Abstract
Background
Host genetic characteristics and environmental factors interactions may play a crucial role in cervical carcinogenesis. We investigated the impact of functional genetic variants of four xenobiotic-metabolizing genes (AhR, CYP1A1, GSTM1, and GSTT1) on cervical cancer development in Tunisian women.
Methods
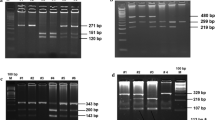
The AhR gene polymorphism was analyzed using the tetra-primer ARMS-PCR, whereas the CYP1A1 polymorphism genotypes were identified by PCR-RFLP. A multiplex ligation-dependent polymerase chain reaction approach was applied for the analysis of GSTM1 and GSTT1 polymorphisms.
Results
The homozygous A/A genotype of the AhR gene (rs2066853) and the heterozygous T/C genotype of the CYP1A1 SNP (CYP1A1-MspI) appeared to be associated with an increased risk of cervical tumorigenesis (ORa = 2.81; ORa = 5.52, respectively). Furthermore, a significantly increased risk of cervical cancer was associated with the GSTT1 null genotype (ORa = 2.65). However, the null GSTM1 genotype showed any significant association with the risk of cervical cancer compared to the wild genotype (ORa = 1.18; p = 0.784). Considering the combined effect, we noted a significantly higher association with cancer risk for individuals with at least two high-risk genotypes of CYP1A1/GSTT1 (ORa = 4.2), individuals with at least two high-risk genotypes of CYP1A1/GSTT1/AhR (ORa = 11.3) and individuals with at least two high-risk genotypes of CYP1A1/GSTM1/GSTT1/AhR exploitation low-risk genotype as a reference.
Conclusion
This study indicated that the single-gene contribution and the combined effect of xenobiotic-metabolizing gene polymorphisms (AhR, CYP1A1-MspI, GSTM1, and GSTT1) may have a considerable association with increased cervical cancer risk.
Similar content being viewed by others
Data Availability
Not applicable.
Abbreviations
- SNP:
-
Single Nucleotide Polymorphism.
- AhR:
-
Aryl hydrocarbon Receptor.
- CYP1A1:
-
Cytochrome P450 1A1.
- GSTM1:
-
Glutathione S-transferase Mu 1.
- GSTT1:
-
Glutathione S-transferase Theta 1.
- TAD:
-
Transactivation Domain.
- TBP:
-
TATA-binding Protein.
- WHO:
-
World Health Organization.
- FIGO:
-
International Federation of Gynecology and Obstetrics.
- SIL:
-
Squamous Intraepithelial lesion.
- LSIL:
-
Low grade Squamous Intraepithelial lesion.
- HSIL:
-
High grade Squamous Intraepithelial lesion.
- ICC:
-
Invasive Cervical Cancer.
- SCC:
-
Squamous Cell Carcinoma.
- AC:
-
Cervical Adenocarcinoma.
- HPV:
-
Human Papillomavirus.
- PAH:
-
Polycyclic aromatic hydrocarbon.
- Pap test:
-
Papanicolaou test.
- PCR:
-
Polymerase Chain Reaction.
- ARMS-PCR:
-
Amplification Refractory Mutation System.
- PCR-RFLP:
-
PCR-Restriction Fragment Length Polymorphism.
- SPSS:
-
Statistical Package for the Social Sciences.
- ORa :
-
Adjusted Odd Ratio.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. C A Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Ferlay J, Ervik M, Lam F, Colombet M, Mery L, PiñerosM, Znaor A, Soerjomataram I, Bray F. Global Cancer Observatory: Cancer Today.Lyon, France: International Agency for Research on Cancer 2020. Available from: https://gco.iarc.fr/today, accessed [05 October 2022].
