Abstract
Context
Habitat connectivity can stabilise animal populations by facilitating immigration and genetic exchange, but it increases the risk of infectious diseases being spread by hosts. Chytridiomycosis caused by Batrachochytrium salamandrivorans (Bsal) threatens European salamander diversity. The extent to which the connectivity of populations of fire salamanders (Salamandra salamandra) contributes to the spread of Bsal remains unclear.
Objectives
We analysed the impact of habitat connectivity of fire salamanders on the spread of Bsal. Moreover, we show how local salamander abundance is associated with habitat connectivity over a five-year period.
Methods
We developed fire salamander habitat suitability models (HSMs) for the Eifel area (Germany), currently considered the core of the range of Bsal in Europe. Habitat models were used to calculate pairwise resistance between salamander occurrences to test whether Bsal presence and salamander abundance were associated with habitat connectivity.
Results
Fire salamanders are widely distributed in the Eifel. Solid bedrock and topographic positioning were important predictors of stream suitability as breeding habitats, while deciduous forests and grassland cover determined overall fire salamander habitat suitability along with breeding habitat suitability. Bsal-positive salamander occurrences were better-connected than Bsal-negative or untested occurrences. Nevertheless, fire salamander larvae were more abundant in well-connected sites.
Conclusion
The connection of salamander populations by suitable habitat seems to support local salamander abundance while facilitating the spread of Bsal. In situ conservation measures counteracting host species connectivity to interrupt Bsal transmission pathways must be implemented with caution, as they may weaken the demographic advantages of connectivity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amphibians are among the most threatened taxonomic groups, with approximately one third of the known species threatened with extinction (Stuart et al. 2004; Bellard et al. 2016; IUCN 2020). Emerging fungal diseases are among the main drivers of amphibian declines and extinctions worldwide (Wake and Vredenburg 2008). In particular, the fungal pathogen Batrachochytrium dendrobatidis (Bd), causing chytridiomycosis in frogs, has negative impacts on anuran populations globally (Skerratt et al. 2007). The related and invasive B. salamandrivorans (Bsal) causes chytridiomycosis mainly in newts and salamanders (Urodela) and can be seen as a severe threat to European salamander diversity (Martel et al. 2013; Spitzen-van der Sluijs et al. 2016; Yap et al. 2017), as infections are fatal in many European species (Martel et al. 2014).
Bsal was first detected in the Netherlands and Belgium, where it led to serious declines in fire salamander populations in the wild (Martel et al. 2013; Spitzen-van der Sluijs et al. 2013; Stegen et al. 2017). Since then, Bsal has spread quickly throughout Europe and is continuously detected in new sites, especially in Germany but also in Spain (e.g., Martel et al. 2020; Schmeller et al. 2020; Thein et al. 2020; Lötters et al. 2020a). An important characteristic for the spread of Bsal may be its ability to build encysted spores that can potentially persist in environmental reservoirs (Stegen et al. 2017; Mosher et al. 2018). The spores are probably transported by natural and anthropogenic vectors and can stick to amphibian skin or the feet of waterfowl (Stegen et al. 2017).
Monitoring and epidemiological modelling of infected fire salamander populations suggest that Bsal can induce local extinction within one season even at low host density (Schmidt et al. 2017; Stegen et al. 2017). Therefore, few mitigation options are available once Bsal has emerged (Stegen et al. 2017; Canessa et al. 2018; Thomas et al. 2019) and a strict implementation of biosecurity protocols in the field and in captivity to reduce human-mediated dispersal of spores is vital (EFSA et al. 2018; Scheele et al. 2019). To develop in situ measures that interrupt pathogen spread (e.g. by fencing) or mitigate disease severeness (e.g. by habitat management), a comprehensive understanding of the host-pathogen-environment interaction triangle is necessary. In frog species affected by Bd, pathogenicity can vary depending on environmental factors, such as water temperature and salinity (Heard et al. 2015, 2018); thus, habitat management addressing those factors may help to provide refugia from severe chytridiomycosis (Heard et al. 2018; Scheele et al. 2019). In such cases, infected amphibian populations can be stabilised by connectivity to environmental refugia, e.g. when local extinctions due to Bsal outbreaks are mediated by recolonization from Bsal free refugia habitats that harbour salamander source populations. However, refugia that reduce pathogenicity have not yet been reported in urodeles across the range of the invasive Bsal strain in Europe (Beukema et al. 2020).
It is difficult to assess the role of host population connectivity in spreading infectious diseases and for metapopulation persistence. In his models, Hess (1996) stressed an increased extinction risk of well-connected host populations affected by disease. However, the demographic advantages of immigration and recolonization are likely to outweigh the risk of the spread of diseases in most systems (McCallum and Dobson 2002; Heard et al. 2015). In the Bsal-fire salamander system, little is known about the role of salamanders as Bsal vectors, and empirical, spatiotemporal data on disease and host metapopulation dynamics are missing (Schmidt et al. 2017). A concentric, continuous spread mediated by fire salamanders can be expected in homogenous habitats with constant host densities (Vredenburg et al. 2010; Schmidt et al. 2017), but those conditions are unrealistic in the landscape matrix of forest fragments, grasslands, intensively used croplands and settlements typically found in Central Europe. For instance, in the Netherlands, an uninfected subpopulation of fire salamanders has been discovered at a distance of only 800 m from the Bsal index site, with no obvious barriers to amphibian migration present (Martel et al. 2013; Spitzen-van der Sluijs et al. 2018). Although the study of single populations on local scales is important to understand how Bsal spreads, these findings might not represent the role of host population connectivity on Bsal distribution at the landscape level (Canessa et al. 2018).
Germany can be considered the current hot spot of the invasive range of Bsal in Europe, and most records lie within the Eifel region (Lötters et al. 2020a). In the northern Eifel, Bsal was first detected in 2015 (Dalbeck et al. 2018), but reports on fire salamander mass mortality in the region predate more than ten years (Lötters et al. 2020b). Despite the presence of Bsal in the region, fire salamanders are (still) widely distributed and inhabit largely forested areas as well as smaller woodland fragments.
Here, we analysed the influence of fire salamander habitat connectivity on (i) the presence of Bsal and (ii) the local fire salamander abundance over a five-year period in the northern Eifel region. As anthropogenic land use can interrupt functional connectivity between geographically close populations (Marsh and Trenham 2001; Nowakowski et al. 2013), we used circuit theory (McRae 2006) to estimate the pairwise landscape resistance between fire salamander occurrences. Furthermore, we analysed the influence of habitat connectivity on local salamander abundance derived from standardized larval monitoring to assess whether local populations still benefit from connectivity at the current stage of pathogen invasion. We hypothesized that Bsal will occur more frequently in well-connected fire salamander populations, as local populations are more likely to come into contact with infected conspecifics. However, we also hypothesized that fire salamander abundance is higher in large and unfragmented habitats. This study helps to inform conservation actions targeting salamander habitat connectivity at the core of the known invasive range of Bsal in Europe.
Methods
Study system
The study took place in the northern part of the Eifel, a low mountain range spanning over the German federal states North Rhine-Westphalia and Rhineland-Palatinate as well as eastern Belgium. In this landscape, extended forests and a high density of near natural creeks that serve as potential sites for the deposition of salamander larvae and their development until metamorphosis (hereafter “breeding habitat”) are characteristic features. A landscape segment of approximately 30 × 40 km was chosen as the study area as it covers the most intensively sampled parts of the northern Eifel since 2014 (Lötters et al. 2020a). Here, yearly precipitation and mean annual temperature range from 1280 mm to 6.5 °C in the most elevated (~ 693 m) areas to 664 mm and 10.24 °C in the lowest parts (115 m) of the study area (Fick and Hijmans 2017).
Salamander presence, abundance and Bsal screening
The data on Bsal distribution as well as fire salamander abundance and distribution used in this study come from three independent sampling activities, which were conducted with different levels of standardization.
-
(i)
Bsal screening: We used a subset of 39 Bsal screening sites reported by Lötters et al. (2020a) that lay within our study area (for details, see Lötters et al. 2020a). Sampled individuals were assigned to a central point per valley (pers. commun. Dalbeck), which resulted in a relatively coarse resolution of Bsal occurrences (minimum distance between screening area centroids: 562 m). Bsal screening survey effort was neither recorded nor standardized.
-
(ii)
Larvae removal sampling: To assess the local abundance of salamanders, standardized removal sampling of salamander larvae was conducted between 2015 and 2019. Larvae were searched for 135 min per year and site in a stream segment of 75 m and captured individuals were kept in a small plastic cup until the end of the sampling process to avoid double counts (for details, see Wagner et al. 2020a). Because not all sites in the study area were continuously sampled, we used a subset of 30 sites that were visited in at least three years between 2015 and 2019 (SI1).
