Abstract
Two new somatostatin analogs with a characteristic part of the sequence -c(Cys-Phe-Trp-Lys-Thr-Cys)- and with two histidine and two aspartic acid moieties in their structures were synthesized and analyzed in terms of their coordination abilities with copper (II) ions. Both peptides bind Cu(II) effectively. Ligands form 4N complexes with \(\left\{ {{\text{N}}_{\text{Im}} ,{ 3} {\text{N}}_{\text{amide}}^{ - } } \right\}\) binding mode in a basic range of pH. But in spite of very similar sequences of the two peptides a significant difference in the effectiveness of the binding of copper (II) ions was observed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many previous studies show that since the discovery of somatostatin and its analogues interest in these peptides is still unabated. They are used in a new way of cancer treatment: peptide receptor radionuclide therapy (PRRT) which is a very promising method used to treat patients with neuroendocrine tumors. In PRRT, somatostatin analog is connected with radionuclides by using a chelator like DOTA or TETA and a linker (Teunissen et al. 2005).
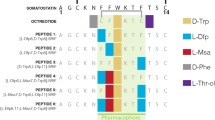
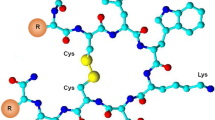
The subjects of the presented study are two somatostatin analogs with the part of the sequence: -c(Cys-Phe-Trp-Lys-Thr-Cys)- which is characteristic for the peptide hormone. Moreover, analyzed peptides have two histidine and two aspartic acid moieties in the peptide chains (Fig. 1). Four amino acids occurring between two cysteinyl moieties are responsible for biological activities and interactions with somatostatin receptors in this group of compounds (Pawlikowski and Melen-Mucha 2004). Furthermore, the presence of His and Asp amino acid residues, which are effective donors for metal ions, especially for copper (II) (Sovago et al. 2006, 2012) allows to create a binding site for a metal ion in the peptide structure. This way of the metal ion binding would allow to eliminate a linker and a chelator from precursors of radiopharmaceuticals potentially useful in PRRT (Fig. 2).
Our previous studies of linear somatostatin analogs show that analyzed ligands bind copper (II) ions effectively. But, unfortunately, cyclic complexes that were formed had structures not similar to natural hormone peptides with disulfide bridge between two cysteinyl moieties or the cyclic complex did not dominate in the physiological range of pH (Marchewka et al. 2012; Marciniak et al. 2012, 2014). Therefore, the analysis of cyclic ligands is the next step in searching of new somatostatin analogs.
Materials and Methods
Synthesis of the Peptides
Peptides were synthesized by the standard manual Fmoc solid-phase peptide synthesis method on the Fmoc-Rink Amide MBHA resin (0.65 mM/g, Iris Biotech GmbH). Synthesis was carried out in single-use plastic reactors (Intavis GmBH). Functional groups in the side chains of the amino acids used for the synthesis were protected as follows: Asp(OtBu), Cys(Acm), His(Trt), Lys(Boc), Thr(tBu), Trp(Boc). Subsequent Fmoc-protected amino acids (3 eq) were attached by using 3 eq PyBOP (1H-benzotriazol-1-yloxy)(tri-1-pyrrolidinyl)phosphonium hexafluorophosphate as a coupling reagent in the presence of N-hydroxybenzotriazole (3 eq) and diisopropylethylamine (6 eq) for 2 h at room temperature. Fmoc protecting groups were removed by 25 % piperidine in dimethylformamide. Acetylation of the N-terminal amino group was performed on the resin by 1:1 mixture of acetic anhydride and 0.4 M N-methylmorpholine in dimethylformamide. Final cleavage of the peptides was achieved by “Reagent K” (81.5 % trifluoroacetic acid, 5 % phenol, 5 % thioanisole, 5 % water, 2.5 % ethanedithiol, 1 % triisopropylsilane) in 2 h at room temperature. Crude peptides, with Cys residues still protected by Acm groups, were precipitated by cold diethylether, washed with ether, dissolved in water and lyophilized.
In order to remove Acm protecting groups from the side chains of Cys residues and to form a disulfide bridge, peptides were dissolved in 80 % acetic acid (2 mL of acetic acid for 2 mg of a peptide) and 20 eq of I2 dissolved in a small volume of 80 % acetic acid was added. The reaction was performed under nitrogen and its progress was monitored by ESI mass spectrometry until no more unoxidized peptide was observed. Mixture was diluted with the same amount of water and excess of I2 was extracted using CCl4. Water fraction containing peptide was lyophilized.
