Abstract
To develop a DNA aptamer-based PET tracer for imaging of glioblastoma. 5 mM of NOTA-AS1411, 60-min, and 37 °C were selected as the optimal condition for 64Cu radiolabeling of AS1411. 64Cu-NOTA-AS1411 remained stable in PBS and 100% mouse serum for at least six hours. From the PET images, 64Cu-NOTA-AS1411 tended to be excreted out through the kidneys and there was high tracer accumulation in the bladder. There was a higher tumor uptake in the AS1411 group than that in the control group. 64Cu-NOTA-AS1411 is a suitable potential PET tracer for imaging murine glioblastoma.
Similar content being viewed by others
Explore related subjects
Find the latest articles, discoveries, and news in related topics.Avoid common mistakes on your manuscript.
Introduction
Ever since their discovery, DNA or RNA-based aptamers have been designed to act against a wide variety of targets present on the surface of cells, ranging from small molecules [1, 2], oligonucleotides [3, 4], peptides [5,6,7], to complex proteins [8,9,10]. DNA/RNA aptamers have been incredibly useful in biomedical applications, such as biosensing [11, 12], biomarker discovery [13, 14], medical imaging [15,16,17], and drug delivery [18, 19]. Among all DNA/RNA aptamers developed or in development, more than ten DNA/RNA aptamers are currently in clinical trials [20,21,22,23,24]. Thus far, U.S. Food and Drug Administration (FDA) has approved one RNA aptamer for the treatment of age-related macular degeneration [25].
Due to their great target affinity, enhanced heat stability, and excellent biocompatibility, aptamers prove promising for molecular imaging. Last two decades, we have witnessed the development of aptamer-based molecular imaging using various imaging modalities [26], such as fluorescent imaging [27, 28], ultrasound molecular imaging [29, 30], magnetic resonance imaging [31, 32], and nuclear medicine imaging techniques [33,34,35] including positron emission tomography (PET) and single-photon emission computed tomography (SPECT). Nuclear medicine imaging utilizes probes carrying radioactive isotopes for imaging, generally named radiopharmaceuticals or radiotracers. Once administrated into living organisms, PET or SPECT scanners may find the signal emitted from such probes. The signal can later be reconstructed for data analysis and allow for subsequent analysis to identify diseased tissues via tracer uptake quantification at different time points after administration. PET imaging detects positron-emitting (β+) radionuclides (such as 11C, 13N, 18F, 64Cu, 68Ga) and offers higher imaging resolution and improved quantifiability than SPECT imaging. Based on the advantages of nuclear medicine imaging and aptamer, Li et al. [36] conducted 64Cu radiolabeling of AS1411 with four different chelators. In vivo PET/CT imaging showed that the molecular probe obtained using chelator CB-TE2A showed significant uptake at the tumor and also exhibited lower liver uptake and higher tumor to background contrast. 64Cu-CB-TE2A-AS1411 probe is suitable for lung cancer imaging.
Herein, we developed a DNA aptamer (AS1411)-based PET tracer for imaging of glioblastoma. AS1411 was functionalized with NOTA chelator and 64Cu (half-life 12.7 h) was later labeled for PET imaging. We systematically investigated different labeling conditions to optimize the labeling efficiency. The obtained tracer, 64Cu-NOTA-AS1411 was further used for imaging of U87MG tumor-bearing mice. Uptake of 64Cu-NOTA-AS1411 and 64Cu-NOTA-CS (control sequence) was studied in U87MG tumor-bearing mice using PET. Results showed that AS1411 showed significantly higher affinity than the control ssDNA, based on semiquantitative PET analysis.
Materials and methods
General
AS1411 aptamer (5′-GGTGGTGGTGGTTGTGGTGGTGGTGG-3′) [37] in the form of single-strand DNA (ssDNA) was purchased from Sangon Biotech (Shanghai) Co., Ltd. (China), from which a control sequence (CS) with the same number of DNA bases but a random sequence was also obtained. Amine group was designed at the 5’ end of AS1411 and CS.