-
(iii)
Larvae presence-only surveys: We surveyed streams across the whole study area for the presence of fire salamander larvae to assess the overall (remaining) distribution of the species. These streams predominantly included first- and second-order streams in or close to forest areas, where predatory fish are unlikely to occur (Thiesmeier 2004). A torch (flashlight) was used to break water surface reflections to visually detect larvae, and potential shelters like stones and leaf litter in the water were systematically checked. Searches during rainfall were avoided to maximise the detectability of larvae. Each creek or creek segment survey was conducted once or twice in 2018 and/or 2019, with a maximum sampling effort of 30 min per person and year (total number of surveys = 246). Survey localities were recorded using a Garmin GPSMAP 64 S device. During all sampling activities, a strict hygiene protocol, including the disinfection of all material that came into contact with amphibians or their habitat, was followed (van Rooij et al. 2017).
Habitat suitability models
As sampling effort of salamander larvae surveys was not constant and the surveyed localities were complemented by opportunistic larvae records spanning from 2014 to 2019, including 30 removal sampling sites and larvae localities provided by the “Biologische Station im Kreis Düren”, fire salamander absence could not be confirmed. Therefore, a presence-only method, the maximum entropy species distribution modelling (MaxEnt), was used to map suitable salamander habitats (Phillips et al. 2006). In comparative studies, MaxEnt is ranked among the best performing presence-only methods (e.g., Giovanelli et al. 2010) since it can handle comparatively small sets of species occurrence data (Wisz et al. 2008; Aguirre-Gutiérrez et al. 2013). Presence data were contrasted to the environmental background sampled at 10,000 randomly distributed points, and raw MaxEnt outputs were interpreted as a relative measurement of habitat suitability (Merow et al. 2013).
Species’ environmental requirements can change according to life stages or seasonality, leading to spatially or temporally different habitat uses (Mateo Sánchez et al. 2014). Modelling frameworks can thus be split into multiple models that use different scales and predictors (Bani et al. 2015; Frans et al. 2018; Goudarzi et al. 2021). Therefore, fire salamander HSMs were split into (i) a spatially restricted breeding habitat and (ii) a whole life cycle habitat model, following the framework of Bani et al. (2015). The breeding habitat model covered a 30 m buffer along the digitised streams. Most streams surveyed were already available as digitised linear features (vector data) in the digital landscape model “DLM50” (status: 2019, see Geobasis NRW 2019a for details), but another 50 first-order creeks that were not included were manually digitised using satellite imagery and water flow accumulation. Following the recommendation of Phillips et al. (2009), the extent of the breeding model was restricted to streams with a maximum distance of 50 m to forest fragments with a minimum size of 1 ha, as larvae presence surveys were based on the same criteria. Background sampling of the whole life cycle models was not spatially restricted since most of the study area (82%) lies within a reasonable postnatal dispersal distance of 500 m to potential breeding habitats (Klewen 1985; Schulte et al. 2007; Merow et al. 2013).
Processing of fire salamander occurrence data
In presence-only habitat suitability models, it is vital to account for sampling bias (Dudík et al. 2005; Kramer-Schadt et al. 2013; Yackulic et al. 2013). Among others, spatial autocorrelation can effectively be reduced by filtering occurrence data with a minimum distance or a grid for systematic sampling (Veloz 2009; Syfert et al. 2013; Fourcade et al. 2014). Therefore, we thinned fire salamander larvae localities (n = 250) in the R environment (R Core Team 2019), using the package spThin (Aiello-Lammens et al. 2015). The algorithm retains the maximum number of observations by randomly deleting points within a user-defined buffer. We used a buffer of 500 m to delete double observations at the same stream from different years. Additionally, Bsal occurrences were considered separated on a similar scale, but postnatal salamander dispersal can still be expected over such distances (Klewen 1985; Schulte et al. 2007; Hendrix et al. 2017). This resulted in a spatially balanced set of 154 points.
Candidate predictors—breeding habitat
Choosing biologically meaningful predictors is vital in HSMs (e.g., Phillips et al. 2006; Guisan et al. 2017; Fourcade et al. 2018). As the breeding model extent was already restricted based on land cover criteria, we exclusively included topographic and geological predictors describing microclimate, stream size and structure. Fire salamanders prefer to deposit their larvae in small creeks close to the spring, which can be measured using stream order, depth or width (Thiesmeier 2004; Manenti et al. 2009; Bani et al. 2015). We used water flow accumulation instead, which additionally accounts for the risk of downstream drift after heavy rainfall (Thiesmeier 2004; Bani et al. 2015). We used bedrock type as an approximation of the structural characteristics of the stream. Due to the unavailability of fine scaled empirical climate data, we used the topographic positioning index (TPI) and elevation as proxy for the local microclimate, which is a limiting factor in urodele core habitats (Wagner et al. 2020b; Goudarzi et al. 2021). Elevation, water flow accumulation and TPI were derived from a digital elevation model (status: 2019, 1 m resolution, see Geobasis NRW 2019b for details). We categorised bedrock using geological maps “IS GK 100 DS” (status: 2016, available as vector data at a scale of 1:100,000, see Geobasis NRW 2019c for details) based on its structural characteristics: loose geological layers, such as sand, gravel and silt, and solid bedrocks. Environmental data were processed to raster format with a cell size of 5 × 5 m using ArcMapv.10.7 (ESRI 2019).
Candidate predictors—whole life cycle habitat
In the whole life cycle model, we included the mean of the modelled breeding habitat suitability in a radius of 500 m as a measure of potentially available breeding habitats for adult salamanders. As topography was already included in the breeding model, whole life cycle models were built using land cover predictors only. We categorised land cover based on the “LBM-DE2018” landscape model (available as vector data, processed to 5 × 5 m raster, for details see BKG 2019) into grassland, settlement, lentic waterbodies (predominantly storage reservoirs) and coniferous, mixed and deciduous forest. To reduce model complexity, we did not include cropland because it barely occurs within potential salamander corridors in the chosen study area. Roads were available as linear elements (vector data, Geobasis NRW 2019a). Therefore, we added a buffer of 5 m around them, categorised them as settlements and processed them to a 5 × 5 m raster. Borders of land cover types were smoothed by calculating the mean proportional cover in a radius of 500 m to account for the spatial ecology of adult salamanders that deposited the recorded larvae (Schulte et al. 2007; Manenti et al. 2009; Bani et al. 2015). However, as potential barriers to fire salamander migration and dispersal are relevant on smaller scales, we calculated settlement and waterbody cover in a radius of 30 m.
Modelling process and predictor selection
To account for multicollinearity, predictors with correlation coefficients r ≥ 0.8 and VIF ≥ 5 need to be excluded (Pradhan 2016, see SI2). Therefore, we removed the proportion of sandy and loose bedrock in the breeding model. For the whole life cycle model, forest types were strongly correlated. We excluded coniferous forest as mixed and deciduous forests are the preferred terrestrial habitats of fire salamanders in central Europe (Thiesmeyer 2004). For model evaluation, MaxEnt v. 3.4.0. was called from the R environment using the ENMeval package v. 1.0 (Muscarella et al. 2014). Occurrence data were separated into a training and a test dataset for cross-validation using randomised 10-folded splitting. For background sampling, the same random points were used for the training and testing data following Merow et al. (2013). In both modelling steps, we used and compared various transformations of the predictors, so-called feature types, including linear[L], quadratic[Q], hinge[H] and product[P] features. Moreover, regulation multiplier (RM) values, which are used to control model overfitting, gradually increased (see SI3 for all candidate models). Candidate models were ranked by the Akaike Information Criterion corrected for small sample sizes (AICc) (Warren and Seifert 2011; Katz and Zellmer 2018).
Because breeding habitat suitability was included as a predictor in the whole life cycle models, only one of two candidate breeding habitat models with ΔAICc ≤ 2 was chosen based on the biological plausibility of the underlying response curves (SI4). In contrast, we selected two different scenarios of habitat suitability throughout the life cycle to be used in the subsequent resistance modelling. These models should represent the best fitting (low AICc) but also biologically distinct scenarios. The latter was based on niche overlap in geographic space (Schoener’s D), which measures the deviation between each raster cell of the habitat suitability predictions. Raw MaxEnt outputs of the two selected whole life cycle models were projected on the whole study area and used without further transformations or definition of absolute barriers as conductance surfaces in resistance models.