Peptides were purified by semipreparative HPLC using Varian ProStar apparatus equipped with TOSOH Bioscience C18 column (300 Å, 21.5 mm, 10 μm beads) and 220 nm UV detector. Water–acetonitrile gradients containing 0.1 % TFA at a flow rate of 7 ml/min. were used for the purifications. Final purity of the lyophilized peptides was >95 % by analytical HPLC (Thermo Separation Product; column: Vydac Protein RP C18, 250 Å, 4.6 mm, 5 μm; linear gradient 0–100 % B in 60 min., solvent A—0.1 % TFA in water, solvent B—0.1 % TFA in 80 % acetonitrile:water solution, UV detection at 220 nm). Chemical identity of the ligands was confirmed by ESI–MS on a Bruker micrOTOF-Q or Bruker apex ultra mass spectrometer. Analytical data of the synthesized peptides are given in Table 1.
Potentiometric Measurements
Potentiometric measurements were carried out using Molspin pH-meter system with Mettler Toledo InLab 422 semimicro combined electrode at 25 °C calibrated in hydrogen ion concentration using HCl (Irving et al. 1967). The ligands concentration was 8 × 10−4 mol/L and pH-metric titrations were performed in 0.1 mol/L KCl solution using sample volumes of 1.2 mL. Alkali (NaOH) was added by using a 0.25 ml micrometer syringe. The concentration of NaOH was 0.1 mol/L. Stability constants and stoichiometry of the complexes were calculated from titration curves using the SUPERQUAD program (Gans et al. 1985). The pH range where precipitation was observed was omitted during the calculations. Due to this fact some stability constants were only estimated, which is indicated in the “Results” section.
Spectroscopic Measurements
Visible spectra of complexes were recorded at 25 °C on Varian Carry 50 Bio spectrophotometer. The electron paramagnetic resonance (EPR) spectra were recorded on Bruker ELEXSYS E500 CW-EPR, X-Band spectrometer, equipped with ER 036TM NMR Teslameter and E 41 FC frequency counter. The EPR simulated spectra and all EPR parameters were obtained by the Biomolecular EPR Spectroscopy Software of Wilfred R. Hagen (Hagen 2009). Circular dichroism (CD) spectra were recorded on Jasco J-1500 magnetic circular dichroism spectrometer in 230–800 nm range. The same concentrations were used for both spectroscopic and potentiometric studies.
Fluorescence Measurements
Cary Eclipse fluorescence spectrophotometer was used for fluorescence measurements with excitations at 280 nm for both peptides. The concentrations for peptides solutions were: 6.6 × 10−5 mol/L for Ac[D1,9H2,10]c(SST) and 6.7 × 10−5 mol/L for Ac[H1,9D2,10]c(SST). Measurements were made at 25 °C. The titration was performed at pH 8.0 and 11.0 as a function of metal concentration. The measurements were done for pure ligands and for solutions with ligand to metal ratios in the range of 1:0.1–1:4.0.
Results and Discussion
Both studied peptides have five protonation constants assigned to two Asp, two His and one Lys amino acid residues (Tables 2, 4) and they are comparable to those found in the literature (Holm et al. 1996).
Analyzed somatostatin analog Ac[D1,9H2,10]c(SST) starts copper (II) ion binding around pH 4 and creates seven complexes (Fig. 3a). In the first one, CuH2L metal ion is probably coordinated by one nitrogen atom deriving from His residue. EPR parameters [A|| = 147 (cm−1) and g|| = 2.34, Table 3] confirm the existence of 1N complex. Moreover, the logβ* = 4.25 (where logβ* = logβ CuH2L − logβ H2L ) supports additional involvement of the carboxylate group from the Asp side chain. This value is different than logβ* for complexes where metal ion is coordinated only by imidazole nitrogen (Kapinos et al. 1998; Marciniak et al. 2014). Next form, CuHL, dominates in the system between pH 5.5 and 6 (Fig. 3a). The value of logβ* CuHL–HL = 6.03 may be connected with the involvement of the second imidazole donor in copper (II) binding (Brasuń et al. 2007). In this conditions CuL complex also exists in the system. The value of logβ = 8.78 (Table 2) may support involvement of the first amide nitrogen donor from the peptide chain (Fig. 3a). Unfortunately, above pH 5 precipitation was observed in the solution and it continued to pH 7.5. Molecules have a neutral charge there and this could be the reason of the precipitation (Hashempour et al. 2010). Therefore, determination of spectroscopic parameters for these two forms was not possible.