Sany SB, Hashim R, Rezayi M, Rahman MA, Razavizadeh BB, Abouzari-lotf E (2015) Karlen D. I. Integrated ecological risk assessment of dioxin compounds. Environ Sci Pollut Res Int 22:11193–11208. https://doi.org/10.1007/s11356-015-4511-x
Ruiz-Vera T, Pruneda-Alvarez LG, Ochoa-Martinez AC, Ramirez-Garcia-Luna JL, Pierdant-Perez M, Gordillo-Moscoso AA, Perez-Vazquez FJ (2015) Perez-Maldonado I. N. Assessment of vascular function in Mexican women exposed to polycyclic aromatic hydrocarbons from wood smoke. Environ Toxicol Pharmacol 40:423–429. https://doi.org/10.1016/j.etap.2015.07.014
Patrizi B, Siciliani de Cumis M, Viciani S, D’Amato F Dioxin and Related Compound Detection: Perspectives for Optical Monitoring.International journal of molecular sciences2019: 20(11):2671. https://doi.org/10.3390/ijms20112671
Kerkvliet N (2002) I. Recent advances in understanding the mechanisms of TCDD immunotoxicity. Int Immunopharmacol 2:277–291. https://doi.org/10.1016/s1567-5769(01)00179-5
Furue M, Ishii Y, Tsukimori K, Tsuji G Aryl Hydrocarbon Receptor and Dioxin-Related Health Hazards-Lessons from Yusho.International journal of molecular sciences2021: 22(2):708. https://doi.org/10.3390/ijms22020708
Vogel C, Van Winkle LS, Esser C, Haarmann-Stemmann T (2020) The aryl hydrocarbon receptor as a target of environmental stressors - Implications for pollution mediated stress and inflammatory responses. Redox Biol 34:101530. https://doi.org/10.1016/j.redox.2020.101530
Li H, Luo L, Wang D, Duan J, Zhang R (2020) Lack of association between multiple polymorphisms in aryl hydrocarbon receptor (AhR) gene and cancer susceptibility. Environ Health Prev Med 25:79. https://doi.org/10.1186/s12199-020-00907-z
Hayashi S, Kiyohara C (1995) Polymorphisms of human Ah receptor gene are not involved in lung cancer. Pharmacogenetics 5:151–158. https://doi.org/10.1097/00008571-199506000-00003
Luo C, Zou P, Ji G, Gu A, Zhao C (2013) The aryl hydrocarbon receptor (AhR) 1661G > A polymorphism in human cancer: a meta-analysis. Gene 513(1):225–230. https://doi.org/10.1016/j.gene.2012.09.050
Wong JM, Harper PA, Meyer UA, Bock KW, Morike K, Lagueux J, Ayotte P, Tyndale RF, Sellers EM, Manchester DK (2001) Okey A. B. Ethnic variability in the allelic distribution of human aryl hydrocarbon receptor codon 554 and assessment of variant receptor function in vitro. Pharmacogenetics 11:85–94. https://doi.org/10.1097/00008571-200102000-00010
Mescher M, Haarmann-Stemmann T (2018) Modulation of CYP1A1 metabolism: From adverse health effects to chemoprevention and therapeutic options. Pharmacol Ther 187:71–87. https://doi.org/10.1016/j.pharmthera.2018.02.012
Chatterjee A, Gupta S (2018) The multifaceted role of glutathione S-transferases in cancer. Cancer Lett 433:33–42. https://doi.org/10.1016/j.canlet.2018.06.028
Hirata H, Hinoda Y, Okayama N, Suehiro Y, Kawamoto K, Kikuno N, Rabban JT, Chen LM, Dahiya R (2007) CYP1A1, SULT1A1, and SULT1E1 polymorphisms are risk factors for endometrial cancer susceptibility. Cancer 112(9):1964–1973. https://doi.org/10.1002/cncr.23392
Rajagopal T, Seshachalam A, Rathnam KK, Jothi A, Talluri S, Venkatabalasubramanian S, Dunna NR. Impact of xenobiotic-metabolizing gene polymorphisms on breast cancer risk in South Indian women.Breast cancer research and treatment2021: 186(3):823–837. https://doi.org/10.1007/s10549-020-06028-z
