Abstract
Different SARS-CoV-2 new variants emerged and spread during the past few months, sparking infections and death counts. The new variant B.1.617 (delta variant) sparked in India in the past few months, causing the highest records. The B.1.617 variant of SARS-CoV-2 has the double mutations E484Q and L452R on its spike Receptor Binding Domain (RBD). The first mutation is like the reported South African and the Brazilian variants (501.V2 and B.1.1.248). This mutation lies in the region C480-C488, which we predicted before to be recognized by the host-cell receptor; Glucose Regulated Protein 78 (GRP78). In the current study, we test the binding affinity of the host-cell receptor GRP78 to the delta variant spike RBD using molecular docking and molecular dynamics simulations of up to 100 ns. Additionally, the ACE2-RBD is tested by protein–protein docking. The results reveal equal average binding affinities of the GRP78 against wildtype and delta variant spikes. This supports our previous predictions of the contribution of GRP78 in SARS-CoV-2 spike recognition as an auxiliary route for entry.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
According to the john-Hopkins COVID-19 counter, India reported the highest daily new record worldwide in the number of infections on May 6, 2021. This spark in the highly contagious virus was attributed to the new variant B.1.617 (delta strain). This variant has double mutations, E484Q and L452R, that are suggested to be crucial for viral recognition because it is found in the RBD of the viral spike. The mutant at the 484 position of the spike was reported before in other variants of SARS-CoV-2 such as B.1.1.248 (beta variant) and 501.V2 (gamma variant), in which it was E484K (Ibrahim et al. 2021). In addition, some cities in India were suffering from limited hospital beds, medicines, and oxygen supplies, leading to a death count surge last year (Explainer: What we know about the Indian variant as coronavirus sweeps South Asia; The effects of virus variants on COVID-19 vaccines: WHO 2021).
Fear is now facing the world due to the massive spread of the delta variant (Explainer: What we know about the Indian variant as coronavirus sweeps South Asia). Studying the mutations that emerged in the spike RBD is essential due to its involvement in vaccine recognition (The effects of virus variants on COVID-19 vaccines: WHO 2021). In previous studies, we reported the efficiency dependence of the SARS-CoV-2 variant on its recognition behavior either by the primary recognizing receptor Angiotensin-Converting Enzyme 2 (ACE2) and the cell-surface-GRP78 (CS-GRP78) (Elfiky and Ibrahim 2021a). Additionally, we reported the mutation at position 488 of the spike would affect its binding to ACE2 by breaking a salt bridge (E484-K31) found in the wildtype SARS-CoV-2 spike (Ibrahim et al. 2021).
Previous prediction studies reported the incorporation of the cell-surface receptor (CS-GRP78) in SARS-CoV-2 recognition and possibly facilitating its internalization into the human alveolar cells (Ibrahim et al. 2020; Elfiky 2020a). The predicted recognition site on the spike of SARS-CoV-2 lies in its RBD (C480–C488). This binding was also indicated for other viruses, including Zika, Ebola, Human papillomavirus, and the MERS-CoV (Elfiky 2020b, c; Elfiky and Ibrahim 2021b; Chu et al. 2018). Additionally, GRP78 was suggested as a possible link between COVID-19 and Mucormycosis (Elgohary et al. 2021). Recently an experimental study using Vero E6-ACE2 cells was conducted by Carlos et al. that confirmed the association of GRP78 with both SARS-CoV-2 and its primary receptor ACE2 (Carlos et al. 1978). They concluded that the spike protein's receptor-binding domain β (SBD β) is the docking platform for GRP78. Furthermore, they reported that GRP78 is important for the cell-surface localization of ACE2. Additionally, the humanized monoclonal antibody (hMAb159) reduced SARS-CoV-2 entry through the spike by decreasing Cs-GRP78 and Cs-ACE2, thus inhibiting SARS-CoV-2 infectivity in vitro (Carlos et al. 1978).
Computational predictions proved their crucial role in COVID-19 fighting (Mahmud et al. 2021; Sonousi et al. 2021; Gyebi et al. 2021; Wang 2020). In the current study, molecular dynamics simulation for the spike RBD of the delta variant was performed, followed by protein–protein docking to test the efficacy of the binding of the spike to both human cell-surface receptors GRP78 and ACE2.
