Abstract
Background
Small auxin-up RNA (SAUR) genes form a wide family supposedly involved in different physiological and developmental processes in plants such as leaf senescence, auxin signaling and transport, hypocotyl development and tolerance to abiotic stresses. The transcription of SAUR genes is quickly induced by auxins, a group of phytohormones of major importance on embryo development. To better understand the distribution and expression profile of such still not explored family in Coffea sp., especially during the development of somatic embryogenesis (SE), SAUR members were characterized in silico using the available Coffea canephora genome data and analyzed for gene expression by RT-qPCR in C. arabica embryogenic samples.
Methods and results
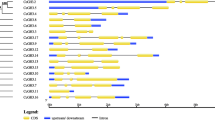
Over C. canephora genome 31 CcSAURs were distributed by 11 chromosomes. Out of these 31 gene members, 5 SAURs were selected for gene expression analysis in C. arabica embryogenic materials. CaSAUR12 and CaSAUR18 were the members highly expressed through almost all plant materials. The other genes had more expression in at least one of the developing embryo stages or plantlets. The CaSAUR12 was the only member to exhibit an increased expression in both non-embryogenic calli and the developing embryo stages.
Conclusion
The identification of SAUR family on C. canephora genome followed by the analysis of gene expression profile across coffee somatic embryogenesis process on C. arabica represents a further additional step towards a better comprehension of molecular components acting on SE. Along with new research about this gene family such knowledge may support studies about clonal propagation methods via somatic embryogenesis to help the scientific community towards improvements into coffee crop.





Similar content being viewed by others
Data availability
All data for gene and amino acid sequences referred to species used into the SAUR family genomic structure relationship and phylogenetic tree are available on the Supplementary material (Supplementary data S1 and S2, respectively). Embryogenic and non-embryogenic calli herein used were obtained under in vitro culture conditions previously described in the materials and methods. The primer sequences for amplification of the CaSAUR genes are also available at the Supplementary material (Supplementary Table S1). In case of any information required for reproduction of the experimental data is not present elsewhere within publication content, please directly contact the corresponding author on reasonable request.
Code availability
Not applicable.
References
Fehér A, Pasternak TP, Dudits D (2003) Transition of somatic plant cells to an embryogenic state. Plant Cell Tiss Org 74:201–228. https://doi.org/10.1023/A:1024033216561
Staritsky G (1970) Embryoid formation in callus tissues of coffee. Acta Bot Neerl 19:509–514. https://doi.org/10.1111/j.1438-8677.1970.tb00677.x
Etienne H, Breton D, Breitler J-C, Bertrand B, Déchamp E, Awada R, Marraccini P, Léran S, Alpizar E, Campa C, Courtel P, Georget F, Ducos J-P (2018) Coffee somatic embryogenesis: how did research, experience gained and innovations promote the commercial propagation of elite clones from the two cultivated species? Front Plant Sci 9:1630. https://doi.org/10.3389/fpls.2018.01630
Campos NA, Panis B, Carpentier SC (2017) Somatic embryogenesis in coffee: the evolution of biotechnology and the integration of omics technologies offer great opportunities. Front Plant Sci. https://doi.org/10.3389/fpls.2017.01460
Ikeuchi M, Favero DS, Sakamoto Y, Iwase A, Coleman D, Rymen B, Sugimoto K (2019) Molecular mechanisms of plant regeneration. Annu Rev Plant Biol 70:377–406. https://doi.org/10.1146/annurev-arplant-050718-100434
Wójcik AM, Wójcikowska B, Gaj MD (2020) Current perspectives on the auxin-mediated genetic network that controls the induction of somatic embryogenesis in plants. Int J Mol Sci 21:1333. https://doi.org/10.3390/ijms21041333
Silva AT, Barduche D, do Livramento KG, Ligterink W, Paiva LV, (2014) Characterization of a putative Serk-Like Ortholog in embryogenic cell suspension cultures of Coffea arabica L. Plant Mol Biol Rep 32:176–184. https://doi.org/10.1007/s11105-013-0632-x
