Abstract
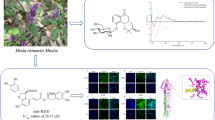
Stimulator of interferon gene (STING) plays critical roles in the cytoplasmic DNA-sensing pathway and in the induction of inflammatory response. Aberrant cytoplasmic DNA accumulation and STING activation are implicated in numerous inflammatory and autoimmune diseases. Here, we reported the discovery of a series of thiazolecarboxamide-based STING inhibitors through a molecular planarity/symmetry disruption strategy. The privileged compound 15b significantly inhibited STING signaling and suppressed immune-inflammatory cytokine levels in both human and murine cells. In vivo experiments demonstrated 15b effectively ameliorated immune-inflammatory cytokines upregulation in MSA-2-stimulated and Trex1-D18N mice. Furthermore, compound 15b exhibited enhanced efficacy in suppressing interferon-stimulated gene 15 (ISG15), a critical positive feedback regulator of STING. Overall, compound 15b deserves further development for the treatment of STING-associated inflammatory and autoimmune diseases.
Graphical abstract
















Similar content being viewed by others
Abbreviations
- STING:
-
Stimulator of interferon gene
- IFN:
-
Interferon
- cGAS:
-
Cyclic GMP-AMP synthase
- cGAMP:
-
Cyclic GMP-AMP
- CDN:
-
Cyclic dinucleotide
- ER:
-
Endoplasmic reticulum
- TBK1:
-
TANK-binding kinase 1
- IRF3:
-
Interferon regulatory factor 3
- IL:
-
Interleukin
- ISG:
-
Interferon-stimulated gene
- CXCL10:
-
C-X-C motif chemokine ligand 10
- SAVI:
-
STING-associated vasculopathy with onset in infancy
- PTM:
-
Post-translational modifications
- AGS:
-
Aicardi-Goutières syndrome
- SLE:
-
Systemic lupus erythematosus
- AKI:
-
Acute kidney injury
- PROTAC:
-
Proteolysis-targeting chimera
- IC50 :
-
Half-maximum inhibitory concentration
- THP1:
-
Tohoku hospital pediatrics 1
- CTD:
-
C-terminal domain
- BMDM:
-
Mouse bone marrow-derived macrophage
- MEF:
-
Mouse embryonic fibroblast
- PCR:
-
Polymerase chain reaction
- TNF:
-
Tumor necrosis factor
- CCK-8:
-
Cell counting kit-8
- ELISA:
-
Enzyme-linked immunosorbent assay
- CC50 :
-
Concentration of 50% cytotoxicity
- AUC:
-
Area under the curve
- Cmax :
-
Maximum concentration
- MRT:
-
Mean residence time
- equiv:
-
Equivalent
- DMF:
-
N,N-Dimethylformamide
- DCM:
-
Dichloromethane
- HATU:
-
2-(7-Azabenzotriazol-1-yl)-N,N,N′,N′-tetramethyluronium hexafluorophosphate
- DIPEA:
-
N,N-Diisopropylethylamine
- TBSCl:
-
tert-Butyldimethylsilyl chloride
- MsCl:
-
methanesulfonyl chloride
- GAPDH:
-
glyceraldehyde 3-phosphate dehydrogenase
- SDS-PAGE:
-
sodium dodecyl sulfate – polyacrylamide gel electrophoresis
- NC:
-
nitrocellulose
- BSA:
-
bovine serum albumin
- TBST:
-
tris-buffered saline + tween 20
References
Ishikawa H, Barber GN (2008) STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455:674–678. https://doi.org/10.1038/nature07317
Soulat D, Bürckstümmer T, Westermayer S, Goncalves A, Bauch A, Stefanovic A, Hantschel O, Bennett KL, Decker T, Superti-Furga G (2008) The DEAD-box helicase DDX3X is a critical component of the TANK-binding kinase 1-dependent innate immune response. EMBO J 27:2135–2146. https://doi.org/10.1038/emboj.2008.126
Zhong B, Yang Y, Li S, Wang YY, Li Y, Diao F, Lei C, He X, Zhang L, Tien P, Shu HB (2008) The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29:538–550. https://doi.org/10.1016/j.immuni.2008.09.003
Ablasser A, Hornung V (2013) DNA sensing unchained. Cell Res 23:585–587. https://doi.org/10.1038/cr.2013.28
Lioux T, Mauny MA, Lamoureux A, Bascoul N, Hays M, Vernejoul F, Baudru AS, Boularan C, Lopes-Vicente J, Qushair G, Tiraby G (2016) Design, synthesis, and biological evaluation of novel cyclic adenosine-inosine monophosphate (cAIMP) analogs that activate stimulator of interferon genes (STING). J Med Chem 59:10253–10267. https://doi.org/10.1021/acs.jmedchem.6b01300
