Abstract
C3H10T1/2, a mouse mesenchymal stem cell line, is a well-known in vitro model of chondrogenesis that can be easily employed to recapitulate some of the mechanisms intervening in this process. Moreover, these cells can be used to validate the effect of candidate molecules identified by high throughput screening approaches applied to the development of targeted therapy for human disorders in which chondrogenic differentiation may be involved, as in conditions characterized by heterotopic endochondral bone formation. Chondrogenic differentiation of C3H10T1/2 cells can be monitored by applying quantitative polymerase chain reaction (qPCR), one of the most sensitive methods that allows detection of small dynamic changes in gene expression between samples obtained under different experimental conditions. In this work, we have used qPCR to monitor the expression of specific markers during chondrogenic differentiation of C3H10T1/2 cells in micromass cultures. Then we have applied the geNorm approach to identify the most stable reference genes suitable to get a robust normalization of the obtained expression data. Among 12 candidate reference genes (Ap3d1, Csnk2a2, Cdc40, Fbxw2, Fbxo38, Htatsf1, Mon2, Pak1ip1, Zfp91, 18S, ActB, GAPDH) we identified Mon2 and Ap3d1 as the most stable ones during chondrogenesis. ActB, GAPDH and 18S, the most commonly used in the literature, resulted to have an expression level too high compared to the differentiation markers (Sox9, Collagen type 2a1, Collagen type 10a1 and Collagen type 1a1), therefore are actually less recommended for these experimental conditions. In conclusion, we identified nine reference genes that can be equally used to obtain a robust normalization of the gene expression variation during the C3H10T1/2 chondrogenic differentiation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
C3H10T1/2 cells are a mouse cell line functionally similar to mesenchymal stem cells [1] that can be induced to differentiate towards chondrocyte, osteoblast and adipocyte lineages when cultured in presence of specific culture media [2,3,4]. In particular, these cells represent an established cellular model to simulate in vitro chondrogenesis, because they do not spontaneously differentiate under normal culture conditions, but can be induced by cellular condensation (high density micromass culture) and treatment with bone morphogenetic protein 2 (BMP2) [3, 5,6,7,8].
Due to limitations of primary cell lines and complexity of animal models, in the last decades, various in vitro cellular models with chondrogenic capabilities have been established to have cost effective, reproducible and simple experimental systems available to study the mechanisms that underlie chondrogenesis [8]. These in vitro models can be successfully applied also to recapitulate some of the crucial events that might be deregulated in pathological conditions.
Fibrodysplasia Ossificans Progressiva (FOP, MIM135100) is one of the most severe and disabling disorder due to extra-skeletal ossification (heterotopic ossification, HO). The disease is caused by mutations of the ACVR1 gene encoding a type I receptor for bone morphogenetic proteins (BMPs), and is characterized by ectopic bone formation affecting skeletal muscles, ligaments and tendons [9, 10]. Early FOP lesions are characterized by local inflammatory response, muscle degeneration and fibro-proliferative reaction, followed by mesenchymal condensation, chondrogenesis and bone neoformation. Therefore, cartilage differentiation can be considered a crucial step in FOP pathogenesis. Regarding the origin of cells undergoing the aberrant differentiation process, the source of the progenitors involved is still actively investigated and debated, and according to recent studies is likely to be heterogeneous [11,12,13,14,15].
So far, no specific treatment is available to cure FOP and prevent the occurrence of the ossification flares. However, in the last years many effort have been devoted to study the cellular and molecular mechanisms altered in the disease and the advancements of the research in the field has provided targets to develop specific therapies by both drug discovery or drug repositioning approaches (for a revision see [11, 16]).
In vitro cellular models like C3H10T1/2 cells represent an essential tool to evaluate compounds identified by high throughput screening (HTS) of large collections of small molecules to find out candidate “hits” to be further developed as targeted therapy in diseases in which chondrogenesis represents a crucial step as in FOP. Indeed, primary screenings provide a list of candidate molecules that need to be validated in reliable secondary assays to confirm their effect on defined targets and to investigate pathways involved in their activity.
The progression of chondrogenic differentiation in C3H10T1/2 micromass cultures can be monitored by applying specific histological stainings such as Alcian Blue or Toluidin Blue, able to detect the deposition of extra-cellular matrix glycosaminoglycans; or by verifying the expression of differentiation markers through immunohistochemical assays with specific antibodies; by using quantitative real-time PCR (qPCR) to evaluate changes in the expression of differentiation markers at different stages of the process.
qPCR is definitely one of the most sensitive methods that allows detection of small dynamic changes in gene expression between samples, therefore it is mandatory to proceed with extreme care in every step of the process, from sample preparation to data interpretation.