Silva AT, Barduche D, do Livramento KG, Paiva LV, (2015) A putative BABY BOOM-like gene (CaBBM) is expressed in embryogenic calli and embryogenic cell suspension culture of Coffea arabica L. In Vitro Cell Dev-Pl 51:93–101. https://doi.org/10.1007/s11627-014-9643-z
Torres LF, Diniz LEC, Do Livramento KG, Freire LL, Paiva LV (2015) Gene expression and morphological characterization of cell suspensions of Coffea arabica L. cv. Catiguá MG2 in different cultivation stages. Acta Physiol Plant. https://doi.org/10.1007/s11738-015-1924-6
Su YH, Liu YB, Zhang XS (2011) Auxin-cytokinin interaction regulates meristem development. Mol Plant 4:616–625. https://doi.org/10.1093/mp/ssr007
Pinto RT, Freitas NC, Máximo WPF, Cardoso TB, Prudente DO, Paiva LV (2019) Genome-wide analysis, transcription factor network approach and gene expression profile of GH3 genes over early somatic embryogenesis in Coffea spp. BMC Genomics 20:812. https://doi.org/10.1186/s12864-019-6176-1
McClure BA, Guilfoyle T (1989) Rapid redistribution of auxin-regulated RNAs during gravitropism. Science 243:91–93. https://doi.org/10.1126/science.11540631
Wu J, Liu S, He Y, Guan X, Zhu X, Cheng L, Wang J, Lu G (2012) Genome-wide analysis of SAUR gene family in Solanaceae species. Gene 509:38–50. https://doi.org/10.1016/j.gene.2012.08.002
Jain M, Tyagi AK, Khurana JP (2006) Genome-wide analysis, evolutionary expansion, and expression of early auxin-responsive SAUR gene family in rice (Oryza sativa). Genomics 88:360–371. https://doi.org/10.1016/j.ygeno.2006.04.008
Wang P, Lu S, Xie M, Wu M, Ding S, Khaliq A, Ma Z, Mao J, Chen B (2020) Identification and expression analysis of the small auxin-up RNA (SAUR) gene family in apple by inducing of auxin. Gene 750:144725. https://doi.org/10.1016/j.gene.2020.144725
Park JE, Kim YS, Yoon HK, Park CM (2007) Functional characterization of a small auxin-up RNA gene in apical hook development in Arabidopsis. Plant Sci 172:150–157. https://doi.org/10.1016/j.plantsci.2006.08.005
Zhang H, Yu Z, Yao X, Chen J, Chen X, Zhou H, Lou Y, Ming F, Jin Y (2021) Genome-wide identification and characterization of small auxin-up RNA (SAUR) gene family in plants: evolution and expression profiles during normal growth and stress response. BMC Plant Biol 21:4. https://doi.org/10.1186/s12870-020-02781-x
Yin H, Li M, Lv M, Hepworth SR, Li D, Ma C, Li J, Wang SM (2020) SAUR15 promotes lateral and adventitious root development via activating H+-ATPases and auxin biosynthesis. Plant Physiol 184:837–851. https://doi.org/10.1104/pp.19.01250
Wang X, Yu R, Wang J, Lin Z, Han X, Deng Z, Fan L, He H, Deng XW, Chen H (2020) The Asymmetric expression of SAUR genes mediated by ARF7/19 promotes the gravitropism and phototropism of plant hypocotyls. Cell Rep 31:107529. https://doi.org/10.1016/j.celrep.2020.107529
Wong JH, Klejchová M, Snipes SA, Nagpal P, Bak G, Wang B, Dunlap S, Park MY, Kunkel EN, Trinidad B, Reed JW (2021) SAUR proteins and PP2C. D phosphatases regulate H+-ATPases and K+ channels to control stomatal movements. Plant Physiol 185:256–273. https://doi.org/10.1093/plphys/kiaa023
Spartz AK, Ren H, Park MY, Grandt KN, Lee SH, Murphy AS, Sussman MR, Overvoorde PJ, Gray WM (2014) SAUR inhibition of PP2C-D phosphatases activates plasma membrane H+-ATPases to promote cell expansion in Arabidopsis. Plant Cell 26:2129–2142. https://doi.org/10.1105/tpc.114.126037
Li ZG, Chen HW, Li QT, Tao JJ, Bian XH, Ma B, Zhang WK, Chen SY, Zhang JS (2015) Three SAUR proteins SAUR76, SAUR77 and SAUR78 promote plant growth in Arabidopsis. Sci Rep. https://doi.org/10.1038/srep12477
Spartz AK, Lor VS, Ren H, Olszewski NE, Miller ND, Wu G, Spalding EP, Gray WM (2017) Constitutive expression of Arabidopsis SMALL AUXIN UP RNA19 (SAUR19) in tomato confers auxin-independent hypocotyl elongation. Plant Physiol 173:1453–1462. https://doi.org/10.1104/pp.16.01514
Denoeud F, et al. (2014) The coffee genome provides insight into the convergent evolution of caffeine biosynthesis. Science 345:1181–1184. DOI: https://doi.org/10.1126/science.1255274 Coffee Genome Hub (http://coffee-genome.org/). Cited 10 March 2020.
