Abstract
Background
Low light is a primary regulator of chrysanthemum growth. Our aim was to analyse the different transcriptomic responses of two Chrysanthemum morifolium cultivars to low light.
Methods and results
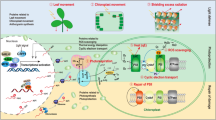
We conducted a transcriptomic analysis of leaf samples from the ‘Nannonggongfen’ and ‘Nannongxuefeng’ chrysanthemum cultivars following a 5-day exposure to optimal light (70%, control [CK]) or low-light (20%, LL) conditions. Gene Ontology (GO) classification of upregulated genes revealed these genes to be associated with 11 cellular components, 9 molecular functions, and 15 biological processes, with the majority being localized to the chloroplast, highlighting the role of chloroplast proteins as regulators of shading tolerance. Downregulated genes were associated with 11 cellular components, 8 molecular functions, and 16 biological processes. Heat map analyses suggested that basic helix–loop–helix domain genes and elongation factors were markedly downregulated in ‘Nannongxuefeng’ leaves, consistent with the maintenance of normal stem length, whereas no comparable changes were observed in ‘Nanonggongfen’ leaves. Subsequent qPCR analyses revealed that phytochrome-interacting factors and dormancy-associated genes were significantly upregulated under LL conditions relative to CK conditions, while succinate dehydrogenase 1, elongated hypocotyls 5, and auxin-responsive gene of were significantly downregulated under LL conditions.
Conclusions
These findings suggest that LL plants were significantly lower than those of the CK plants. Low-light tolerant chrysanthemum cultivars may maintain reduced indole-3-acetic acid (IAA) and elongation factor expression as a means of preventing the onset of shade-avoidance symptoms.




Similar content being viewed by others
Data availability
All data are fully available without restriction.
References
Deng YM, Li CC, Shao QS et al (2012) Differential responses of double petal and multi petal jasmine to shading: I. Photosynthetic characteristics and chloroplast ultrastructure. Plant Physiol Biochem 55:93–102. https://doi.org/10.1016/j.plaphy.2012.03.006
Zheng L, Labeke MCV (2017) Chrysanthemum morphology, photosynthetic efficiency and antioxidant capacity are differentially modified by light quality. J Plant Physiol 213:66–74. https://doi.org/10.1016/j.jplph.2017.03.005
Parlitz S, Kunze R, Mueller RB, Balazadeh S (2011) Regulation of photosynthesis and transcription factor expression by leaf shading and re-illumination in Arabidopsis thaliana leaves. J Plant Physiol 168:1311–1319. https://doi.org/10.1016/j.jplph.2011.02.001
Dutta S, Cruz JA, Imran SM, Chen J, Kramer DM, Osteryoung KW (2017) Variations in chloroplast movement and chlorophyll fluorescence among chloroplast division mutants under light stress. J Exp Bot 68:3541–3555. https://doi.org/10.1093/jxb/erx203
Li Y, Xin G, Wei M, Shi Q, Yang F, Wang X (2017) Carbohydrate accumulation and sucrose metabolism responses in tomato seedling leaves when subjected to different light qualities. Sci Hortic 225:490–497. https://doi.org/10.1016/j.scienta.2017.07.053
Xu PL, Guo YK, Bai JG, Shang L, Wang XJ (2008) Effects of long-term chilling on ultrastructure and antioxidant activity in leaves of two cucumber cultivar under low light. Physiol Plant 132:467–478. https://doi.org/10.1111/j.1399-3054.2007.01036.x
Somporn C, Kamtuo A, Theerakulpisut P, Siriamornpun S (2012) Effect of shading on yield, sugar content, phenolic acids and antioxidant property of coffee beans (Coffea Arabica L. cv. Catimor) harvested from north-eastern Thailand. J Sci Food Agr 92:1956–1963. https://doi.org/10.1002/jsfa.5568