Sun L, Wu J, Du F, Chen X, Chen ZJ (2013) Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339:786–791. https://doi.org/10.1126/science.1232458
Burdette DL, Monroe KM, Sotelo-Troha K, Iwig JS, Eckert B, Hyodo M, Hayakawa Y, Vance RE (2011) STING is a direct innate immune sensor of cyclic di-GMP. Nature 478:515–518. https://doi.org/10.1038/nature10429
Danilchanka O, Mekalanos JJ (2013) Cyclic dinucleotides and the innate immune response. Cell 154:962–970. https://doi.org/10.1016/j.cell.2013.08.014
Sun W, Li Y, Chen L, Chen H, You F, Zhou X, Zhou Y, Zhai Z, Chen D, Jiang Z (2009) ERIS, an endoplasmic reticulum IFN stimulator, activates innate immune signaling through dimerization. Proc Natl Acad Sci U S A 106:8653–8658. https://doi.org/10.1073/pnas.0900850106
Abe T, Barber GN (2014) Cytosolic-DNA-mediated, STING-dependent proinflammatory gene induction necessitates canonical NF-κB activation through TBK1. J Virol 88:5328–5341. https://doi.org/10.1128/jvi.00037-14
Yum S, Li MH, Fang Y, Chen ZJ (2021) TBK1 recruitment to STING activates both IRF3 and NF-κB that mediate immune defense against tumors and viral infections. Proc Natl Acad Sci U S A 118:e2100225118. https://doi.org/10.1073/pnas.2100225118
Lin C, Kuffour EO, Fuchs NV, Gertzen CGW, Kaiser J, Hirschenberger M, Tang X, Xu HC, Michel O, Tao R, Haase A, Martin U, Kurz T, Drexler I, Görg B, Lang PA, Luedde T, Sparrer KMJ, Gohlke H, König R, Münk C (2023) Regulation of STING activity in DNA sensing by ISG15 modification. Cell Rep 42:113277. https://doi.org/10.1016/j.celrep.2023.113277
Decout A, Katz JD, Venkatraman S, Ablasser A (2021) The cGAS-STING pathway as a therapeutic target in inflammatory diseases. Nat Rev Immunol 21:548–569. https://doi.org/10.1038/s41577-021-00524-z
MacLauchlan S, Kushwaha P, Tai A, Chen S, Manning C, Swarnkar G, Abu-Amer Y, Fitzgerald KA, Sharma S, Gravallese EM (2023) STING-dependent interferon signatures restrict osteoclast differentiation and bone loss in mice. Proc Natl Acad Sci U S A 120:e2210409120. https://doi.org/10.1073/pnas.2210409120
Frémond ML, Hadchouel A, Berteloot L, Melki I, Bresson V, Barnabei L, Jeremiah N, Belot A, Bondet V, Brocq O, Chan D, Dagher R, Dubus JC, Duffy D, Feuillet-Soummer S, Fusaro M, Gattorno M, Insalaco A, Jeziorski E, Kitabayashi N, Lopez-Corbeto M, Mazingue F, Morren MA, Rice GI, Rivière JG, Seabra L, Sirvente J, Soler-Palacin P, Stremler-Le Bel N, Thouvenin G, Thumerelle C, Van Aerde E, Volpi S, Willcocks S, Wouters C, Breton S, Molina T, Bader-Meunier B, Moshous D, Fischer A, Blanche S, Rieux-Laucat F, Crow YJ, Neven B (2021) Overview of STING-associated vasculopathy with onset in infancy (SAVI) among 21 patients. J Allergy Clin Immunol Pract 9:803–818.e811. https://doi.org/10.1016/j.jaip.2020.11.007
Liu Y, Jesus AA, Marrero B, Yang D, Ramsey SE, Sanchez GAM, Tenbrock K, Wittkowski H, Jones OY, Kuehn HS, Lee CR, DiMattia MA, Cowen EW, Gonzalez B, Palmer I, DiGiovanna JJ, Biancotto A, Kim H, Tsai WL, Trier AM, Huang Y, Stone DL, Hill S, Kim HJ, St Hilaire C, Gurprasad S, Plass N, Chapelle D, Horkayne-Szakaly I, Foell D, Barysenka A, Candotti F, Holland SM, Hughes JD, Mehmet H, Issekutz AC, Raffeld M, McElwee J, Fontana JR, Minniti CP, Moir S, Kastner DL, Gadina M, Steven AC, Wingfield PT, Brooks SR, Rosenzweig SD, Fleisher TA, Deng Z, Boehm M, Paller AS, Goldbach-Mansky R (2014) Activated STING in a vascular and pulmonary syndrome. N Engl J Med 371:507–518. https://doi.org/10.1056/NEJMoa1312625
Wardlaw CP, Petrini JHJ (2022) ISG15 conjugation to proteins on nascent DNA mitigates DNA replication stress. Nat Commun 13:5971. https://doi.org/10.1038/s41467-022-33535-y