Materials and Methods
Protein Data Bank (PDB) database was used to download the solved structures of spike receptor binding domain (RBD) (PDB ID: 6M17 chain E), ACE2 (PDB ID: 6M17 chain B), and GRP78 (PDB ID: 5E84, chain A). Two point mutations (E484Q and L452R) were administered in the spike RBD using the CHARMM-GUI webserver (Jo et al. 2008, 2014). RBD, ACE2, and GRP78 structures were prepared for docking using HADDOCK V2.4 webserver by removing unnecessary molecules such as water and other ligands (except for the oligosaccharides), while missing hydrogen atoms were added. Active sites for each protein were retrieved from literature, for GRP78: T428, V429, V432, T434, F451, S452, V457, and I459 (Yang et al. 2015), while for spike RBD: C480-C488 (against GRP78) and K417, Y453, Q474, F486, Q498, T500, and N501Y (against ACE2) and for ACE2: Q24, D30, H34, Y41, Q42, M82, K353, and R357 (Yan et al. 2020). The easy interface of HADDOCK V2.4 was used, and the remaining settings were set as default.
NAMD V2.13 was utilized to perform Molecular Dynamic Simulation (MDS). CHARMM-GUI web server was used to prepare the RBD and GRP78-RBD complex (produced from HADDOCK) necessary files for MDS (Jo et al. 2008; Phillips et al. 2005; Brooks et al. 2009; Lee et al. 2016). Both systems were solvated in the TIP3P water model, salt concentration was set to 0.154 M NaCl, and the temperature was set to 310 K. Time step was set to 2 fs, and the systems were minimized for 20,000, and 10,000 steps for RBD and GRP78-RBD complex, respectively. This is followed by an equilibration run for one ns in a constant number of atoms, constant pressure, and constant temperature (NPT ensemble) for both systems. The pressure was maintained at 1 atm using a Langevin piston, while the temperature was maintained at 310 K using Langevin dynamics. The systems were allowed to explore their conformational spaces for 100 ns in a constant number of atoms, constant volume, and constant temperature (NVT ensemble) for the production runs. After completing the MDS for the RBD, the equilibrated trajectories were clustered using TTClust V 4.8.3 python library (Tubiana et al. 2018). The number of clusters was determined automatically using the elbow method implemented in the TTClust library. Two clusters were obtained, and for each cluster, a representative frame was selected by the library. HADDOCK V 2.4 was used to dock each representative frame with the GRP78 solved structure using the previously mentioned active sites. Protein–Ligand Interaction Profiler (PLIP) was used to detect the number and types of interaction between the RBD and GRP78 for each S RBD-GRP78 complex produced (Salentin et al. 2015). The same protocol was pursued for S RBD (WT and delta)-ACE2 complex to test its binding affinities.
Results and Discussion
The mutated isoform of the SARS-CoV-2 spike (E484Q and L452R) is built and optimized, then subjected to a 100 ns MDS production run alongside the wildtype RBD to prepare the structures for the docking study. The dynamics are performed to explore the possible conformational space of the mutated spike prior to the docking study. The Root Mean Square Deviation (RMSD) in Å, the Radius of Gyration (RoG) in Å, and the Surface Accessible Surface Area (SASA) in Å2 are plotted in Fig. 1A and B. As reflected from the plots, the systems were equilibrated (RMSD is flattened) after the first 10 ns of the simulation, with an average value for the RMSD (blue line) of 2.66 ± 0.68 and 3.82 ± 0.54 Å for the wild type and delta RBD, respectively. The systems are equilibrated and stable, as reflected in the values of the RoG (orange line) and SASA (gray line). The average RoG is 17.9 ± 0.16 Å and 17.7 ± 0.12 Å, while SASA has average values of 11,092 ± 260 Å2 and 11,207 ± 295 Å2 for the wildtype and delta RBD, respectively.
