Abstract
Chloroplast is a crucial organelle for plant photosynthesis and maintaining normal life activities in higher plants. Although some genes related to chloroplast development and pigment synthesis have been identified or cloned in rice, little is known about the relationship between these genes and abiotic stress response. In this study, we identified a novel mutant white stripe leaf 12 (wsl12) affecting pigment synthesis, chloroplast development and abiotic stress response in rice. The mutant phenotype was obvious at seeding and tillering stages and in response to the temperature change. Genetic analysis of reciprocal crosses between wsl12 and wild-type plants showed that wsl12 was a recessive mutant in a single nuclear locus. Map-based cloning revealed that the WSL12 locus encoded OsNDPK2, one of the three nucleoside diphosphate kinases (OsNDPKs). WSL12 expressed in all tested tissues, while it highly expressed in leaves and young tissues. The WSL12 protein localized to the chloroplast. The wsl12 mutant showed higher superoxide anion level and enhanced sensitivity to abscisic acid (ABA) and salinity. The transcription pattern of many genes involved in chlorophyll biosynthesis, ABA synthesis, light signaling pathway, reactive oxygen species-scavenging pathway and the other two OsNDPKs was altered in the wsl12 mutant. These results indicate that the OsNDPK2 encoded by WSL12 plays an important role in chloroplast development and chlorophyll biosynthesis by regulating the expression levels of related genes. In addition, WSL12 also affects the response to abiotic stress, such as ABA and salinity in rice, and is beneficial to molecular breeding of stress tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The transformation from proplastids to photosynthetically active chloroplasts is regulated by plastid and nuclear genes. The chloroplast proteins essential for chloroplast development and functions are mainly encoded by nuclear genes (Chen et al. 2010). Chloroplast only encodes about 100 genes because it is a semiautonomous organelle (Delannoy et al. 2009). The mutation of these genes may result in the generation of many mutants with visible leaf discoloration and abnormal seedling viability (Zhen et al. 2014; Guo et al. 2015). The mutants with abnormal pigment contents or deficient in chloroplast development are ideal materials for studying the underlying mechanisms that regulate chlorophyll biosynthesis and chloroplast development in rice. Many of these genes have been identified, including OsCHLH, Chlorina 1, Chlorina 9, V1, V2, V3, ST1, OsPPR1 and WSL (Jung et al. 2003; Zhang et al. 2006; Kusumi et al. 2011; Sugimoto et al. 2007; Yoo et al. 2009; Gothandam et al. 2005; Tan et al. 2014).
Nucleoside diphosphate kinases (NDPKs) are highly conserved enzymes that catalyze the transfer of terminal phosphoryl group from nucleoside triphosphate (NTP) to nucleoside diphosphate through a ping-pong mechanism (Parks and Agarwal 1973). In animals, NDPKs participate in cell proliferation, differentiation, invasion and motility (Keim et al. 1992; Kantor et al. 1993). In plants, the main function of NDPKs is to maintain the balance between cellular NTPs and nucleoside diphosphates and provide NTPs for biosynthesis except for adenosine triphosphate (Roberts et al. 1997; Bernard et al. 2000). In addition to the housekeeping role, previous researches demonstrate that NDPKs are involved in other processes, such as phytochrome-mediated light signaling (Choi et al. 1999), UV-B signaling (Zimmermann et al. 1999), wounding (Harris et al. 1994), heat-shock response (Escobar Galvis et al. 2001), H2O2-mediated MAPK signaling (Moon et al. 2003) and auxin-mediated response (Choi et al. 2005). Plant NDPKs have been classified into three distinct types (type I–III) based on their amino acid sequences. All NDPKs sequences share a common catalytic histidine residue, and several of them exhibit an N-terminal extension for organellar targeting (Lascu and Gonin 2000; Dorion and Rivoal 2015). AtNDPK2 is considered to be linked with chloroplast function, oxidative stress and auxin signaling (Dorion and Rivoal 2015). Higher expression level of AtNDPK2 is found in leaves and inflorescences, but lower or absent in roots (Verslues et al. 2007; Hammargren et al. 2007). Previous studies have shown that AtNDPK2 activity can be increased by binding to phytochrome A (Choi et al. 1999; Shen et al. 2005). The Atndpk2 mutant showed deficient in cotyledon opening and greening in both red and far-red lights (Choi et al. 1999). These results indicate that AtNDPK2 participates in the photomorphogenesis of leaves. NDPK2 can also regulate auxin-mediated responses for plant growth and development mainly through changing the expression of auxin-related genes to affect auxin transport (Choi et al. 2005). Among the three types of NDPKs, only NDPK2 is considered associated with reactive oxygen species (ROS) signaling and oxidative stress management. The Atndpk2 mutant has higher ROS and H2O2 levels compared to its wild-type plants (Moon et al. 2003; Verslues et al. 2007). Under stress conditions, AtNDPK2 is involved in oxidative stress signaling by interacting with AtMPK3 and AtMPK6, two H2O2-activated Arabidopsis thaliana mitogen-activated protein kinases (Moon et al. 2003). AtNDPK2 also participates in salt stress signaling by interacting with class 3 sucrose nonfermenting 1-related kinase (SOS2) and catalase (CAT; Verslues et al. 2007). AtNDPK2 can regulate the activity of G protein since AtNDPK2 directly interacts with small G-proteins and controls their activities. It has been proposed that NDPK2 may be a missing link between the phytochrome-mediated light signaling and G protein-mediated signaling (Shen et al. 2008).
Many studies related to NDPK2 have been carried out in the dicotyledon model plant, and both the structure and function of AtNDPK2 have been well characterized in previous studies (Dorion and Rivoal 2015). However, little is known about the function of NDPK2 in rice (Oryza sativa L.). In this study, we investigated a white stripe leaf 12 (wsl12) mutant, which exhibits white stripe leaf at seedling and tillering stages, and the phenotype is also affected by temperature. Genetic analysis of reciprocal crosses between wsl12 and wild-type plants showed that wsl12 was a recessive mutant in a single nuclear locus. Map-based cloning revealed that WSL12 encodes the nucleoside diphosphate kinase 2 (OsNDPK2). Further researches demonstrate that OsNDPK2 plays an important role in chlorophyll biosynthesis, chloroplast development and abiotic stress response. In addition, the transcription levels of the genes associated with abscisic acid (ABA) synthesis, ROS-scavenging pathway and light signaling pathway were changed in the wsl12 mutant, which suggests the function of WSL12 is pleiotropic.
