Abstract
Burkholderia cepacia complex (BCC) is a group of gram-negative bacteria composed of at least 20 different species that cause diseases in plants, animals as well as humans (cystic fibrosis and airway infection). Here, we analyzed the proteomic data of 47 BCC strains by classifying them in three groups. Phylogenetic analyses were performed followed by individual core region identification for each group. Comparative analysis of the three individual core protein fractions resulted in 1766 ortholog/proteins. Non-human homologous proteins from the core region gave 1680 proteins. Essential protein analyses reduced the target list to 37 proteins, which were further compared to a closely related out-group, Burkholderia gladioli ATCC 10,248 strain, resulting in 21 proteins. 3D structure modeling, validation, and druggability step gave six targets that were subjected to further target prioritization parameters which ultimately resulted in two BCC targets. A library of 12,000 ZINC drug-like compounds was screened, where only the top hits were selected for docking orientations. These included ZINC01405842 (against Chorismate synthase aroC) and ZINC06055530 (against Bifunctional N-acetylglucosamine-1-phosphate uridyltransferase/Glucosamine-1-phosphate acetyltransferase glmU). Finally, dynamics simulation (200 ns) was performed for each ligand–receptor complex, followed by ADMET profiling. Of these targets, details of their applicability as drug targets have not yet been elucidated experimentally, hence making our predictions novel and it is suggested that further wet-lab experimentations should be conducted to test the identified BCC targets and ZINC scaffolds to inhibit them.
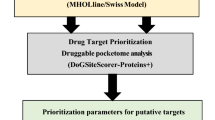
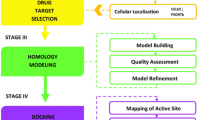
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Burkholderia cepacia complex (BCC) represents a group of aerobic gram-negative bacillus of Betaproteobacteria, posing a serious threat not only to the humans, but also to the plant and animal population [1]. In humans, BCC is regarded as challenging opportunistic pathogens especially in immunocompromised individuals and in patients with genetic disorders like cystic fibrosis (CF). A group of 20 + closely related bacterial species form BCC group with up to 78% identical genomes whose size ranges from 7 to 9 MB. The genes are typically arranged in three chromosomes and multiple plasmids, which enables more flexibility in gene gain/loss to support its disease virulence and biology [2]. BCC species form biofilms in vitro and in vivo in the lungs of patients with CF, protecting them from antibiotic drugs, and sustaining continual infection. Compared to planktonic samples, the minimum inhibitory concentrations (MIC) of most β-lactam agents are significantly higher toward BBC due to the protective effect of their biofilm. Other factors include (i) the putative expression of virulent genes regulated by quorum sensing, (ii) the production of exopolysaccharide associated to mucous phenotypes that promote the escape of host response, and (iii) the production of lipopolysaccharide that contributes to tissue damage [3]. Infection diffusion from person to person has been reported; thereby, many hospitals, clinics, and campuses have adopted severe isolation measures to counter the onsets caused by BCC. Infected individuals are often treated in area separated from non-infected patients to limit disease spread, as BCC infection can lead to a rapid decline in pulmonary function and cause mortality. The pathogenicity of BBC in patients with chronic granulomatous disease appears to be dependent on their ability to resist the killing of non-oxidative neutrophils and the neutrophil necrosis induction [2,3,4]. BCC also causes prolonged respiratory infection among individuals with CF, and may cause pneumonia, septicemia, and soft tissue eruptions among individuals with chronic granulomatous disease [5]. Resistance to β-lactam agents is most often promoted by inducible chromosomal β- lactamases or efflux pumps. The capacity of novel β-lactamase inhibitors to reestablish the in vitro action of ceftazidime suggests a cumulative effect of β-lactamase and efflux pumps in clinical BCC samples [6, 7]. Less commonly, resistance is due to plasmid-mediated β-lactamases of the TEM class (cephalosporinases) or modifications in the penicillin-binding proteins. Intrinsic resistance of BBC strains to aminoglycosides and polymyxin results from the decreased site-specific binding of these cationic drugs to lipopolysaccharide, which has the effect to reduce outer membrane permeability, and efflux pump activation. One specific species B. vietnamiensis, is susceptible to aminoglycosides, yet resistant to other polycationic antimicrobials. In CF patients with pulmonary infection induced by B. vietnamiensis, the resistance to aminoglycoside or azithromycin treatment has been connected with the emergence of aminoglycoside elimination by means of induction of active drug efflux [8].
The vulnerability testing to antimicrobial drug combinations of BCC strains isolated from CF patients that developed resistance to single drug therapy has been extensively performed although evidence of clinical efficacy is still lacking. MIC testing using combinations of two drugs seems to have limited efficacy in blind treatment for CF patients with broad resistance to BCC. The combinations of epsilometer tests or e-test strips as well as the breakpoint combination vulnerability testing are simpler and faster approaches that are considered for screening the efficacy of two-drug combinations against BCC. Though they have good association, they still have clinical limitations [9,10,11]. Recently, combination of three drugs, i.e., tobramycin, meropenem, with a third mediator being either piperacillin–tazobactam, ceftazidime, trimethoprim–sulfamethoxazole, or amikacin, was effective in vitro against half of 47 multidrug-resistant BCC isolates producing biofilms. Improvement of new agents with in vitro and in vivo activity against BCC for patient therapy is still challenging [12,13,14,15].
Reverse vaccinology includes bioinformatics and structural biology methods that were developed after the revolution of next-generation genome sequencing technologies (NGS) for rapid identification of novel therapeutics by mining the enormous quantity of prokaryotic genome data. Other approaches like subtractive microbial genomics and differential genome analysis were also implemented for the identification of molecular targets in different human pathogens like M. tuberculosis, Burkholderia pseudomallei, Helicobacter pylori, Pseudomonas aeruginosa, Neisseria gonorrhea Corynebacterium pseudotuberculosis, and Salmonella typhi, among others [16,17,18]. The main idea is to find therapeutic targets that are keys for the pathogen survival and that do not possess any homologous pair in the host, i.e., specific to the pathogen. As a consequence, the drug inhibitors specific for these protein targets are expect to carry main benefits to patient by maximizing therapy effectiveness and minimizing collateral toxicity due to off-target activity. Some pathogen proteins might show a certain degree of homology to their respective host proteome, yet they might be assigned as potential targets for structure-based specific inhibitor development given site-specific differences in active site amino acid residues in the pathogen protein or other druggable pockets. For such studies, a prerequisite is the availability of completely sequenced genomes of target pathogens for data screening. The Burkholderia genome database (www.burkholderia.com) includes all updates concerning BCC genome annotations, curation, and comparison, thereby, providing a great asset for the CF research community [19, 20].
In this report, the genomic data of multiple BCC strains were mined for novel broad-spectrum druggable protein targets together with the identification of novel drug-like inhibitors from the ZINC15 database (http://www.zinc15.docking.org), with a goal to encompass the BCC pathogenesis. Additionally, it might open new insights for finding novel therapeutics against cystic fibrosis.