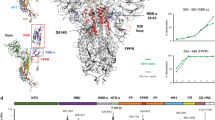
Molecular dynamics simulation of the SARS-CoV-2 spike RBD wild type (A) and delta variant (B). The Root Mean Square Deviation (RMSD) (blue line), Radius of Gyration (RoG) (orange line), and Surface Accessible Surface Area (SASA) (gray line) versus the simulation time. (C) the per-residue Root Mean Square Fluctuation (RMSF) for the WT (blue line) and Indian (delta) (orange line) variants of SARS-CoV-2 spike RBD. The highly fluctuating regions are marked by the colored rectangles on the RMSF curves and colored cartoons in the structure. The mutations L452R and E484Q are marked on the RMSF curve and by red sticks on the structure
After the MDS, TTClust is utilized to cluster the trajectories using the elbow methods, and we come up with five clusters representing the 1000 trajectories during the 100 ns MDS. We select a representative conformation from each cluster to test the GRP78 binding using HADDOCK 2.4. Tables 1 and 2 summarize the interactions established for each cluster representative docked with GRP78 along with the HADDOCK scores. The mutated residues R452 and Q484 are shown in red color in Table 2. The average HADDOCK score for the delta mutant isoform of the spike RBD is − 72.58 ± 9.5. This score is almost the same as the wildtype (WT) RBD docking score against GRP78 (− 74.3 ± 0.9) (Elfiky and Ibrahim 2021a). The two mutated residues (R452 and Q484) have an impact on the binding of the spike to GRP78, especially Q484. In the five different conformations, Q484 contributed at least one H-bond to the GRP78 RBDβ residues in four conformations. Bold residues in Tables 1 and 2 represent the active residues from both GRP78 and spike selected by HADDOCK to be flexible during searching for the best binding mode. In delta RBD, the most-reported residues from the spike to form H-bonds are N481 (8), S477 (4), and N487 (4), while the residues that form both H-bonds and hydrophobic interactions are F486 (10), Q484 (8), V483 (6), and Y489 (4). On the other hand, the wildtype RBD shows a slightly higher average number of H-bonds (8 ± 1.2) compared to the delta RBD (6.6 ± 1.6) but a slightly lower average number of hydrophobic contacts (3.3 ± 1.3) compared to the delta RBD (5.0 ± 1.1). The most reported residues that form interactions with GRP78 in the case of the wildtype RBD are T428 (5), V429 (5), G430 (4), T434 (4), Q449 (4), and T458 (4).
Figure 1C shows the per-residue Root Mean Square Fluctuations (RMSF) for the wildtype spike (WT) (blue line) and the delta variant spike (orange line) after 100 ns MDS runs. The structure of the spike RBD is shown in green cartoons. For the delta variant, the most fluctuating regions (RMSF < 3 Å) are depicted in different colors in the structure and marked on the RMSF curve as well. Two regions show high fluctuations, the yellow region (P384-F392) and the blue region (Q474–Q488), with RMSF reaching 4.3 and 5 Å, respectively. The C-terminal residue (orange sticks) also shows high RMSF (6.4 Å), while the N-terminal residue C336 (magenta sticks) is stabilized by the formed H-bond (dashed-yellow line) to G339. The dashed-green region marks the GRP78 recognition site (C480-C488) on the RMSF curve. This region fluctuates in the delta variant (RMSF of up to 5.00 Å) compared to the WT RBD (RMSF less than 2.26 Å). The mutation E484Q (red stick) also lies in this region, which may be the reason for the increased flexibility of this blue loop.
We reported in a previous study on the SARS-CoV-2 spike recognition site by host cell surface GRP78 (Ibrahim et al. 2020). This recognition site lies in the spike's receptor-binding domain (RBD), the same domain that binds human ACE2 (Elfiky 2020a; Elfiky et al. 2021a). New experimental work by Carlos et al. supported our prediction. At the same time, the cover artwork of the journal of biological chemistry for the July 2021 issue shows how the recognition occurs between GRP78 and the spike (Carlos et al. 2021). We run MDS for 100 ns for the complex formed between the GRP78 and the delta variant spike RBD in the current study. Figure 2A and B show the RMSD in Å, the RoG in Å, and the SASA in Å2 versus time in ns for the wildtype RBD-GRP78 complex (A) and delta RBD-GRP78 complex (B). As reflected from the plots, the systems are equilibrated at the middle of the simulation with an average value for the RMSD (blue line) of 6.3 Å and 9.2 Å for the wild type RBD-GRP78 and delta RBD-GRP78 complexes, respectively. The systems are equilibrated and stable as reflected also from the RoG (orange line) and SASA (gray line) values. The average RoG are 34.8 Å and 38.0 Å, while SASA has average values of 42,407 Å2and 42,000 Å2 for the wild type RBD-GRP78 and delta RBD-GRP78 complexes, respectively. The values of the RMSD, RoG, and SASA of the complexes (Fig. 2) are larger than that of the spike RBD alone (Fig. 1). This is due to complexity of the system in the case of GRP78-spike RBD compared to RBD alone.