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Kalyaanamoorthy S, Minh BQ, Wong TKF, Haeseler A, Jermiin LS (2017) ModelFinder: Fast model selection foraccurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Nguyen LT, Schmidt HA, Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins 64:643–651. https://doi.org/10.1002/prot.21018
Sahu SS, Loaiza CD, Kaundal R (2020) Plant-mSubP: a computational framework for the prediction of single- and multi-target protein subcellular localization using integrated machine-learning approaches. AoB Plants. https://doi.org/10.1093/aobpla/plz068
Armenteros JJA, Sønderby CK, Sønderby SK, Nielsen H, Winther O (2017) DeepLoc: prediction of protein subcellular localization using deep learning. Bioinformatics 33:3387–3395. https://doi.org/10.1093/bioinformatics/btx431
Gasteiger E, Hoogland C, Gattiker A, Duvaud Se, Wilkins MR, Appel RD, Bairoch A (2005) Protein Identification and Analysis Tools on the ExPASy Server, In: Walker JM (ed) The Proteomics Protocols Handbook, Humana Press, Totowa Doi: https://doi.org/10.1385/1-59259-890-0:571
Teixeira JBJ, Junqueira CS, Pereira AJPC, Mello RIS, Silva APD, Mundim DA (2004) Multiplicação clonal de café (Coffea arábica L.) via embriogênese somática. Embrapa Recursos Genéticos e Biotecnologia. https://ainfo.cnptia.embrapa.br/digital/bitstream/CENARGEN/24709/1/doc121.pdf. Cited 1 June 2018
Boxtel JV, Berthouly M (1996) High frequency somatic embryogenesis from coffee leaves. Plant Cell Tiss Org 44:7–17. https://doi.org/10.1007/BF00045907
Zamarripa A, Ducos JP, Bollon H, Dufour M, Pétiard V (1991) Production d’embryons somatiques de caféier en milieu liquide: effets densité d’inoculation et renouvellement du milieu. Café, Cacao, Thé 35:233–244
Maciel ALR, Rodrigues FA, Pasqual M, Carvalho CHS (2016) Large-scale, high-efficiency production of coffee somatic embryos. Crop Breed Appl Biot 16:102–107. https://doi.org/10.1590/1984-70332016v16n2a16
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45. https://doi.org/10.1093/nar/29.9.e45
Freitas NC, Barreto HG, Fernandes-Brum CN, Moreira RO, Chalfun A, Paiva LV (2017) Validation of reference genes for qPCR analysis of Coffea arabica L. somatic embryogenesis-related tissues. Plant Cell Tiss Org 128:663–678. https://doi.org/10.1007/s11240-016-1147-6
Ferreira DF (2014) Sisvar: a Guide for its Bootstrap procedures in multiple comparisons. Ciênc Agrotec 38:109–112. https://doi.org/10.1590/S1413-70542014000200001
Bai QS, Hou D, Li L, Cheng Z, Ge W, Liu J, Li X, Mu S, Gao J (2017) Genome-wide analysis and expression characteristics of small auxin-up RNA (SAUR) genes in moso bamboo (Phyllostachys edulis). Genome 60:325–336. https://doi.org/10.1139/gen-2016-0097
Stortenbeker N, Bemer M (2019) The SAUR gene family: the plant’s toolbox for adaptation for adaptation of growth and development. J Exp Bot 70:17–27. https://doi.org/10.1093/jxb/ery332
Chae K, Isaacs CG, Reeves PH, Maloney GS, Muday GK, Nagpal P, Reed JW (2012) Arabidopsis SMALL AUXIN UP RNA63 promotes hypocotyl and stamen filament elongation. Plant J 71:684–697. https://doi.org/10.1111/j.1365-313X.2012.05024.x
Spartz AK, Lee SH, Wenger JP, Gonzalez N, Itoh H, Inzé D, Peer WA, Murphy AS, Overvoorde PJ, gray WM, (2012) The SAUR19 subfamily of SMALL AUXIN UP RNA genes promote cell expansion. Plant J 70:978–990. https://doi.org/10.1111/j.1365-313X.2012.04946.x
Stamm P, Kumar PP (2013) Auxin and gibberellin responsive Arabidopsis SMALL AUXIN UP RNA36 regulates hypocotyl elongation in the light. Plant Cell Rep 32:759–769. https://doi.org/10.1007/s00299-013-1406-5
Michalczuk L, Cooke TJ, Cohen JD (1992) Auxin levels at different stages of carrot somatic embryogenesis. Phytochemistry 31:1097–1103. https://doi.org/10.1016/0031-9422(92)80241-6