Bifunctional chelator 2-S-(4-isothiocyanatobenzyl)-1,4,7-triazacyclononane-1,4,7-triacetic acid (p-SCN-Bn-NOTA) was purchased from Macrocyclics, Inc. (Plano, TX, USA). Dulbecco’s Modified Eagle’s Medium (DMEM) and fetal bovine serum (FBS) were purchased from Thermo Fisher Technology (China) Co., Ltd.
Other reagents were purchased from Sinopharm Chemical Reagent Co., Ltd., and used as received, unless stated otherwise.
Bifunctional chelator conjugation
To achieve 64Cu radiolabeling of ssDNA, p-SCN-Bn-NOTA was conjugated to the amine group at the end of ssDNA. SsDNA (AS1411 or CS) was dissolved in 1 × PBS (phosphate buffered saline) to achieve a final concentration of 0.1 mM. 0.5–1 mg of p-SCN-Bn-NOTA was weighed, dissolved in DMSO, and then added to the ssDNA solution, of which the final pH was adjusted to 9.0–9.2 using a carbonate buffer. The reaction was allowed for 2 h with constant shaking under room temperature. p-SCN-Bn-NOTA conjugated ssDNA (NOTA-AS1411 or NOTA-CS) was purified with a NAP-5 column (GE Healthcare, Fairfield, CT, USA) with 1 × PBS as the elution buffer and DNA concentration of each fraction was examined with UV–VIS spectroscopy (Agilent Cary 60, Agilent Technologies, Santa Clara, CA, USA).
64Cu Radiolabeling of NOTA-ssDNA
64Cu was produced by the PETtrace cyclotron (GE Healthcare) via the 64Ni(p,n)64Cu reaction. 64Cu in 0.1 M HCl was added to 100 μL of sodium acetate (0.05 M, pH 5.5) and further mixed with 500 μL of NOTA-ssDNA. The reaction was performed at a certain temperature for a certain amount of time with constant shaking. Radiolabeling efficiency was determined by thin layer chromatography (TLC) at different time points during the labeling reaction. Sodium EDTA solution (0.05 M) was used as the mobile phase. The reaction solution was purified by a NAP-5 column and the fraction with the highest radioactivity, which was 64Cu-NOTA-ssDNA, was used for stability tests and animal studies.
In vitro stability test
To investigate in vitro stability, 64Cu-NOTA-AS1411 was added to 1 × PBS and 100% mouse serum for incubation under 37 °C, and the radiochemical purity was determined by TLC at different time points.
Glioma tumor model establishment
Human malignant glioblastoma cells (U87MG) were purchased from Fuheng Biotechnology Co., Ltd. (Shanghai, China) and cultured in DMEM supplemented with 10% FBS at 37 °C in a humidified incubator with 5% CO2.
About 5 × 106 U87MG cells in 100 μL 1 × PBS were subcutaneously injected into the right flank of each athymic mouse (4–6 weeks old, female, Beijing Vital River Laboratory Animal Technology Co., Ltd.). The mice were subjected to subsequent PET imaging when the tumors reached 0.8–1.0 cm in diameter.
The animal study protocol had been approved by the Medical Ethics Committee of Inner Mongolia Medical University (No. YKD202001143). The animal experiments were performed in accordance with the ‘Guide for the Care and Use of Laboratory Animals’ (Institute of Laboratory Animal Resources, Commission on Life Sciences, National Research Council, ISBN: 0-309-58869-3, 140 pages, 1996).
PET imaging of 64Cu-NOTA-AS1411
1.85–3.7 MBq (50–100 µCi) of 64Cu-NOTA-AS1411 (or 64Cu-NOTA-CS) was intravenously injected into U87MG tumor-bearing mice. PET scans were performed with Micro-PET/CT (SIEMENS Inveon MM, Siemens Ltd., Munich, Germany) at 0.5 and 3 h after injection. Region-of-interest (ROI) analysis was performed to determine time-activity curves for different organs after decay correction. Tracer accumulation in each organ or tissue was denoted as the percentage of injected dose per gram of tissue (%ID/g).
Results and discussion
64Cu radiolabeling of AS1411
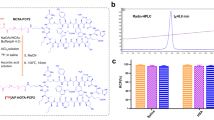
To label AS1411 with 64Cu, AS1411 was modified first with a bifunctional chelator p-SCN-Bn-NOTA. NOTA-AS1411was successfully obtained through p-SCN-Bn-NOTA being conjugated to the amine group at the end of AS1411, which was shown in Fig. 1.
