Abstract
Background
Intramuscular fat content, an important meat quality trait, strongly affects flavor, juiciness, and tenderness. Sex hormones regulate lipid metabolism, and female hormones stimulate fat deposition, thereby making the female chickens always fatter than males. In this study, the effect of sex on IMF deposition was screened following transcriptomics in chickens.
Methods and results
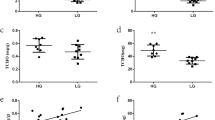
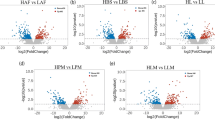
Results confirmed significantly higher IMF content of 150-day female chickens as compared to the male chickens. The female chickens manifested higher serum TG, LDL-C, and VLDL, and significantly lower HDL-C contents than male chickens. Moreover, differential expression of genes involved in lipid metabolism were obtained in the muscle and liver between female and male chicken, which could partly interpret the possible reasons for the sex-mediated differences of IMF content. Cellular results revealed that inhibition of PLIN2 significantly inhibited chicken preadipocyte proliferation and induces apoptosis of preadipocytes, as well as promoted adipocyte differentiation.
Conclusions
According to our results, PLIN2 may be considered as a molecular marker for poultry meat quality and applying this gene in early breed selection.








Similar content being viewed by others
Data availability
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation. All the data are available in the SRA database with accession number PRJNA732829.
References
Petracci M, Mudalal S, Soglia F, Cavani C (2015) Meat quality in fast-growing broiler chickens. Worlds Poult Sci J 71(2):363–374
Petracci M, Cavani C (2012) Muscle growth and poultry meat quality issues. Nutrients 4(1):1–12. https://doi.org/10.3390/nu4010001
Li J, Yang C, Peng H, Yin H, Wang Y, Hu Y, Yu C, Jiang X, Du H, Li Q, Liu Y (2019) Effects of slaughter age on muscle characteristics and meat quality traits of Da-Heng meat type birds. Animals. https://doi.org/10.3390/ani10010069
Liu L, Liu X, Cui H, Liu R, Zhao G, Wen J (2019) Transcriptional insights into key genes and pathways controlling muscle lipid metabolism in broiler chickens. BMC Genomics 20(1):863. https://doi.org/10.1186/s12864-019-6221-0
Hocquette JF, Gondret F, Baéza E, Médale F, Jurie C, Pethick DW (2010) Intramuscular fat content in meat-producing animals: development, genetic and nutritional control, and identification of putative markers. Animal 4(2):303–319. https://doi.org/10.1017/s1751731109991091
Warner RD, Greenwood PL, Pethick DW, Ferguson DM (2010) Genetic and environmental effects on meat quality. Meat Sci 86(1):171–183. https://doi.org/10.1016/j.meatsci.2010.04.042
Zerehdaran S, Vereijken AL, van Arendonk JA, van der Waaijt EH (2004) Estimation of genetic parameters for fat deposition and carcass traits in broilers. Poult Sci 83(4):521–525. https://doi.org/10.1093/ps/83.4.521
Gawrieh S (2015) Sex hormones, sex hormone-binding globulin, and liver fat: which came first, the chicken or the egg? Clin Gastroenterol Hepatol 13(9):1694–1696. https://doi.org/10.1016/j.cgh.2015.04.182
Chen X, Geng Z, Niu J (2017) Gene expression and plasma lipid content in relation to intramuscular fat in Chinese indigenous Wuhua chicken. J Appl Poult Res 26(3):391–400
Saez G, Davail S, Gentès G, Hocquette JF, Jourdan T, Degrace P, Baéza E (2009) Gene expression and protein content in relation to intramuscular fat content in Muscovy and Pekin ducks. Poult Sci 88(11):2382–2391. https://doi.org/10.3382/ps.2009-00208
Yin L, Yu L, Zhang L, Ran J, Li J, Yang C, Jiang X, Du H, Hu X, Liu Y (2019) Transcriptome analysis reveals differentially expressed genes and pathways for oviduct development and defense in prelaying and laying hens. Am J Reprod Immunol 82(3):e13159. https://doi.org/10.1111/aji.13159