Resistance modelling
Habitat connectivity was assessed by calculating the pairwise resistances between 167 fire salamander occurrences in Circuitscape v. 4.0 (McRae et al. 2013). Compared to other approaches, such as the least cost path algorithm, circuit theory reflects diffuse dispersal, similar to random walk theory in ecology (McRae and Beier 2007). Due to the high uncertainty about possible pathways, such an approach seems more suitable to model disease transmission by hosts (Nobert et al. 2016). To reduce computing time, suitability maps were upscaled to 20 × 20 m resolution by averaging. This resulted in a resistance surface with 5,519,281 grid cells in total. We added a buffer of random resistance spanning over 20% of the maximum width of the study area to account for edge effects (Koen et al. 2010, 2014). We used random resistances as the buffer exceeded the political border of Germany, beyond which environmental data was not available in the same format and resolution. Salamander occurrences defined as nodes included the centroids of Bsal screening areas (tested nodes = 39) and those larval localities used to build the HSMs (nodes not tested for Bsal, n = 128) that had no screening centroid within a radius of 500 m.
Habitat connectivity and Bsal presence
As our independent variables (pairwise resistances) referred to pairs of sites while our dependent variables (Bsal presence, larvae abundance) referred to single sites, we derived three node-based measures of connectivity for each site (cf. Koen et al. 2016) to predict the presence of Bsal: (1) the Euclidean distance and (2) the resistance to a single neighbouring (geographically closest) Bsal outbreak site (DBsal and RBsal), (3) the summed up resistance to (a) all (n = 166) and the neighbouring (b) 32, (c) 16 and (d) 8 tested and untested salamander occurrences (RSum, Fig. 1). We included untested localities in the latter variable to measure the sites general connectivity within the salamander metapopulation, as the resource intensive Bsal screening was only conducted in 39 out of 167 available salamander localities (see Fig. 2). Additionally, we tested for potential anthropogenic vectors by including the predicted number of recreational visitors per year and ha (Schägner et al. 2016) and the local density of trails in a radius of 500 m (provided by Beukema et al. 2021).
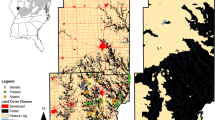
Node-based parameters of salamander habitat connectivity derived from pairwise resistances. For RBsal, the resistance to the geographically closest known Bsal outbreak site was extracted. For RSum, the pairwise resistances to all (n = 166) and to the geographically closest 8, 16 and 32 neighbours were summed up as a measure of (local) connectivity within the salamander metapopulation. All resistances were modelled twice based on two different habitat suitability models used as conductance surface (L1 and LQH3, see Fig. 2)
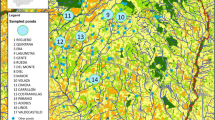
Current density based on the LQH3 (left) and L1 (right) whole life cycle habitat suitability models used as conductance surfaces in circuit theory-based connectivity analyses. Circles (39) indicate Bsal screening area centroids (see Lötters et al. 2020a). Triangles (128) indicate larvae localities without Bsal screening centroids within a radius of 500 m. Squares include larvae removal sampling sites (30). Note that 15 out of 30 larvae removal sampling sites were screened for Bsal as well
To assess the parameter importance for Bsal presence, we first built Generalized Linear Models (GLMs) using the glmulti package (Calcagno and de Mazancourt 2010). Models were fitted using maximum likelihood estimation since our primary goal was to derive importance across multiple parameter combinations. As the variables RSum and RBsal (Fig. 1) were highly correlated (see SI5), each node-based resistance parameter was combined with trail and visitor density in a separate call (n = 11). In the next step, the averaged effect across all candidate models per call (n = 64) was calculated using the coef.glmulti function (Burnham and Anderson 2002; Johnson and Omland 2004). We did not include random effects to account for potential non-independence of neighbouring sites since no spatial autocorrelation in the models’ residuals was detected using Moran’s I test (see SI5). As we could not define Bsal absence for most of the known salamander localities, the above-described modelling steps were then applied to a second subset of Bsal-tested sites only. However, no interactions could be modelled for this subset due to the reduced sampling size of 39 sites (potential overparameterization). All predictors were normalized with an orderNorm transformation using the bestNormalize package (Peterson and Peterson 2020) prior to the analysis.
Habitat connectivity and salamander abundance
The rounded mean number of larvae per monitoring site (n = 30) was used as a measure of local salamander abundance. Because of overdispersion in the count data, we built univariate negative binomial regression models (nbGLMs) to assess the effect of RBsal and RSum on local salamander abundance using the MASS package. RBsal was set to “0” for monitoring sites that were also screened for Bsal (15 out of 30) and found to be positive (n = 4). Models were ranked by AICc to identify the best predictor. Again, no spatial autocorrelation was detected in the model residuals.
Results
Salamander and Bsal presence
Fire salamanders were still widely distributed in forest areas of the northern Eifel (Fig. 2), as we found larvae in 169 of our 246 larvae presence-only surveys in 2018 and 2019. No or few larvae were found in elevated areas dominated by coniferous forest, such as the upper “Weiße Wehe” valley (northern central region, Fig. 2) and the “Erkensruhr” valley (southwestern region). Additionally, no larvae were found in most streams sampled in the “Sauerbach” valley (southern central region). Here, Bsal was detected on newts in a pond system at the stream source as well as on adult salamanders downstream. In total, 17 Bsal-positive sites that met the criteria of recent fire salamander records within a 500 m radius were used as nodes in the following resistance models.
Breeding habitat suitability
The two top-ranked breeding habitat models with ΔAICc ≤ 2 (i.e., L0.5 [linear features, regulation multiplier = 0.5] and P0.5 [product features]) barely differed in their predictive ability or level of overparameterization according to AUC (0.794 and 0.799), average difference in AUC of models built with test data (0.021, Var: 0.01 and 0.024, Var: 0.01, respectively) or omission rate (both 0.007, Var: <0.001; see SI3 for all candidate models). Therefore, we relied on the response curves of the models (SI4) for final model selection. We selected the L0.5 (linear feature only, regulation multiplier = 0.5) breeding habitat model since no response to the proportion of solid bedrock was visible in the P0.5 model. In the selected L0.5 model, breeding habitat suitability decreased with increasing flow accumulation, elevation and TPI, while an increasing amount of solid bedrock led to increased predicted suitability values. TPI was by far the most important variable in terms of percent contribution (49.1%) and permutation importance (45.3%). Nevertheless, solid bedrock, water flow accumulation and elevation contributed considerably to the predicted suitability (Table 1). In particular, permuting water flow accumulation values worsened the predictive performance, indicating the high importance of this environmental parameter for salamander breeding habitats.
Whole life cycle habitat suitability
For the whole salamander life cycle, the top-ranked model (LQH3: linear, quadratic and hinge features, regulation multiplier = 3) was more complex than the selected breeding model (see SI3 for all candidate models). The predictive ability was good (AUC = 0.846, average or MTP = 0.013 (Var: 0.001)), and the AUC values of models built with test localities slightly dropped to 0.832 (Var: 0.004) on average, indicating a moderate level of overparameterization. The mean breeding habitat suitability and deciduous and mixed forest cover were by far the most important variables, as they contributed to more than 80% of the model variance (Table 2). As a second, biologically distinct (ΔSchoeners D = 0.735, see SI3) scenario, we selected a model using an RM-value of 1 and linear features only (AUC = 0.808, average or MTP = 0.006 (Var: <0.001), average ΔAUCtest = 0.017 (Var. 0.007)).
Compared to the top-ranked model, grassland cover replaced breeding habitat suitability as the most influential predictor. Waterbody and settlement cover, potentially interrupting salamander functional connectivity, were of minor importance in both models. The underlying response curves (SI4) of the two selected whole life cycle models were similar except for deciduous and mixed forest cover. In the LQH3 model, the underlying curve was hump-shaped, and the predicted suitability value peaked at approximately 50% cover of deciduous and mixed forest in a radius of 500 m. Regardless, areas completely covered by this forest type were more suitable than areas with different land cover. In the L1 model, the predicted suitability values monotonically increased with increasing forest cover. In both selected whole life cycle models, increasing grassland, settlement and waterbody cover led to a reduction, while mean breeding habitat suitability led to an increase in predicted habitat suitability values.