Materials and methods
Retrieval of proteome datasets
BCC strains (n = 47) were randomly included in this study at the time this project was performed, and their complete genomes/proteomes (referred hereafter interchangeably) were retrieved from the ftp server of NCBI (ftp://ftp.ncbi.nih.gov/genomes/Bacteria) [21]. All strains were classified in three groups based on their sample collection source: (a) strains isolated from human sources; (b) strains from plant sources; and (c) strains from environmental sources. The data from NCBI were cross-checked with the Genome OnLine Database—GOLD (www.gold.jgi.doe.gov) [22]. Figure 1 describes the hierarchical steps followed for subtractive genome analyses to rank putative novel drug and vaccine targets. All redundant genomes and protein sequences were removed, and only genome representing each strain was retained. Also, draft and incomplete genomes were excluded to ensure analysis standardization. Tables 1, 2, 3, and 4 give elementary genome information regarding bacterial strains, bioproject ID, replicons, genome size, predicted proteins, chromosome/plasmid accession numbers, GC content (%), total genes and pseudogenes, tRNA, rRNA, and disease data.
Phylogenetic analyses
Phylogenetic analyses for BCC could be performed using multiple genes including 16S rRNA, recA, gyrB, rpoB, acdS, atpD, gltB, and lepA genes, employing both maximum likelihood and neighbor-joining (NJ) approaches [23], for convenience, we selected the housekeeping gene 16S rRNA. To find a phylogeny inference, i.e., a hypothetical chart representation and not definitive facts of evolutionary relationships among organisms, multi-fasta files containing sequences of 16S rRNA gene [(cytosine967-C5)-methyltransferase (WP_027788851.1)] from all forty-seven strains (combined and individual tree construction for each environmental, plants, human groups), were prepared and subjected to the Molecular Evolutionary Genetics Analysis software (MEGA7) (www.megasoftware.net) [24, 25]. The pattern of branching reflects how species are evolved from a series of ordinary ancestors. MEGA7 creates a multiple alignment file (fasta.txt) and forwards it to a multiple sequence alignment tool (ClustalW) for phylogeny inference. The NJ trees were constructed based on Tamura–Nei distances, using 1000 bootstrap replicates [26].
BCC core genome and non-host homology
To find the core genome of a specific species individually or as whole for all of the three BCC groups, EDGAR software was used. EDGAR is an Efficient Database framework for comparative Genome Analyses that uses BLAST score Ratios and is freely available (https://www.uni-giessen.de/fbz/fb08/Inst/bioinformatik/software/EDGAR) [27, 28]. This framework aims at high-throughput comparative genomics analyses of large groups of related genomes by clustering orthologous genes and then classify these genes as core genes or singletons. PATRIC, Pathosystem Resource Integration Center (https://www.patricbrc.org) was employed for circular genome comparison that has integrated data analysis tools performing a wide range of bioinformatics analyses related to biomedical research on bacterial infectious diseases [29]. Furthermore, the identified core genomes/proteomes of human, plant, and environmental BCC groups were compared with each other using the NCBI-BLASTp program (www.ncbi.nlm.nih.gov) (default threshold) to acquire a single-core region for all of the 47 strains. The core file was cross-checked with Burkholderia cenocepacia HI2424 (human host) to verify the core proteome data using BLASTp (all-against-all, e-value ≤ 0.0001, bit score ≥ 200) [16, 30].
Next, the NCBI-BLASTp was re-employed to compare the BCC representative core proteomes with the human RefSeq proteome for non-homology analyses to avoid off-targeting using the default parameters (e-value ≤ 0.0001, bit score 200, and identity ≥ 25%) [17]. For non-host homology analyses, only the human host proteome was considered because of the complexity and unavailability of plant and environmental host genomes in the public databases.
Gene essentiality, subcellular localization, and virulence analyses
The minimal set of genes that are essential to support cellular life are called essential genes. DEG datasets are now accessible in the literature and comprises all essential genes for bacteria, archaea, and eukaryotes. To identify essential genes in BCC, the core non-host homologous proteins were considered as query data versus the subject data of DEG essential genes in the BLASTp (e-value ≤ 0.0001 and identity ≥ 25%) (28). The non-redundant and non-homologous proteins were further investigated for subcellular location to check the exoproteome and secretome of the BCC using PsortB (www.psort.org/psortb/) [31] and Cello2GO (www.cello.life.nctu.edu.tw.cello2go/) [32], both of these tools are based on vector machine and suffix tree algorithm features. The exoproteome and secretome are viewed as source of vaccine candidates because of their continuous contact with biotic and abiotic elements in the extracellular environment. Subcellular localization of proteins was performed. Both tools predicted same number of proteins from outer membrane and extracellular locations and were further evaluated for their involvement in virulence [31, 33]. Virulent factors are proteins involved in disease intensity, which are associated to microbial pathogenesis. This step is important because antigenic/virulent proteins could serve worthy vaccine candidates since they intervene serious flagging pathways in the host cells and might potentially activate host immune system in contrast to non-virulent proteins. Additionally, the mixtures of virulent proteins from a pathogen induce improved host protection when challenged with the respective microbial infection. Extracellular and surface membrane proteins retrieved from the previous screening step were compared to the Virulent Factor Database-VFDB (www.mgc.ac.cn/VFs/) to extract informational index of active proteins. Proteins having bit score ≥ 100 and identity of ≥ 50% were considered as potential virulent proteins, 40% and 60% sequence identity values are normally sufficient for functional transfer, whereas at domain level it ranges from 50 to 70% and sometimes even 80% [34, 35].
3D structure prediction and targets prioritization
For a putative protein to be an attractive druggable target, several prioritization parameters are considered as pathogenic markers essential for the microbe such as subcellular localization, protein–protein interaction (ppi), molecular weight, and druggability analyses. Those fulfilling the required criteria were considered as drug targets. For modeling 3D structures, core proteins were submitted to the MHOLline suite (www.mholline.lncc.br) as adapted by Hassan et al. [16] that integrates HMMTOP (a tool for prediction of transmembrane helices and proteins topology), BLAST, (BATS) Blast Automatic Targeting for Structures), MODELLER (comparative homology modeling tool for protein 3D structure prediction), and PROCHECK (checks the stereochemical quality of a protein structure by analyzing residue-by-residue the geometry and overall protein structure), among others. From the MHOLline summary file, only G2 group sequences (including very high- and high-quality sequences) were selected for 3D structure prediction, for cross-checking these were further subjected to the Swiss-Model database (www.swissmodel.expasy.org/) that uses the query sequence against the SWISS-MODEL template library using BLAST and HHBlits searches [36]. The structure qualities, Q-mean values, and Ramachandran scores were evaluated via PROCHECK [37] and, PyMOL (v2.3) (https://pymol.org/2/) [38] and UCSF Chimera [39] were used for visualization. The molecular weights (MW) of potential targets were calculated using ExPASy (https://web.expasy.org/compute_pi/) [40] and were classified accordingly. The STRING protein–protein interaction network and function enrichment analyses tool (https://string-db.org) was used to identify the interactome of target proteins [41]. For druggable pockets, a well-established fact is that the druggability of a protein 3D structure determines their efficiency to bind a drug-like molecule/ligand. To this end, the core essential non-homologous target proteins were submitted to the DoGSiteScorer (https://proteins.plus) [42]. The DoGSiteScorer is a pocket recognition and examination tool to compute the druggability of protein cavities. The druggability score ranges from 0 to 1, a standard value closer to 1 designate a highly druggable protein cavity, i.e., the cavities that are expected to bind ligands with high affinity. For further validation, the list of final targets was then subjected to the Target-Pathogen Database (http://target.sbg.qb.fcen.uba.ar) to prioritize drug targets in pathogens. This Database also focuses on the structural druggability, essentiality, and metabolic role of proteins.