Wang Y, Ghaffari N, Johnson CD, Braga-Neto UM, Wang H, Chen R, Zhou H (2011) Evaluation of the coverage and depth of transcriptome by RNA-Seq in chickens. BMC Bioinformatics 12(10):S5. https://doi.org/10.1186/1471-2105-12-s10-s5
Jeong J, Bong J, Kim GD, Joo ST, Lee HJ, Baik M (2013) Transcriptome changes favoring intramuscular fat deposition in the longissimus muscle following castration of bulls. J Anim Sci 91(10):4692–4704. https://doi.org/10.2527/jas.2012-6089
Miao X, Luo Q, Qin X, Guo Y, Zhao H (2015) Genome-wide mRNA-seq profiling reveals predominant down-regulation of lipid metabolic processes in adipose tissues of Small Tail Han than Dorset sheep. Biochem Biophys Res Commun 467(2):413–420. https://doi.org/10.1016/j.bbrc.2015.09.129
Ye M, Zhou B, Wei S, Ding M, Lu X, Shi X, Ding J, Yang S, Wei W (2016) Transcriptomic analysis identifies candidate genes related to intramuscular fat deposition and fatty acid composition in the breast muscle of squabs (Columba). G3 (Bethesda, Md) 6(7):2081–2090. https://doi.org/10.1534/g3.116.029793
Ramayo-Caldas Y, Mach N, Esteve-Codina A, Corominas J, Castelló A, Ballester M, Estellé J, Ibáñez-Escriche N, Fernández AI, Pérez-Enciso M, Folch JM (2012) Liver transcriptome profile in pigs with extreme phenotypes of intramuscular fatty acid composition. BMC Genomics 13:547. https://doi.org/10.1186/1471-2164-13-547
Shi H, Wang Q, Wang Y, Leng L, Zhang Q, Shang Z, Li H (2010) Adipocyte fatty acid-binding protein: an important gene related to lipid metabolism in chicken adipocytes. Comp Biochem Physiol B 157(4):357–363. https://doi.org/10.1016/j.cbpb.2010.08.005
Royan M, Navidshad B (2016) Peroxisome proliferator-activated receptor gamma (PPARγ), a key regulatory gene of lipid metabolism in chicken. Worlds Poult Sci J 72(04):773–784
Cui H, Zhao G, Liu R, Zheng M, Chen J, Wen J (2012) FSH stimulates lipid biosynthesis in chicken adipose tissue by upregulating the expression of its receptor FSHR. J Lipid Res 53(5):909–917
Cui HX, Liu RR, Zhao GP, Zheng MQ, Chen JL, Wen J (2012) Identification of differentially expressed genes and pathways for intramuscular fat deposition in pectoralis major tissues of fast-and slow-growing chickens. BMC Genomics 13:213. https://doi.org/10.1186/1471-2164-13-213
Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA (2003) DAVID: database for annotation, visualization, and integrated discovery. Genome Biol 4(5):P3
Zhang M, Li DH, Li F, Sun JW, Jiang RR, Li ZJ, Han RL, Li GX, Liu XJ, Kang XT, Sun GR (2018) Integrated analysis of MiRNA and genes associated with meat quality reveals that Gga-MiR-140-5p affects intramuscular fat deposition in chickens. Cell Physiol Biochem 46(6):2421–2433. https://doi.org/10.1159/000489649
Zhang R, Lin Y, Zhi L, Liao H, Zuo L, Li Z, Xu Y (2017) Expression profiles and associations of adiponectin and adiponectin receptors with intramuscular fat in Tibetan chicken. Br Poult Sci 58(2):151–157. https://doi.org/10.1080/00071668.2016.1268252
Guo-Bin C, Li-Li L, Xue-Yu Z, Ke-Hua W, Guo-Hong C (2010) Development rule of intramuscular fat content in chicken. J Anim Vet Adv 9(2):297–298
Yang Y, Wen J, Fang GY, Li ZR, Liu J (2015) The effects of raising system on the lipid metabolism and meat quality traits of slow-growing chickens. J Appl Anim Res 43(2):1–6
Yan J, Liao K, Wang T, Mai K, Xu W, Ai Q (2015) Dietary lipid levels influence lipid deposition in the liver of large yellow croaker (Larimichthys crocea) by regulating lipoprotein receptors, fatty acid uptake and triacylglycerol synthesis and catabolism at the transcriptional level. PLoS ONE 10(6):e0129937. https://doi.org/10.1371/journal.pone.0129937
Leveille GA, Romsos DR, Yeh Y, Ohea EK (1975) Lipid biosynthesis in the chick. A consideration of site of synthesis, influence of diet and possible regulatory mechanisms. Poult Sci 54(4):1075–1093. https://doi.org/10.3382/ps.0541075