MIQE (minimum information for publication of quantitative real-time PCR experiments) guidelines help to define the most important steps from RNA to qPCR to minimize errors and the minimum set of information required to evaluate the reliability of qPCR data [17]. According to these guidelines, after accurate evaluation of RNA quality and cDNA synthesis, it is essential to rely on the use of solid reference genes. Moreover, it is highly recommended to use at least two different reference genes in order to normalize the expression of mRNA under analysis.
Nevertheless, few studies report a systematic screening of panels of reference genes before the choice of the best candidates to be used in relationship to the cell type, differentiation stage and experimental conditions [18]. Very frequently, only a single gene is applied to normalize, and the most commonly used remain Glyceraldehyde 3-phosphate dehydrogenase (GAPDH), β-actin (ActB) and 18S ribosomal RNA also in C3H10T1/2 differentiation studies [19,20,21].
Reference genes stability can be evaluated by several methods. The most popular rely on the use of dedicated programs such as geNorm [22], BestKeeper [23], and NormFinder [24] that are based on different algorithms, but all evaluate the variance in Cq values (quantification cycle, the cycle at which fluorescence from amplification exceeds the background fluorescence) of candidate reference genes across different samples, cell types, experimental conditions, etc.
In the current work, PrimerDesign geNormPLUS kit was employed to screen a panel of 12 reference genes in order to select the best candidates to be applied in monitoring the chondrogenic differentiation process in C3H10T1/2 cells.
To this aim, C3H10T1/2 were cultured in the appropriate conditions to induce chondrogenesis and used to select the best reference genes to obtain a reliable expression profile of differentiation markers which could be applied in different experimental settings.
Materials and methods
Cell culture
C3H10T1/2 cells were purchased from ATCC cell biology collection (C3H/10T1/2, Clone 8, ATCC® CCL-226™). Cells were used between the 5th and 15th passage and were routinely cultured in complete medium consisting of MEM with Earle’s Salts (Euroclone® S.p.a) supplemented with 10% fetal bovine serum (FBS, Gibco, ThermoFisher Scientific), 2 mM Glutamine, 100 U/ml Penicillin, 0.1 mg/ml Streptomycin (Euroclone® S.p.a). Cells were cultured at 37 °C in a humidified atmosphere with 5% CO2.
C3H10T1/2 and chondrocyte-like differentiation
In order to induce chondrocyte-like differentiation, micromass cultures were obtained as described by Denker et al., 1999 [3]. Briefly, cells were trypsinized and 107 cells/ml were resuspended in Ham’s F12 medium with 10% fetal bovine serum. 10 µl drop of cell suspension was placed in the center of a well in a standard 24-well plate (Euroclone® S.p.a). The cells were allowed to adhere for 2–3 h at 37 °C and 5% CO2, and then 500 µl of medium containing 100 ng/ml of BMP2 (Peprotech) was added to each well. Medium was replaced every 3 days.
To check chondrocyte differentiation, cells were either processed for RNA extraction or for Alcian Blue staining to verify glycosaminoglycans deposition. To this aim, micromass cultures were rinsed with Dulbecco’s phosphate-buffered saline (DPBS without Ca and Mg, Euroclone® S.p.a), fixed for 10 min in 4% Paraformaldehyde solution (Santa Cruz Biotechnology, Inc.) and stained overnight with a mixture containing 0.05% w/v Alcian Blue 8GX (Sigma-Aldrich), 0.2 M sodium acetate buffer, pH5.8 and 0.5 M MgCl2.