To improve the labeling efficiency and obtain the best radiochemical purity, we planned to conduct several experiments by variating from the aspects of DNA concentration, reaction time, and reaction temperature. Results are shown in Fig. 2.
According to the previous results of our research, the concentration of ssDNA is one of the important reaction parameters to improve the labeling efficiency in the study of labeling ssDNA with 18F or 99mTc. Thus, in this study, for 64Cu radiolabeling of AS1411, we set five different ssDNA concentrations at 0.1, 0.5, 1, 5, and 10 mM with the reaction time being 60 min and the reaction temperature being 37 °C. When araising from 0.1 to 10 mM of AS1411 concentration, the labeling efficiency improved from 5% to almost 90%, indicative of the positive correlation between radiochemical yield (RCY) and precursor concentration (AS1411), as shown in Fig. 2a.
The reaction time and reaction temperature also played pivotal roles in the 64Cu radiolabeling of AS1411. As shown in Fig. 2b, the labeling efficiency increased in function of time with 5 mM of NOTA-AS1411 and 37 °C. However, compared with the 60-min reaction, the labeling efficiency of 120-min reaction did not significantly improve (62.03 ± 2.61 at 60 min vs 63.03 ± 2.97 at 120 min). The higher reaction temperature proved beneficial for the labeling efficiency from 54.50 ± 3.31 at 25 °C to 71.73 ± 2.23 at 90 °C, which were shown in Fig. 2c. After considering various important reaction parameters, we selected 5 mM of NOTA-AS1411, 60-min reaction time, and 37 °C as the condition for 64Cu radiolabeling of AS1411, which was similar to the labeling condition reported in the literature [36]. The reaction solution was purified by a NAP-5 column, and the specific activity of 64Cu-NOTA-AS1411 was in the range of 0.8–1.7 mCi/nmol.
In vitro stability test
The as-prepared 64Cu-NOTA-AS1411 was further tested to validate the radiolabeling stability in 1 × PBS and 100% mouse serum. As shown in Fig. 3, 64Cu-NOTA-AS1411 proved stable in either buffer for at least six hours as evidenced by the minimal leaking of free 64Cu.
PET imaging of 64Cu-NOTA-AS1411
To validate the DNA aptamer tracer for PET imaging of glioblastoma, we prepared mice bearing U87MG subcutaneous tumors on their right shoulder. To better highlight the specificity of the DNA aptamer AS1411, we also radiolabeled CS to serve as a control. As shown in Fig. 4a, we could find that 64Cu-NOTA-ssDNA tended to be excreted out through the kidneys and we saw high tracer accumulation in the bladder in both groups (51.20 ± 44.30% ID/g for AS1411 vs 37.10 ± 22.70% ID/g for CS, at 0.5 h after injection). We found a higher tumor uptake from 1.50 ± 0.12% ID/g at 0.5 h to 1.80 ± 0.14% ID/g at 3 h in the AS1411 group, while in the control group, the tumor uptakes were 0.90 ± 0.24% ID/g at 0.5 h and 0.80 ± 0.15% ID/g at 3 h after injection, which was shown in Fig. 4b. From the ratio of radioactive uptake in tumor and heart, the ratio of AS1411 group (0.71 at 0.5 h and 1.50 at 3 h) was significantly higher than that of CS group (0.25 at 0.5 h and 0.36 at 3 h) at both time points, suggesting that compared with 64Cu-NOTA-CS, more 64Cu-NOTA-AS1411 infiltrated into the tumor tissue from the blood, and could be specifically concentrated in the tumor tissue. As to tumor-to-background ratios, tumor/muscle ratios in AS1411 group (4.33 at 0.5 h and 10.72 at 3 h) were also higher than that in CS group (0.91 at 0.5 h and 0.92 at 3 h), further demonstrating the specificity of AS1411 for glioblastoma. We also found that CS showed a higher uptake in the liver (3.70 ± 0.81% ID/g for AS1411 vs 6.70 ± 0.51% ID/g for CS, at 0.5 h after injection), while AS1411 cleared out of the body relatively faster (3.00 ± 1.00% ID/g for AS1411 vs 9.60 ± 1.70% ID/g for CS, at 3 h after injection in the kidneys). The liver uptake significantly increased the background signal of CS group and produced too many noises to delineate the tumor. In the reported literature [36], 64Cu-CB-TE2A-AS1411, prepared with CB-TE2A as the chelator, showed the highest tumor/background ratio at 1 h after injection. The majority of 64Cu-CB-TE2A-AS1411 was accumulated in the liver with slow clearance from kidney, which was similar to the results in this study.