Materials and methods
Plant materials and growing conditions
The rice wsl12 mutant was obtained from the mutagenized populations of the japonica rice cultivar Wuyujing7 treated by EMS solution. The wsl12 was crossed with the wild-type Wuyujing7 for genetic analysis. For fine mapping WSL12 locus, a F2 population derived from the cross between wsl12 and the indica cultivar 93-11 was used. Rice plants were grown in the paddy fields in Hangzhou (30°N latitude, summer season) and Sanya (18°N latitude, winter season) or in plant incubators (Panasonic, MLR-352H-PC). In the temperature experiments, seedlings were grown in incubators (12-/12-h light/dark; light intensity 300 μmol m−2 s−1) at the constant temperature of 22, 26, 30 and 34 °C conditions, respectively.
Pigment content measurement
Chlorophyll and carotenoid concentrations were detected according to the method described by Arnon (1949) and Wellburn (1994). Leaf samples (0.15 g, fresh weight) were cut into segments, immersed in 10 ml 80 % acetone and incubated at 26 °C in dark for 24 h. The optical density of sample solutions was measured by ultraviolet spectrophotometer (DU800, Beckman, USA) with 663 nm (the maximum absorption peak of chlorophyll a), 645 nm (the maximum absorption peak of chlorophyll b) and 470 nm (the maximum absorption peak of carotenoid). Each sample was measured by three biological repeats.
Transmission electron microscopy analysis of chloroplast ultrastructure
The leaves from the wsl12 mutant and wild-type plants were cut into small pieces, placed in a phosphate buffer containing 2.5 % glutaraldehyde (pH = 7.2) and then vacuumed in a vacuum pumping machine until the leaves sank. The transmission electron microscopy samples were treated according to the method described by Gothandam et al. (2005). The processed samples were examined with a transmission electron microscope (Hitachi H-7650, Tokyo, Japan).
NBT staining
Superoxide anion was detected by NBT staining according to the method described by Thordal-Christensen et al. (1997). The fully-expanded third leaves from wsl12 mutant and wild-type plants were detached and soaked in NBT solution (0.5 mg/ml, NBT powder was dissolved with K3PHO4 solution, pH = 7.6) in the dark for 5 h.
Genetic analysis and map-based cloning of WSL12
A total of 118 polymorphic markers scattered across all rice chromosomes and a DNA bulk pool from 33 individuals with the mutant phenotype of F2 population were used for preliminary linkage analysis. Subsequently, the fine mapping of WSL12 was performed using 2257 plants with recessive phenotype. Simple sequence repeat markers were obtained from the Gramene database (http://www.Gramene.org). New polymorphic markers were designed using the Primer Premier 5.0 software based on the sequence differences between the japonica rice cultivar Nipponbare and the indica cultivar 93-11. DNA was extracted from fresh rice leaves using the CTAB method (Murray and Thompson 1980). PCR-based molecular markers used to identify the genotypes of these progenies are listed in Additional file 2: Table S1. The 20 μl PCR mixture consisted of 50 ng template DNA, 2.0 μl 10× PCR buffer, 0.2 mM dNTP, 0.2 μM of each primer and 0.2 U Taq DNA polymerase. The PCR procedure was as follows: initial denaturation step at 94 °C/4 min; followed by 40 cycles of 94 °C/30 s, annealing (temperature changes were based on primers, usually 55 °C) for 30 s, 72 °C/30 s; and a final extension step at 72 °C/10 min. Products were separated with 4 % agarose gels, stained by GelRed and visualized by UV radiation.
Vector construction and transformation
A 9826-bp wild-type genomic DNA fragment containing the entire WSL12 coding region, a 2074-bp upstream sequence and a 1641-bp downstream region was amplified by PCR using the primer pair LC1. PCR products were digested with KpnI and XbaI. The target fragment was purified and then inserted into the binary vector pCAMBIA1300. The recombinant plasmid, named pWSL12 and pCAMBIA1300 (pCK, contained the promoter of WSL12 and part of the code region), was introduced into the calluses generated from mature seed embryos of wsl12 mutant using Agrobacterium-mediated transformation (Hiei et al. 1994). The 1.9-kb sequence of WSL12 promoter was amplified by PCR using the primer pair LGUS. PCR products were digested with EcoRI and BglII. The fragment was purified and then inserted into the binary vector pCAMBIA1305. The recombinant plasmid was used to transform Wuyujing7 following the protocol described above. The positive transgenic plants were visualized using GUS staining assay (Scarpella et al. 2003). For over-expression vector construction, the entire cDNA of WSL12 was amplified using the primer pair LOE. PCR products were digested with KpnI and XbaI. The target fragment was inserted into the vector pCAMBIA1300s (35S promoter was inserted into pCAMBIA1300) and then transformed wsl12. Primers used for vector construction are listed in Additional file 2: Table S2.
RNA preparation and quantitative real-time PCR
The total RNA of various samples from wsl12 and wild-type plants was extracted using the MicroRNA Extraction kit (Axygen, http://www.axygenbio.com), and the first-strand cDNA was synthesized using the ReverTra Ace qPCR RT kit (Toyobo Co. Ltd.; http://www.toyobo.cn/) following the manufacturer’s instructions. The expression levels of WSL12 and the genes associated with chlorophyll synthesis, chloroplast development, ABA synthesis and abiotic stress were assessed using quantitative real-time PCR (qRT-PCR). OsActin1 (Os03g0718100) was used as the internal control. All relevant primers are listed in Additional file 2: Table S3. QRT-PCR was conducted on Applied Biosystems 7900HT. Each 10 μl reaction system consisted of 1 μl total cDNA, 5 μl 2× SYBR Green PCR Master Mix (TaKaRa Co. Ltd., Otsu, Japan; http://www.takara.com.cn) and 4 μl of 1 μM primers. QRT-PCR conditions were as follows: 95 °C/5 min, followed by 40 cycles of 95 °C/30 s, 60 °C/30 s and 72 °C/30 s. For each sample, qRT-PCR results were measured by four technical replicates and three biological replicates. The 2−ΔΔCT method was adopted to analyze the relative expression levels of relevant genes (Livak and Schmittgen 2001), and two-tailed t test was used to detect statistically significant differences.
Subcellular location of WSL12 in rice protoplast
To investigate the subcellular localization of WSL12 protein, the WSL12 full-length cDNA without termination codon was cloned into the AHLG (reconstructed from pCAMBIA1300) binary vector. WSL12 sequence was placed upstream of GFP coding sequence. The fusion plastid and the control plastid (empty vector) were transformed into rice protoplasts according to the protocols described previously (Chen et al. 2006). Fluorescence signals were visualized with confocal fluorescence scanning microscope (LSM780, Carl Zeiss, Germany).