Quantitative PCR primers
In this study, we used the Mouse geNormPLUS kit (PrimerDesign) that includes a list of 12 reference genes (Ap3d1, Csnk2a2, Cdc40, Fbxw2, Fbxo38, Htatsf1, Mon2, Pak1ip1, Zfp91, 18S, ActB, GAPDH) selected for their high stability level in different biological samples and treatment conditions. Oligonucleotides detect all the transcript variants and are PrimerDesign proprietary information (See Table S1 for accession numbers and anchor nucleotide of the assays). Primers sequences of the other genes used in this work are shown below and were selected from the literature [25, 26] to produce mouse-specific amplicons. Primer sequences: Sox9, Fwd GAGCCCGATCTGAAGAAGGA, Rev GCTTGACGTGCGGCTTGTTC (151 bp, [25]); Collagen type 2a1, (Col2a1) Fwd CACCAAATTCCTGTTCAGCC, Rev TGCACGAAACACACTGGTAAG (124 bp, [25]); Collagen type 10a1, (Col10a1) Fwd TTCTGCTGCTAATGTTCTTGACC, Rev GGGATGAAGTATTGTGTCTTGGG (115 bp, [26]); Collagen type 1a1, (Col1a1) Fwd ATGCCGCGACCTCAAGATG, Rev TGAGGCACAGACGGCTGAGTA (153 bp, [26]).
RNA extraction and cDNA synthesis
Total RNA was isolated from T0, T1, T3, T6 and T13 time points by using the RNeasy Plus Mini kit (Qiagen) according to the manufacturers' protocol. RNA quality and concentration were established by electrophoresis on 1% Agarose gel (Fig S1) and by nanodrop spectrophotometer (ThermoFisher Scientific). All RNA samples displayed 260/280 ratios > 2.0 and first strand cDNA was obtained from 1 µg RNA, using the iScript reverse transcription supermix for RT-qPCR (Bio-Rad), according to the manufacturers’ instructions. cDNA preparations were diluted six-fold prior to quantitative PCR analysis.
Quantitative PCR (qPCR)
Gene expression profiles were evaluated through qPCR using the SYBER Green system, (IQ™ SYBER® Green Supermix, Bio-Rad). Reactions were prepared in duplicate in 20 µl volumes using 5 µl of diluted cDNA (around 25 ng cDNA assuming 1:1 synthesis) per well. qPCR was performed on the IQ5 instrument from Bio-Rad with the following protocol: 95 °C for 3 min, 40 amplification cycles: denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s and extension at 72 °C for 40 s. The presence of single specific amplification product was checked by melting curve analysis and template-free controls in all runs performed. All samples were quantified in the same run for a given reference gene and reaction efficiencies were estimated by standard dilution curve and analysis of individual traces.
Quantification cycle (Cq) values for each replicate reaction were extracted from IQ5 analysis and analyzed through the qBasePLUS software (Biogazelle, https://www.qbaseplus.com/) to calculate geNorm values. Details about geNorm algorithm are described in Vandesompele et al., 2002 [22]. Relative expression of differentiation markers was calculated by the ∆Ct Method, derived as a modification of the 2−∆∆Ct (Livak) approach [27]. The applied ∆Ct Method uses the difference between reference and target Ct values for each sample, (Ratio (reference/target) = 2 Ct(reference)−Ct(target)).
Statistical analysis
The results obtained from reference genes study were analyzed by the qBasePLUS software (Biogazelle). Experiments to evaluate gene expression profile by qPCR were performed from three independent biological replicates and the results were represented as average ± Standard Deviation of gene relative expression. The unpaired two-tailed Student’s t-test (GraphPad, https://www.graphpad.com/quickcalcs/ttest1) was applied to verify statistical significance of the observed variation. Significant variations were defined as *P < 0.05.
Results
Chondrogenic differentiation of C3H10T1/2
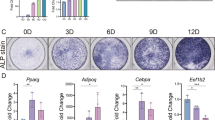
C3H10T1/2 micromass cell cultures were induced to differentiate with BMP2, starting from day 1 to day 13. In order to evaluate the differentiation process, micromass cultures were stained with Alcian Blue, specific for the deposition of the matrix glycosaminoglycans, after 1, 3, 6, and 13 days of culture in inductive medium (Fig. 1). The expression level of markers associated with chondro-differentiation and cartilage formation (Sox9, Col2a1, Col10a1 and Col1a1) was also monitored (Fig. S2).
Study of 12 candidate reference genes
In parallel, cells cultured in the same conditions and blocked at the different time points were used to identify the best reference genes suitable to study gene expression variation during differentiation of C3H10T1/2. RNA quality and concentration (Fig. S1) were checked for all the samples deriving from two different groups of experiments and the expression of a panel of 12 reference genes was evaluated. qPCR results were processed with the IQ5 software and Cq values were exported to be analysed with qbase+ software (Table S2). Amplification efficiency, calculated from standard curves was between 1.9 and 2.10 for all genes, corresponding to 90–110% which represents the recommended range (Table S1).