Hou K, Wu W, Gan SS (2013) SAUR36, a SMALL AUXIN UP RNA gene, is involved in the promotion of leaf senescence in Arabidopsis. Plant Physiol 161:1002–1009. https://doi.org/10.1104/pp.112.212787
Qiu T, Qi M, Ding X, Zheng Y, Zhou T, Chen Y, Han N, Zhu M, Bian H, Wang J (2020) The SAUR41 subfamily of SMALL AUXIN UP RNA genes is abscisic acid inducible to modulate cell expansion and salt tolerance in Arabidopsis thaliana seedlings. Ann Bot 125:805–819. https://doi.org/10.1093/aob/mcz160
He Y, Liu Y, Li M, Lamin-Samu AT, Yang D, Yu X, Izhar M, Jan I, Ali M, Lu G (2021) The Arabidopsis SMALL AUXIN UP RNA32 protein regulates ABA-mediated responses to drought stress. Front Plant Sci 12:259. https://doi.org/10.3389/fpls.2021.625493
Markakis MN, Boron AK, Van Loock B, Saini K, Cirera S, Verbelen J-P, Vissenberg K (2013) Characterization of a Small Auxin-Up RNA (SAUR)-Like Gene Involved in Arabidopsis thaliana Development. PLoS ONE. https://doi.org/10.1371/journal.pone.0082596
Sagee O, Riov J, Goren R (1990) Ethylene-enhanced catabolism of [C]indole-3-acetic acid to indole-3-carboxylic acid in citrus leaf tissues. Plant Physiol 92:54–60. https://doi.org/10.1104/pp.92.1.54
Márquez-López RE, Pérez-Hernández C, Ku-González Á, Galaz-Ávalos RM, Loyola-Vargas VM (2018) Localization and transport of indole-3-acetic acid during somatic embryogenesis in Coffea canephora. Protoplasma 255:695–708. https://doi.org/10.1007/s00709-017-1181-1
Acknowledgements
We would like to thank the National Council for Scientific and Technological Development (CNPq) from Ministry of Science and Technology, Brazil, and the Foundation of Support Research of the State of Minas Gerais (FAPEMIG) for the financial support provided to this study. The Coordination for the Improvement of Higher Education Personnel (CAPES) from Ministry of Education, for the scholarships awarded to the authors. We also thank the Federal University of Lavras (UFLA) for providing the infrastructure and equipment to accomplish this research, including real time PCR machine, computers and servers for bioinformatics analyses, which is supported by Ministry of Science and Technology and Ministry of Education.
Funding
FAPEMIG, CNPq, and CAPES. Conselho Nacional de Desenvolvimento Científico e Tecnológico,Coordenação de Aperfeiçoamento de Pessoal de Nível Superior,Fundação de Amparo à Pesquisa do Estado de Minas Gerais
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Leandro Eugenio Cardamone Diniz and Luciano Vilela Paiva get funding acquisition for the financial support from funding agencies. Fabiana Couto Zanin and Natália Chagas Freitas conducted in vitro culture experiments and gene expression analyses through RT-qPCR. Wesley Pires Flausino Máximo and Renan Terassi Pinto carried out the studies about the phylogenetic relationship, genomic structure and the statistical analysis for RT-qPCR data. Fabiana Couto Zanin, Natália Chagas Freitas, Renan Terassi Pinto, and Wesley Pires Flausino Máximo contributed to writing the manuscript. All authors contributed equally to revising the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Ethical approval
Not applicable.
Consent to participate
All authors are consent about their participation at the manuscript.
Consent for publication
All authors are consent about publishing this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zanin, F.C., Freitas, N.C., Pinto, R.T. et al. The SAUR gene family in coffee: genome-wide identification and gene expression analysis during somatic embryogenesis. Mol Biol Rep 49, 1973–1984 (2022). https://doi.org/10.1007/s11033-021-07011-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-07011-7