Results
Phenotypic characteristics of the wsl12 mutant
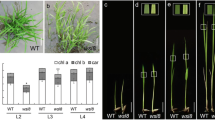
The wsl12 mutant showed white stripe leaf phenotype when the first leaf fully developed (Fig. 1a). But these abnormal leaves gradually reverted to green along with the plant development. The newly development leaves after the third leaf stage displayed green color (Fig. 1b). More interesting, abnormal leaves also appeared at tillering stage. Three types of leaves could be observed in the wsl12 mutant plants at tillering stage (Fig. 1c, e, f): Almost whole white leaf (ml1) mainly distributed on the new leaves of new tillers (Fig. 1e), white stripe leaf (ml2) and green leaf (ml3) (Fig. 1f). The ml1 leaves gradually turned to green, and then only ml3 leaves could be observed at mature stage (Fig. 1d). The panicles of wsl12 also showed abnormal phenotype at initial heading stage and then turned to normal. The wsl12 mutant showed lower photosynthetic rate compared with that of wild-type plants (Additional file 1: Fig. S1). Compared with the wild type, the wsl12 mutant exhibited significant differences in the major agronomic traits, including plant height, seed-setting rate, heading date and the number of tillers per plant (Additional file 1: Fig. S2). The plant height of wsl12 was approximately 76.7 ± 1.11 cm, which was 9.25 % shorter than the wild type (83.8 ± 1.1 cm). The seed-setting rate of wsl12 was about 56.5 ± 3.5 %, which was much lower than that of the wild type (96 ± 0.03 %). The wsl12 mutant exhibited delayed flowering time and less tillers per plant. There was no significant difference between wsl12 and wild-type plants in panicle length, the number of primary/secondary branches and 1000-grain weight (Additional file 1: Fig. S2). Previous studies showed that leaf color was closely linked with photosynthetic pigments and chloroplast development (Murchie et al. 2005). Our results indicated that the concentrations of chlorophyll a (Chla), chlorophyll b (Chlb) and carotenoid in ml1 and ml2 leaves decreased significantly compared with that in the wild type, whereas the pigment concentrations of ml3 leaves were almost normal (Fig. 1g). Transmission electron microscopy analysis showed the lamellar structure of chloroplasts in the wild type was well developed (Fig. 2a, d). But in the wsl12 mutant, there was no chloroplast or chloroplast-like organelle in ml1 leaves (Fig. 2b). Although the chloroplast number and size between ml3 and wild type did not show significant difference (Fig. 2c), there was no well-organized lamellar structure in the ml3 leaves (Fig. 2e). These observations indicate that the wsl12 phenotypes are closely associated with chloroplast development and chlorophyll synthesis.
Phenotypic characterization of wild type and the wsl12 mutant. Phenotypes of wild-type (left) and wsl12 mutant (right) plants at two-leaf stage (a), four-leaf stage (b), tillering stage (c) and filling stage (d). The red and white arrows show normal and white stripe region of leaves in wild type and wsl12 mutant, respectively. e New tiller of wsl12 mutant, white arrow shows the new leaf. f Three leaf types of the wsl12 mutant at tillering stage. g Pigment content in leaves of wild-type and wsl12 mutant at tillering stage. Chlorophyll a: chl a, chlorophyll b: chl b, carotenoid: car. Data represent mean ± SD from three independent biological replicates. ml1: white leaf in wsl12, ml2: white stripe leaf in wsl12, ml3: green leaf in wsl12. Bars 5 cm in a, b, e and f and 10 cm in c and d
Transmission electron microscopic images of cells from wild type and the wsl12 mutant at tillering stage. a Cells in leaf of wild type. b Cells in white sectors of leaf in wsl12 mutant. c Cells in green sectors of wsl12 mutant. d Chloroplast from wild type. e Chloroplast in green sectors of the mutant. Bars 5 μm in a–c and 0.5 μm in d and e. C chloroplast, N nucleus
The phenotype of wsl12 is affected by temperature
To examine whether the pigment content in the mutant wsl12 is affected by temperature, we compared the pigment concentrations between the wsl12 mutant and wild-type plants grown at the constant temperature of 22, 26, 30, and 34 °C conditions, respectively. At 22 °C condition, the leaves of wsl12 mutant exhibited severe albino phenotype at three-leaf stage (Additional file 1: Fig. S3a), and a large decrease in pigment concentrations compared with that of wild-type plants (Additional file 1: Fig. S3e). When the seedlings were exposed to 26 °C condition, wsl12 mutant showed much lower albino degree and pigment concentrations increased (Additional file 1: Fig. S3b, f). When seedlings were grown at 30 °C condition, there was almost no difference between wsl12 mutant and wild-type plants (Additional file 1: Fig. S3c, g). Interestingly, we could easily detect their difference at 34 °C (Additional file 1: Fig. S3d, i). These results demonstrate that the wsl12 phenotype is strongly affected by temperature.
The single-base deletion of WSL12 is responsible for the wsl12 phenotype
To determine whether the wsl12 phenotype is controlled by a single gene, we performed genetic analysis of reciprocal crosses between wsl12 and the japonica cultivars Wuyujing7. The results showed that wsl12 was a recessive mutant, based on examination of individuals of the F2 segregating population. The ratio of individuals with normal phenotype to mutant phenotype was 3:1 (χ2 = 0.358 < χ2 0.05,1 = 3.84), indicating that the wsl12 phenotype was controlled by a single recessive nuclear gene.