GeNorm study was applied following the instruction of the software. GeNorm M value represents reference gene stability and according to the geNorm manual, candidate genes with lower M value have to be considered more stable. The graph starts with the least stable gene on the left side and ends with the most stable gene on the right part, with a threshold of 0.5 between stable and unstable genes. Therefore, all the tested reference genes appeared to be stable upon the different conditions and thus suitable for gene expression analysis at different time points of C3H10T1/2 chondrogenic differentiation (Fig. 2a). However, the best candidate reference genes identified were Mon2 and Ap3d1, whereas the less stable were ActB and GAPDH, curiously among the most commonly used in the literature. The geNorm V value suggests the minimum number of reference genes required for normalization. The threshold value is 0.15, below which the inclusion of an additional control is not required [22]. As shown in Fig. 2b, all the reference genes combination has Vn/n+1 < 0.15 (n = optimal number of reference genes), suggesting that a combination of two reference genes was sufficient for a correct normalization.
GeNorm analysis of 12 candidate reference genes. a geNorm M value indicates reference genes stability. The genes with geNorm M value below 0.5 can be considered for the normalization and the lower is the value, the more stable are the genes. b GeNorm V value represents the pairwise variation (n/n + 1) and indicates the minimum number of reference genes required for normalization. Values below 0.15 indicate the optimal number (n) of reference genes to be considered. In this study, a combination of two genes was sufficient for a correct normalization, V2/3 = 0.082
Expression profile of the differentiation markers normalized against the different reference genes
To validate geNorm analysis and reference genes selection, C3H10T1/2 micromass cultures were harvested at time 0 and after 6 days of chondrogenic differentiation and processed for qPCR assay.
The expression level of markers associated with chondro-differentiation and cartilage formation (Sox9, Col2a1, Col10a1) was obtained by the ∆Ct Method. In this evaluation, various reference genes were employed, including all single candidate reference genes, a combination of the most stable ones identified in this study, calculated by the geometric average of Mon2&Ap3d1, Ap3d1&Fbxw2, Mon2&Ap3d1&Fbxw2 (indicated as M&A, A&F, M&A&F), and the geometric average of all 12 reference genes Ct values, (designed as ‘All’). As shown in Fig. 3 up-regulation of the chondrogenic genes Sox9, Col2a1, Col10a1 was demonstrated by all normalization strategies. As a control, we evaluated also the expression of Col1a1, a gene not associated with cartilage differentiation and expected to be downregulated in micromass cultures induced by BMP2 [28]. As shown in Fig. 3, the decrease in Col1a1 expression upon differentiating conditions does not appear significant when ActB is applied as normalizing gene, compared to ‘All’ and to other single or combined reference genes.
Fold-change expression of chondrogenic differentiation markers, normalized against different reference genes after 6 days of differentiation. aSox9, bCol2a1, cCol10a1, dCol1a1. Relative expression was calculated by ∆Ct Method and histograms represent average ± Standard Deviation of three independent biological replicates. The statistical significance was evaluated by Student’s T-test analysis with *P < 0.05
Comparison of relative expression of 12 candidate reference genes
In order to select the proper reference genes with transcription level similar to that of target genes, Ct values deriving from qPCR assay on RNA samples at T0 and T6, were normalized against Fbxo38, which showed the lowest expression level.
As reported in Fig. 4, relative expression of most of the reference genes have similar magnitude, whereas ActB, GAPDH and 18S showed the highest expression in C3H10T1/2 cells.
Relative expression of the 12 reference genes. Cells were analysed after 6 days of differentiation. Gene expression values were calculated using ∆Ct Method against Fbxo38, that showed the lowest expression. Histograms represent average and standard deviation of three independent biological replicates. Y axis was on a log10 scale
This analysis suggested the exclusion of ActB, GAPDH and 18S because their transcription levels were too far from that of target genes and this might affect the accuracy of the normalization process.
Discussion
qPCR is one of the most sensitive method to evaluate the expression level of genes of interest under different experimental conditions, and thus in order to obtain reliable results it is crucial to select the suitable reference genes [26, 29,30,31,32,33,34].
In the last years, several algorithms were developed to identify the optimal reference genes in different cell types/tissues or treatment conditions. GeNorm [22], BestKeeper [23], and NormFinder [24] are the most frequently applied to this purpose and are reported to usually show similar results [35, 36].