Virtual screening
A ZINC library of 12,000 drug-like compounds was retrieved from the ZINC database (www.zinc.docking.org) and employed in virtual screening against the final set of predicted putative BCC targets using Molecular Operating Environment (MOE v2016.11) [43, 44]. MOE performs accurate prediction of binding affinities through a predefined algorithm and scoring to classify best docking orientations between receptor and ligand in a ligand–receptor interaction analysis. In MOE, docking and visualization were performed according to a slightly modified protocol by Basharat et al. [20], the parameters used were as follows: placement = triangle matcher, rescoring 1 = London dG, refinement = forcefield, rescoring 2 = affinity dG. All docked ZINC compounds were arranged in ascending order according to their binding energies and those with least energy of ligand–receptor complex was considered as top conformation. Compounds that were able to pass Lipinski’s drug-like test and had minimum energy were selected as suitable inhibitors. Top two ligands for each target protein were selected among the 20 best ranked compounds. Each top ligand was among those having the maximum number of hydrogen bonds (H-bonds) and the lowest ligand–receptor energy S scores.
ADMET and MD simulation studies and binding free energy calculations
Among the top 10 hits that had minimum energy and were able to pass Lipinski’s drug-like test were selected as suitable inhibitors. ADME/Tox analysis and skin permeation/other physicochemical values were calculated using ADMET prediction server (http://lmmd.ecust.edu.cn/admetsar2) and Swiss ADME (http://www.swissadme.ch/), respectively [45, 46] to validate the parameters for suitable drug/binding candidates. The structure of each ligand was optimized before docking analyses by calculating charges, structure correction if required, applying force field (MMFF94x) and minimizing energy.
Molecular dynamics simulation (MD) was done considering the top best drug complexed with the predicted target protein and was subjected to NAMD package (v2.14 GPU36) using the CHARMM36m force field [47,48,49]. The particle mesh Ewald (PME) method evaluated long-range Coulombic interactions. The integration time step was set to 2 fs (femto seconds). The production simulations were performed in the NPT ensemble (constant number of particles, pressure, and temperature) (p = 1.01325 bar and T = 300 K), using the Langevin dynamics. Na+ and Cl− ions, corresponding to a physiological concentration of 150 mM, were placed in the simulation box to set the ionic strength and neutralize the systems. After 10,000 steps (20 ps) of minimization, the complexes were equilibrated for 135,000 steps (270 ps). The production simulations last 200 ns and the trajectories from MD were analyzed using MD analysis software [47, 48], while interactions were calculated with PLIP (v2.1.6) software [49].
The MM/PBSA method (molecular mechanics poisson–boltzmann surface area) is one of the most widely adopted approaches for calculating binding free energies (ΔGbind) of ligands bound to biomolecule receptors after molecular docking or dynamics. MM/PBSA was employed for binding free energy calculations that are performed in three steps, Molecular Mechanics (MM), Poisson–Boltzmann (PB) (or generalized Born (GB), and Surface Area solvation (SA) before the summation is used to estimate the binding energy [50]. ΔGbind were done using the MM/PBSA as implemented in the CaFE package, a plugin of VMD software [51, 52].
Results
Overview of B. cepacia complex and genome statistics
The genomic data of BCC obtained from NCBI were tabulated in three different groups (G) including the environment samples (Table 1), the plant samples (Table 2), and human samples (Table 3). In the environmental samples, the number of proteins/genes comprises 5515/5629 to 7396/7590 and %GC content from 65.73 to 67.68%. The Burkholderia vietnamiensis G4 ATCC 53,617, the only strain in this group has five plasmids, while the Burkholderia pseudomultivorans SUB-INT23-BP2 have only one plasmid. In the plant samples, the number of proteins/genes are from 6449/5908 to 7354/7524 and the %GC content are 66.60–67.09%. Five out of seven in plant group had a single plasmid. On the other hand, in human group, the number of proteins/genes was between 5561/5717 and 7425/7898 with a %GC content between 66.31% and 67.31% and only 8 strains have 1 plasmid. The table includes information regarding the disease type, their prospective hosts, source of sample isolation, and country information emphasizing the global prevalence of BCC pathogenesis and their broad-spectrum host diversity.
Phylogenetic analyses of BCC
Phylogenetic tree is a graph display of the evolutionary relationship among taxa under comparison. The basic assumption of a phylogenetic investigation is that all taxa (leaves of the tree) are related among them through homologous relationships with hypothetical ancestors that compose the internal vertices of the tree. Here, we compared the phylogenetic relationships between BCC species or strains based on the nucleotide sequences of the 16S rRNA (cytosine967-C5)-methyltransferase gene, though, as aforementioned, many other genic combinations have also been used, other studies also describe the cluster and distinct lineages formation in detail based on NJ as well as ML phylogeny analyses [23]. Through MEGA7, multiple phylogenetic trees were constructed, where all the 47 strains are split in two clusters (Fig. 2A–D), where interestingly BCC strains from all sources are mixed in the two clusters. Inside MEGA7, one can download the sequences into the Alignment Explorer and retrieve the unaligned sequences in FASTA format by selecting Export Alignment from the Data menu. The multiple alignment could be followed as per study requirement using tools like Muscle, T-coffee, and Clustal Omega available at EBBL-EBI (https://www.ebi.ac.uk/Tools/msa/) and their output files could be used as input files in the MEGA program (41). MEGA7 uses ClustalW together with the neighbor-joining (NJ) method and a focus was made to elucidate the ancestral relationship in Genus Burkholderia of all different strains isolated from diverse sources and locations worldwide. It is assumed that a more closely or distinctly relatedness among various BCC strains could aid in designing future projects, where a genus size could vary by the addition of new variants and, hence, would give an insight into their pan and core genomes as well as therapeutic targets identification for a broad spectrum of BCC pathogens.