Liu Y, Xu LZ (2012) Meta-analysis of association between GSTM1gene polymorphism and cervical cancer. Asian Pac J Top Med 5:480–484. https://doi.org/10.1016/S1995-7645(12)60083-2
Leme CVD, Raposo LS, Ruiz MT, Galbiatti A, Maniglia J (2010) GSTM1 and GSTT1 genes analysis in head and neck cancer patients. Res Assoc Med Bras 56:299–303. https://pubmed.ncbi.nlm.nih.gov/20676536/
Zivkovic M, Bubic M, Kolakovic A, Dekleva M, Stankovic G, Stankovic A, Djuric T. The association of glutathione S-transferase T1 and M1 deletions with myocardial infarction.Free radical research2021: 55(3):267–274. https://doi.org/10.1080/10715762.2021.1931166
Morris K (2013) Revising the Declaration of Helsinki. The lancet 381(9881):1889–1890. https://doi.org/10.1016/s0140-6736(13)60951-4
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16:1215–1218. https://doi.org/10.1093/nar/16.3.1215
Baily LR, Roodi N, Verrier CS, Yee CJ, Dupont WD, Parl FF (1998) Breast cancer and CYP1A1, GSTM1 and GSTT1 polymorphisms: evidence of a lack of association in Caucasians and African Americans. Cancer Res 58:65–70. https://pubmed.ncbi.nlm.nih.gov/9426059/
Mittal B, Bhattacharya S (2013) Associations of CYP1A1, GSTM1 and GSTT1 polymorphisms with lung cancer susceptibility in a northern Indian population. Asian Pac J Cancer Prey 14:3345–3349. https://doi.org/10.7314/apjcp.2013.14.5.3345
Babekir EA, Abdelateif NM, Adelrahim SO, Hassan R, Ibrahim IK (2019) GSTM1 and GSTT1 Polymorphisms and Susceptibility to Acute Myeloid Leukemia: A Case-Control Study of the Sudanese Population. A P J C B 4:7–10. https://doi.org/10.31557/APJCB.2019.4.1.7
Al-Koofee D. A., Mobarak S. M. PRIMER1: A network service for tetra-arms PCR primer design based on well-known dbSNP. Research Journal of Pharmacy and Technology 2018:11(8): 3633–3637. https://doi.org/10.5958/0974-360X.2018.00669.8
Patel H, Jeve YB, Sherman SM, Moss EL. (2016) Knowledge of human papillomavirus and the human papillomavirus vaccine in European adolescents: a systematic review. Sex Transm Infect 92(6):474–479. https://doi.org/10.1136/sextrans-2015-052341
I. A. R. C. Monographs on the evaluation of carcinogenic risks to humans. Polycyclic aromatic compound. Part 1.Chemical, environmental and experimental data:32. https://www.ncbi.nlm.nih.gov/books/NBK419306/?report=reader
Nebert DW. (2017). Aryl hydrocarbon receptor (AHR): “pioneer member” of the basic-helix/loop/helix per-Arnt-sim (bHLH/PAS) family of “sensors” of foreign and endogenous signals. Prog Lipid Res 67:38–57. https://doi.org/10.1016/j.plipres.2017.06.001
Sangrajrang S, Sato Y, Sakamoto H, Ohnami S, Laird NM, Khuhaprema T, Brennan P (2009) Boffetta P., Yoshida T.Genetic polymorphisms of estrogen metabolizing enzyme and breast cancer risk in Thai women. Int J Cancer 125:837–843. https://doi.org/10.1002/ijc.24434
Gu A, Ji G, Jiang T, Lu A, You Y, Liu N, Luo C, Yan W, Zhao P (2012) Contributions of aryl hydrocarbon receptor genetic variants to the risk of glioma and PAH-DNA adducts. Toxicol Sci 128:357–364. https://doi.org/10.1093/toxsci/kfs158
Cheng T, Gamage S, Hewage D, Lu CT, Aktar S, Gopalan V, Lam A (2022) K. AHR gene expression and the polymorphism rs2066853 are associated with clinicopathological parameters in colorectal carcinoma. Hum Pathol 122:50–59. https://doi.org/10.1016/j.humpath.2022.02.001
Cotterchio M, Boucher BA, Manno M, Gallinger S, Okey AB, Harper PA (2008) Red meat intake, doneness, polymorphisms in genes that encode carcinogen-metabolizing enzymes, and colorectal cancer risk. Cancer Epidemiol Biomarkers Prev 11:3098–3107. https://doi.org/10.1158/1055-9965.EPI-08-0341