To the aim of using C3H10T1/2 cells as an in vitro model of chondro-like differentiation [3], we applied the geNorm software to identify the most stable references genes suitable for normalization during the in vitro differentiation process of these cells. GeNorm has been developed according to the normalization strategy described by Vandesompele et al., 2002 [22], conceived to identify the most stably expressed control genes in a given set of tissues, and to determine the minimum number of genes required to calculate a reliable normalization factor.
In our study, we used a commercial kit of 12 murine ‘house-keeping’ genes, specifically designed by PrimerDesign for geNorm studies, and we performed two biological replicates of C3H10T1/2 chondrogenic differentiation after 1, 3, 6, and 13 days.
The first parameter generated by the geNormPLUS analysis is geNorm M, which evaluates expression stability of candidate reference genes between samples. M value represents the average pairwise variation of a particular gene with all the other control genes: the lowest M value indicates the most stable gene expression. The cut-off value is 0.5 and all the genes we analyzed in this study could be considered potential reference genes.
According to the MIQE guidelines, an accurate normalization process relies on the use of two or more reference genes and geNorm V value (pairwise variation, V(n/n+1) coefficient) helps to determine the number of them required for optimal normalization. The suitable number of reference genes “n” is found when V(n/n+1) drops below 0.15.
As shown in Fig. 2, V2/3 was less than 0.15 (V2/3 = 0.082) and two reference genes were sufficient for a correct normalization, ideally Mon2 and Ap3d1.
To validate geNorm analysis, we evaluated the expression levels of markers specific for chondrogenic differentiation of C3H10T1/2 cells at T0 and after 6 days of micromass culture in inductive medium containing BMP2. Sox9, Col2a1 and Col10a1 showed high expression level after BMP2 treatment and all normalization strategies showed comparable expression level. In particular, the combination of Mon2 and Ap3d1, Ap3d1 and Fbxw2, and Mon2, Ap3d1 and Fbxw2, gave similar results to the normalization obtained with ‘All’, which is considered as the most accurate strategy.
Furthermore, it is known that for accurate gene expression studies the expression level of reference genes should be similar to that of target genes [26, 37]. Therefore, we compared the expression profile of all the 12 reference genes in our cell model. Our analysis suggested that ActB, GAPDH and especially 18S genes should not be considered for normalization because their expression level significantly diverges from that of the selected differentiation markers in C3H10T1/2 cells.
Moreover, the use of ActB as reference gene failed to efficiently detect the expected down-regulation of Col1a1 expression in differentiating conditions [28].
This result suggested the exclusion of ActB, one of the most applied in expression studies, from the best reference genes group, because during the evaluation of treatment effects, also small gene expression differences may become relevant. All the other reference genes and combination analysed in this work, showed similar expression levels and pass our validation.
Conclusions
qPCR is a potent method to study gene expression profiles in different experimental settings. Reliability of results is strongly dependent on the normalization procedure; reference genes have to be carefully selected for every tissue/cell types and for the different experimental conditions. We applied the geNorm analysis to select the best reference genes in a widely used in vitro model of chondrogenesis. According to our analysis, the most commonly used reference genes, such as ActB, GAPDH and 18S, are not actually the best candidates for normalization. We suggest to use a different combination of at least two genes selected from the panel we tested, such as Mon2 and Ap3d1 when normalizing gene expression profiles during chondrogenesis in C3H10T1/2 cells.