A Phylogenetic tree of 47 BCC species and strains based on the 16S rRNA (cytosine967-C5)-methyltransferase gene. B Phylogenetic tree of 19 BCC strains isolated from human sources based on the 16S rRNA (cytosine967-C5)-methyltransferase gene. C Phylogenetic tree of seven BCC strains isolated from plant sources based on the 16S rRNA (cytosine967-C5)-methyltransferase gene. D Phylogenetic tree of 21 BCC strains isolated from environmental sources based on the 16S rRNA (cytosine967-C5)-methyltransferase gene
Prediction of intraspecies core genome with EDGAR and PATRIC
EDGAR comparative genomics workflow was employed to identify core genomes of BCC strains isolated from human and plant samples. At this stage, the plant core genome was identified but excluded to avoid complications with host homology analyses to be performed in the forthcoming step. The individual core files comprised of 2495 and 2371 proteins for plant and human isolates, respectively, sowing a close comparison of the BCC strains isolated from completely different sources. Furthermore, PATRIC resulted bidirectional BLASTp genome comparison in circular graphs for representative human isolates (Fig. 3). PATRIC has the limitation that it can analyze only 10 genomes at a time; hence the dataset of nineteen genomes isolated from humans was divided into two sets, whereas plant isolates were analyzed separately. The proteome comparison tool of the PARIC workflow readily identified insertions and deletions in up to ten genomes among a user-selected reference genome and other nine as target genomes. The reference genome can possibly be (a) a user-private genome in PATRIC, (b) an annotated genome outside the PATRIC, (c) any genome that is publicly available in PATRIC, or (d) possibly a genome feature group containing a set of proteins saved in PATRIC. The tool performing the proteome comparison follows the basic principle of RAST tool that is based on the sequence-based comparison, coloring each gene based on protein similarity after using BLASTp [53]. Later, each of the gene is marked as either a unique, a unidirectional or bidirectional best hit after comparing to the selected reference genome. The output file includes a whole-genome schematic circular view that is colored after running BLASTp and provides an insight into the circular interactive graphical representation of the alignment of genes and other genomic data. All the tracks on the genome viewer are shown as concentric rings that are arranged from outermost to innermost including position, contigs/chromosomes, CDS forward and reverse, non-CDS features, GC content, and GC skew, along with other details given in the PATRIC user manual available at PATRIC homepage. In, general, the conserved and missing genomic regions among different BCC strains are prominent for viewing.
Core proteome, host non-homology, and essential genes identification
The core genome files from plant and human isolates were compared with BLASTp (e-value ≤ 0.0001) to find the conserved genome between the BCC species that parasite both phyla, which resulted in a single-core genome file containing 1766 genes (Fig. 4). This file was, then, compared by BLASTp (e-value ≤ 0.0001, bit score = 100 and identity ≥ 25%) to the human proteome (host for potential therapy against BCC) to filter out potential homologous protein with humans from the core BCC. Through this step, we identified 1680 proteins specific of the core BCC that did not show significant homology with humans. This step aimed to minimize unwanted toxic effect of drugs for humans resulting from cross-reactivity between BCC protein targets and the human proteome.
Distribution of intra/interspecies orthologs in BCC human and plant isolates. Venn diagram representation showing the individual intraspecies core genome/proteome of BCC isolates from human sources (left) and from plant sources (right), whereas the central part of the diagram represents the interspecies core genome/proteome of BCC genus (only selected species in this work)
To find pathogen essential genes out of 1680 core file, the amino acid data files of prokaryotes, eukaryotes, and archaea were retrieved from the DEG database followed by a comparison with the set of BCC core proteins using BLASTp (e-value ≤ 0.0001 and identity ≥ 25%). Essential genes/proteins comprise of a minimal set of genes/proteins that are essential for survival of living cell in their environment. The essential genes among the BCC core (1680 genes) were found to be only 37, which we considered as putative therapeutic targets. To improve the reliability of these proteins as potential broad-spectrum therapeutic targets, we compared the file of 37 BCC proteins with Burkholderia gladioli ATCC 10,248 (BLASTp, e-value ≤ 0.0001), another closely related BCC species. This filtering further drop downed the BCC core, essential , and non-host homologous target proteins to 21.
Comparative subcellular localization
The 21 putative therapeutic targets of BCC were shared among the Burkholderia genus, we further anticipated the comparative subcellular localization of these proteins using Cello2GO and were cross-checked using PSORTb tool. The tool represents a number of analytical modules for the prediction of target protein localization based on localization scores. These scores represent the confidence values for each of the localization sites, a site having a score > 7.5, that site and its score are returned as predicted localization. Occasionally, more than one site returns high scores that means that the protein under study may have multiple localization sites. This step is important to identify the exact location of targeted proteins and classify them as secreted, cytoplasmic, putative surface exposed (PSE), and transmembrane proteins (according to signal peptides, retention signals, and transmembrane helices in accordance to the biological role they play during cell growth, replication, and host interaction).
3D structure prediction via high-throughput comparative homology modeling
The 21 proteins submitted to MHOLline yielded an output file of different groups of target protein sequences comprising G0, G1, G2, and G3 groups where only the G2 group could be proceeded for further analyses. The MHOLline generates these four groups depending on alignment quality parameters of input sequences for finding template structures in the PDB database using integrated BLASTp and BATS tools. Based on initial alignment, the grouping of input sequences in G2 follow strict parameters; e-value ≤ 10 −5, identity ≥ 25%, and Length Variation Index (LVI) ≤ 0.7 (the coverage scores in the MHOLline, i.e., LVI ≤ 0.1 is equivalent to ≥ 90%, an identity coverage between query and target protein) thus G2 is the only group that comply with MHOLline criterion and was used for further analyses. The G2 group comprises further seven subgroups (BATS analyses) where only the first four groups (good, medium to good, high, and very high-quality sequences) were included in this study. In subgroups, only 13 distinct quality model sequences were found, for 12 sequences the 3D homology structures were effectively predicted by MHOLline via integrated MODELLER software. The output summary file summarizes information regarding all steps of the MHOLline workflow where among the selected four subgroups of G2 contained structural data for six sequences only both from high and very high-quality subgroups, thereby reducing the putative target list to six from 21 (Table 4). The PROCHECK values from the MHOLline suite, showing the stereochemical qualities of the constructed models were further checked for the final set of 6 BCC targets.
3D structure comparison, virulent factors, and interactome analyses
The MHOLline structures were cross-checked with SWISS-MODEL structures as both MHOLline and SWISS-MODEL employee Modeler software for 3D structure prediction. Qualities of these structures (PROCHECK values) were almost the same, over 90% of the amino acid residues of target proteins were in the most favored regions of the Ramachandran plot. However, in some cases of low-quality 3D structures from MHOLline, the SWISS-MODEL gave better results to ensure the success of future docking analyses. Furthermore, targets were subjected to VFDB that analyzed and reported all as virulent proteins (Table 5). Virulence factors (VFs) refer to the properties of pathogenic bacteria in terms of gene products that make them capable to establish an internal or external interaction with a host cell, proliferate and enhance their potential to cause a disease. These VFs include bacterial toxins, cell surface proteins, cell surface glycoproteins, bacterial protection proteins and hydrolytic proteins, among others. The VFDB uses VFanalyzer for systematic screening of virulence factors in bacterial complete as well as draft genomes by not using simple BLAST searches, rather it first constructs orthologous groups within the query genome and pre-analyzed reference genomes from VFDB to avoid potential false positive results due to genes/proteins paralogues. The next step is an iterative and exhaustive sequence similarity searches among the hierarchical pre-build datasets of VFDB to accurately and specifically identify VFs in query strains. At the end, VFanalyzer achieve relatively high specificity and sensitivity through a context-based data refinement process for VFs encoded by gene clusters [34]. BCC target proteins having bit score ≥ 100 and identity of ≥ 50% were identified as potential VFs and were rendered for further physicochemical extrapolations.
