in cancer. Cancer Manag Res 11:10327–10335. https://doi.org/10.2147/CMAR.S231971
Vilming Elgaaen B, Olstad OK, Haug KBF, Brusletto B, Sandvik L, Staff AC, Gautvik KM, Davidson B (2014) Global miRNA expression analysis of serous and clear cell ovarian carcinomas identifies differentially expressed miRNAs including miR-200c-3p as a prognostic marker. BMC Cancer 14:1–13. https://doi.org/10.1186/1471-2407-14-80
Sujamol S, Vimina ER, Krishnakumar U (2021) Improving recurrence prediction accuracy of Ovarian Cancer using multi-phase feature selection methodology. Appl Artif Intell 35:206–226. https://doi.org/10.1080/08839514.2020.1854988
LI R, WU H, JIANG H, WANG Q, DOU Z, MA H, YUAN YANS, YANG C, KONG N B (2020) FBLN5 is targeted by microRNA-27a-3p and suppresses tumorigenesis and progression in high-grade serous ovarian carcinoma. Oncol Rep 44:2143–2151. https://doi.org/10.3892/or.2020.7749
Ferlay J, Colombet M, Soerjomataram I, Mathers C, Parkin DM, Piñeros M, Znaor A, Bray F (2019) Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int J Cancer 144:1941–1953. https://doi.org/10.1002/ijc.31937
Acknowledgements
We are thankful to the Danish CancerBiobank (Bio- and GenomeBank, Denmark - RBGB) and the Danish Gynecologic Cancer Database (DGCD) for making samples and data available for use in this study. This work was founded by: The Mermaid Foundation, URL: http://www.mermaidprojektet.dk/ (PHDP, JLJ, CKH and EVH received the funding), Danish Cancer Research Foundation: URL:http://www.dansk-kraeftforsknings-fond.dk/ (EVH received the funding), and Herlev Hospital Research Council, URL: https://www.herlevhospital.dk/forskning/ (EVH received the funding).
Funding
This study was funded by the The Mermaid Foundation, URL: http://www.mermaidprojektet.dk/ (PHDP, JLJ, CKH and EVH received the funding), Danish Cancer Research Foundation, URL:http://www.dansk-kraeftforsknings-fond.dk/ (EVH received the funding) as well as Herlev Hospital Research Council https://www.herlevhospital.dk/forskning/ (EVH received the funding).
Open access funding provided by Royal Library, Copenhagen University Library
Author information
Authors and Affiliations
Contributions
J.L.J., E.V.H., C.K.H., P.H.D.P. designed the study. E.V.H. and C.K.H. provided the materials and data. P.H.D.P. and J.L.J. performed the data acquisition, processing, and analyses. All authors interpreted the data. P.H.D.P. and J.L.J. wrote the manuscript. All authors have reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was approved by the Danish National Committee for Research Ethics (approval codes KF01-227/03 and KF01-143/04) and all patients have provided written informed consent. The study followed the guideline laid out by the Declaration of Helsinki.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Petersen, P.H., Lopacinska-Jørgensen, J., Høgdall, C.K. et al. Identification of stably expressed microRNAs in plasma from high-grade serous ovarian carcinoma and benign tumor patients. Mol Biol Rep 50, 10235–10247 (2023). https://doi.org/10.1007/s11033-023-08795-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08795-6