Li S, Wang Y, Wang Y (2022) Advances in genetic mechanism and clinical research of ADAR1-related neurological diseases. Fudan Univ J Med Sci 49:265–269. https://doi.org/10.3969/j.issn.1672-8467.2022.02.015
Ramantani G, Kohlhase J, Hertzberg C, Innes AM, Engel K, Hunger S, Borozdin W, Mah JK, Ungerath K, Walkenhorst H, Richardt HH, Buckard J, Bevot A, Siegel C, von Stülpnagel C, Ikonomidou C, Thomas K, Proud V, Niemann F, Wieczorek D, Häusler M, Niggemann P, Baltaci V, Conrad K, Lebon P, Lee-Kirsch MA (2010) Expanding the phenotypic spectrum of lupus erythematosus in Aicardi-Goutières syndrome. Arthritis Rheum 62:1469–1477. https://doi.org/10.1002/art.27367
Gao X, Yang X, Chen H (2021) Aicardi-Goutières syndrome caused by TREX1 gene variation: a case report. J Clin Pediatr 39:542–545. https://doi.org/10.3969/j.issn.1000-3606.2021.07.015
Rice GI, Rodero MP, Crow YJ (2015) Human disease phenotypes associated with mutations in TREX1. J Clin Immunol 35:235–243. https://doi.org/10.1007/s10875-015-0147-3
Lee-Kirsch MA, Gong M, Chowdhury D, Senenko L, Engel K, Lee YA, de Silva U, Bailey SL, Witte T, Vyse TJ, Kere J, Pfeiffer C, Harvey S, Wong A, Koskenmies S, Hummel O, Rohde K, Schmidt RE, Dominiczak AF, Gahr M, Hollis T, Perrino FW, Lieberman J, Hübner N (2007) Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 are associated with systemic lupus erythematosus. Nat Genet 39:1065–1067. https://doi.org/10.1038/ng2091
Maekawa H, Inoue T, Ouchi H, Jao TM, Inoue R, Nishi H, Fujii R, Ishidate F, Tanaka T, Tanaka Y, Hirokawa N, Nangaku M, Inagi R (2019) Mitochondrial damage causes inflammation via cGAS-STING signaling in acute kidney injury. Cell Rep 29:1261–1273.e1266. https://doi.org/10.1016/j.celrep.2019.09.050
Shi L, Zha H, Pan Z, Wang J, Xia Y, Li H, Huang H, Yue R, Song Z, Zhu J (2023) DUSP1 protects against ischemic acute kidney injury through stabilizing mtDNA via interaction with JNK. Cell Death Dis 14:724. https://doi.org/10.1038/s41419-023-06247-4
Gong W, Lu L, Zhou Y, Liu J, Ma H, Fu L, Huang S, Zhang Y, Zhang A, Jia Z (2021) The novel STING antagonist H151 ameliorates cisplatin-induced acute kidney injury and mitochondrial dysfunction. Am J Physiol-renal 320:F608–F616. https://doi.org/10.1152/ajprenal.00554.2020
Chung KW, Dhillon P, Huang S, Sheng X, Shrestha R, Qiu C, Kaufman BA, Park J, Pei L, Baur J, Palmer M, Susztak K (2019) Mitochondrial damage and activation of the STING pathway lead to renal inflammation and fibrosis. Cell Metab 30:784–799.e785. https://doi.org/10.1016/j.cmet.2019.08.003
Liu J, Yuan L, Ruan Y, Deng B, Yang Z, Ren Y, Li L, Liu T, Zhao H, Mai R, Chen J (2022) Novel CRBN-recruiting proteolysis-targeting chimeras as degraders of stimulator of interferon genes with in vivo anti-inflammatory efficacy. J Med Chem 65:6593–6611. https://doi.org/10.1021/acs.jmedchem.1c01948
Siu T, Altman MD, Baltus GA, Childers M, Ellis JM, Gunaydin H, Hatch H, Ho T, Jewell J, Lacey BM, Lesburg CA, Pan BS, Sauvagnat B, Schroeder GK, Xu S (2019) Discovery of a novel cGAMP competitive ligand of the inactive form of STING. ACS Med Chem Lett 10:92–97. https://doi.org/10.1021/acsmedchemlett.8b00466
Hong Z, Mei J, Li C, Bai G, Maimaiti M, Hu H, Yu W, Sun L, Zhang L, Cheng D, Liao Y, Li S, You Y, Sun H, Huang J, Liu X, Lieberman J, Wang C (2021) STING inhibitors target the cyclic dinucleotide binding pocket. Proc Natl Acad Sci U S A 118:e2105465118. https://doi.org/10.1073/pnas.2105465118
Li S, Hong Z, Wang Z, Li F, Mei J, Huang L, Lou X, Zhao S, Song L, Chen W, Wang Q, Liu H, Cai Y, Yu H, Xu H, Zeng G, Wang Q, Zhu J, Liu X, Tan N, Wang C (2018) The cyclopeptide Astin C specifically inhibits the innate immune CDN sensor STING. Cell Rep 25:3405–3421.e3407. https://doi.org/10.1016/j.celrep.2018.11.097