Zhuo Z, Lamont SJ, Lee WR, Abasht B (2015) RNA-seq analysis of abdominal fat reveals differences between modern Commercial broiler chickens with high and low feed efficiencies. PLoS ONE 10(8):e0135810. https://doi.org/10.1371/journal.pone.0135810
Wang X, Wei D, Song Z, Jiao H, Lin H (2012) Effects of fatty acid treatments on the dexamethasone-induced intramuscular lipid accumulation in chickens. PLoS ONE 7(5):e36663. https://doi.org/10.1371/journal.pone.0036663
Dominique H, John CM, Bernard L (1984) Plasma lipoprotein profile in fasted and refed chickens of two strains selected for high or low adiposity. J Nutr 6:1112–1121
Hermier D (1997) Lipoprotein metabolism and fattening in poultry. J Nutr 127(5):805s–808s. https://doi.org/10.1093/jn/127.5.805S
Griffin JD, Lichtenstein AH (2013) Dietary cholesterol and plasma lipoprotein profiles: randomized-controlled trials. Curr Nutr Rep 2(4):274–282. https://doi.org/10.1007/s13668-013-0064-0
PO, VA, EO (2016) Comparative evaluation of cholesterol content and storage quality of chicken and quail eggs. World J Nutr Health 4(1):5–9
Ye Y, Lin S, Mu H, Tang X, Ou Y, Chen J, Ma Y, Li Y (2014) Analysis of differentially expressed genes and signaling pathways related to intramuscular fat deposition in skeletal muscle of sex-linked dwarf chickens. Biomed Res Int 2014:724274. https://doi.org/10.1155/2014/724274
Liu L, Cui H, Fu R, Zheng M, Liu R, Zhao G, Wen J (2017) The regulation of IMF deposition in pectoralis major of fast- and slow- growing chickens at hatching. J Anim Sci Biotechnol 8:77. https://doi.org/10.1186/s40104-017-0207-z
Leveille GA, O’Hea EK, Chakbabarty K (1968) In vivo lipogenesis in the domestic chicken. Proc Soc Exp Biol Med Soc Exp Biol Med (New York, NY) 128(2):398–401. https://doi.org/10.3181/00379727-128-33022
Li H, Ma Z, Jia L, Li Y, Xu C, Wang T, Han R, Jiang R, Li Z, Sun G, Kang X, Liu X (2016) Systematic analysis of the regulatory functions of microRNAs in chicken hepatic lipid metabolism. Sci Rep 6:31766. https://doi.org/10.1038/srep31766
Li H, Wang T, Xu C, Wang D, Ren J, Li Y, Tian Y, Wang Y, Jiao Y, Kang X, Liu X (2015) Transcriptome profile of liver at different physiological stages reveals potential mode for lipid metabolism in laying hens. BMC Genomics 16:763. https://doi.org/10.1186/s12864-015-1943-0
Gan L, Liu Z, Cao W, Zhang Z, Sun C (2015) FABP4 reversed the regulation of leptin on mitochondrial fatty acid oxidation in mice adipocytes. Sci Rep 5:13588. https://doi.org/10.1038/srep13588
Storch J, Thumser AE (2000) The fatty acid transport function of fatty acid-binding proteins. Biochim Biophys Acta 1486(1):28–44. https://doi.org/10.1016/s1388-1981(00)00046-9
He J, Tian Y, Li J, Shen J, Tao Z, Fu Y, Niu D, Lu L (2013) Expression pattern of L-FABP gene in different tissues and its regulation of fat metabolism-related genes in duck. Mol Biol Rep 40(1):189–195. https://doi.org/10.1007/s11033-012-2048-3
Gao GL, Na W, Wang YX, Zhang HF, Li H, Wang QG (2015) Role of a liver fatty acid-binding protein gene in lipid metabolism in chicken hepatocytes. Genet Mol Res 14(2):4847–4857. https://doi.org/10.4238/2015.May.11.17
Lee SY, Nagy BP, Brooks AR, Wang DM, Paulweber B, Levy-Wilson B (1996) Members of the caudal family of homeodomain proteins repress transcription from the human apolipoprotein B promoter in intestinal cells. J Biol Chem 271(2):707–718. https://doi.org/10.1074/jbc.271.2.707
Morais S, Monroig O, Zheng X, Leaver MJ, Tocher DR (2009) Highly unsaturated fatty acid synthesis in Atlantic salmon: characterization of ELOVL5- and ELOVL2-like elongases. Mar Biotechnol (NY) 11(5):627–639. https://doi.org/10.1007/s10126-009-9179-0
Lavoie HA, King SR (2009) Transcriptional regulation of steroidogenic genes: STARD1, CYP11A1 and HSD3B. Exp Biol Med (Maywood) 234(8):880–907. https://doi.org/10.3181/0903-mr-97
Widmann P, Nuernberg K, Kuehn C, Weikard R (2011) Association of an ACSL1 gene variant with polyunsaturated fatty acids in bovine skeletal muscle. BMC Genet 12:96. https://doi.org/10.1186/1471-2156-12-96