Habitat connectivity and Bsal occurrence
The resistance to the neighbouring Bsal outbreak site (RBsal) was the most important predictor for Bsal presence and sites that had higher RBsal were less likely to be Bsal positive in both datasets, including all (Table 3) and tested sites only (Table 4). Likewise, the general connectivity within the salamander metapopulation measured by RSum was higher in Bsal-positive sites. Euclidean distance to the neighbouring Bsal outbreak site (DBsal) as well as visitor and trail density were of little importance compared to RBsal and RSum, irrespective of the scaling (number of neighbours) or resistance surfaces (L or LQH) used (see SI6). However, the interaction of trail density and the summed-up resistance to the closest 16 neighbouring salamander occurrences (RSum16 LQH) was of considerable importance (ranked 4th ). Interactions between visitor density and node-based connectivity measures were less important [DBsal (km): Visitors was ranked 14th ]. Overall parameter importance was moderate, especially when Bsal-tested sites were used exclusively (maximum importance = 0.59, see Table 4) and no interactions could be modelled due to the reduced sampling size.
Habitat connectivity and salamander abundance
The mean number of larvae strongly differed between the monitoring sites (0.3 to 138.3, see SI1) and was best explained by the resistance to the neighbouring 32 salamander occurrences (RSum32 LQH, Table 5). Larvae abundance increased with local habitat connectivity, as fewer larvae were found in sites with high summed-up resistances (Fig. 3). Likewise, the resistance to the neighbouring Bsal site was lower in sites with many larvae, even though models using RBsal as predictor showed worse predictive ability according to AICc and the models’ standard errors.
Projected effect of the resistance to the geographically closest know Bsal outbreak site (RBsal, left) and the sum of resistances to 32 neighbouring salamander occurrences (RSum, right) on the rounded mean salamander larvae abundance between 2015 and 2019 in 30 monitoring sites using univariate nbGLMs.
Discussion
Emerging infectious diseases can spread through animal populations in multiple ways, but direct transmission by host migration and dispersal is often assumed to be the most relevant way of infecting individuals (White et al. 2000; Altizer et al. 2011; Schmidt et al. 2017). Among all European tailed amphibians, the European fire salamander (Salamandra salamandra) has been the species impacted most by the spread of Bsal. Adult salamanders are highly susceptible, and infections are lethal in most cases. Population crashes through Bsal infection have repeatedly been documented, for instance, in the Netherlands (Spitzen-van der Sluijs et al. 2013) or the Ruhr district in Germany (Schulz et al. 2020). Most studies dealing with Bsal infection dynamics assume salamander individual-based disease transmission (Schmidt et al. 2017; Canessa et al. 2018), and migrating and dispersing infected salamanders likely represent an important natural vector. However, there is poor evidence of the extent of their contribution to the distributional pattern of Bsal observed at the landscape level. In this study, Bsal occurrence was positively associated with fire salamander habitat connectivity, which indicates that Bsal spread via salamander movement seems to be an important factor in large, unfragmented habitats (cf. Beukema et al. 2021). However, salamander larvae, and therefore also adults, were more abundant in well-connected areas.
Habitat suitability models
Due to Bsal-induced salamander absences, the equilibrium state assumption of the HSMs used as conductance surfaces might be violated (cf. Sandvoß et al. 2020). However, salamander larvae were found in most surveyed habitats in the northern Eifel. Breeding habitat models were built along digitised streams in forest areas only, since breeding in small, stagnant water bodies is of minor importance for Eifel populations compared to lowland populations, and it has thus far not been reported for Eifel populations (Dalbeck and Thiesmeier 2011; Hendrix et al. 2017). Puddles and pools close to streams might be used occasionally, but larvae were almost exclusively found in flowing waters. Small-scale data on the structural and biotic characteristics of streams, such as watercourse heterogeneity, stream depth and width, number of pools, substrate, macrozoobenthos communities or periphyton, were not available for the whole study area, which could improve breeding habitat model precision (Manenti et al. 2009; Wagner et al. 2020b). Instead, water flow accumulation, solid bedrock, elevation and topographic positioning were found to be suitable alternatives to predict the streams’ topographic and structural suitability.
In their terrestrial habitat, fire salamanders are mostly found in deciduous and mixed forests, while plantations of coniferous trees, such as spruce (Picea abies), are largely avoided (Joly 1968; Thiesmeier 2004). Overall, this expectation was fulfilled, but in the top-ranked whole life cycle model (LQH3), areas dominated by deciduous and mixed forest were predicted to be (moderately) less suitable than areas with intermediate deciduous and mixed forest cover. This might be due to the typical position of streams at the border of two different land cover types, and consequently, many larvae were located at forest edges. To account for this artefact, a second, biologically distinct whole life cycle model (L1) was used, predicting a monotonous relation to deciduous and mixed forest cover, which seems biologically more accurate (Joly 1968; Thiesmeier 2004). Nevertheless, validation with spatially independent test data is needed (Veloz 2009) to project these models to landscapes where salamanders inhabit other habitat types, such as railway embankments (Thiesmeier 2004).
Habitat connectivity and Bsal occurrence
In our connectivity models, we assumed salamanders to migrate preferentially along areas of high projected habitat suitability, as suggested by Bani et al. (2015) and Goudarzi et al. (2021). However, microclimatic factors influencing the rate of water loss can be relevant for salamander migration within suitable habitats (Peterman et al. 2014; Nowakowski et al. 2015). Salamanders might use suboptimal habitats for compensatory movement (Peterman et al. 2014) and, compared to occurrence, effective dispersal can be driven by divergent landscape characteristics (Cushman et al. 2013). Notwithstanding, we assumed that our nodes-based resistances derived from HSMs are best suited to explain the spread of Bsal, as these measure the continuous availability of salamanders for Bsal (cf. Fig. 3). This assumption is also supported by the minor importance of the geographical distance to neighbouring Bsal sites (DBsal) in our models, highlighting that resistance to salamander movement rather than pure distance to Bsal occurrence drives the distribution of Bsal. Moreover, unfragmented habitats of high quality supported higher salamander abundances, which should accelerate the spread of Bsal as well. In contrast to other studies (e.g. Beukema et al. 2021), we did not determine Bsal absence. The detectability of chytrids depends on seasonality, the local demographic structure of hosts, and human activities (Adams et al. 2010; Mosher et al. 2018; Bozzuto and Canessa 2019; Beukema et al. 2021), but most importantly, it depends on the number of animals tested (DiGiacomo and Koepsell 1986). The latter was biased in Bsal-positive sites in our study region (see Suppl. 3 in Lötters et al. 2020a) and by omitting untested sites (128 out of 167), large parts of the known salamander distribution would be ignored. Therefore, we modelled both tested and untested sites, but still found Bsal presence to be best explained by the resistance to neighbouring Bsal outbreak sites (RBsal), followed by the sites’ connectivity within the local metapopulation (RSum) in both datasets.
Human activities can obscure natural patterns of pathogen spread (Greer and Collins 2008; Adams et al. 2010; Padgett-Flohr and Hopkins 2010; Cohen et al. 2016). For instance, fragmentation and road density may facilitate Bsal spread, e.g., via touristic activities in the Eifel (Allan et al. 2003; Jousimo et al. 2014; Paquette et al. 2014; Beukema et al. 2021). Despite that, trail and visitor density were poor predictors for Bsal presence in our study, indicating that anthropogenic vectors play a minor role for the spread of Bsal at least on local scales. Our findings thereby emphasize the potential of chytrids to afflict amphibians especially in pristine, unfragmented habitats (Lötters et al. 2009; Becker and Zamudio 2011).
As Bsal occurrence is not restricted to the range of fire salamanders in Central Europe, many other pathways for Bsal dispersal must exist (Stegen et al. 2017; Canessa et al. 2018; Dalbeck et al. 2018; Beukema et al. 2021). Nearly all native amphibian species can come into contact with fire salamanders (Dalbeck et al. 2007; Dalbeck and Thiesmeier 2011), and in the northern Eifel, Bsal was found on the skin of smooth, palmate, alpine and crested newts (Lissotriton vulgaris, L. helveticus, Ichthyosaura alpestris and Triturus cristatus) in proximity and distant from fire salamander habitats (Lötters et al. 2020a). Infected newts suffer less from infection and may migrate longer distances compared to fire salamanders (Kovar et al. 2009), which could make them important Bsal vectors as well (Canessa et al. 2018; Spitzen-van der Sluijs et al. 2018; Beninde et al. 2021). As the modelled salamander habitat suitability partly entails suitable newt habitats, salamander connectivity likely overlaps with newt connectivity. Moreover, anurans, waterfowl or crayfish can be reservoirs and vectors for chytrid spores (McMahon et al. 2013; Stegen et al. 2017). Environmental transmission, e.g. by spores floating in streams, may have contributed to the pattern found in this study (Stegen et al. 2017; Mosher et al. 2018), but former studies indicate that biotic factors primarily drive the distribution of pathogens (Cohen et al. 2016; Spitzen-van der Sluijs et al. 2018).