We constructed the mapping population by crossing the wsl12 mutant with the indica cultivar 93-11. Bulked segregant analysis (BSA) was used to produce a primary map of WSL12. PCR genotyping was carried out using a bulk DNA pool from 33 wsl12/93-11 F2 individuals with the mutant phenotype, and 118 molecular markers scattered on all of the rice chromosomes were used to determine the approximate map position of WSL12. WSL12 was primarily located on the long arm of chromosome 12, closely linked to SSR marker RM1246 (Fig. 3a). To further fine map WSL12, we designed new markers next to RM1246 based on the sequence difference between japonica rice variety Nipponbare and indica variety 93-11 (http://www.gramene.org/resources/). The polymorphism primers were subsequently used to screen the genotypes of 2, 257 individuals with mutant phenotype and then mapped WSL12 between two markers, L-22 and L-17, within a 31-kb physical interval (Fig. 3b). There were four predicted ORFs according to the Rice Genome Annotation Project (http://rice.plantbiology.msu.edu/index.shtml; Fig. 3b). Sequence analysis revealed only a single-base deletion in the third ORF (LOC_Os12g36194, OsNDPK2) of the wsl12 mutant. Bioinformatic analysis showed that WSL12 consisted of 6,111 nucleotides, contained seven exons (Fig. 3c) and encoded a 331 amino acid protein. Compared with the wild type, the deletion was in the third exon and caused a frameshift mutation which resulted in premature translation termination (Fig. 3c, Addition file 1: Fig. S4). Genetic complementation verified the identity of the WSL12 candidate gene. We introduced the complementary plastid containing the entire OsNDPK2 ORF, and pCK vector as a negative control, into the wsl12 mutant. The phenotype of wsl12 was restored to normal in the positive transgenic plants (Fig. 3d), whereas all pCK lines failed to recover the wild-type phenotype, which demonstrated that the wsl12 phenotype can specifically rescue by introducing the OsNDPK2 genomic fragment (Fig. 3d, e). In addition, we detected the pigment content of wild-type and complementary plastid transgenic plants and found that the pigment content in the wild-type and positive transgenic plants was higher than that of pCK transgenic lines (Fig. 3f). Moreover, over-expressing WSL12 in wsl12 mutant plants showed that the mutant phenotype could be restored to normal phenotype (Addition file 1: Fig. S5). These results suggest that LOC_Os12g36194 is responsible for the wsl12 phenotype.
Map-based cloning of WSL12 and transgenic complementation of the wsl12 mutant. a WSL12 was initially mapped to a 2.5-cM interval on chromosome 12 between markers L-6 and L-1. b WSL12 locus was narrowed to a 31-kb region between markers L-22 and L-17 on BAC clone OJ1123_B09. Four open reading frames (ORFs) were predicted in the mapped region. c Structure of WSL12 (ORF3), ATG, TGA represent for start and stop codon, black boxes indicate exons, and lines between black boxes indicate introns. Black pane shows the right translation way, one single nucleotide deletion result in premature transcription termination (red pane). d Phenotypes of wild-type, wsl12, transgenic-positive plants and transgenic plants carried negative controlled vector (pCK). Scale bar 2 cm. e Expression analysis of WSL12 in young leaves of wild-type, pCK, and two complemented lines (Line1, Line2) by qRT-PCR. f Total pigment content of young leaves from wild-type, pCK, and two complemented lines. Data represent mean ± SD based on three independent biological experiments
Expression pattern of WSL12
Quantitative real-time PCR (qRT-PCR) was used to analyze the expression pattern of WSL12. We examined the expression levels of WSL12 transcripts in root, culm, leaf blade, leaf sheath and immature panicle. The expression level of WSL12 was higher in leaves (Fig. 4a). β-glucuronidase (GUS) staining showed the same results as that of qRT-PCR. GUS staining signals appeared in the entire tissues in normal field conditions (Fig. 4b, c), which indicated that WSL12 displayed as constitutive expression pattern in rice. Since the albino phenotype was more obvious in newly developed leaves at seedling and tillering stages (Fig. 1a, e), we detected the expression levels of WLS12 between new and old tissues. Our results demonstrated that WSL12 was highly expressed in young tissues (Fig. 4d). We also examined the expression levels of WSL12 in the leaves of plants grown under different temperatures. We found that the expression levels of WSL12 were higher in plants grown at 22 and 34 °C than those grown at 26 and 30 °C (Fig. 4f). These result indicated that WSL12 may be responsible for the temperature stress.
Expression pattern analysis and subcellular localization of WSL12. a QRT-PCR estimation of the relative expression level of WSL12. RNA was isolated from root (R), culm (C), leaf blade (L), leaf sheath (S) and young panicle (P) of wild-type plants. b–c WSL12 expression was revealed by β-glucuronidase (GUS) staining in WSL12 promoter-GUS transgenic plants. d QRT-PCR analysis of relative expression levels of WSL12 in leaves and roots at different development stages. R1, R2 and R3 represent roots of wild-type at 3, 7 and 30 days after germination, respectively. e Main leaves of 4-week-old wild-type plants grown at Hangzhou field conditions. f Expression levels of WSL12 in leaves of wild-type plants and the mutant under temperature treatment. Bars 1 cm in b and e and 5 mm in c. Data represent mean ± SD based on three independent biological replicates, and asterisk indicates a significant difference (Student’s t test, *P < 0.05; **P < 0.01). Rice protoplasts cell expressing empty GFP vector. g GFP florescence. h Bright-field image of GFP. i Merged image of g and h. Rice protoplasts cell expressing the WSL12-GFP fusion protein. j GFP florescence. k Auto-fluorescence of chloroplasts. l Merged image of j and k. Scale bar = 5 μm
Chloroplast localization protein WSL12 regulates the expression of the genes associated with chlorophyll synthesis and chloroplast development
To examine the subcellular location of WSL12, we constructed the WSL12-GFP fusion vector and introduced it into rice protoplast cells. The green fluorescent signals of WSL12-GFP were colocalized with the auto-fluorescent signals of chloroplasts, suggesting that WSL12 located in the chloroplasts (Fig. 4g–l). The expression levels of genes associated with chlorophyll synthesis and chloroplast development determine pigment content and chloroplast developmental status. The expression patterns of these genes were examined in wsl12 and wild-type plants under normal field conditions. They were glutamyl-tRNA reductase (HEMA1), Mg-chelatase subunit H (CHLH), 3, 8-divinyl protochlorophyllide a 8-vinyl reductase (DVR), pchlide oxidoreductase (PORA), chlorophyllide a oxygenase (CAO1), chlorophyll synthase (YGL1), small subunit of rubisco (rbcS), light-harvesting Chl a/b-binding protein of PSII, photosynthesis-related (cab1R) and cab2R. The expression levels of these genes were significantly decreased in the wsl12 plants (Additional file 1: Fig. S6). The other genes related to photosynthesis including psaA, psbA (encoding two reaction center polypeptides) and rbcL (encoding the large subunit of rubisco) were also down-regulated in the wsl12 plants. But rpoA and rpoB (two genes related to RNA polymerase) were significantly up-regulated in the wsl12 plants (Additional file 1: Fig. S6). These results demonstrate that WSL12 plays an important role in regulating the expression of genes associated with chlorophyll biosynthesis, chloroplast development and photosynthesis.