ProtParam is a molecular weight calculator for biological macromolecules, an important step toward target prioritization in accordance to the Lipinski rule of five (Table 5) [54]. The STRING aims at collecting scores and integrating all publicly available sources of protein–protein interaction information, and to complement these data with computational predictions. The predicted interactome based on the six protein targets enabled to detect the protein neighbors that showed maximum interactions (minimum three interactions) (Fig. 5). All putative targets were found to have more than 10 interactions but Bamb_0438 (RS01725_chromosome_1 30S ribosomal protein S10), and Bamb_0648 (RS03735_chromosome_1 Aminodeoxychorismate synthase component I) showed even more interactions. Bamb_0648 is a heterodimeric complex, which provides important physiological functions unique to plants, bacteria, fungi and certain parasites and due to its absence in plant and animal hosts make it an excellent target for antimicrobial agents and herbicides [55].
STRING analysis of protein–protein interactions for the six putative protein targets. Green lines connect proteins which are associated by recurring neighborhood method of the STRING database; blue connections are inferred by phylogenetic co-occurrence, and red lines indicate gene-fusion events; line thickness is a rough indicator for the strength of the association; purple lines indicate experimental evidence; yellow lines show text mining indication; black lines denote co-expression evidence; light blue lines represent database evidence. The colorful circles denote nodes while the lines are for edges
Deciphering of druggability and druggable pockets
The information acquired from 3D structures and druggability studies are essential features for drug development to inhibit pathogen targets. According to DoGSiteScorer (Fig. 6A–F), all target proteins were found highly druggable. Meanwhile the Target-Pathogen Database, a bioinformatics approach to prioritize drug targets in a pathogen, was also consulted in order to re-check and compare the druggability and other biochemical functions. DoGSiteScorer uses a “Difference of Gaussian” filter to detect potential binding pockets (grid-based method) depending solely on the protein 3D structure, splitting them into sub-pockets. Protein global properties are calculated describing the size, shape and chemical features of the predicted sub-pockets and assign a druggability score to each sub-pocket, based on a linear combination of the three descriptors describing volume, hydrophobicity and enclosure. Furthermore, another subset of specific descriptors is added in a support vector machine (libsvm) to predict the druggability score of sub-pocket/s [42]. Target proteins with a score ≥ 0.8 were predicted as highly druggable on a scale ranging from 0 to 1. The different colors of pockets refer to the druggability score, i.e., a green colored pocket represent high druggability whereas a red colored pocket represent low druggability, the other colored pockets lie between these druggability scores and colored pockets, respectively. The information of putative cavities with their corresponding druggability scores are given in Table 5.
A–F Cartoon representation of three-dimensional models of BCC target proteins together with identification of druggable pockets via the DoGSiteScorer server. A pocket having a score closer to 1 is regarded as highly druggable and vice versa, on a 0–1 scale. A 30S ribosomal protein S10; B 50S ribosomal protein L6; C Aminodeoxychorismate synthase component I; D 50S ribosomal protein L5; E Bifunctional N- acetylglucosamine-1-phosphate uridyltransferase/glucosamine-1-phosphate acetyltransferase; F Chorismate synthase
Virtual screening and molecular docking analyses
The functional characterization of six targets using UniProt, KEGG, ExPASy, and InterProScan databases aided in selecting final targets by emphasizing on their prospective roles in different metabolic pathways, thus, resulted in only two proteins; BCEN2424_RS14895, Bifunctional N-acetylglucosamine-1-phosphate uridyltransferase/Glucosamine-1-phosphate acetyltransferase glmU (PDB template: 2W0W), and BCEN2424_RS07230, Chorismate synthase aroC (PDB template: 1UM0), which were rendered to virtual screening (VS) using a library of 12,000 drug-like compounds (ZINC15 database) via the MOE program. For VS, we only selected the protein targets that already had ligand compounds in their 3D templates, already retrieved from PDB database. The resulting lists contained the best hits for each putative target protein. The interactions within the active site of PDB target-ligand complex structure were checked and, then, were followed by docking analyses selecting only the specific residues involved in the putative target activity. The lower energy scores of the MOE program indicates a better ligand–protein binding complex formation compared to high energy values. In this work, the best hits from the VS step were docked, each having 15 poses, for the identification of best ranked ligands. For both BCEN2424_RS14895 and BCEN2424_RS07230 the top 10 best ranked hits, their energy values, and 2D interactions details are tabulated, respectively (Tables 6, 7), while 2D interactions are shown only for the top 1 compound for both targets. Figure 7 illustrates the interactions of ZINC06055530 into the druggable cavity of BCEN2424_RS14895 (Glucosamine-1-phosphate acetyltransferase glmU), interacting with two glycine residues (Gly9 and Gly99) through hydrogen bonding with the minimum possible binding energy value (−6.8601) as compared to other nine best hits across the column (Table 6). Gly9 made an arene–hydrogen interaction while among other interactions, three residues were basic (shown in blue circle in Fig. 7) and one acidic (shown in red circle in Fig. 7). For ZINC01405842 interaction with aroC, six basic residues made interaction (shown in blue circle in Fig. 7), while no acidic residue made any interaction. Several hydrogen bonds were observed (Lys49, Ser126, Ser127, Arg299), where Lys49 was representative of an arene–cation interaction. Overall, energy values were lower for aroC interaction with ZINC01405842, compared to GlmU interaction with ZINC06055530 compound.
Two-dimensional (2D) representation of drug–protein target interactions using MOE. Top best ranked docked compounds with best possible orientations for ZINC06055530 and ZINC01405842 in the most druggable cavity of bifunctional N-acetylglucosamine-1-phosphate uridyltransferase/glucosamine-1-phosphate acetyltransferase (BCEN2424_RS14895) and Chorismate synthase (BCEN2424_RS07230), respectively
ADMET Profiling, MD simulation, and binding free energy calculations
For the selected compounds, pharmacokinetics and pharmacology properties (absorption, distribution, metabolism, and excretion referred to as ADME, were studied to check higher penetration and least side effects to human and other hosts, if any. Most of them were substrates for P-glycoprotein whereas some of these compounds showed blood–brain barrier permeability or mutagenicity, and also they did not show maximum inhibition of cytochromes. Those predicted positive for mutagenicity, it is presumed that they do not cause mutations in the host DNA replication or translation processes. Most of the compounds showed the least acute oral toxicity to humans. Since the top 10 hits were selected, other compounds from the remaining 8 inhibitors could possibly be selected; in case, some are hazardous to human or other hosts. The drug-like compounds mined in this study as potential inhibitor candidates were found to be active, safe, and have not previously been studies as anti-BCC and would require laboratory validations (Tables 8, 9).