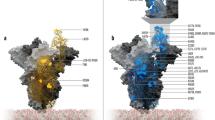
Molecular dynamics simulation of the RBD-GRP78 complexes. A and B The Root Mean Square Deviation (RMSD) (blue line), Radius of Gyration (RoG) (orange line), and Surface Accessible Surface Area (SASA) (gray line) versus the simulation time for the wild type RBD-GRP78 and delta RBD-GRP78 complexes. C The per-residue Root Mean Square Fluctuation (RMSF) for the WT, UK, and Indian (delta) variants of SARS-CoV-2 spike RBD -GRP78 complexes. The highly fluctuating regions are marked by the colored rectangles on the RMSF curves and colored cartoons in the structure. D The superposition of the solved structure of SARS-CoV-2 Spike (PDB ID: 6VYB) and the complex of RBD (delta)-GRP78 we modeled
Additionally, the RMSF (in Å) of the GRP78-spike delta RBD complex is depicted in Fig. 2C (gray line) alongside the wildtype (WT) and the beta (UK) strain RBD-GRP78 complexes (blue and orange lines, respectively). Five regions of the GRP78 (green cartoon) are found to be highly flexible in the delta spike RBD-GRP78 complex (RMSF < 5 Å). These include; F45-G58 (orange), F266-K326 (magenta), L480-I494 (yellow), D511-I522 (blue), and L561-S604 (red). These regions are declared on the RMSF curve with colored rectangles and depicted in the structures by colored cartoons. Noticeably, the yellow and blue regions (L480-I494 and D511-I522) are the loops that are involved in the interaction with the RBD of the spike. These two regions are more flexible in the UK and the Indian variants of the RBD-GRP78 complexes (orange and gray curves) compared to the WT RBD-GRP78 complex (blue curve). This increased flexibility may be a reason for the increased susceptibility of the delta RBD to be recognized by different host-cell receptors.
We superimposed the generated model of the RBD (delta) -GRP78 complex with the solved structure of the full-length spike of SARS-CoV-2 (PDB ID: 6VYB) (see Fig. 2D). GRP78 is shown in the magenta cartoon, while the superimposed RBDs are shown in the cyan cartoon. The spike homotrimer is shown in green, cyan, and magenta cartoons. This model represents the recognition of the virus by the host cell surface GRP78 (Ibrahim et al. 2020; Elfiky 2020a; Elfiky et al. 2021a). As reflected from the superposition, the two RBDs (solved structure and the interaction model with GRP78) coincide with each other except for the GRP78 recognition loop (C480–C488), which is missing in the solved structure. This loop is highly flexible and hence missing in the electron density map. The flexibility of this region is the highest in the delta RBD (gray) compared to the wildtype (blue) and UK variant (orange), as shown in the RMSF in Fig. 2C.
On the other hand, the recognition of ACE2 to SARS-CoV-2 Spike RBD delta is tested using the same protocol. Tables 3 and 4 show the detailed interactions established upon docking the ACE2 (PDB ID: 6M17 chain B) against the RBD WT (Table 3) and RBD delta (Table 4) utilizing the HADDOCK 2.4 web server. Bold residues indicate the active residues used to drive the docking in HADDOCK. At least 9 H-bonds and three hydrophobic contacts are established between the two proteins (GRP78 and the WT and delta RBDs), with a salt bridge formed in some conformations. The mutated residues in the delta strain found in the spike RBD (L452R and E484Q) do not contribute to the interaction with ACE2 except in one conformation in which Q484 is involved in H-bond formation, and R452 is involved in a salt bridge. The average docking score for the five conformations is − 106.52 ± 6.9, which is 18.4% higher than the docking score of the WT RBD to ACE2 (Ibrahim et al. 2021). This means that the binding affinity of the ACE2 to the delta variant of SARS-CoV-2 is about 18% lesser than its binding affinity against the WT SARS-CoV-2 spike.