The wsl12 mutant is sensitive to ABA and salinity stresses
Under normal field conditions, we found that there was significant difference between wsl12 and wild-type plants in the germination time. The wsl12 mutant seeds germinated later than wild-type seeds. In previous studies, abnormal ABA concentrations that can affect seed germination have been reported. We detected the ABA concentration in wsl12 and wild-type plants. The ABA concentration of wsl12 mutant decreased obviously compared with wild-type plants (Additional file 1: Fig. S7a). The transcription levels of the genes related to ABA synthesis were decreased in the wsl12 mutant (Additional file 1: Fig. S7b). Therefore, we speculated that WSL12 might regulate ABA response in rice. Seed germination tests showed that the germination rate of wsl12 mutant was lower than that of wild-type plants with exogenous 2 μM ABA (Fig. 5c). High concentration of ABA (4 μM) could inhibit the germination of wild-type (50 % germination rate) and mutant seeds (20 % germination rate), while the inhibition was much stronger in wsl12 seeds (Fig. 5d). Besides, ABA-induced inhibition on shoot and root length was stronger in wsl12 mutant compared with wild-type plants when exogenous ABA concentration increased from 2 to 4 μM (Additional file 1: Fig. S7c–e). These observations demonstrate that the wsl12 mutant is sensitive to ABA-mediated inhibition at the seed germination and seedling stage (Fig. 5a–d; Additional file 1: Fig. S7). Thus, WSL12 can significantly affect ABA response and biosynthesis.
Effect of ABA and NaCl on the growth of plants. Germination rate of wild-type and wsl12 seeds in the absence (a) and presence of 1 μM (b), 2 μM (c) or 4 μM (d) of ABA. Germination rate of wild-type, wsl12 and over-expression seeds in the absence (e) and presence of 100 mM (f), 150 mM (g) or 200 mM (h) of NaCl. The germination was identified as the coleoptiles were grown at least 5 mm long. Data represent mean ± SD based on three independent biological replicates, and asterisk indicates a statistically significant difference (Student’s t test, *P < 0.05; **P < 0.01). The X axis means germination rate, and Y axis represents days
Since salinity response is closely associated with ABA in plant (Tan et al. 2014), we examined the salinity response in wsl12 mutant and WSL12 over-expression plants by adding excessive NaCl to the 1/2 MS medium. The wsl12 mutant showed delayed seed germination, but there was no significant difference between wild-type and WSL12 over-expression plants in the germination time without additional NaCl (Fig. 5e). The germination time of WSL12 over-expression plants was earlier than that of wild-type plants and wsl12 mutant with 200 mM NaCl (Fig. 5h). However, the germination rate of wsl12 mutant decreased with excessive NaCl (Fig. 5f–h). In contrast, the germination rate of WSL12 over-expression plants was higher than the wild-type at 200 mM NaCl condition (Fig. 5h). These results demonstrate that wsl12 mutant is more sensitive to NaCl.
The superoxide level is high in wsl12 mutant
ABA is an important hormone regulating plant growth and development. Previous studies showed that ABA can enhance the tolerance of plants against stress through inducing the expression of antioxidant enzymes including superoxide dismutase (SOD), CAT and ascorbate peroxidase (APX; Kaminaka et al. 1999; Anderson et al. 1994). Thus, we investigated the expression levels of these genes and oxidative status of cells in the leaves of wsl12 mutant and wild-type plants at seedling stage. Nitrotetrazolium blue chloride (NBT) staining showed that the oxidative status in wsl12 leaves was more severe than that of wild type (Additional file 1: Fig. S8a). The expression levels of APX1, APX2 and CatA were down-regulated, while the transcription levels of AOX1a and SODA1 were up-regulated in wsl12 mutant (Additional file 1: Fig. S8b). These results indicate that WSL12 participates in reactive oxygen species-scavenging pathway by regulating the expression of these genes. AtNDPK2 can directly interact with phytochrome A since it acts as a positive signal transducer in the phytochrome signaling pathway (Choi et al. 1999). Thus, we detected the expression levels of three phytochrome genes (PHYA, PHYB and PHYC) and the other two OsNDPKs (OsNDPK1 and OsNDPK3). We found that the expression levels of the three phytochrome genes and OsNDPK1 were down-regulated in wsl12 mutant (Additional file 1: Fig. S8c). However, the expression level of OsNDPK3 which involved in the signaling for stress response was significantly increased in wsl12 mutant (Escobar Galvis et al. 2001; Liu et al. 2015), which might compensate for the function of OsNDPK2.
Discussions
The diverse phenotypes in rice mutants affecting chlorophyll synthesis and chloroplast development are characterized by seedling viability and/or leaf coloration, such as virescent (v), stripe (st), albino, chlorina, zebra and yellow-variegated leaves (Jung et al. 2003). V1 encodes a chloroplast-located protein NUS1 which is essential for establishing rice plastid genetic system during early development (Kusumi et al. 2011). V2 encodes a guanylate kinase located in both plastid and mitochondria and functions in chloroplast differentiation (Sugimoto et al. 2004, 2007). V3 and ST1 encode the large and small subunits of ribonucleotide reductase, respectively. They are mainly involved in DNA synthesis and repair during early leaf development (Yoo et al. 2009). WSL encodes a pentatricopeptide repeat protein which is required for the accumulation of plastids in ribosomes and affects the splice of chloroplast rpl2 transcripts (Tan et al. 2014). The wsl, v1, v2, v3 and str1 mutants are all sensitive to temperature. The phenotypes of wsl plants are more obvious in low temperature (Tan et al. 2014). The leaves of v1 and v2 mutants become chlorotic when grown under restrictive temperatures (≤20 °C), but show nearly deep green color at the temperature ≥30 °C. On the contrary, v3 and st1 produce bleached leaves at the constant temperature of 20 or 30 °C, but generate almost green leaves under the temperature between 20 and 30 °C (Kusumi et al. 2011; Sugimoto et al. 2007; Yoo et al. 2009). The wsl12 mutant, also a temperature-sensitive mutant, showed conspicuous abnormal leaves under the constant temperature of 22 or 34 °C. However, at the temperature of 30 °C, the leaves of wsl12 mutant were almost as green as wild-type plants (Additional file 1: Fig. S3c). These results show that the phenotype of wsl12 mutant is strongly affected by temperature. We can observe white stripe leaves in newly developed leaves of wsl12 mutant (Fig. 1a, e). The seed-setting rate of wsl12 mutant was significantly decreased, which may due to weakened photosynthesis caused by the abnormal chloroplast structure (Fig. 2, Additional file 1: Fig. S1). The panicles of wsl12 mutant also showed white-green phenotype at initial heading stage, demonstrating that WSL12 also affects the chloroplast development in parenchyma cells of palea and lemma.