We acknowledge that due to our multistep modelling framework, prediction biases may be multiplied in the final GLM’s (compound error), especially since we combined results of survey activities that yield differently resolved data on Bsal and salamander larvae occurrence. However, larvae localities included in our habitat and connectivity models represented one of several possible sets of points, as larvae could typically be found over a few hundred meters of the streams. This also counteracts potential bias due to the upscaling of resistance data, as bigger pixels still included streams with larval presence. Additionally, as we modelled occurrence, our models should be less sensitive to uncertainties and biases in the predictors of the first modelling step compared to quantitative analysis like modelling Bsal prevalence. Even though we selected the two most distinct (Schoender’D) habitat suitability models (SI3) as resistances surfaces, predictors showed consistent qualitative responses (SI4). Therefore, major compound errors leading to misidentified key parameters for Bsal occurrence seem unlikely. Still, the final estimates of regression parameters likely have higher standard errors than indicated in our tables due to our stepwise framework.
Habitat connectivity and salamander abundance
Despite Bsal presence being associated with well-connected salamander habitats, these habitats generally supported higher salamander abundance. Large populations that typically occupy extensive, unfragmented habitat patches are more resilient and less likely to become extinct because local extinctions can be buffered by immigration (Hanski and Simberloff 1997; Marsh and Trenham 2001; Jousimo et al. 2014; Heard et al. 2015). Nevertheless, drastic declines in salamander abundance have been observed at sites in the study area and elsewhere (Spitzen-van der Sluijs et al. 2013; Dalbeck et al. 2018), and slightly fewer larvae were found at Bsal outbreak sites in the Eifel (Wagner et al. 2020a). Fire salamander populations may be able to persist at low densities despite Bsal presence in large, unfragmented forest areas where recolonisation is facilitated (cf. Schmidt et al. 2017). Additionally, Bsal primarily affects reproductive adults (Stegen et al. 2017), and fire salamanders in Germany mostly start to reproduce in their 5th year at the earliest (Thiesmeier 2004). Therefore, facilitated population recovery due to increased connectivity can be time-lagged.
Implication for mitigation efforts
As eradication attempts of Bsal in the northern Eifel or comparable regions are logistically unfeasible and likely to fail (Bosch et al. 2015), in situ conservation efforts need to focus on population and habitat management that support the resilience of afflicted salamanders and newts (Heard et al. 2018; Scheele et al. 2019). In situ conservation strategies, such as fencing, include measures that counteract salamander connectivity (Thomas et al. 2019), so it is highly important to gauge their demographic consequences. Here, it has been demonstrated that isolation attempts seeking to interrupt Bsal transmission mediated by amphibians could potentially counteract demographic advantages of connectivity (McCallum and Dobson 2002; Heard et al. 2015). However, long-term demographic consequences are difficult to predict, and overall advantages of salamander habitat connectivity can be expected only if Bsal-induced mortality appears in a patchy pattern that is outweighed by increased recruitment and recolonisation from neighbouring areas (cf. Heard et al. 2015; Heard et al. 2018).
Climatic or other environmental gradients, such as water salinity or temperatures that exceeds the pathogen’s physiological niche or optimum, can contribute to natural refugia from chytrids that harbour source populations (Puschendorf et al. 2009; Doddington et al. 2013; Heard et al. 2015; Urban et al. 2015; Nowakowski et al. 2016; Hettyey et al. 2019), but no such refugia habitats from Bsal have been detected thus far (Beukema et al. 2020). Future research must identify mismatches between salamander and Bsal niches that can be exploited by feasible in situ conservation measures to amplify the resilience of salamander metapopulations in regions with extended and well-connected habitats, such as the northern Eifel (Campbell Grant et al. 2018; Scheele et al. 2019). This requires a comprehensive and spatiotemporally explicit research approach that assesses local salamander population dynamics and Bsal occupancy depending on reservoir host abundance, landscape connectivity, physiochemical parameters and microbiota communities (Campbell Grant et al. 2018; Bozzuto and Canessa 2019).
Data availability
Salamander occurrences, geological, land cover and topographical data and the associated R code used to build habitat suitability models and GLMs are available at https://doi.org/10.5281/zenodo.7759814.
References
Adams MJ, Chelgren ND, Reinitz D, Cole RA, Rachowicz LJ, Galvan S, McCreary B, Pearl CA, Bailey LL, Bettaso J, Bull EL, Leu M (2010) Using occupancy models to understand the distribution of an amphibian pathogen, Batrachochytrium dendrobatidis. Ecol Appl 20(1):289–302
Aguirre-Gutiérrez J, Carvalheiro LG, Polce C, van Loon EE, Raes N, Reemer M, Biesmeijer JC (2013) Fit-for-purpose: species distribution model performance depends on evaluation criteria—dutch hoverflies as a case study. PLoS ONE 8(5):e63708
Aiello-Lammens ME, Boria RA, Radosavljevic A, Vilela B, Anderson RP (2015) spThin: an R package for spatial thinning of species occurrence records for use in ecological niche models. Ecography 38(5):541–545
Allan BF, Keesing F, Ostfeld R (2003) Effect of forest fragmentation on lyme disease risk. Conserv Biol 17(1):267–272
Altizer S, Bartel R, Han BA (2011) Animal migration and infectious disease risk. Science 331(6015):296–302
Bani L, Pisa G, Luppi M, Spilotros G, Fabbri E, Randi E, Orioli V (2015) Ecological connectivity assessment in a strongly structured fire salamander (Salamandra salamandra) population. Ecol Evol 5(16):3472–3485
Becker CG, Zamudio KR (2011) Tropical amphibian populations experience higher disease risk in natural habitats. PNAS 108(24):9893–9898
Bellard C, Genovesi P, Jeschke JM (2016) Global patterns in threats to vertebrates by biological invasions. Proc Biol Sci. https://doi.org/10.1098/rspb.2015.2454
Beninde J, Keltsch F, Veith M, Hochkirch A, Wagner N (2021) Connectivity of Alpine newt populations (Ichthyosaura alpestris) exacerbates the risk of Batrachochytrium salamandrivorans outbreaks in european fire salamanders (Salamandra salamandra). Conserv Genet 22(4):653–659
Beukema W, Erens J, Schulz V, Stegen G, Spitzen-van der Sluijs A, Stark T, Laudelout A, Kinet T, Kirschey T, Poulain M, Miaud C, Steinfartz S, Martel A, Pasmans F (2021) Landscape epidemiology of Batrachochytrium salamandrivorans: reconciling data limitations and conservation urgency. Ecol Appl 31(5):e02342
Beukema W, Pasmans F, van Praet S, Ferri-Yáñez F, Kelly M, Laking AE, Erens J, Speybroeck J, Verheyen K, Lens L, Martel A (2020) Microclimate limits thermal behaviour favourable to disease control in a nocturnal amphibian. Ecol Lett 24(1):27–37
BKG - Bundesamt für Kartographie und Geodäsie (2019) Digitales Landbedeckungsmodell für Deutschland, Stand 2018 (LBM-DE2018). Data available at https://gdz.bkg.bund.de/index.php/default/digitales-landbedeckungsmodell-fur-deutschland-stand-2018-lbm-de2018.html
Bosch J, Sanchez-Tomé E, Fernández-Loras A, Oliver JA, Fisher MC, Garner TWJ (2015) Successful elimination of a lethal wildlife infectious disease in nature. Biol Lett. https://doi.org/10.1098/rsbl.2015.0874