Conclusively, the binding affinity of the delta strain spike RBD against the host cell receptors ACE2 and GRP78 is reduced in the former but maintained in the latter. This reflects the increased contribution of GRP78 in viral recognition in the delta RBD versus the wildtype RBD. This increased contribution of the GRP78 recognition was reported for the other variants of SARS-CoV-2 compared to the wildtype RBD (Elfiky and Ibrahim 2021a, 2022; Ibrahim et al. 2021). It appears that in the new variants, the virus increases its ability to recognize different host-cell receptors to increase its transmissibility. Therefore, we could combat the delta strain by targeting these receptors with inhibitors to reduce the probability of virus entry and vaccines that detect their binding sites on the viral spike (Elfiky et al. 2021b; Elfiky 2021; Elshemey et al. 2022).
Conclusion
SARS-CoV-2 delta strain is more contagious than the WT strain raising fear of the effectiveness of the current vaccination strategy. Furthermore, India reported the highest daily new infection due to this strain in May 2021. Therefore, it is essential to check for the entry mechanism of this strain, aiming to stop or decelerate the infection rate. The current study tested the potential of the main entry receptors, ACE2 and GRP78, in viral recognition. GRP78 shows the same binding affinity to RBD of the delta strain, while ACE2 affinity is slightly reduced. This congeals the effectiveness of using anti-ACE2 and anti-GRP78 as a possible route for viral fighting in the new starins of SARS-CoV-2.
Data Availability
Data is available upon request from the corresponding author.
References
Brooks BR, Brooks CL 3rd, Mackerell AD Jr, Nilsson L, Petrella RJ, Roux B et al (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30(10):1545–1614
Carlos AJ, Ha DP, Yeh D-W, Van Krieken R, Tseng C-C, Zhang P et al (1978) American Academy of Pediatrics. Committee on Environmental Hazards. PCBs Breast Milk. Pediatrics 62(3):407
Carlos AJ, Ha DP, Yeh D-W, Van Krieken R, Gill P, Machida K et al (2021) GRP78 binds SARS-CoV-2 spike protein and ACE2 and GRP78 depleting antibody blocks viral entry and infection in vitro. bioRxiv. https://doi.org/10.1101/2021.01.20.427368
Chu H, Chan CM, Zhang X, Wang Y, Yuan S, Zhou J et al (2018) Middle East respiratory syndrome coronavirus and bat coronavirus HKU9 both can utilize GRP78 for attachment onto host cells. J Biol Chem 293(30):11709–11726
Elfiky AA (2020a) SARS-CoV-2 spike-heat shock protein A5 (GRP78) recognition may be related to the immersed human coronaviruses. Front Pharmacol 11:577467
Elfiky AA (2020b) Ebola virus glycoprotein GP1-host cell-surface HSPA5 binding site prediction. Cell Stress Chaperones 25(3):541–548
Elfiky AA (2020c) Human papillomavirus E6: Host cell receptor, GRP78, binding site prediction. J Med Virol 92(12):3759–3765
Elfiky AA (2021) Natural products may interfere with SARS-CoV-2 attachment to the host cell. J Biomol Struct Dyn 39(9):3194–3203
Elfiky AA, Ibrahim IM (2021a) Host-cell recognition through GRP78 is enhanced in the new UK variant of SARS-CoV-2, in silico. J Infect 82(5):186–230
Elfiky AA, Ibrahim IM (2021b) Zika virus envelope—heat shock protein A5 (GRP78) binding site prediction. J Biomol Struct Dyn 39(14):5248–5260
Elfiky AA, Ibrahim IM (2022) Host-cell recognition through Cs-GRP78 is enhanced in the new Omicron variant of SARS-CoV-2, in silico structural point of view. J Infect 84(5):722–746
Elfiky AA, Ibrahim IM, Ismail AM, Elshemey WM (2021a) A possible role for GRP78 in cross vaccination against COVID-19. J Infect 82(2):282–327
Elfiky AA, Ibrahim IM, Amin FG, Ismail AM, Elshemey WM (2021b) COVID-19 and cell stress. In: Rezaei N (ed) Coronavirus disease—COVID-19. Springer, Cham, pp 169–178
Elgohary AM, Elfiky AA, Barakat K (2021) GRP78: a possible relationship of COVID-19 and the mucormycosis; in silico perspective. Comput Biol Med 139:104956