Next complex, CuH−1L, dominates in pH 7.5 (Fig. 3a). The stoichiometry of this form suggests the loss of 6 protons from the ligand molecule. In these conditions lysyl residue is still protonated, so it can be concluded that besides two Asp and two His residues, two amide nitrogen atoms are deprotonated. However, owing to the location of both imidazole donors in the peptide chain, together with deprotonation of two amide donors, the discussed complex may be characterized by \(\left\{ {{\text{N}}_{\text{Im}} ,{\text{ 2N}}_{\text{amide}}^{ - } } \right\}\) binding mode and thereby metal ion is coordinated here only by three nitrogen atoms. The second His residue did not bind metal ion. Spectroscopic results confirm the existence of 3N complex (Table 2). The experimental value of λ max = 582 nm is in a good agreement with the theoretical λ max = 584 nm calculated for two amides, one imidazole nitrogen and one oxygen from water molecule donor sets (Prenesti et al. 1999). Furthermore, the appearance of two positive bands on CD spectrum at 247 and 323 nm confirms the binding of copper (II) ion by both: imidazole and amide nitrogen atoms, respectively (Table 2). Value of logK = 7.18 obtained from potentiometric results also suggest the involvement of the amide nitrogen atom in the metal ion coordination (Table 2) (Kallay et al. 2009; Marciniak et al. 2014).
Around pH 7 CuH−2L complex appears in the system. The logK value of proton dissociation and formation of this complex is equal to 8.21 and supports binding of the next amide nitrogen and formation of the square planar complex with the \(\left\{ {{\text{N}}_{\text{Im}} ,{\text{ 3N}}^{ - }_{\text{amide}} } \right\}\) binding mode (Orfei et al. 2003). The EPR parameters obtained for the system at pH 9 [A|| = 199 (cm−1) and g|| = 2.194, Table 3] confirm coordination of four nitrogen atoms to the metal ion (Peisach and Blumberg 1974). Moreover, the components of g tensor (gz ≫ gy = gx > 2.0023) show the axial symmetry and the characteristic \({\text{A}}_{\text{II}}^{\text{Cu}} \, \gg \,{\text{A}}_{ \bot }^{\text{Cu}}\) splitting pattern suggests the dx2−y2 ground state (Godlewska et al. 2013; Pap et al. 2011). The comparison of the theoretical λmax = 523 nm (Prenesti et al. 1999) together with the experimental λmax = 525 nm (Table 2) in absorption spectrum also confirms \(\left\{ {{\text{N}}_{\text{Im}} ,{\text{ 3N}}^{ - }_{\text{amide}} } \right\}\) binding mode for this complex. Moreover, positive CT transition at 322 nm (Table 2) shows major involvement of amide nitrogen atoms in the metal ion binding.
Above pH 9 next two protons dissociate from the molecule and CuH−3L and CuH−4L complexes are formed (Fig. 3a). However, no changes in spectroscopic parameters are observed. The logK value of formation of CuH−3L complex (10.19, Table 2) suggests proton dissociation from Lys residues. The creation of the CuH−4L may be a result of the loss of a proton from the second nitrogen atom in the imidazole ring from the histidine moiety involved in copper ion coordination, what was observed in the case of other His-peptides (Brasuń et al. 2007).
The second somatostatin analog, Ac[H1,9D2,10]c(SST), has different position of His and Asp moieties in the peptide chain in comparison with the previously described peptide. It creates seven types of complexes with copper(II) ions (Fig. 3b). CuH3L coexists in the system with the next one CuH2L and with the uncoordinated metal ion so it was not possible to describe it using spectroscopic methods. However, analysis of the potentiometric results may give some suggestion. The value of logβ* = 3.01, where logβ* = logβ CuHnL − logβ HnL , suggests the {NIm} binding mode (Kapinos et al. 1998; Marciniak et al. 2014). Moreover, the logK = 4.45, for the reaction: CuH3L → CuH2L is comparable to the protonation constants for deprotonation of one carboxylate group from Asp residues (logK = 4.53) without binding to the metal ion (Table 4).
LogK = 5.24 for the reaction of CuHL formation suggests proton dissociation from histidine moiety (Table 4). Stoichiometry of the described form indicates the loss of protons from two His residues. Therefore, {2NIm} coordination mode is possible here. The value of logβ* = logK CuHL–logβ HL = 6.55, which is similar to logβ* for the other complexes with the same coordination mode in systems described in the literature (Brasuń et al. 2007; Kotynia et al. 2014) confirms this type of copper (II) binding.