Ong WWS, Dayal N, Chaudhuri R, Lamptey J, Sintim HO (2023) STING antagonists, synthesized via Povarov-Doebner type multicomponent reaction. Rsc Medicinal Chemistry 14:1101–1113. https://doi.org/10.1039/d3md00061c
Chang J, Hou S, Yan X, Li W, Xiao J (2023) Discovery of novel STING inhibitors based on the structure of the mouse STING agonist DMXAA. Molecules 28:2096. https://doi.org/10.3390/molecules28072906
Haag SM, Gulen MF, Reymond L, Gibelin A, Abrami L, Decout A, Heymann M, van der Goot FG, Turcatti G, Behrendt R, Ablasser A (2018) Targeting STING with covalent small-molecule inhibitors. Nature 559:269–273. https://doi.org/10.1038/s41586-018-0287-8
Humphries F, Shmuel-Galia L, Jiang Z, Zhou JY, Barasa L, Mondal S, Wilson R, Sultana N, Shaffer SA, Ng SL, Pesiridis GS, Thompson PR, Fitzgerald KA (2023) Targeting STING oligomerization with small-molecule inhibitors. Proc Natl Acad Sci U S A 120:e2305420120. https://doi.org/10.1073/pnas.2305420120
Barasa L, Chaudhuri S, Zhou JY, Jiang ZZ, Choudhary S, Green RM, Wiggin E, Cameron M, Humphries F, Fitzgerald KA, Thompson PR (2023) Development of LB244, an irreversible STING antagonist. J Am Chem Soc 145:20273–20288. https://doi.org/10.1021/jacs.3c03637
Zhu Z, Johnson RL, Zhang Z, Herring LE, Jiang G, Damania B, James LI, Liu P (2023) Development of VHL-recruiting STING PROTACs that suppress innate immunity. Cell Mol Life Sci 80:149. https://doi.org/10.1007/s00018-023-04796-7
Luo Q, Wang Y, Hou Z, Liang H, Tu L, Xing Y, Wan C, Liu J, Wang R, Zhu L, Han W, Wu J, Lu F, Yin F, Li Z (2024) Covalent PROTAC design method based on a sulfonyl pyridone probe. Chem Commun (Cambridge, U K) 60:686–689. https://doi.org/10.1039/d3cc05127g
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Ishikawa M, Hashimoto Y (2011) Improvement in aqueous solubility in small molecule drug discovery programs by disruption of molecular planarity and symmetry. J Med Chem 54:1539–1554. https://doi.org/10.1021/jm101356p
Lohning AE, Levonis SM, Williams-Noonan B, Schweiker SS (2017) A practical guide to molecular docking and homology modelling for medicinal chemists. Curr Top Med Chem 17:2023–2040. https://doi.org/10.2174/1568026617666170130110827
Marchese Robinson RL, Geatches D, Morris C, Mackenzie R, Maloney AGP, Roberts KJ, Moldovan A, Chow E, Pencheva K, Vatvani DRM (2019) Evaluation of force-field calculations of lattice energies on a large public fataset, assessment of pharmaceutical relevance, and comparison to density functional theory. J Chem Inf Model 59:4778–4792. https://doi.org/10.1021/acs.jcim.9b00601
Acknowledgements
This work was supported by China Postdoctoral Science Foundation (Certificate Number: 2023T160663), Institutes for Drug Discovery and Development, Chinese Academy of Sciences (No. SIMM0320231002), the Natural Science Foundation of China for Innovation Research Group (81821005), the Shanghai Municipal Science and Technology Major Project, the Collaborative Innovation Cluster Project of Shanghai Municipal Commission of Health and Family Planning (2020CXJQ02), and the Shandong Laboratory Program (SYS202205).
Author information
Authors and Affiliations
Contributions
ZJ and YZ contributed equally to this article. ZJ and YZ: Investigation, Methodology, Writing – original draft. XL: Investigation, Methodology. MG and WD: Conceptualization, Supervision, Writing – review & editing. ZX and HZ: Conceptualization, Investigation, Supervision, Writing – review & editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jin, Z., Zhang, Y., Luo, X. et al. Design, synthesis, and evaluation of thiazolecarboxamide derivatives as stimulator of interferon gene inhibitors. Mol Divers (2024). https://doi.org/10.1007/s11030-024-10860-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-024-10860-6