Chen D, Tian T, Wang H, Liu H, Hu Z, Wang Y, Liu Y, Ma H, Fan W, Miao R, Sun W, Wang Y, Qian J, Jin L, Wei Q, Shen H, Huang W, Lu D (2009) Association of human aryl hydrocarbon receptor gene polymorphisms with risk of lung cancer among cigarette smokers in a Chinese population. Pharmacogenet Genomics 19:25–34. https://doi.org/10.1097/FPC.0b013e328316d8d8
Long JR, Egan KM, Dunning L, Shu XO, Cai Q, Cai H, Zheng Y (2006) Population-based case-control study of AhR (aryl hydrocarbon receptor) and CYP1A2 polymorphisms and breast cancer risk. Pharmacogenet Genomics 16:237–243. https://doi.org/10.1097/01.fpc.0000189803.34339.ed
Budhwar S, Bahl C, Sharma S, Singh N, Behera D (2018) Role of sequence variations in AhR gene towards modulating smoking-induced lung cancer susceptibility in North Indian population: a multiple interaction analysis. Curr Genomics 19:313–326. https://doi.org/10.2174/1389202918666170915160606
Zhang DS, Lin GF, Ma QW, Shen JH (2002) Non-association of aryl hydrocarbon receptor genotypes with susceptibility to bladder cancer in Shanghai population. Acta Pharmacol Sin 23:188–192. https://pubmed.ncbi.nlm.nih.gov/11866883/
Gao M, Li Y, Xue X, Long J, Chen L, Shah W, Kong Y, Impact(2014) of AhR, CYP1A1 and GSTM1 genetic polymorphisms on TP53 R273G mutations in individuals exposed to polycyclic aromatic hydrocarbons. Asian Pac. J. Cancer Prev. : 15: 2699–2705. https://doi.org/10.7314/APJCP.2014.15.6.2699
Ramadoss P, Marcus C, Perdew GH (2005) Role of the aryl hydrocarbon receptor in drug metabolism. Expert Drug Metab Toxicol 1:9–21. https://doi.org/10.1517/17425255.1.1.9
Aftabi Y, Colagar AH, Mehrnejad F. An in-silico approach to investigate the source of the controversial interpretations about the phenotypic results of the human AhR-gene G1661A polymorphism.J. Theor. Biol.2016: 393:1–15. https://doi.org/10.1016/j.jtbi.2016.01.001
Smart J, Daly AK (2000) Variation in induced CYP1A1 levels: Relationship to CYP1A1, Ah receptor and GSTM1 polymorphisms. Pharmacogenetics 10:11–24. https://doi.org/10.1097/00008571-200002000-00003
Shiizaki K., Kawanishi M., Yagi T. Modulation of benzo[a]pyrene–DNA adduct formation by CYP1 inducer and inhibitor. Genes and Environ 2017: 39: 14. https://doi.org/10.1186/s41021-017-0076-x
Ding G., Xu W., Liu H., Zhang M., Huang Q., Liao Z. CYP1A1MspI polymorphism is associated with prostate cancer susceptibility: evidence from a meta-analysis. Mol. Biol. Rep. 2013: 40: 3483-3491 https://doi.org/10.1007/s11033-012-2423-0
Xia L., Gao J., Liu Y., Wu K. Significant association between CYP1A1 T3801C polymorphism and cervical neoplasia risk: a systematic review and meta-analysis. Tumor Biol. 2013: 34: 223-230. https://doi.org/10.1007/s13277-012-0542-9
Ding B., Sun W., Han S., Cai Y., Ren M., Shen Y. Cytochrome P450 1A1 gene polymorphisms and cervical cancer risk: a systematic review and meta-analysis. Medicine 2018: 97(13): e0210. https://doi.org/10.1097/MD.0000000000010210
Juárez-Cedillo T., Vallejo M., Fragoso J. M., Hernández-Hernandez D. M., Rodríguez-Pérez J. M., Sánchez- García S., del Carmen García-Peña M., García-Carrancá A., Mohar-Betancourt A., Granados J., Vargas-Alarcón G. The risk of developing cervical cancer in Mexican women is associated to CYP1A1MspI polymorphism. Eur. J. Cancer 2007: 43(10): 1590-1595. https://doi.org/10.1016/j.ejca.2007.03.025