Bozzuto C, Canessa S (2019) Impact of seasonal cycles on host-pathogen dynamics and disease mitigation for Batrachochytrium salamandrivorans. Glob Ecol Conserv 17:e00551
Burnham KP, Anderson DR (2002) Model selection and multimodel inference. Springer, New York
Calcagno V, de Mazancourt C (2010) Glmulti: an R package for easy automated model selection with (generalized) linear models. J Stat Softw 34:1–29
Campbell Grant EH, Adams MJ, Fisher RN, Grear DA, Halstead BJ, Hossack BR, Muths E, Richgels KLD, Russell RE, Smalling KL, Waddle JH, Walls SC, LeAnn White C (2018) Identifying management-relevant research priorities for responding to disease-associated amphibian declines. Glob Ecol Conserv 16:e00441
Canessa S, Bozzuto C, Campbell Grant EH, Cruickshank SS, Fisher MC, Koella JC, Lötters S, Martel A, Pasmans F, Scheele BC, Spitzen-van der Sluijs A, Steinfartz S, Schmidt BR, McCallum H (2018) Decision-making for mitigating wildlife diseases: from theory to practice for an emerging fungal pathogen of amphibians. J Appl Ecol 55(4):1987–1996
Cohen JM, Civitello DJ, Brace AJ, Feichtinger EM, Ortega CN, Richardson JC, Sauer EL, Liu X, Rohr JR (2016) Spatial scale modulates the strength of ecological processes driving disease distributions. PNAS 113(24):e3359–e3364
Cushman SA, McRae B, Adriaensen F, Beier P, Shirley M, Zeller K (2013) Biological corridors and connectivity [Chapter 21]. In: Macdonald DW, Willis KJ (eds) Key topics in conservation biology 2. Wiley-Blackwell, Hoboken, pp 384–404
Dalbeck L, Düssel-Siebert H, Kerres A, Kirs K, Koch A, Lötters S, Ohlhof D, Sabino-Pinto J, Preißler K, Schulte U, Schulz V, Steinfartz S, Veith M, Vences M, Wagner N, Wegge J (2018) Die Salamanderpest und ihr Erreger Batrachochytrium salamandrivorans (Bsal): aktueller stand in Deutschland. Z Feldherpetol 25(1):1–22
Dalbeck L, Thiesmeier B (eds) (2011) Feuersalamander—Salamandra salamandra. In: Hachtel M, Bußmann M (eds) Handbuch der Amphibien und Reptilien Nordrhein-Westfalens, 1st edn. Z Feldherpetol - Suppl 16/1. Laurenti, Bielefeld
Dalbeck L, Lüscher B, Ohlhoff D (2007) Beaver ponds as habitat of amphibian communities in a central european highland. Amphibia-Reptilia 28(4):493–501
DiGiacomo RF, Koepsell TD (1986) Sampling for detection of infection or disease in animal populations. JAVMA 189(1):22–23
Doddington BJ, Bosch J, Oliver JA, Grassly NC, Garcia G, Schmidt BR, Garner TWJ, Fisher MC (2013) Context-dependent amphibian host population response to an invading pathogen. Ecology 94(8):1795–1804
Dudík M, Phillips SJ, Schapire RE (2005) Correcting sample selection bias in maximum entropy density estimation. NIPS 18:323–330
ESRI (2019) ArcGis desktop: version 10.7. Environmental Systems Research Institute, Redlands
Fick SE, Hijmans RJ (2017) WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int J Climatol 37(12):4302–4315
Fourcade Y, Besnard AG, Secondi J (2018) Paintings predict the distribution of species, or the challenge of selecting environmental predictors and evaluation statistics. Glob Ecol Biogeogr 27(2):245–256
Fourcade Y, Engler JO, Rödder D, Secondi J (2014) Mapping species distributions with MAXENT using a geographically biased sample of presence data: a performance assessment of methods for correcting sampling bias. PLoS ONE 9(5):e97122
Frans VF, Augé AA, Edelhoff H, Erasmi S, Balkenhol N, Engler JO, McPherson J (2018) Quantifying apart what belongs together: a multi-state species distribution modelling framework for species using distinct habitats. Methods Ecol Evol 9(1):98–108
Geobasis NRW (2019a) Digitales Landschaftsmodell 50 NW (DLM50). Data downloaded from https://www.geoportal.nrw/opendatadownloadclient. Accessed 20 Aug 2019
Geobasis NRW (2019b) Digitales Geländemodell - Gitterweite 1 m (XYZ). Data downloaded from https://www.geoportal.nrw/opendatadownloadclient. Accessed 22 Aug 2019
Geobasis NRW (2019c) IS GK 100 DS - Informationssystem Geologische Karte von Nordrhein-Westfalen 1:100.000. Data downloaded from https://www.geoportal.nrw/opendatadownloadclient. Accessed 27 Aug 2019
Giovanelli JGR, de Siqueira MF, Haddad CFB, Alexandrino J (2010) Modeling a spatially restricted distribution in the Neotropics: how the size of calibration area affects the performance of five presence-only methods. Ecol Model 221(2):215–224
Goudarzi F, Hemami M-R, Malekian M, Fakheran S, Martínez-Freiría F (2021) Species versus within-species niches: a multi-modelling approach to assess range size of a spring-dwelling amphibian. Sci Rep 11(1):597
Greer AL, Collins JP (2008) Habitat fragmentation as a result of biotic and abiotic factors controls pathogen transmission throughout a host population. J Anim Ecol 77(2):364–369
Guisan A, Thuiller W, Zimmermann NE (2017) Habitat suitability and distribution models: with applications in R. Ecology, biodiversity and conservation. Cambridge University Press, Cambridge
Hanski I, Simberloff D (1997) The metapopulation approach: its history, conceptual domain, and application to conservation. In: Hanski I, Gilpin ME (eds) Metapopulation biology: ecology, genetics, and evolution. Academic Press, San Diego, pp 5–25
Heard GW, Scroggie MP, Ramsey DSL, Clemann N, Hodgson JA, Thomas CD (2018) Can habitat management mitigate disease impacts on threatened amphibians? Conserv Lett 11(2):e12375
Heard GW, Thomas CD, Hodgson JA, Scroggie MP, Ramsey DSL, Clemann N (2015) Refugia and connectivity sustain amphibian metapopulations afflicted by disease. Ecol Lett 18(8):853–863
Hendrix R, Schmidt BR, Schaub M, Krause ET, Steinfartz S (2017) Differentiation of movement behaviour in an adaptively diverging salamander population. Mol Ecol 26(22):6400–6413
Hess G (1996) Disease in metapopulation models: implications for conservation. Ecology 77(5):1617–1632
Hettyey A, Ujszegi J, Herczeg D, Holly D, Vörös J, Schmidt BR, Bosch J (2019) Mitigating disease impacts in amphibian populations: capitalizing on the thermal optimum mismatch between a pathogen and its host. Front Ecol Evol 7:356
IUCN (2020) The IUCN red list of threatened species. https://www.iucnredlist.org/. Accessed 9 Feb 2020)
Johnson JB, Omland KS (2004) Model selection in ecology and evolution. Trends Ecol Evol 19:101–108
Joly J (1968) Données écologiques sur la salamandre tachetée, Salamandra salamandra. Ann Sci Nat Zool 12:301–366
Jousimo J, Tack AJM, Ovaskainen O, Mononen T, Susi H, Tollenaere C, Laine A-L (2014) Disease ecology. Ecological and evolutionary effects of fragmentation on infectious disease dynamics. Science 344(6189):1289–1293
Katz TS, Zellmer AJ (2018) Comparison of model selection technique performance in predicting the spread of newly invasive species: a case study with Batrachochytrium salamandrivorans. Biol Invasions 20(8):2107–2119
Klewen R (1985) Untersuchungen zur Ökologie und Populationsbiologie des Feuersalamanders (Salamandra salamandra terrestris) an einer isolierten Population im Kreis Paderborn. Abh Landesmus Naturk Münster 47:1–51
Koen EL, Garroway CJ, Wilson PJ, Bowman J (2010) The effect of map boundary on estimates of landscape resistance to animal movement. PLoS ONE 5(7):e11785
Koen EL, Bowman J, Sadowski C, Walpole AA (2014) Landscape connectivity for wildlife: development and validation of multispecies linkage maps. Methods Ecol Evol 5(7):626–633
Koen EL, Bowmann J, Wilson PJ (2016) Node-based measures of connectivity in genetic networks. Mol Ecol Resour 16:69–79
Kovar R, Brabec M, Bocek R, Vita R (2009) Spring migration distances of some central European amphibian species. Amphibia-Reptilia 30(3):367–378