Cui JX, Zeng YQ, Wang H, Chen W, Du JF, Chen QM, Hu YX, Yang L (2011) The effects of DGAT1 and DGAT2 mRNA expression on fat deposition in fatty and lean breeds of pig. Livest Sci 140(1–3):292–296
Qiu F, Xie L, Ma JE, Luo W, Zhang L, Chao Z, Chen S, Nie Q, Lin Z, Zhang X (2017) Lower expression of SLC27A1 enhances intramuscular fat deposition in chicken via down-regulated fatty acid oxidation mediated by CPT1A. Front Physiol 8:449. https://doi.org/10.3389/fphys.2017.00449
Brasaemle DL (2007) Thematic review series: adipocyte biology: the perilipin family of structural lipid droplet proteins: stabilization of lipid droplets and control of lipolysis. J Lipid Res 48(12):2547–2559. https://doi.org/10.1194/jlr.R700014-JLR200
Bildirici I, Schaiff WT, Chen B, Morizane M, Oh SY, O’Brien M, Sonnenberg-Hirche C, Chu T, Barak Y, Nelson DM, Sadovsky Y (2018) PLIN2 is essential for trophoblastic lipid droplet accumulation and cell survival during hypoxia. Endocrinology 159(12):3937–3949. https://doi.org/10.1210/en.2018-00752
Tsai TH, Chen E, Li L, Saha P, Lee HJ, Huang LS, Shelness GS, Chan L, Chang BH (2017) The constitutive lipid droplet protein PLIN2 regulates autophagy in liver. Autophagy 13(7):1130–1144. https://doi.org/10.1080/15548627.2017.1319544
Dahlhoff M, Camera E, Picardo M, Zouboulis CC, Chan L, Chang BH (1830) Schneider MR (2013) PLIN2, the major perilipin regulated during sebocyte differentiation, controls sebaceous lipid accumulation in vitro and sebaceous gland size in vivo. Biochim Biophys Acta 10:4642–4649. https://doi.org/10.1016/j.bbagen.2013.05.016
Li S, Raza SHA (2020) Overexpression of PLIN1 promotes lipid metabolism in bovine adipocytes. Animals. https://doi.org/10.3390/ani10111944
Raciti GA, Fiory F, Campitelli M, Desiderio A, Spinelli R, Longo M, Nigro C, Pepe G, Sommella E, Campiglia P, Formisano P, Beguinot F, Miele C (2018) Citrus aurantium L. dry extracts promote C/ebpβ expression and improve adipocyte differentiation in 3T3-L1 cells. PLoS ONE 13(3):0193704. https://doi.org/10.1371/journal.pone.0193704
Kruidering M, Evan GI (2000) Caspase-8 in apoptosis: the beginning of “the end”? IUBMB Life 50(2):85–90. https://doi.org/10.1080/713803693
Porter AG, Jänicke RU (1999) Emerging roles of caspase-3 in apoptosis. Cell Death Differ 6(2):99–104. https://doi.org/10.1038/sj.cdd.4400476
Würstle ML, Laussmann MA, Rehm M (2012) The central role of initiator caspase-9 in apoptosis signal transduction and the regulation of its activation and activity on the apoptosome. Exp Cell Res 318(11):1213–1220. https://doi.org/10.1016/j.yexcr.2012.02.013
Chiou SK, Rao L, White E (1994) Bcl-2 blocks p53-dependent apoptosis. Mol Cell Biol 14(4):2556–2563. https://doi.org/10.1128/mcb.14.4.2556
Funding
This research was funded by Sichuan Science and Technology Program (Grant No. 2021YFYZ0031), the Innovation Key Laboratory of Sichuan Province (Grant No. 2017JZ0033) and Key Technology Support Program of Sichuan Province (Grant No. 2018NZDZX0004).
Author information
Authors and Affiliations
Contributions
Conceptualization, JL and CY; methodology, JL and PR; software, JL and ZL; investigation, JL, CY, DZ and LW; resources, CY and XJ; writing—original draft preparation, JL; writing—review and editing, CY, XJ, PR and YL; supervision, XJ and YL; project administration, YL; funding acquisition, YL and CY All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical statement
The animal study was reviewed and approved by the Ethics Committee for Animal Experiments of Sichuan agricultural university.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, J., Yang, C., Ren, P. et al. Transcriptomics analysis of Daheng broilers reveals that PLIN2 regulates chicken preadipocyte proliferation, differentiation and apoptosis. Mol Biol Rep 48, 7985–7997 (2021). https://doi.org/10.1007/s11033-021-06831-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06831-x