Map-based cloning and complementary experiment demonstrate that WSL12 encodes OsNDPK2. WSL12 expressed in various tissues and highly expressed in leaves and young tissues (Fig. 4a–d), which can explain that the albino phenotype of new leaves in the wsl12 mutant was more obvious. Contradictory results have been reported on the cellular location of NDPK2. Although many data and results showed that NDPK2 should be targeted to the chloroplast (Yang and Lamppa 1996; Bovet et al. 1999), AtNDPK2 was falsely considered to have nuclear and cytoplasmic locations in Arabidopsis (Zimmermann et al. 1999; Choi et al. 1999). Later reports thoroughly revisited the location of AtNDPK2 and demonstrated that AtNDPK2 was exclusively targeted to chloroplasts (Bölter et al. 2007; Jaedicke et al. 2011). They consider that previous unreasonable results may be caused by the use of inadequate constructs for detecting the location of AtNDPK2. In this study, the fluorescent signals of WSL12-GFP were colocalized with the auto-fluorescent signals of chloroplasts, which confirmed that OsNDPK2 targeted to chloroplasts (Fig. 4g–l). The expression levels of genes associated with chlorophyll biosynthesis and chloroplast development were down-regulated, except for rpoA and rpoB (Additional file 1: Fig. S6), which indicated that WSL12 was required for the normal transcription of these genes. The expression pattern of rpoA and rpoB was different from other chlorophyll biosynthetic genes, but the mechanism that caused the difference needs to be further researched. In addition, the mutation of wsl12 repressed the transcription of genes involved in photosynthesis, such as psaA, psbA, rbcL and rbcS, which may be the underlying mechanism of the low photosynthetic rate in wsl12.
Various environmental stresses, including extreme temperature, salt, drought, excessive light and osmotic shock, can cause high ROS levels. ROS is an important cellular regulator for stress response and oxidative cell death (Moon et al. 2003). Chloroplasts, the major cellular component for photosynthesis, are highly exposed to ROS damage since photosynthesis is a source of ROS and H2O2. Therefore, chloroplast is also involved in triggering ROS/redox-dependent signaling (Petrov and Van Breusegem 2012; Kangasjärvi et al. 2012). Many evidences have confirmed that AtNDPK2 is involved in ROS signaling and oxidative stress management (Yang et al. 2003). Over-expressing AtNDPK2 induces the expression of multiple antioxidant genes, which can enhance the resistance to oxidative stress (Yang et al. 2003; Kim et al. 2009). This protective role against oxidative stress makes NDPK2 a useful target for plant biotechnology. Over-expression AtNDPK2 in plants can enhance tolerance against multiple stresses (Kim et al. 2009; Moon et al. 2003). In our study, over-expressing OsNDPK2 could enhance rice germination rate under salt stress (Fig. 5), which may be a useful germplasm resource with high tolerance against environment stress such as salinity for rice molecular breeding.
In recent years, the area of arable land is decreased because of environment pollution. With the increasing saline-alkali soil and drought land, food production faces the risk of reduction. Some experts concerned that global warming also threatened grain output. Previous studies and this work proved that NDPK2 is involved in oxidative stress regulation and G protein regulation pathway (Shen et al. 2008). Some G protein-related genes in rice have been cloned. For example, a key enhancing yield gene DEP1 or nutrient-efficient gene DEP1-1 has been identified and participates in G protein activity. It can provide a breeding strategy to improve new varieties for environmentally sustainable increases in nitrogen use efficiency and rice grain yield (Sun et al. 2014). These findings provide an important genetic information for improving crop yield and abiotic stress tolerance via molecular breeding application of NDPK2 gene (Guo and Ye 2014; Qian et al. 2016).
The wsl12 mutant showed higher superoxide anion level compared with the wild-type plants (Additional file 1: Fig. S8a). But not all of the genes related to anti-oxidate were down-regulated (Additional file 1: Fig. S8b). We speculated that other genes, such as OsNDPK3 which was highly expressed in wsl12 mutant, might involve in the regulation of cellular redox state. We observed conspicuous phenotypic difference between wsl12 and wild-type plants at the temperature of 22 and 34 °C, but not at 30 °C (Additional file 1: Fig. S3). The expression level of WSL12 in wild-type plants under extreme temperature was higher than that under mild temperature, which is consistent with the function of WSL12 since temperature stress could cause high ROS levels. ROS could accumulate in the cells of wsl12 mutant, suggesting that these cells may partially dead due to the loss function of WSL12 (Fig. 2b).
Previous reports demonstrate that AtNDPK2 is an upstream component of the phytochrome-mediated light signaling pathway in Arabidopsis thaliana (Choi et al. 1999; Shen et al. 2005). In wsl12 mutant, three phytochrome genes (PHYA, PHYB and PHYC) were all down-regulated (Additional file 1: Fig. S8c). Previous studies show that phytochrome B regulates chlorophyll synthesis and chloroplast development and also affects ABA metabolism (Zhao et al. 2012; Gu et al. 2012). Thus, further studies were necessary to confirm whether the expression change in the genes related to chlorophyll synthesis was directly caused by the loss function of WSL12. Actually, the ABA concentration of wsl12 mutant decreased obviously compared with wild-type plants (Additional file 1: Fig. S7a), but the mutant plants were more sensitive to ABA-induced inhibition (Fig. 5a–d; Additional file 1: Fig. S7c-e). This may because exogenous ABA can induce hydrogen peroxide (H2O2) and superoxide radical (O2 −) production (Pei et al. 2000; Jiang and Zhang 2001), but the wsl12 mutant had lower ROS tolerance because of the loss function of WSL12. This demonstrated that the extent of high ROS level repressing plants growth was more serious in wsl12 than in wild-type plants. In addition, auxin and ROS are interconnected in a complex signaling network regulating plant growth and development (Tognetti et al. 2012). It is possible that the phenotype of wsl12 mutant is affected by auxin signaling and ROS pathways. In summary, all these discoveries suggest that WSL12 is a multifunctional protein involved in chlorophyll synthesis, cell signaling and stress responses. Further studies of WSL12 should focus on elucidating the complex network among ROS, phytochrome-mediated and auxin-mediated signal transduction pathways.