Elshemey WM, Elfiky AA, Ibrahim IM, Elgohary AM (2022) Interference of Chaga mushroom terpenoids with the attachment of SARS-CoV-2; in silico perspective. Comput Biol Med 145:105478
Gyebi GA, Elfiky AA, Ogunyemi OM, Ibrahim IM, Adegunloye AP, Adebayo JO et al (2021) Structure-based virtual screening suggests inhibitors of 3-chymotrypsin-like protease of SARS-CoV-2 from Vernonia amygdalina and Occinum gratissimum. Comput Biol Med 136:104671
Ibrahim IM, Abdelmalek DH, Elshahat ME, Elfiky AA (2020) COVID-19 spike-host cell receptor GRP78 binding site prediction. J Infect 80(5):554–562
Ibrahim IM, Elfiky AA, Elgohary AM (2021) Recognition through GRP78 is enhanced in the UK, South African, and Brazilian variants of SARS-CoV-2; an in silico perspective. Biochem Biophys Res Commun 562:89–93
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29(11):1859–1865
Jo S, Cheng X, Islam SM, Huang L, Rui H, Zhu A et al (2014) CHARMM-GUI PDB manipulator for advanced modeling and simulations of proteins containing nonstandard residues. Adv Protein Chem Struct Biol 96:235–265
Lee J, Cheng X, Swails JM, Yeom MS, Eastman PK, Lemkul JA et al (2016) CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12(1):405–413
Mahmud S, Elfiky AA, Amin A, Mohanto SC, Rahman E, Acharjee UK et al (2021) Targeting SARS-CoV-2 nonstructural protein 15 endoribonuclease: an in silico perspective. Future Virol 16(7):467–474
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E et al (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26(16):1781–1802
Salentin S, Schreiber S, Haupt VJ, Adasme MF, Schroeder M (2015) PLIP: fully automated protein–ligand interaction profiler. Nucleic Acids Res 43(W1):W443–W447
Sonousi A, Mahran HA, Ibrahim IM, Ibrahim MN, Elfiky AA, Elshemey WM (2021) Novel adenosine derivatives against SARS-CoV-2 RNA-dependent RNA polymerase: an in silico perspective. Pharmacol Rep 73(6):1754–1764
The effects of virus variants on COVID-19 vaccines: WHO (2021). https://www.who.int/news-room/feature-stories/detail/the-effects-of-virus-variants-on-covid-19-vaccines?gclid=Cj0KCQjw-LOEBhDCARIsABrC0TljpxT8YVr5pRKK0rRSFjMMEnW6MSGa7zz-htFxKobT0vLHUpsXfa8aAhYdEALw_wcB. Accessed 1 May 2021
Tubiana T, Carvaillo JC, Boulard Y, Bressanelli S (2018) TTClust: a versatile molecular simulation trajectory clustering program with graphical summaries. J Chem Inf Model 58(11):2178–2182
Wang J (2020) Fast identification of possible drug treatment of coronavirus disease-19 (COVID-19) through computational drug repurposing study. J Chem Inf Model 60(6):3277–3286
Yan R, Zhang Y, Li Y, Xia L, Guo Y, Zhou Q (2020) Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367(6485):1444–1448
Yang J, Nune M, Zong Y, Zhou L, Liu Q (2015) Close and allosteric opening of the polypeptide-binding site in a human Hsp70 chaperone BiP. Structure 23(12):2191–2203
Acknowledgements
Bibliotheca Alexandrina HPC in Alexandria, Egypt, is utilized to perform the MDS calculations.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). Cairo University funded the research through the COVID-19 fund received by Abdo Elfiky.
Author information
Authors and Affiliations
Contributions
AE own the research idea, wrote the manuscript, and draws figures. II performed the calculations. AE revised the manuscript and draw a conclusion. All the authors approve the final version of the document.
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare no competing interest in this work.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Ethical Approval
Not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Elfiky, A.A., Ibrahim, I.M. & Elgohary, A.M. SARS-CoV-2 Delta Variant is Recognized Through GRP78 Host-Cell Surface Receptor, In Silico Perspective. Int J Pept Res Ther 28, 146 (2022). https://doi.org/10.1007/s10989-022-10450-w
Accepted:
Published:
DOI: https://doi.org/10.1007/s10989-022-10450-w