To study the complex stability, properties like free binding energy, root-mean-square deviation (RMSD), number of interactions, root-mean-square fluctuation (RMSF), and the radius of gyration (Rg) are very useful for studying the stability of the complexes. Most of the energy values are negative that indicates a favorable ligand–protein complex formation.
Table 10 shows the free binding energy and contribution of different mechanisms to it. Negative free binding energy implies favorable complex formation. In this case, only complex 01 formation has negative energy, and then turns out to be favorable. Nevertheless, it is positively smaller, and the contribution of Coulomb and van der Waals interaction is stronger in this complex than in complex 01.
The latter is confirmed accounting the interactions made during the simulation time. The number of hydrogen bonds (H-bond), hydrophobic contacts, salt bridge, π-π stacking, and π-cation interactions through the simulation were determined for each complex using the PLIP software. Form Fig. 8, we can see that the total interaction number made by 02 is more than twice than 01 but they are made by less residues. In both cases, almost all interaction types are present in each complex.
Figure 9 shows that the RMSD for both complexes attain stability after the first 50 ns of the simulation showing with little high values around 8 and 5 Å for system 01 and 02, respectively. It is well known that sometimes, the rigid-body alignment is not rich enough and, the RMSD and RMSF will increase for all atoms, overestimating them and neglecting important fluctuations associated with biological function if there are small portions of the complex with high mobility [56].
A measure of the residue fluctuation is the root-mean-square fluctuation (RMSF) parameter. Figure 10 show the RMSF calculated for both complexes. Similar to the results for RMSD, the 02 complex shows lower values for all the residues responsible for the interactions (see figure PLIP Interaction figure), indicating that the ligand makes the enzyme less flexible.
A measurement of how compact the complex is, can be verified by calculating the radius of gyration (Rg). The complex 01 shows the lowest and most stable value of Rg than the complex 02 with oscillations around 1.5 Å. On the other hand, complex 02 shows a decreasing value with simulation time (Fig. 11).
Discussion
In this work, an attempt was made to highlight the differences of genome architecture through Pangenomics including rRNA, tRNA, pseudogenes, % GC content and size of different BCC strains isolated from plant, human and environmental sources followed by identification of therapeutic protein targets in a step-by-step manner. BCC forms a group of related bacterial species that are capable of imparting cystic fibrosis and aerosol-based contamination. Bacterial genotyping showed that infections by Burkholderia cepacia strains are more specific in cystic fibrosis patients, which denotes a greater propagation capacity among these patients. Based on the genomic information, we constructed different phylogenetic trees to check the ancestral relationship of all different isolates of BCC complex on an individual and combined basis. The combined phylogenetic tree showed that strains isolated from plant, human, and environmental sample sources represent different branch position when compared to the individual trees, which might implicate that BCC strains isolated from different sources might be able to evolve differently under specific circumstances and cause diseases in several different hosts. Due to the fact that BCC is responsible for a wide range of diseases from nosocomial infection in CF patients to rice root infection in plant, among other, a focus was made on finding the minimal set of genes/proteins shared by all the strains (core genome/proteomes) included in this study. Keeping a stringent criterion for core genome identification for a large number of bacterial genomes drastically reduces the totality of core protein targets in comparison to the pan genome that increase with increase in number of genomic datasets under study. This core data although shared by BCC strains might also share similarity with their host genomes/proteomes in terms of homology, therefore it is very important to filter such data at this stage by comparing to their respective host genomes. Since the NCBI database provide more information about viral and prokaryotic genomes and up to certain degree of eukaryotic organisms, the host homology in this study was restricted only keeping in mind the human as the major host of utmost importance. Following these filtering steps reduces the number of genes/proteins in the dataset under analyses that was further reduced after subjecting to gene essentiality step where only those genes/proteins are selected that are vital for the survival of the microorganisms.
To reduce the cost and development time of BCC drugs, virtual screening (VS) of a large number of drug libraries is now extensively used. Only few compounds have yet been discovered by virtual screening analyses that might interact with the BCC protein structures. Despite the fact that structural information was computationally predicted and could, therefore, differ from experimental facts, we constructed the 3D models of target proteins by comparative homology modeling using experimental templates obtained from the PDB databank. No compound is reported so far mentioning interaction to our identified target protein structures via high-throughput virtual screening methods. Therefore, in the current study, the top ten ligands were selected on the basis of their binding affinities after the screening of a library of 12,000 drug-like molecules. Among them, the best ligands were selected according to their high binding affinity and minimal energy scores for ligand–receptor interaction. The information given here might further aid in designing bench experiments for antibiotic and vaccine development. The putative BCC protein candidates that were identified here are key therapeutic targets for a number of reasons given below separately.
BCEN2424_RS14895_glmU
Bifunctional N-acetylglucosamine-1-phosphate uridyltransferase/glucosamine-1-phosphate acetyltransferase is an essential precursor of peptidoglycan and rhamnose-GlcNAc linker region of the mycobacterial cell wall. The pathway for UDP-GlcNAc biosynthesis is significantly different in eukaryotes and prokaryotes. Since in vitro experiments showed that glmU is essential for bacterial cell wall, we assume that it is a potential drug target in BCC. It is also reported as a drug candidate for tuberculosis [57]. The best interacting leads are shown along with their ZINC IDs, minimized energy, number of interactions, and interacting residues. ZINC06269029 was predicted as the top-ranked molecule interacting with Gly9 and Gly99 residues in the binding site of glmU (Table 4, Fig. 7).
BCEN2424_RS07230_aroC
Chorismate synthase catalyzes the formation of Chorismate, the last step of the shikimate pathway. Chorismate is a branch-point metabolite used in the synthesis of aromatic amino acids, p-aminobenzoic acid, folate, and other cyclic metabolites such as ubiquinone. Shikimate pathways are present only in plants, fungi, and bacteria, making these pathway enzymes possible targets for herbicides, antibiotics, and antifungals. Chorismate synthase from Mycobacterium tuberculosis is also considered to be a potential therapeutic target [58]. A comparison between model template structures was made and Lys49, Ser126, Ser127, and Arg299 residues are shown with top ZINC-selected docked compound (Table 5, Fig. 7).
Conclusions
In this report, we performed a series of in silico analyses using a number of bioinformatics tools that led us to identify novel therapeutic targets for the first time in BCC. Furthermore, some of the BCC targets identified here were already reported experimentally, which validate our methodology. We believe that the set of target proteins proposed here is worthy for future in vitro and in vivo experimentation for drugs and vaccine development. Furthermore, the set of integrated techniques used here is/could be extended to the search of therapeutic targets in a number of other pathogens [59].