References
Anderson MD, Prasad TK, Martin BA, Stewart CR (1994) Differential gene expression in chilling-acclimated maize seedlings and evidence for the involvement of abscisic acid in chilling tolerance. Plant Physiol 105(1):331–339
Arnon DI (1949) Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol 24(1):1–15
Bernard MA, Ray NB, Olcott MC, Hendricks SP, Mathews CK (2000) Metabolic functions of microbial nucleoside diphosphate kinases. J Bioenerg Biomembr 32(3):259–267
Bölter B, Sharma R, Soll J (2007) Localisation of Arabidopsis NDPK2—revisited. Planta 226(4):1059–1065
Bovet L, Meylan-Bettex M, Eggman T, Martinoia E, Siegenthaler PA (1999) CDP phosphotransferase activity in spinach intact chloroplasts: possible involvement of nucleoside diphosphate kinase II. Plant Physiol Biochem 37(9):645–652
Chen S, Tao L, Zeng L, Vega-Sanchez ME, Umemura K, Wang GL (2006) A highly efficient transient protoplast system for analyzing defence gene expression and protein–protein interactions in rice. Mol Plant Pathol 7(5):417–427
Chen M, Galvao RM, Li MN, Burger B, Bugea J, Bolado J, Chory J (2010) Arabidopsis HEMERA/pTAC12 initiates photomorphogenesis by phytochromes. Cell 141(7):1230–1240
Choi G, Yi H, Lee J, Kwon YK, Soh MS, Shin BC, Luka Z, Hahn TR, Song PS (1999) Phytochrome signalling is mediated through nucleoside diphosphate kinase 2. Nature 401(6753):610–613
Choi G, Kim JI, Hong SW, Shin B, Choi G, Blakeslee JJ, Murphy AS, Seo YW, Kim K, Koh EJ, Song PS, Lee H (2005) A possible role for NDPK2 in the regulation of auxin-mediated responses for plant growth and development. Plant Cell Physiol 46(8):1246–1254
Delannoy E, Le Ret M, Faivre-Nitschke E, Estavillo GM, Bergdoll M, Taylor NL, Pogson BJ, Small I, Imbault P, Gualberto JM (2009) Arabidopsis tRNA adenosine deaminase arginine edits the wobble nucleotide of chloroplast tRNAArg (ACG) and is essential for efficient chloroplast translation. Plant Cell 21(7):2058–2071
Dorion S, Rivoal J (2015) Clues to the functions of plant NDPK isoforms. Naunyn-Schmiedeberg’s Arch Pharmacol 388(2):119–132
Escobar Galvis ML, Marttila S, Håkansson G, Forsberg J, Knorpp C (2001) Heat stress response in pea involves interaction of mitochondrial nucleoside diphosphate kinase with a novel 86-kilodalton protein. Plant Physiol 126(1):69–77
Gothandam KM, Kim ES, Cho HJ, Chung YY (2005) OsPPR1, a pentatricopeptide repeat protein of rice is essential for the chloroplast biogenesis. Plant Mol Biol 58(3):421–433
Gu JW, Zhang F, Zhao J, Zhou JZ, Qian FQ, Yan LH, Zang X, Xian XZ (2012) Light signals mediated by phytochrome B affect abscisic acid pathway in rice (in Chinese). Chin Sci Bull (Chin Ver) 57(25):2371–2379
Guo LB, Ye GY (2014) Use of major quantitative trait loci to improve grain yield of rice. Rice Sci 21(2):65–82
Guo M, Li RD, Yao J, Zhu J, Fan XY, Wang W, Tang SZ, Gu MH, Yan CJ (2015) RL3(t), responsible for leaf shape formation, delimited to a 46-kb DNA fragment rice. Rice Sci 22(1):44–48
Hammargren J, Sundström J, Johansson M, Bergman P, Knorpp C (2007) On the phylogeny, expression and targeting of plant nucleoside diphosphate kinases. Physiol Plant 129(1):79–89
Harris N, Taylor JE, Roberts JA (1994) Isolation of a mRNA encoding a nucleoside diphosphate kinase from tomato that is up-regulated by wounding. Plant Mol Biol 25(4):739–742
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6(2):271–282
Jaedicke K, Rösler J, Gans T, Hughes J (2011) Bellis perennis: a useful tool for protein localization studies. Planta 234(4):759–768
Jiang M, Zhang J (2001) Effect of abscisic acid on active oxygen species, antioxidative defence system and oxidative damage in leaves of maize seedlings. Plant Cell Physiol 42(11):1265–1273
Jung KH, Hur J, Ryu CH, Choi Y, Chung YY, Miyao A, Hirochika H, An G (2003) Characterization of a rice chlorophyll-deficient mutant using the T-DNA gene-trap system. Plant Cell Physiol 44(5):463–472
Kaminaka H, Morita S, Tokumoto M, Masumura T, Tannaka K (1999) Differential gene expressions of rice superoxide dismutase isoforms to oxidative and environmental stress. Free Radic Res 31(Suppl):219–225
Kangasjärvi S, Neukermans J, Li SC, Aro EM, Noctor G (2012) Photosynthesis, photorespiration, and light signalling in defence responses. J Exp Bot 63(4):1619–1636
Kantor JD, McCormick B, Steeg PS, Zetter BR (1993) Inhibition of cell motility after nm23 transfection of human and murine tumor cells. Cancer Res 53(9):1971–1973
Keim D, Hailat N, Melhem R, Zhu XX, Lascu I, Veron M, Strahler J, Hanash SM (1992) Proliferation-related expression of p19/nm23 nucleoside diphosphate kinase. J Clin Invest 89(3):919–924
Kim YH, Lim S, Yang KS, Kim CY, Kwon SY, Lee HS, Wang X, Zhou Z, Ma DF, Yun DJ, Kwak SS (2009) Expression of Arabidopsis NDPK 2 increases antioxidant enzyme activities and enhances tolerance to multiple environmental stresses in transgenic sweetpotato plants. Mol Breed 24(3):233–244
Kusumi K, Sakata C, Nakamura T, Kawasaki S, Yoshimura A, Iba K (2011) A plastid protein NUS1 is essential for build-up of the genetic system for early chloroplast development under cold stress conditions. Plant J 68(6):1039–1050
Lascu I, Gonin P (2000) The catalytic mechanism of nucleoside diphosphate kinases. J Bioenerg Biomembr 32(3):237–246
Liu H, Weisman D, Tang L, Tan L, Zhang WK, Wang ZH, Huang YH, Lin WX, Liu XM, Colón-Carmona A (2015) Stress signaling in response to polycyclic aromatic hydrocarbon exposure in Arabidopsis thaliana involves a nucleoside diphosphate kinase, NDPK-3. Planta 241(1):95–107
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25(4):402–408
Moon H, Lee B, Choi G, Shin D, Prasad DT, Lee O, Kwak SS, Kim DH, Nam J, Bahk J, Hong JC, Lee SY, Cho MJ, Lim CO, Yun DJ (2003) NDP kinase 2 interacts with two oxidative stress-activated MAPKs to regulate cellular redox state and enhances multiple stress tolerance in transgenic plants. Proc Natl Acad Sci USA 100(1):358–363