Similarly to Ac[D1,9H2,10]c(SST), when CuL complex with neutral charge exists in the system, precipitation was observed. Therefore, determination of spectroscopic parameters for CuL was not possible. Only on the basis of the stoichiometry of the described complex it can be concluded that the first amide nitrogen is involved in the metal ion coordination.
While pH increases, next three complexes: CuH−1L, CuH−2L; and CuH−3L appear in the system (Fig. 3b). Coordination modes for these forms are the same as in case of Ac[D1,9H2,10]c(SST) analog, what is confirmed by potentiometric and spectroscopic parameters (Tables 4, 5).
It is difficult to define which histidine moieties are involved in the coordination of copper (II) ion in the particular complexes. In case of 4 N complexes with {NIm, 3N−} coordination modes two options for each of the peptides are possible (Fig. 4). However, in view of the creation of (6,5,5)-chelate rings in case of copper (II) ion coordination by histidine moieties on C-terminal of peptides chains, this way of binding seems to be energetically more favorable than the second one where (7,5,5)-chelate rings are created (Sovago et al. 2012). Therefore, coordination by His10 in Ac[D1,9H2,10]c(SST) and His9 in Ac[H1,9D2,10]c(SST) is much more probable.
Tryptophan is responsible for the phenomenon of fluorescence in both analyzed peptides (Lakowicz 2006). In the manner of the metal ion binding for CuH−2L complexes proposed above no atoms derived from this amino acid residue are involved in the coordination of Cu(II). Therefore, study of the fluorescence quenching in analyzed systems could be an indication that described binding manners are possible. Analysis in pH 8.0 and 11.0 show that in both peptides quenching of fluorescence is very similar (Fig. 5). In the analyzed conditions in both systems the same types of complexes with the same coordination modes are formed. In all cases almost 100 % of fluorescence is quenched. It means that tryptophan residues are located on the surface of the complexes and are not protected from the quencher by the rest of the molecule. It can be concluded that fragments of the peptide chains with tryptophan residues have similar spatial arrangement in both ligands. Therefore, the proposed coordination modes with involvement of histidine moieties on C-terminal are possible.
Fluorescence quenching was also analyzed by Stern–Volmer equation (Lakowicz 2006) (Fig. 6). Obtained plots indicate mixed mechanism of quenching: static and dynamic.
Figure 7 shows the comparison of the efficiency in copper (II) ion binding between the two analyzed peptides. Both ligands have a very similar construction but Ac[H1,9D2,10]c(SST) coordinates copper (II) ion much better than the second one in the whole analyzed range of pH. It is probably related to the earlier (in lower pH) coordination of amide nitrogen atoms in case of Ac[H1,9D2,10]c(SST)/Cu2+ system (first amide nitrogen coordinates to copper (II) ions in CuL complexes in both systems). Moreover, the ligand with histidine moiety in the first position coordinates the metal ion by amide nitrogen from Cys8 in CuH−1L complex. This is the first amino acid which is involved in the coordination and it occurres within the cyclic structure. In case of Ac[D1,9H2,10]c(SST) similar situation is observed later in the CuH−2L complex. It can also affect the difference in the effectiveness in copper (II) ion coordination.
Presented results demonstrated that small modifications within the peptide sequence can significantly affect efficiency of the metal ion binding.
Conclusion
In this paper we have shown that new somatostatin analogs with the characteristic part of the sequence -c(Cys-Phe-Trp-Lys-Thr-Cys)- and with two histidine and two aspartic acid moieties in their structures bind copper(II) ions effectively. Both ligands have very similar construction but Ac[H1,9D2,10]c(SST) coordinates copper (II) ion much better than the second one in the whole analyzed range of pH. It was shown that small modifications within the peptide sequence can significantly affect efficiency of the metal ion binding. These results may be an introduction to the discovery of new precursors of pharmaceuticals useful in the PRRT.