Kramer-Schadt S, Niedballa J, Pilgrim JD, Schröder B, Lindenborn J, Reinfelder V, Stillfried M, Heckmann I, Scharf AK, Augeri DM, Cheyne SM, Hearn AJ, Ross J, Macdonald DW, Mathai J, Eaton J, Marshall AJ, Semiadi G, Rustam R, Bernard H, Alfred R, Samejima H, Duckworth JW, Breitenmoser-Wuersten C, Belant JL, Hofer H, Wilting A, Robertson M (2013) The importance of correcting for sampling bias in MaxEnt species distribution models. Divers Distrib 19(11):1366–1379
Lötters S, Kielgast J, Bielby J, Schmidtlein S, Bosch J, Veith M, Walker SF, Fisher MC, Rödder D (2009) The link between rapid enigmatic amphibian decline and the globally emerging chytrid fungus. EcoHealth 6(3):358–372
Lötters S, Wagner N, Albaladejo G, Böning P, Dalbeck L, Düssel H, Feldmeier S, Guschal M, Kirst K, Ohlhoff D, Preißler K, Reinhardt T, Schlüpmann M, Schulte U, Schulz V, Steinfartz S, Twietmeyer S, Veith M, Vences M, Wegge J (2020) The amphibian pathogen Batrachochytrium salamandrivorans in the hotspot of its european invasive range: past—present—future. Salamandra 56(3):173–188
Lötters S, Veith M, Wagner N, Martel A, Pasmans F (2020b) Bsal-driven salamander mortality pre-dates the european index outbreak. Salamandra 56(3):239–242
Manenti R, Ficetola GF, Bernardi FD (2009) Water, stream morphology and landscape: complex habitat determinants for the fire salamander Salamandra salamandra. Amphibia-Reptilia 30:7–15
Marsh DM, Trenham PC (2001) Metapopulation dynamics and amphibian conservation. Conserv Biol 15(1):40–49
Martel A, Blooi M, Adriaensen C, van Rooij P, Beukema W, Fisher MC, Farrer RA, Schmidt BR, Tobler U, Goka K, Lips KR, Muletz C, Zamudio KR, Bosch J, Lötters S, Wombwell E, Garner TWJ, Cunningham AA, van der Sluijs A, Salvidio S, Ducatelle R, Nishikawa K, Nguyen TT, Kolby JE, van Bocxlaer I, Bossuyt F, Pasmans F (2014) Recent introduction of a chytrid fungus endangers Western Palearctic salamanders. Science 346(6209):630–631
Martel A, Spitzen-van der Sluijs A, Blooi M, Bert W, Ducatelle R, Fisher MC, Woeltjes A, Bosman W, Chiers K, Bossuyt F, Pasmans F (2013) Batrachochytrium salamandrivorans sp. nov. causes lethal chytridiomycosis in amphibians. PNAS 110(38):15325–15329
Martel A, Vila-Escale M, Fernández‐Giberteau D, Martinez‐Silvestre A, Canessa S, van Praet S, Pannon P, Chiers K, Ferran A, Kelly M, Picart M, Piulats D, Li Z, Pagone V, Pérez‐Sorribes L, Molina C, Tarragó‐Guarro A, Velarde‐Nieto R, Carbonell F, Obon E, Martínez‐Martínez D, Guinart D, Casanovas R, Carranza S, Pasmans F (2020) Integral chain management of wildlife diseases. Conserv Lett 13(2):e12707
Mateo Sánchez MC, Cushman SA, Saura S (2014) Scale dependence in habitat selection: the case of the endangered brown bear (Ursus arctos) in the cantabrian range (NW Spain). Int J Geogr Inf Sci 28(8):1531–1546
McCallum H, Dobson A (2002) Disease, habitat fragmentation and conservation. Proc Biol Sci 269(1504):2041–2049
McMahon TA, Brannelly LA, Chatfield MWH, Johnson PTJ, Joseph MB, McKenzie VJ, Richards-Zawacki CL, Venesky MD, Rohr JR (2013) Chytrid fungus batrachochytrium dendrobatidis has non-amphibian hosts and releases chemicals that cause pathology in the absence of infection. PNAS 110(1):210–215
McRae BH (2006) Isolation by resistance. Evolution 60(8):1551–1561
McRae BH, Beier P (2007) Circuit theory predicts gene flow in plant and animal populations. PNAS 104(50):19885–19890
McRae BH, Shah V, Mohapatra T (2013) Circuitscape 4.0 user guide. The Nature Conservancy. http://www.circuitscape.org
Merow C, Smith MJ, Silander JA (2013) A practical guide to MaxEnt for modeling species’ distributions: what it does, and why inputs and settings matter. Ecography 36(10):1058–1069
Mosher BA, Huyvaert KP, Bailey LL (2018) Beyond the swab: ecosystem sampling to understand the persistence of an amphibian pathogen. Oecologia 188(1):319–330
Muscarella R, Galante PJ, Soley-Guardia M, Boria RA, Kass JM, Uriarte M, Anderson RP, McPherson J (2014) ENMeval: an R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol Evol 5(11):1198–1205
Nobert BR, Merrill EH, Pybus MJ, Bollinger TK, Hwang YT, McCallum H (2016) Landscape connectivity predicts chronic wasting disease risk in Canada. J Appl Ecol 53(5):1450–1459
Nowakowski AJ, Jiménez BO, Allen M, Diaz-Escobar M, Donnelly MA (2013) Landscape resistance to movement of the poison frog, Oophaga pumilio, in the lowlands of northeastern Costa Rica. Anim Conserv 16(2):188–197
Nowakowski AJ, Veiman-Echeverria M, Kurz DJ, Donnelly MA (2015) Evaluating connectivity for tropical amphibians using empirically derived resistance surfaces. Ecol Appl 25(4):928–942
Nowakowski AJ, Whitfield SM, Eskew EA, Thompson ME, Rose JP, Caraballo BL, Kerby JL, Donnelly MA, Todd BD (2016) Infection risk decreases with increasing mismatch in host and pathogen environmental tolerances. Ecol Lett 19(9):1051–1061
Padgett-Flohr GE, Hopkins RL (2010) Landscape epidemiology of Batrachochytrium dendrobatidis in central California. Ecography 33(4):688–697
Panel on Animal Health and Welfare (AHAW), More S, Angel Miranda M, Bicout D, Bøtner A, Butterworth A, Calistri P, Depner K, Edwards S, Garin-Bastuji B, Good M, Michel V, Raj M, Saxmose Nielsen S, Sihvonen L, Spoolder H, Stegeman JA, Thulke H‐H, Velarde A, Willeberg P, Winckler C, Baláž V, Martel A, Murray K, Fabris C, Munoz‐Gajardo I, Gogin A, Verdonck F, Gortázar Schmidt C (2018) Risk of survival, establishment and spread of Batrachochytrium salamandrivorans (Bsal) in the EU. EFSA J 16(4):2581
Paquette SR, Talbot B, Garant D, Mainguy J, Pelletier F (2014) Modelling the dispersal of the two main hosts of the raccoon rabies variant in heterogeneous environments with landscape genetics. Evol Appl 7(7):734–749
Peterman WE, Connette GM, Semlitsch RD, Eggert LS (2014) Ecological resistance surfaces predict fine-scale genetic differentiation in a terrestrial woodland salamander. Mol Ecol 23(10):2402–2413
Peterson RA, Peterson MRA (2020) Package ‘bestNormalize’. Normalizing transformation functions. R package version, 1
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Model 190(3–4):231–259
Phillips SJ, Dudik M, Elith J, Graham CH, Lehmann A, Lethwick J, Ferrier S (2009) Sample selection bias and presence-only distribution models: implications for background and pseudo-absence data. Ecol Apl 19(1):181–197
Pradhan P (2016) Strengthening MaxEnt modelling through screening of redundant explanatory bioclimatic variables with variance inflation factor analysis. Researcher. https://doi.org/10.7537/marsrsj080516.05
Puschendorf R, Carnaval AC, VanDerWal J, Zumbado-Ulate H, Chaves G, Bolaños F, Alford RA (2009) Distribution models for the amphibian chytrid batrachochytrium dendrobatidis in Costa Rica: proposing climatic refuges as a conservation tool. Divers Distrib 15(3):401–408
R Core Team (2019) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Sandvoß M, Wagner N, Lötters S, Feldmeier S, Schulz V, Steinfartz S, Veith M (2020) Spread of the pathogen Batrachochytrium salamandrivorans and large-scale absence of larvae suggest unnoticed declines of the european fire salamander in the southern Eifel mountains. Salamandra 56(3):215–226
Schägner JP, Brander L, Maes J, Paracchinia ML, Hartje V (2016) Mapping recreational visits and values of European National Parks by combining statistical modelling and unit value transfer. J Nat Conserv 31:71–84
Scheele BC, Foster CN, Hunter DA, Lindenmayer DB, Schmidt BR, Heard GW (2019) Living with the enemy: facilitating amphibian coexistence with disease. Biol Conserv 236:52–59
Schmeller DS, Utzel R, Pasmans F, Martel A (2020) Batrachochytrium salamandrivorans kills alpine newts (Ichthyosaura alpestris) in southernmost Germany. Salamandra 56(3):230–232