Murchie EH, Hubbart S, Peng S, Horton P (2005) Acclimation of photosynthesis to high irradiance in rice: gene expression and interactions with leaf development. J Exp Bot 56(411):449–460
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8(19):4321–4325
Parks RE Jr, Agarwal RP (1973) Nucleoside diphosphokinases. Enzymes 8(Part A):307–334
Pei ZM, Murata Y, Benning G, Thomine S, Klüsener B, Allen GJ, Grill E, Schroeder JI (2000) Calcium channels activated by hydrogen peroxide mediate abscisic acid signaling in guard cells. Nature 406(6797):731–734
Petrov VD, Van Breusegem F (2012) Hydrogen peroxide—a central hub for information flow in plant cells. Aob Plants, pls014
Qian Q, Guo LB, Smith SM, Li JY (2016) Breeding high-yield superior-quality hybrid super-rice by rational design. Natl Sci Rev. doi:10.1093/nsr/nww006
Roberts JKM, Aubert S, Gout E, Bligny R, Douce R (1997) Cooperation and competition between adenylate kinase, nucleoside diphosphokinase, electron transport, and ATP synthase in plant mitochondria studied by 31P-Nuclear Magnetic Resonance. Plant Physiol 113(1):191–199
Scarpella E, Rueb S, Meijer AH (2003) The RADICLELESS1 gene is required for vascular pattern formation in rice. Development 130(4):645–658
Shen Y, Kim JI, Song PS (2005) NDPK2 as a signal transducer in the phytochrome-mediated light signaling. J Biol Chem 280(7):5740–5749
Shen Y, Han YJ, Kim JI, Song PS (2008) Arabidopsis nucleoside diphosphate kinase-2 as a plant GTPase activating protein. BMB REP 41(9):645–650
Sugimoto H, Kusumi K, Tozawa Y, Yazaki J, Kishimoto N, Kikuchi S, Iba K (2004) The virescent-2 mutation inhibits translation of plastid transcripts for the plastid genetic system at an early stage of chloroplast differentiation. Plant Cell Physiol 45(8):985–996
Sugimoto H, Kusumi K, Noguchi K, Yano M, Yoshimura A, Iba K (2007) The rice nuclear gene, VIRESCENT 2, is essential for chloroplast development and encodes a novel type of guanylate kinase targeted to plastids and mitochondria. Plant J 52(3):512–527
Sun HY, Qian Q, Wu K, Luo JJ, Wang SS, Zhang CW, Ma YF, Liu Q, Huang XZ, Yuan QB, Han RX, Zhao M, Dong GJ, Guo LB, Zhu XD, Gou ZH, Wang W, Wu YJ, Lin HX, Fu XD (2014) Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat Genet 46(6):652–656
Tan JJ, Tan ZH, Wu FQ, Sheng PK, Heng YQ, Wang XH, Ren YL, Wang JL, Guo XP, Zhang X, Cheng ZJ, Jiang L, Liu XM, Wang HY, Wan JM (2014) A novel chloroplast-localized pentatricopeptide repeat protein involved in splicing affects chloroplast development and abiotic stress response in rice. Mol Plant 7(8):1329–1349
Thordal-Christensen H, Zhang Z, Wei Y, Collinge DB (1997) Subcellular localization of H2O2 in plants: H2O2 accumulation in papillae and hypersensitive response during the barley-powdery mildew interaction. Plant J 11(6):1187–1194
Tognetti VB, Mühlenbock P, Van Breusegem F (2012) Stress homeostasis-the redox and auxin perspective. Plant, Cell Environ 35(2):321–333
Verslues PE, Batelli G, Grillo S, Agius F, Kim YS, Zhu J, Agarwal M, Katiyar-Agarwal S, Zhu JK (2007) Interaction of SOS2 with NDPK2 and catalases reveals a point of connection between salt stress and H2O2 signaling in Arabidopsis thaliana. Mol Cell Biol 27(22):7771–7780
Wellburn AR (1994) The spectral determination of chlorophylls a and b, as well as total carotenoids, using various solvents with spectrophotometers of different resolution. J Plant Physiol 144(3):307–313
Yang LM, Lamppa GK (1996) Rapid purification of a chloroplast nucleoside diphosphate kinase using CoA-affinity chromatography. Biochim Biophys Acta 1294(3):99–102
Yang KA, Moon H, Kim G, Lim CJ, Hong JC, Lim CO, Yun DJ (2003) NDP kinase 2 regulates expression of antioxidant genes in Arabidopsis. P JPN ACAD B-PHYs Biol Sci 79(3):86–91
Yoo SC, Cho SH, Sugimoto H, Li J, Kusumi K, Koh HJ, Iba K, Paek NC (2009) Rice virescent3 and stripe1 encoding the large and small subunits of ribonucleotide reductase are required for chloroplast biogenesis during early leaf development. Plant Physiol 150(1):388–401
Zhang H, Li J, Yoo JH, Yoo SC, Cho SH, Koh HJ, Seo HS, Paek NC (2006) Rice Chlorina-1 and Chlorina-9 encode ChlD and ChlI subunits of Mg-chelatase, a key enzyme for chlorophyll synthesis and chloroplast development. Plant Mol Biol 62(3):325–337
Zhao J, Zhou JJ, Gu JW, Qian FQ, Xie XZ (2012) Phytochrome B positively regulates chlorophyll biosynthesis and chloroplast development in rice. Chin J Rice Sci 26(6):637–642
Zhen XH, Xu JG, Shen WJ, Zhang XJ, Zhang QJ, Lu YG, Chen GX, Gao ZP (2014) Photosynthetic characteristics of flag leaves in rice white stripe mutant 6001 during senescence process. Rice Sci 21(6):335–342
Zimmermann S, Baumann A, Jaekel K, Marbach I, Engelberg D, Frohnmeyer H (1999) UV-responsive genes of Arabidopsis revealed by similarity to the Gcn4-mediated UV response in yeast. J Biol Chem 274(24):17017–17024
Acknowledgments
This research was supported by National Natural Science Foundation of China (31501279, 31271700), China Postdoctoral Science Foundation (2015M570181), National Key technology Research and Development Program (2015BAD01B02) and Shenzhen Scientific and Technological Program (JCYJ20140504111101999, JCYJ20150630165133402). The authors are grateful to the editors and the anonymous reviewers for their valuable comments.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Weijun Ye and Shikai Hu have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Ye, W., Hu, S., Wu, L. et al. White stripe leaf 12 (WSL12), encoding a nucleoside diphosphate kinase 2 (OsNDPK2), regulates chloroplast development and abiotic stress response in rice (Oryza sativa L.). Mol Breeding 36, 57 (2016). https://doi.org/10.1007/s11032-016-0479-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-016-0479-6