References
Ferreira AS, Leitão JH, Sousa SA et al (2007) Functional analysis of Burkholderia cepacia genes bceD and bceF, encoding a phosphotyrosine phosphatase and a tyrosine autokinase, respectively: role in exopolysaccharide biosynthesis and biofilm formation. Appl Environ Microbiol 73:524–534. https://doi.org/10.1128/AEM.01450-06
Mahenthiralingam E, Urban TA, Goldberg JB (2005) The multifarious, multireplicon Burkholderia cepacia complex. Nat Rev Microbiol 3:144–156. https://doi.org/10.1038/nrmicro1085
Lewis ERG, Torres AG (2016) The art of persistence-the secrets to Burkholderia chronic infections. Pathog Dis 74:ftw070. https://doi.org/10.1093/femspd/ftw070
Lipuma JJ (2010) The changing microbial epidemiology in cystic fibrosis. Clin Microbiol Rev 23:299–323. https://doi.org/10.1128/CMR.00068-09
Lipuma JJ (2005) Update on the Burkholderia cepacia complex. Curr Opin Pulm Med 11:528–533. https://doi.org/10.1097/01.mcp.0000181475.85187.ed
Tseng S-P, Tsai W-C, Liang C-Y et al (2014) The contribution of antibiotic resistance mechanisms in clinical Burkholderia cepacia complex isolates: an emphasis on efflux pump activity. PLoS ONE 9:e104986. https://doi.org/10.1371/journal.pone.0104986
Mushtaq S, Warner M, Livermore DM (2010) In vitro activity of ceftazidime+NXL104 against Pseudomonas aeruginosa and other non-fermenters. J Antimicrob Chemother 65:2376–2381. https://doi.org/10.1093/jac/dkq306
Jassem AN, Zlosnik JEA, Henry DA et al (2011) In vitro susceptibility of Burkholderia vietnamiensis to aminoglycosides. Antimicrob Agents Chemother 55:2256–2264. https://doi.org/10.1128/AAC.01434-10
Dales L, Ferris W, Vandemheen K, Aaron SD (2009) Combination antibiotic susceptibility of biofilm-grown Burkholderia cepacia and Pseudomonas aeruginosa isolated from patients with pulmonary exacerbations of cystic fibrosis. Eur J Clin Microbiol Infect Dis 28:1275–1279. https://doi.org/10.1007/s10096-009-0774-9
Manno G, Ugolotti E, Belli ML et al (2003) Use of the E test to assess synergy of antibiotic combinations against isolates of Burkholderia cepacia-complex from patients with cystic fibrosis. Eur J Clin Microbiol Infect Dis 22:28–34. https://doi.org/10.1007/s10096-002-0852-8
Tunney MM, Scott EM (2004) Use of breakpoint combination sensitivity testing as a simple and convenient method to evaluate the combined effects of ceftazidime and tobramycin on Pseudomonas aeruginosa and Burkholderia cepacia complex isolates in vitro. J Microbiol Methods 57:107–114. https://doi.org/10.1016/j.mimet.2003.12.001
Regan KH, Bhatt J (2019) Eradication therapy for Burkholderia cepacia complex in people with cystic fibrosis. Cochrane Database Syst Rev 4:CD009876. https://doi.org/10.1002/14651858.CD009876.pub4
Wang H, Wang H, Yu X et al (2019) Impact of antimicrobial stewardship managed by clinical pharmacists on antibiotic use and drug resistance in a Chinese hospital, 2010–2016: a retrospective observational study. BMJ Open 9:e026072. https://doi.org/10.1136/bmjopen-2018-026072
Martiniano SL, Wagner BD, Brennan L et al (2021) Pharmacokinetics of oral antimycobacterials and dosing guidance for Mycobacterium avium complex treatment in cystic fibrosis. J Cyst Fibros 20:772–778. https://doi.org/10.1016/j.jcf.2021.04.011
van der Meer R, Wilms EB, Sturm R, Heijerman HGM (2021) Pharmacokinetic interactions between ivacaftor and cytochrome P450 3A4 inhibitors in people with cystic fibrosis and healthy controls. J Cyst Fibros 20:e72–e76. https://doi.org/10.1016/j.jcf.2021.04.005
Hassan SS, Tiwari S, Guimarães LC et al (2014) Proteome scale comparative modeling for conserved drug and vaccine targets identification in Corynebacterium pseudotuberculosis. BMC Genom 15(Suppl 7):S3. https://doi.org/10.1186/1471-2164-15-S7-S3
Jamal SB, Hassan SS, Tiwari S et al (2017) An integrative in-silico approach for therapeutic target identification in the human pathogen Corynebacterium diphtheriae. PLoS ONE 12:e0186401. https://doi.org/10.1371/journal.pone.0186401
Radusky LG, Hassan S, Lanzarotti E et al (2015) An integrated structural proteomics approach along the druggable genome of Corynebacterium pseudotuberculosis species for putative druggable targets. BMC Genom 16(Suppl 5):S9. https://doi.org/10.1186/1471-2164-16-S5-S9
Winsor GL, Khaira B, Van Rossum T et al (2008) The Burkholderia genome database: facilitating flexible queries and comparative analyses. Bioinformatics 24:2803–2804. https://doi.org/10.1093/bioinformatics/btn524
Basharat Z, Jahanzaib M, Yasmin A, Khan IA (2021) Pan-genomics, drug candidate mining and ADMET profiling of natural product inhibitors screened against Yersinia pseudotuberculosis. Genomics 113:238–244. https://doi.org/10.1016/j.ygeno.2020.12.015
Sayers EW, Bolton EE, Brister JR et al (2022) Database resources of the national center for biotechnology information. Nucleic Acids Res 50:D20–D26. https://doi.org/10.1093/nar/gkab1112
Mukherjee S, Stamatis D, Bertsch J et al (2021) Genomes OnLine Database (GOLD) vol 8: overview and updates. Nucleic Acids Res 49:D723–D733. https://doi.org/10.1093/nar/gkaa983
Estrada-de los Santos P, Vinuesa P, Martínez-Aguilar L et al (2013) Phylogenetic analysis of burkholderia species by multilocus sequence analysis. Curr Microbiol 67:51–60. https://doi.org/10.1007/s00284-013-0330-9
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Thompson JD, Gibson TJ, Higgins DG (2002) Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinformatics. https://doi.org/10.1002/0471250953.bi0203s00
Dieckmann MA, Beyvers S, Nkouamedjo-Fankep RC et al (2021) EDGAR3.0: comparative genomics and phylogenomics on a scalable infrastructure. Nucleic Acids Res 49:W185–W192. https://doi.org/10.1093/nar/gkab341
Blom J, Kreis J, Spänig S et al (2016) EDGAR 2.0: an enhanced software platform for comparative gene content analyses. Nucleic Acids Res 44:W22-28. https://doi.org/10.1093/nar/gkw255
Wattam AR, Davis JJ, Assaf R et al (2017) Improvements to PATRIC, the all-bacterial bioinformatics database and analysis resource center. Nucleic Acids Res 45:D535–D542. https://doi.org/10.1093/nar/gkw1017
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Yu NY, Wagner JR, Laird MR et al (2010) PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26:1608–1615. https://doi.org/10.1093/bioinformatics/btq249
Yu C-S, Cheng C-W, Su W-C et al (2014) CELLO2GO: a web server for protein subCELlular LOcalization prediction with functional gene ontology annotation. PLoS ONE 9:e99368. https://doi.org/10.1371/journal.pone.0099368
Yu C-S, Lin C-J, Hwang J-K (2004) Predicting subcellular localization of proteins for Gram-negative bacteria by support vector machines based on n-peptide compositions. Protein Sci 13:1402–1406. https://doi.org/10.1110/ps.03479604