Joseph T., Chacko P., Wesley R., Jayaprakash P. G., James F. V., Pillai M. R. Germline genetic polymorphisms of CYP1A1, GSTM1 and GSTT1 genes in Indian cervical cancer: associations with tumour progression, age and human papillomavirus infection. Gynecologic oncology 2006: 101(3): 411-417. https://doi.org/10.1016/j.ygyno.2005.10.033
Abbas M., Srivastava K., Imran M., Banerjee M. Association of CYP1A1 gene variants rs4646903 (T > C) and rs1048943 (A > G) with cervical cancer in a North Indian population. Eur. J. Obstet. Gynecol. Reprod. Biol. 2014: 176: 68-74. https://doi.org/10.1016/j.ejogrb.2014.02.036
Von Keyserling H., Bergmann T., Schuetz M., Schiller U., Stanke J., Hoffmann C., Schneider A., Lahrach H., Dahl A., Kaufmann A. M. Analysis of 4 single nucleotide polymorphisms in relation to cervical dysplasia and cancer development using a high-though put ligation-detection reaction procedure. Int. J. Gynecol. Cancer 2011: 21: 1664- 1671. https://doi.org/10.1097/IGC.0b013e31822b6299
Tan Y. H., Sidik S. M., Syed Husain S. N., Lye M. S., Chong P. P. CYP1A1MspI polymorphism and cervical carcinoma risk in the multi-ethnic population of Malaysia: a case-control study. Asian Pac. J. Cancer Prev. 2016: 17: 57-64. https://doi.org/10.7314/apjcp.2016.17.1.57
Nishino K., Sekine M., Kodama S., Sudo N., Aoki Y., Seki N., Tanaka K. Cigarette smoking and glutathione Stransferase M1 polymorphism associated with risk for uterine cervical cancer. J. Obstet. Gynecol. Res. 2008: 34: 994-1001. https://doi.org/10.1111/j.1447-0756.2008.00798.x
Satinder K., Sobti R. C., Pushpinder K. Impact of single nucleotide polymorphism in chemical metabolizing genes and exposure to wood smoke on risk of cervical cancer in North-Indian women. Exp. Oncol. 2017: 39: 69-74. https://pubmed.ncbi.nlm.nih.gov/28361858/
Agorastos T., Papadopoulos N., Lambropoulos A. F., Chrisafi S., Mikos T., Goulis D. G., Constantinidis T. C., Kotsis A., Bontis J. N. Glutathione-S-transferase M1 and T1 and cytochrome P1A1 genetic polymorphisms and susceptibility to cervical intraepithelial neoplasia in Greek women. Eur. J. Cancer Prev. 2007: 16: 498-504. https://doi.org/10.1097/01.cej.0000243859.99265.92
Matos A., Cindy Castelão C., Pereira da Sliva A., Alho I., Bicho M., Medeiros R., Bicho M. C. Epistatic interaction of CYP1A1 and COMT polymorphisms in cervical cancer. Oxid. Med. Cell. Longev. 2016: 2016: 1-7. https://doi.org/10.1155/2016/2769804
Sneha S., Baker S. C., Green A., Storr S., Aiyappa R., Martin S., Pors K. Intratumoral Cytochrome P450 Expression in Breast Cancer: Impact on Standard of Care Treatment and New Efforts to Develop Tumour-Selective Therapies. Biomedicines 2021: 9(3): 290. https://doi.org/10.3390/biomedicines9030290
Autrup H. Genetic polymorphisms in human xenobiotica metabolizing enzymes as susceptibility factors in toxic response. Mutat. Res. 2000: 464: 65-76. https://doi.org/10.1016/s1383-5718(99)00167-9
Dusinska M., Ficek A., Horska A., Raslova K., Petrovska H., Vallova B., Drlickova M., Wood S. G., Stupakova A., Gasparovic J., Bobek P., Nagyova A., Kovacikova Z., Blazicek P., Liegebel U., Collins A. R. Glutathione Stransferase polymorphisms influence the level of oxidative DNA damage and antioxidant protection in humans. Mutat. Res. 2001: 482: 47-55. https://doi.org/10.1016/s0027-5107(01)00209-3