Geromel C, Ferreira LP, Davrieux F, Guyot B, Ribeyre F, Scholz MBS, Pereira LFP, Vaast P, Pot D, Leroy T, Filho AA, Vieira LGE, Mazzafera P, Marraccini P (2008) Effects of shade on the development and sugar metabolism of coffee (Coffea arabica L.) fruits. Plant Physiol Biochem 46:569–579. https://doi.org/10.1016/j.plaphy.2008.02.006
Hu W, Ma Y, Lv F, Liu J, Zhao W, Chen B, Meng Y, Wang Y, Zhou Z (2016) Effects of late planting and shading on sucrose metabolism in cotton fiber. Environ Exp Bot 131:164–172. https://doi.org/10.1016/j.envexpbot.2016.08.001
Han S, Jiang JF, Li HY, Song AP, Chen SM, Chen FD (2015). The differential response of two chrysanthemum cultivars to shading: photosynthesis, chloroplast and sieve element-companion cell ultrastructure. Hortscience 50(8):1192–1195. https://doi.org/10.21273/HORTSCI.50.8.1192
Han S, Chen SM, Song AP, Liu RX, Li HY, Jiang JF, Chen FD (2017) Photosynthetic responses of Chrysanthemum morifolium to growth irradiance: morphological, anatomical and chloroplast ultrastructure. Phothosynthica 55(1):184–192. https://doi.org/10.1007/s11099-016-0219-5
Sasaki K (2017) Generation of expressed sequence tags for discovery of genes responsible for floral traits of Chrysanthemum morifolium by next-generation sequencing technology. BMC Genom 18:683
Xu YJ, Gao S, Yang YJ, Huang MY, Cheng L, Wei Q, Fei ZJ, Gao JP, Hong B (2013) Transcriptome sequencing and whole genome expression profiling of chrysanthemum under dehydration stress. BMC Genom 14:1–15
Wang H, Wu GX, Zhao BB, Lang ZH, Zhang CY, Wang HY (2016) Regulatory modules controlling early shade avoidance response in maize seedlings. BMC Genom 17:269
Carabelli M, Possenti M, Sessa G, Ruzza V, Morelli G, Ruberti I (2018) Arabidopsis HD-Zip II proteins regulated the exit from proliferation during leaf development in canopy shade. J Exp Bot 9:1–15. https://doi.org/10.1093/jxb/ery331
Xie YR, Liu Y, Wang H, Ma XJ, Wang BB, Wu GX, Wang HY (2018) Phytochrome-interacting factors directly suppress MIR156 expression to enhance shade-avoidance syndrome in Arabidopsis. Nat Commun 8:1–15
Iglesias MJ, Sellaro R, Zurbriggen MD, Casal JJ (2017) Multiple links between shade avoidance and auxin networks. J Exp Bot 69:213–228. https://doi.org/10.1093/jxb/erx295
Yang Y, Jiang ZF, Guo JN (2018) Trancriptomic analyses of Chrysanthemum morifolium under UV-B radiation treatment reveal variations in the metabolisms associated with bioactive components. Ind Crop Prod 124:475–486. https://doi.org/10.1016/j.indcrop.2018.08.011
Lu CF, Pu Y, Liu YT, Li YJ, Qu JP, Huang H, Dai SL (2019) Comparative transcriptomics and weighted gene co-expression correlation network analysis (WGCNA) reveal potential regulation mechanism of carotenoid accumulation in Chrysanthemum×morifolium. Plant Physiol Biochem 142:415–428. https://doi.org/10.1016/j.plaphy.2019.07.023
Liu Y, Liu L, Zhao WQ, Guan ZY, Jian JF, Fang WM, Chen FD (2021) A transcriptional response atlas of Chrysanthemum morifolium to dodder invasion. Environ Exp Bot 181:1–14. https://doi.org/10.1016/j.envexpbot.2020.104272
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Dai YJ, Shen ZG, Liu Y, Wang LL, Hannaway D, Lu HF (2009) Effects of shade treatments on the photosynthetic capacity, chlorophyll fluorescence, and chlorophyll content of Tetrastigma hemsleyanum Diels et Gilg. Environ Exp Bot 65:177–182. https://doi.org/10.1016/j.envexpbot.2008.12.008
Pierik R, Djakovic-Petrovic T, Keuskamp DH, de Wit M, Voesenek LA (2009) Auxin and ethylene regulate elongation responses to neighbor proximity signals independent of gibberellins and Della proteins in Arabidopsis. Plant Physiol 149:1701–1712. https://doi.org/10.1104/pp.108.133496
Liu WG, Ren ML, Liu T, Du YL, Zhou T, Liu XM, Liu J, Hussain S, Yang WY (2018) Effects of shade stress on lignin biosynthesis in soybean stems. J Integr Agr 17:1594–1604