Liu B, Zheng D, Jin Q et al (2019) VFDB 2019: a comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res 47:D687–D692. https://doi.org/10.1093/nar/gky1080
Addou S, Rentzsch R, Lee D, Orengo CA (2009) Domain-based and family-specific sequence identity thresholds increase the levels of reliable protein function transfer. J Mol Biol 387:416–430. https://doi.org/10.1016/j.jmb.2008.12.045
Waterhouse A, Bertoni M, Bienert S et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46:W296–W303. https://doi.org/10.1093/nar/gky427
Morris AL, MacArthur MW, Hutchinson EG, Thornton JM (1992) Stereochemical quality of protein structure coordinates. Proteins 12:345–364. https://doi.org/10.1002/prot.340120407
Schrödinger L, DeLano W. PyMOL, http://www.pymol.org/pymol
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Wilkins MR, Gasteiger E, Bairoch A et al (1999) Protein identification and analysis tools in the ExPASy server. Methods Mol Biol 112:531–552. https://doi.org/10.1385/1-59259-584-7:531
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613. https://doi.org/10.1093/nar/gky1131
Volkamer A, Kuhn D, Rippmann F, Rarey M (2012) DoGSiteScorer: a web server for automatic binding site prediction, analysis and druggability assessment. Bioinformatics 28:2074–2075. https://doi.org/10.1093/bioinformatics/bts310
Sterling T, Irwin JJ (2015) ZINC 15–ligand discovery for everyone. J Chem Inf Model 55:2324–2337. https://doi.org/10.1021/acs.jcim.5b00559
Vilar S, Cozza G, Moro S (2008) Medicinal chemistry and the molecular operating environment (MOE): application of QSAR and molecular docking to drug discovery. Curr Top Med Chem 8:1555–1572. https://doi.org/10.2174/156802608786786624
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Shen J, Cheng F, Xu Y et al (2010) Estimation of ADME properties with substructure pattern recognition. J Chem Inf Model 50:1034–1041. https://doi.org/10.1021/ci100104j
Lee J, Cheng X, Swails JM et al (2016) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12:405–413. https://doi.org/10.1021/acs.jctc.5b00935
Lee J, Hitzenberger M, Rieger M et al (2020) CHARMM-GUI supports the Amber force fields. J Chem Phys 153:035103. https://doi.org/10.1063/5.0012280
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29:1859–1865. https://doi.org/10.1002/jcc.20945
Gowers R, Linke M, Barnoud J et al (2016) MDAnalysis: a python package for the rapid analysis of molecular dynamics simulations. SciPy, Austin, pp 98–105
Michaud-Agrawal N, Denning EJ, Woolf TB, Beckstein O (2011) MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J Comput Chem 32:2319–2327. https://doi.org/10.1002/jcc.21787
Adasme MF, Linnemann KL, Bolz SN et al (2021) PLIP 2021: expanding the scope of the protein–ligand interaction profiler to DNA and RNA. Nucleic Acids Res 49:W530–W534. https://doi.org/10.1093/nar/gkab294
Wang E, Sun H, Wang J et al (2019) End-point binding free energy calculation with MM/PBSA and MM/GBSA: strategies and applications in drug design. Chem Rev 119:9478–9508. https://doi.org/10.1021/acs.chemrev.9b00055
Liu H, Hou T (2016) CaFE: a tool for binding affinity prediction using end-point free energy methods. Bioinformatics 32:2216–2218. https://doi.org/10.1093/bioinformatics/btw215
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Overbeek R, Olson R, Pusch GD et al (2014) The SEED and the rapid annotation of microbial genomes using subsystems technology (RAST). Nucleic Acids Res 42:D206-214. https://doi.org/10.1093/nar/gkt1226
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26. https://doi.org/10.1016/s0169-409x(00)00129-0
He Z, Toney MD (2006) Direct detection and kinetic analysis of covalent intermediate formation in the 4-amino-4-deoxychorismate synthase catalyzed reaction. Biochemistry 45:5019–5028. https://doi.org/10.1021/bi052216p
Irfan M, Tariq M, Basharat Z et al (2022) Genomic analysis of Chryseobacterium indologenes and conformational dynamics of the selected DD-peptidase. Res Microbiol. https://doi.org/10.1016/j.resmic.2022.103990
Martínez L (2015) Automatic identification of mobile and rigid substructures in molecular dynamics simulations and fractional structural fluctuation analysis. PLoS ONE 10:e0119264. https://doi.org/10.1371/journal.pone.0119264
Zhang W, Jones VC, Scherman MS et al (2008) Expression, essentiality, and a microtiter plate assay for mycobacterial GlmU, the bifunctional glucosamine-1-phosphate acetyltransferase and N-acetylglucosamine-1-phosphate uridyltransferase. Int J Biochem Cell Biol 40:2560–2571. https://doi.org/10.1016/j.biocel.2008.05.003
Bulloch EMM, Jones MA, Parker EJ et al (2004) Identification of 4-amino-4-deoxychorismate synthase as the molecular target for the antimicrobial action of (6s)-6-fluoroshikimate. J Am Chem Soc 126:9912–9913. https://doi.org/10.1021/ja048312f
Acknowledgements
We offer our sincere gratitude to Prof. Dr. Atta Ur Rehman, Prof. Dr. Iqbal Chaudhary, and other team members for being the founder and pioneer of JRC Genome Research, ICCBS, UoK. This work was carried out under mutual agreement between CAPES—RJ and CDTS—Fiocruz—RJ, Brazil. Furthermore, we would like to extend our gratitude and acknowledge the support and cooperation of all team members and collaborators. This project was carried out in a mutual collaboration among the Department of Chemistry, Islamia College Peshawar, KP-Pakistan, PCMD- ICCBS, University of Karachi, Sindh-Pakistan and the Brazilian research institute CDTS—Fiocruz, Rio de Janeiro—Brazil. This study was carried out under the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior–Brasil (CAPES)–Finance Code 001 (CAPES-Fiocruz/CDTS).
Funding
This research project did not receive any funding/grant from public, commercial or non-profit sector/organization/s.
Author information
Authors and Affiliations
Contributions
Conceived and designed the project: SSH, RS, SS, CMM. Analyzed the data: SSH, RS, SS, ZB, IC. Wrote the first draft of the manuscript: SSH, RS, JA. Contributed to the writing of the manuscript: JA, IC, RS. Agreed and reviewed the manuscript data and results: AU, JA, MB, ZB, YK, SSH. Jointly prepared the arguments and made critical revision: SSH, RS, AU, MI, MB, ZB. Approval of the final manuscript version. The final manuscript was reviewed and approved by all authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hassan, S.S., Shams, R., Camps, I. et al. Subtractive sequence analysis aided druggable targets mining in Burkholderia cepacia complex and finding inhibitors through bioinformatics approach. Mol Divers 27, 2823–2847 (2023). https://doi.org/10.1007/s11030-022-10584-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10584-5