He H. R., You H. S., Sun J. Y., Hu S. S., Ma Y., Dong Y. L., Lu J. Glutathione S-transferase gene polymorphisms and susceptibility to acute myeloid leukemia: meta-analyses. Jpn. J. Clin. Oncol. 2014: 44(11): 1070-1081. https://doi.org/10.1093/jjco/hyu121
Liu K., Lin X., Zhou Q., Ma T., Han L., Mao G., Chen J., Yue X., Wang H., Zhang L., Jin G., Jiang J., Zhao J., Zou B. The associations between two vital GSTs genetic polymorphisms and lung cancer risk in the Chinese population/ evidence from 71 studies. P. Lo. S. O. N. E. 2014: 9(7): e102372. https://doi.org/10.1371/journal.pone.0102372
Kalacas N. A., Garcia J. A., Ortin T. S., Valdez Jr A., Fellizar A., Ramos M. C., Albano P. M. GSTM1 and GSTT1 genetic polymorphisms and breast cancer risk in selected Filipino cases. Asian Pacific journal of cancer prevention: A. P. J. C. P. 2019: 20(2): 529. https://doi.org/10.31557/APJCP.2019.20.2.529
Sfar S., Saad H., Mosbah F., Chouchane L. Combined effects of the angiogenic genes polymorphisms on prostate cancer susceptibility and aggressiveness. Molecular biology reports 2009: 36(1): 37–45. https://doi.org/10.1007/s11033-007-9149-4
Fan H. H., Li B. Q., Wu K. Y., Yan H. D., Gu M. J., Yao X. H., Dong H. J., Zhang X., Zhu J. H. Polymorphisms of Cytochromes P450 and Glutathione S-Transferases Synergistically Modulate Risk for Parkinson’s Disease. Frontiers in aging neuroscience 2022: 14: 888942. https://doi.org/10.3389/fnagi.2022.888942
Ritambhara Tiwari S., Vijayaraghavalu S., Kumar M. Genetic Polymorphisms of Xenobiotic Metabolizing Genes (GSTM1, GSTT1, GSTP1), Gene-Gene Interaction with Association to Lung Cancer Risk in North India; A Case Control Study. Asian Pacific journal of cancer prevention: APJCP: 2019: 20(9): 2707–2714. https://doi.org/10.31557/APJCP.2019.20.9.2707
Acknowledgements
The authors thank all Tunisian subjects for their participation in this study. This work was supported by the Ministry of Higher Education and Scientific and Technological Research and the Ministry of Public Health of Tunisia. The authors thank Mr. Adel Rdissi and Dr. Manel Ben Taher for the English revision, and Pr. Asma Omezzine for her support in the statistical study.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
Study design: A. H. and A. K., Patient enrollment and sample collection: A. H., N. B., and H. B., Experiments, data collection, and data analysis: A. H., N. B., S.S., F. D., M. D., Z. H., and A. K. Manuscript writing: A. H., S. S., M. D., and A. K., All authors revised the manuscript and approved its final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval:
Approval of the ethical standards of the Ethics Committee for Research in Life Sciences and Health of the ISBM (CER-SVS/isbm 007/2021) was obtained.
Consent to participate:
Informed written consent was obtained from all women according to the ethical standards of the Ethics Committee for Research in Life Sciences and Health of the ISBM (CER-SVS/ISBM).
Consent for publication:
Informed consent was obtained from all participants for whom identifying information is included in this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Helaoui, A., Sfar, S., Boudhiba, N. et al. Association of xenobiotic-metabolizing genes polymorphisms with cervical cancer risk in the Tunisian population. Mol Biol Rep 50, 949–959 (2023). https://doi.org/10.1007/s11033-022-07945-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07945-6