References
Brasuń L, Matera A, Ołdziej S, Światek-Kozłowska J, Messori L, Gabbiani C, Orfei M, Ginanneschi M (2007) The copper(II) coordination abilities of three novel cyclic tetrapeptides with -His-Xaa-His- motif. J Inorg Biochem 101:452–460
Gans P, Sabatini A, Vacca A (1985) SUPERQUAD: an improved general program for computation of formation constants from potentiometric data. J Chem Soc Dalton Trans 6:1195–1200
Godlewska S, Jezierska J, Baranowska K, Augustin E, Dołęga A (2013) Copper(II) complexes with substituted imidazole and chloride ligands: x-ray, UV–Vis, magnetic and EPR studies and chemotherapeutic potential. Polyhedron 65:288–297
Hagen WR (2009) Biomolecular EPR Spectrocopy. CRC Press, Boca Raton
Hashempour M, Razavizadeh H, Rezaie HR, Hashempour M, Ardestani M (2010) Chemical mechanism of precipitate formation and pH effect on the morphology and thermochemical co-precipitation of W-Cu nanocomposite powders. Mater Chem Phys 123:83–90
Holm RH, Kennepohl P, Solomon EI (1996) Structural and functional aspects of metal sites in biology. Chem Rev 96:2239–2314
Irving HM, Miles MG, Pettitt LD (1967) A study of some problems in determining the stoichiometric proton dissociation constants of complexes by potentiometric titrations using a glass electrode. Anal Chim Acta 38:475–488
Kallay C, Varnagy K, Malandrinos G, Hadjiliadis N, Sanna D, Sovago I (2009) Thermodynamic and structural characterization of the macrochelates formed in the reactions of copper(II) and zinc(II) ions with peptides of histidine. Inorg Chim Acta 362:935–945
Kapinos LE, Song B, Siegel H (1998) Metal ion-coordinating properties of imidazole and derivatives in aqueous solution: interrelation between complex stability and ligand basicity. Inorg Chim Acta 280:50–56
Kotynia A, Czyżnikowska Ż, Bielińska S, Szyrwiel Ł, Kamysz W, Malinka W, Brasuń J (2014) The impact of two –GlyProGly– motifs on formation of di-copper complexes by His4-cyclopeptides. New J Chem 38:5198–5206
Lakowicz JR (2006) Principles of fluorescence spectroscopy, 3rd edn. Springer, New York
Marchewka A, Czyżnikowska Ż, Cebrat M, Brasuń J (2012) The structural aspects of the copper(II) binding by the His-analogue of somatostatin. Polyhedron 42:236–242
Marciniak A, Cebrat M, Czyżnikowska Ż, Brasuń J (2012) Novel short-chain analogues of somatostatin as ligand for Cu(II) ions. Role of the metal ion binding on the spatial structure of the ligand. J Inorg Biochem 117:10–17
Marciniak A, Czyżnikowska Ż, Cebrat M, Kotynia A, Brasuń J (2014) Structural aspects of copper(II) binding by a Multi-His analogue of somatostatin. Inorg Chim Acta 416:57–62
Orfei M, Alcaro MC, Marcon G, Chelli M, Ginnaneschi M, Kozlowski H, Brasun J, Messori L (2003) Modeling of copper(II) sites in proteins based on histidyl and glycyl residues. J Inorg Biochem 97:299–307
Pap JS, Kripli B, Bányai V, Giorgi M, Korecz L, Gajda T, Árus D, Kaizer J, Speier G (2011) Tetra-, penta- and hexacoordinate copper(II) complexes with N3 donor isoindoline-based ligands: characterization and SOD-like activity. Inorg Chim Acta 376:158–169
Pawlikowski M, Mełeń-Mucha G (2004) Somatostatin analogs – from New molecules to new applications. Curr Opin Pharmacol 4:608–613
Peisach J, Blumberg WE (1974) Structural implications derived from the analysis of electron paramagnetic resonance spectra of natural and artificial copper proteins. Arch Biochem Biophys 165:691–708
Prenesti E, Daniele PG, Prencipe M, Ostacoli G (1999) Spectrum-structure correlation for visible absorption spectra of copper(II) complexes in aqueous solution. Polyhedron 18:3233–3241
Sovago I, Osz K (2006) Metal ion selectivity of oligopeptides. Dalton Trans 32:3841–3854
Sovago I, Kallay C, Varnagy K (2012) Peptides as complexing agents: factors influencing the structure and thermodynamic stability of peptide complexes. Coord Chem Rev 256:2225–2233
Teunissen JJM, Kwekkeboom DJ, de Jong M, Esser JP, Valkema R, Krenning EP (2005) Peptide receptor radionuclide therapy. Best Pract Res Clin Gastroenterol 19:595–616
Acknowledgments
The presented studies were financially supported by Wroclaw Medical University (ST-705).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Aleksandra Marciniak, Marek Cebrat, and Justyna Brasuń declare that they have no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Marciniak, A., Cebrat, M. & Brasuń, J. The Coordination Abilities of New Cyclic Analogs of Somatostatin. Int J Pept Res Ther 23, 135–143 (2017). https://doi.org/10.1007/s10989-016-9546-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-016-9546-4