References
Reznikoff CA, Brankow DW, Heidelberger C (1973) Establishment and characterization of a cloned line of C3H mouse embryo cells sensitive to postconfluence inhibition of division. Cancer Res 33(12):3231–3238
Ahrens M, Ankenbauer T, Schröder D, Hollnagel A, Mayer H, Gross G (1993) Expression of human bone morphogenetic proteins-2 or -4 in murine mesenchymal progenitor C3H10T1/2 cells induces differentiation into distinct mesenchymal cell lineages. DNA Cell Biol 12(10):871–880
Denker AE, Haas AR, Nicoll SB, Tuan RS (1999) Chondrogenic differentiation of murine C3H10T1/2 multipotential mesenchymal cells: I. Stimulation by bone morphogenetic protein-2 in high-density micromass cultures. Differentiation 64(2):67–76
Fakhry M, Hamade E, Badran B, Buchet R, Magne D (2013) Molecular mechanisms of mesenchymal stem cell differentiation towards osteoblasts. World J Stem Cells 5(4):136
Haas AR, Tuan RS (1999) Chondrogenic differentiation of murine C3H10T1/2 multipotential mesenchymal cells: II. Stimulation by bone morphogenetic protein-2 requires modulation of N-cadherin expression and function. Differentiation 64(2):77–89
Izzo MW, Pucci B, Tuan RS, Hall DJ (2002) Gene expression profiling following BMP-2 induction of mesenchymal chondrogenesis in vitro. Osteoarthritis Cartilage 10(1):23–33
Roy R, Kudryashov V, Doty SB, Binderman I, Boskey AL (2010) Differentiation and mineralization of murine mesenchymal C3H10T1/2 cells in micromass culture. Differentiation 79(4–5):211–217
Takács R, Matta C, Somogyi C, Juhász T, Zákány R (2013) Comparative analysis of osteogenic/chondrogenic differentiation potential in primary limb bud-derived and C3H10T1/2 cell line-based mouse micromass cultures. Int J Mol Sci 14(8):16141–16167
Shore EM, Xu M, Feldman GJ, Fenstermacher DA, Cho TJ, Choi IH et al (2006) A recurrent mutation in the BMP type I receptor ACVR1 causes inherited and sporadic fibrodysplasia ossificans progressiva. Nat Genet 38(5):525-7. Epub 2006 Apr 23. Erratum in: Nat Genet. 2007 Feb;39(2):276
Kaplan FS, Xu M, Seemann P, Connor JM, Glaser DL, Carroll L et al (2009) Classic and atypical fibrodysplasia ossificans progressiva (FOP) phenotypes are caused by mutations in the bone morphogenetic protein (BMP) type I receptor ACVR1. Hum Mutat 30(3):379–390
Cappato S, Giacopelli F, Ravazzolo R, Bocciardi R (2018) The horizon of a therapy for rare genetic diseases: a “druggable” future for fibrodysplasia ossificans progressiva. Int J Mol Sci 19(4):989
Medici D, Olsen BR (2012) The role of endothelial-mesenchymal transition in heterotopic ossification. J Bone Miner Res 27(8):1619–1622
Wosczyna MN, Biswas AA, Cogswell CA, Goldhamer DJ (2012) Multipotent progenitors resident in the skeletal muscle interstitium exhibit robust BMP-dependent osteogenic activity and mediate heterotopic ossification. J Bone Miner Res 27(5):1004–1017
Dey D, Bagarova J, Hatsell SJ, Armstrong KA, Huang L, Ermann J et al (2016) Two tissue-resident progenitor lineages drive distinct phenotypes of heterotopic ossification. Sci Transl Med 23(366):366ra163
Lees-Shepard JB, Goldhamer DJ (2018) Stem cells and heterotopic ossification: lessons from animal models. Bone 109:178–186
Katagiri T, Tsukamoto S, Kuratani M (2018) Heterotopic bone induction via BMP signaling: potential therapeutic targets for fibrodysplasia ossificans progressiva. Bone 109:241–250
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M et al. (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55(4):611–622
Chapman JR, Waldenström J (2015) With reference to reference genes: a systematic review of endogenous controls in gene expression studies. PLoS ONE 10(11):e0141853
Huang X, Xu J, Huang M, Li J, Dai L, Dai K et al. (2014) Histone deacetylase1 promotes TGF-β1-mediated early chondrogenesis through down-regulating canonical Wnt signaling. Biochem Biophys Res Commun 453(4):810–816
Howard M, Tuan RS, Wallis GA (2016) The function and interrelationship between GDF5 and ERG-010 during chondrogenesis in vitro. In Vitro Cell Dev Biol Anim 52(2):182–192
Dai J, Yu D, Wang Y, Chen Y, Sun H, Zhang X et al (2017) Kdm6b regulates cartilage development and homeostasis through anabolic metabolism. Ann Rheum Dis 76(7):1295–1303
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A et al. (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7): research0034–1
Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP (2004) Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol Lett 26(6):509–515