Podolec R, Ulm R (2018) Photoreceptor-mediated regulation of the COP1/SPA E3 ubiquitin ligase. Curr Opin Plant Biol 45:18–25. https://doi.org/10.1016/j.pbi.2018.04.018
Liu WG, Hussain S, Liu T, Zou JL, Ren ML, Zhou T, Liu J, Yang F, Yang WY (2019) Shade stress decreases stem strength of soybean through restraining lignin biosynthesis. J Integr Agr 18:43–53
Tang YJ, Liesche J (2017) The molecular mechanism of shade avoidance in crops-how data from Arabidopsis can help to identify targets for increasing yield and biomass production. J Integr Agr 16:1244–1255. https://doi.org/10.1016/S2095-3119(16)61434-X
Chaiwanon J, Wang WF, Zhu JY, Oh E, Wang ZY (2016) Information integration and communication in plant growth regulation. Cell 164:1257–1268. https://doi.org/10.1016/j.cell.2016.01.044
Li XJ, Chen XJ, Guo X, Yin LL, Ahammed GJ, Xu CJ, Chen KS, Liu CC, Xia XJ, Shi K, Zhou J, Zhou YH, Yu JQ (2016) DWARF overexpression induces alteration in phytohormone homeostasis, development, architecture and carotenoid accumulation in tomato. Plant Biotechnol J 14:1021–1033. https://doi.org/10.1111/pbi.12474
Stamm P, Kumar P (2010) The phytohormone signal network regulating elongation growth during shade avoidance. J Exp Bot 61:2889–2903. https://doi.org/10.1093/jxb/erq147
Gangappa SN, Berriri S, Khuma SV (2017) PIF4 coordinates thermosensory growth and immunity in Arabidopsis. Curr Biol 27(2):243–249. https://doi.org/10.1016/j.cub.2016.11.012
Casal JJ (2013) Photoreceptor signaling network in plant responses to shade. Annu Rev Plant Biol 64:403–427. https://doi.org/10.1146/annurev-arplant-050312-120221
Tao Y, Ferrer JL, Ljunf K (2008) Rapid synthesis of auxin via a new tryptophan-dependent pathway is required for shade avoidance in plants. Cell 133:164–176. https://doi.org/10.1016/j.cell.2008.01.049
Salehin M, Bagchi R, Estelle M (2015) SCFTIR1/AFB-based auxin perception: mechanism and role in plant growth and development. Plant Cell 27(1):9–19
Yang C, Xie F, Jiang YP, Li Z, Huang X, Li L (2018) Phytochrome A negatively regulates the shade avoidance response by increasing auxin/indole acidic acid protein stability. Dev Cell 44:29–41. https://doi.org/10.1016/j.devcel.2017.11.017
Moosavi B, Zhu XL, Yang WC, Yang GF (2020) Molecular pathogenesis of tumorigenesis caused by succinate dehydrogenase defect. Eur J Cell Biol 99(1):1–11. https://doi.org/10.1016/j.ejcb.2019.151057
Huang S, Taylor NL, Ströher E, Fenske R, Millar AH (2013) Succinate dehydrogenase assembly factor 2 is needed for assembly and activity of mitochondrial complex II and for normal root elongation in Arabidopsis. Plant J 73(3):429–441. https://doi.org/10.1111/tpj.12041
Funding
This work was funded by the National Natural Science Foundation of China (31902066), Scientific and Technological Projects of Henan Province (182102110241), and the Henan Provincial Natural Science Foundation, China (182300410058). We would like to thank the Nanjing Agricultural University for providing the chrysanthemum materials and the Program for Science & Technology Innovative Research Team in University of Henan Province (21IRTSTHN025).
Author information
Authors and Affiliations
Contributions
Performed the experiments: SH and QZ; analyzed the data: HW; prepared and wrote the manuscript: SH; conceived and designed the experiment: DP.
Corresponding author
Ethics declarations
Conflict of interest
The authors in this study declared no conflict of any interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Han, S., Zhang, Q., Wang, H. et al. Comparison of the transcriptomic responses of two Chrysanthemum morifolium cultivars to low light. Mol Biol Rep 48, 7293–7301 (2021). https://doi.org/10.1007/s11033-021-06729-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06729-8