Schmidt BR, Bozzuto C, Lötters S, Steinfartz S (2017) Dynamics of host populations affected by the emerging fungal pathogen Batrachochytrium salamandrivorans. R Soc Open Sci 4(3):160801
Schulte U, Küster D, Steinfartz S (2007) A PIT tag based analysis of annual movement patterns of adult fire salamanders (Salamandra salamandra) in a middle european habitat. Amphibia-Reptilia 28:531–536
Schulz V, Schulz A, Klamke M, Preissler K, Sabino-Pinto J, Müsken M, Schlüpmann M, Heldt L, Kamprad F, Enss J, Schweinsberg M, Virgo J, Rau H, Veith M, Lötters S, Wagner N, Steinfartz S, Vences M (2020) Batrachochytrium salamandrivorans in the Ruhr District, Germany: history, distribution, decline dynamics and disease symptoms of the salamander plague. Salamandra 56(3):189–214
Skerratt LF, Berger L, Speare R, Cashins S, McDonald KR, Phillott AD, Hines HB, Kenyon N (2007) Spread of chytridiomycosis has caused the rapid global decline and extinction of frogs. EcoHealth 4(2):125–134
Spitzen-van der Sluijs A, Martel A, Asselberghs J, Bales EK, Beukema W, Bletz MC, Dalbeck L, Goverse E, Kerres A, Kinet T, Kirst K, Laudelout A, Marin da Fonte LF, Nöllert A, Ohlhoff D, Sabino-Pinto J, Schmidt BR, Speybroeck J, Spikmans F, Steinfartz S, Veith M, Vences M, Wagner N, Pasmans F, Lötters S (2016) Expanding distribution of lethal amphibian fungus Batrachochytrium salamandrivorans in Europe. Emerg Infect Dis 22(7):1286–1288
Spitzen-van der Sluijs A, Spikmans F, Bosman W, de Zeeuw M, van der Meij T, Goverse E, Kik M, Pasmans F, Martel A (2013) Rapid enigmatic decline drives the fire salamander (Salamandra salamandra) to the edge of extinction in the Netherlands. Amphibia-Reptilia 34(2):233–239
Spitzen-van der Sluijs A, Stegen G, Bogaerts S, Canessa S, Steinfartz S, Janssen N, Bosman W, Pasmans F, Martel A (2018) Post-epizootic salamander persistence in a disease-free refugium suggests poor dispersal ability of Batrachochytrium salamandrivorans. Sci Rep 8(1):3800
Stegen G, Pasmans F, Schmidt BR, Rouffaer LO, van Praet S, Schaub M, Canessa S, Laudelout A, Kinet T, Adriaensen C, Haesebrouck F, Bert W, Bossuyt F, Martel A (2017) Drivers of salamander extirpation mediated by Batrachochytrium salamandrivorans. Nature 544(7650):353–356
Stuart SN, Chanson JS, Cox NA, Young BE, Rodrigues ASL, Fischman DL, Waller RW (2004) Status and trends of amphibian declines and extinctions worldwide. Science 306(5702):1783–1786
Syfert MM, Smith MJ, Coomes DA (2013) The effects of sampling bias and model complexity on the predictive performance of MaxEnt species distribution models. PLoS ONE 8(2):e55158
Thein J, Reck U, Dittrich C, Martel A, Schulz V, Hansbauer G (2020) Preliminary report on the occurrence of Batrachochytrium salamandrivorans in the Steigerwald, Bavaria, Germany. Salamandra 56(3):227–229
Thiesmeier B (2004) Der Feuersalamander. Suppl 4 der Z Feldherpetol. Laurenti, Bielefeld
Thomas V, Wang Y, van Rooij P, Verbrugghe E, Baláž V, Bosch J, Cunningham AA, Fisher MC, Garner TWJ, Gilbert MJ, Grasselli E, Kinet T, Laudelout A, Lötters S, Loyau A, Miaud C, Salvidio S, Schmeller DS, Schmidt BR, van der Spitzen A, Steinfartz S, Veith M, Vences M, Wagner N, Canessa S, Martel A, Pasmans F (2019) Mitigating Batrachochytrium salamandrivorans in Europe. Amphibia-Reptilia. https://doi.org/10.1163/15685381-20191157
Urban MC, Lewis LA, Fučíková K, Cordone A (2015) Population of origin and environment interact to determine oomycete infections in spotted salamander populations. Oikos 124(3):274–284
Van Rooij P, Pasmans F, Coen Y, Martel A (2017) Efficacy of chemical disinfectants for the containment of the salamander chytrid fungus Batrachochytrium salamandrivorans. PLoS ONE 12(10):e0186269
Veloz SD (2009) Spatially autocorrelated sampling falsely inflates measures of accuracy for presence-only niche models. J Biogeogr 36(12):2290–2299
Vredenburg VT, Knapp RA, Tunstall TS, Briggs CJ (2010) Dynamics of an emerging disease drive large-scale amphibian population extinctions. PNAS 107(21):9689–9694
Wagner N, Lötters S, Dalbeck L, Düssel H, Guschal M, Kirst K, Ohlhoff D, Wegge J, Reinhardt T, Veith M (2020a) Long-term monitoring of european fire salamander populations (Salamandra salamandra) in the Eifel Mountains (Germany): five years of removal sampling of larvae. Salamandra 56(3):243–253
Wagner N, Harms W, Hildebrandt F, Martens A, Ong SL, Wallrich K, Lötters S, Veith M (2020b) Do habitat preferences of european fire salamander (Salamandra salamandra) larvae differ among landscapes? A case study from Western Germany. Salamandra 56(3):254–264
Wake DB, Vredenburg VT (2008) Are we in the midst of the sixth mass extinction? A view from the world of amphibians. PNAS. https://doi.org/10.1073/pnas.0801921105
Warren DL, Seifert SN (2011) Ecological niche modeling in Maxent: the importance of model complexity and the performance of model selection criteria. Ecol Appl 21(2):335–342
White A, Watt AD, Hails RS, Hartley SE (2000) Patterns of spread in insect-pathogen systems: the importance of pathogen dispersal. Oikos 89(1):137–145
Wisz MS, Hijmans RJ, Li J, Peterson AT, Graham CH, Guisan A (2008) Effects of sample size on the performance of species distribution models. Divers Distrib 14(5):763–773
Yackulic CB, Chandler R, Zipkin EF, Royle JA, Nichols JD, Campbell Grant EH, Veran S, O’Hara RB (2013) Presence-only modelling using MAXENT: when can we trust the inferences. Methods Ecol Evol 4(3):236–243
Yap TA, Nguyen NT, Serr M, Shepack A, Vredenburg VT (2017) Batrachochytrium salamandrivorans and the risk of a second amphibian pandemic. EcoHealth 14(4):851–864
Acknowledgements
We gratefully thank Lutz Dalbeck, Vanessa Schulz, Gonzalo Albaladejo, Sönke Twietmeyer and the “Biologische Station im Kreis Düren” for providing us with data on salamander distribution and for their technical and logistical support. Jesse Erens, Wouter Beukema and Juliana Döstel provided data on trail and visitor density from the region. Luciano Bani gave valuable assistance for the habitat suitability and resistance modelling. Nico Spaarkogel and the American Journal Experts (AJE) edited this article.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by projects of the “Bundesamt für Naturschutz” (F + E project: ‘Monitoring und Entwicklung von Vorsorgemaßnahmen zum Schutz vor der Ausbreitung des Chytridpilzes Batrachochytrium salamandrivorans “Bsal” im Freiland’) and the “Deutsche Bundesstiftung Umwelt”. RK appreciates the support of the German Research Foundation (DFG) for funding the sMon working group through the iDiv (DFG FZT 118, 202548816).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or nonfinancial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10980_2023_1636_MOESM1_ESM.pdf
SI1. Removal sampling data extracted from Wagner et al. 2020a. Supplementary material 1 (PDF 219.1 kb)
10980_2023_1636_MOESM2_ESM.pdf
SI2. Spearman rank correlation and Variance Inflation of HSM candidate variables. Supplementary material 2 (PDF 211.3 kb)
10980_2023_1636_MOESM3_ESM.pdf
SI3. Tables with candidate breeding and whole life cycle habitat suitability models. Supplementary material 3 (PDF 281.9 kb)
10980_2023_1636_MOESM5_ESM.pdf
SI5: Person’s correlation coefficients of GLM candidate predictors and Moran’s I test for spatial autocorrelation of the candidate model residuals. Supplementary material 5 (PDF 250.2 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bolte, L., Goudarzi, F., Klenke, R. et al. Habitat connectivity supports the local abundance of fire salamanders (Salamandra salamandra) but also the spread of Batrachochytrium salamandrivorans. Landsc Ecol 38, 1537–1554 (2023). https://doi.org/10.1007/s10980-023-01636-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-023-01636-8