Andersen CL, Jensen JL, Ørntoft TF. (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64(15):5245–5250
Xiong Y, He J, Zhang W, Zhou G, Cao Y, Liu W (2015) Retention of the stemness of mouse adipose-derived stem cells by their expansion on human bone marrow stromal cell-derived extracellular matrix. Tissue Eng Part A 21(11–12):1886–1894
Zhai Z, Yao Y, Wang Y (2013) Importance of suitable reference gene selection for quantitative RT-PCR during ATDC5 cells chondrocyte differentiation. PLoS ONE 8(5):e64786
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)). Method. Methods 25(4):402–408
Carlberg AL, Pucci B, Rallapalli R, Tuan RS, Hall DJ (2001) Efficient chondrogenic differentiation of mesenchymal cells in micromass culture by retroviral gene transfer of BMP-2. Differentiation 67(4–5):128–138
Sugden K, Pariante CM, McGuffin P, Aitchison KJ, D’Souza UM (2010) Housekeeping gene expression is affected by antidepressant treatment in a mouse fibroblast cell line. J Psychopharmacol 24(8):1253–1259
Crowther LM, Wang SC, Eriksson NA, Myers SA, Murray LA, Muscat GE (2011) Chicken ovalbumin upstream promoter-transcription factor II regulates nuclear receptor, myogenic, and metabolic gene expression in skeletal muscle cells. Physiol Genomics 43(4):213–227
Arsenijevic T, Grégoire F, Delforge V, Delporte C, Perret J (2012) Murine 3T3-L1 adipocyte cell differentiation model: validated reference genes for qPCR gene expression analysis. PLoS ONE 7(5):e37517
Zhang J, Tang H, Zhang Y, Deng R, Shao L, Liu Y et al (2014) Identification of suitable reference genes for quantitative RT-PCR during 3T3-L1 adipocyte differentiation. Int J Mol Med 33(5):1209–1218
Hildyard JC, Wells DJ (2014) Identification and validation of quantitative PCR reference genes suitable for normalizing expression in normal and dystrophic cell culture models of myogenesis. PLoS Curr. https://doi.org/10.1371/currents.md.faafdde4bea8df4aa7d06cd5553119a6
Ferraz FB, Fernandez JH (2016) Selection and validation of reference house-keeping genes in the J774A1 macrophage cell line for quantitative real-time PCR. Genet Mol Res 15(1):15017720
McCulloch RS, Ashwell MS, O’Nan AT, Mente PL (2012) Identification of stable normalization genes for quantitative real-time PCR in porcine articular cartilage. J Anim Sci Biotechnol 12(1):36
Wang Q, Ishikawa T, Michiue T, Zhu BL, Guan DW, Maeda H (2012) Stability of endogenous reference genes in postmortem human brains for normalization of quantitative real-time PCR data: comprehensive evaluation using geNorm, NormFinder, and BestKeeper. Int J Legal Med 126(6):943–952
de Kok JB, Roelofs RW, Giesendorf BA, Pennings JL, Waas ET, Feuth T et al (2005) Normalization of gene expression measurements in tumor tissues: comparison of 13 endogenous control genes. Lab Invest 85(1):154–159
Acknowledgements
This study was funded by Fondazione Telethon (Grant No. GGP15196 to RR), by FOP Italia Onlus, “Ricerca corrente” and Italian Ministry of Health.
Author information
Authors and Affiliations
Contributions
SC conceived, designed, performed and analysed qPCR experiments, and wrote the manuscript; FG and LT performed differentiation assays; RR and RB supervised the experiments and manuscript preparation.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2019_4713_MOESM1_ESM.tif
Supplementary material 1—Agarose gel electrophoresis of representative RNA samples deriving from C3H10T1/2 micromasses. Integrity of RNA was represented by the presence of two bands corresponding to 28S and 18S ribosomal subunits. RNA from two different experiments for T0 (Lanes 1 and 2), and from two different experiments for T6 time point (Lanes 3 and 4) are shown (TIF 1228 KB)
11033_2019_4713_MOESM2_ESM.tif
Supplementary material 2—Expression level of the indicated markers after 1, 3, 6 and 13 days of chondrogenic differentiation. (A) Sox9, (B) Col2a1, (C) Col10a1, (D) Col1a1. The expression of each gene has been normalized against the geometric average of all the 12 Reference genes. Histograms represent the average ± Standard Deviation of two independent experiments (TIF 37 KB)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Cappato, S., Giacopelli, F., Tonachini, L. et al. Identification of reference genes for quantitative PCR during C3H10T1/2 chondrogenic differentiation. Mol Biol Rep 46, 3477–3485 (2019). https://doi.org/10.1007/s11033-019-04713-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04713-x