Conclusions
In summary, we prepared a 64Cu-labeled DNA aptamer AS1411 (64Cu-NOTA-AS1411) as a PET tracer for imaging of U87MG in murine cancer models. During the radiolabeling process, we compared and identified the optimal labeling conditions for 64Cu labeling of AS1411 via NOTA chelation. The as-prepared 64Cu-NOTA-AS1411 showed excellent stability in both 1 × PBS and 100% mouse serum, suggesting its potential for in vivo studies. In animal PET imaging, we found that 64Cu-NOTA-AS1411 showed higher tumor uptake and lower liver uptake than its nonspecific counterpart (64Cu-NOTA-CS).
Data availability
All data generated or analysed during this study are included in this published article.
References
Li Y, Zhao Q (2019) Aptamer structure switch fluorescence anisotropy assay for small molecules using streptavidin as an effective signal amplifier based on proximity effect. Anal Chem 91:7379–7384
Deng R, Dong Y, Xia X et al (2018) Recognition-enhanced metastably shielded aptamer for digital quantification of small molecules. Anal Chem 90:14347–14354
Teles FRF, Teles RP, Siegelin Y et al (2011) RNA-oligonucleotide quantification technique (ROQT) for the enumeration of uncultivated bacterial species in subgingival biofilms. Mol Oral Microbiol 26:127–139
Wu Y, Zhang G, Li Y et al (2008) Inhibition of highly pathogenic avian H5N1 influenza virus replication by RNA oligonucleotides targeting NS1 gene. Biochem Bioph Res Co 365:369–374
Wang C-Y, Lin B-L, Chen C-H (2021) Targeted drug delivery using an aptamer against shared tumor-specific peptide antigen of MAGE-A3. Cancer Biol Ther 22:12–18
Agyei D, Pan S, Acquah C et al (2019) Structure-informed detection and quantification of peptides in food and biological fluids. J Food Biochem 43:e12482
Tabarzad M, Jafari M (2016) Trends in the design and development of specific aptamers against peptides and proteins. Protein J 35:81–99
Wan Q, Zeng Z, Qi J et al (2022) Aptamer targets triple-negative breast cancer through specific binding to surface CD49c. Cancers 14:1570
An Y, Li X, Yao F et al (2022) Novel complex of PD-L1 aptamer and albumin enhances antitumor efficacy in vivo. Molecules 27:1482
Lin M, Zhang J, Wan H et al (2021) Rationally designed multivalent aptamers targeting cell surface for biomedical applications. ACS Appl Mater Inter 13:9369–9389
Nakatsuka N, Abendroth JM, Yang K-A et al (2021) Divalent cation dependence enhances dopamine aptamer biosensing. ACS Appl Mater Inter 13:9425–9435
Zhou X, Zhu Q, Yang Y (2020) Aptamer-integrated nucleic acid circuits for biosensing: classification, challenges and perspectives. Biosens Bioelectron 165:112422
Huang J, Chen X, Fu X et al (2021) Advances in aptamer-based biomarker discovery. Front Cell Dev Biol 9:659760
Xiong H, Yan J, Cai S et al (2019) Cancer protein biomarker discovery based on nucleic acid aptamers. Int J Biol Macromol 132:190–202
Koudrina A, DeRosa MC (2020) Advances in medical imaging: aptamer- and peptide-targeted MRI and CT contrast agents. ACS Omega 5:22691–22701
Calzada V (2020) Aptamers in diagnostic and molecular imaging applications. In: Urmann K, Walter J-G (eds) Aptamers in biotechnology. Springer, Cham, pp P141–P160
Röthlisberger P, Gasse C, Hollenstein M (2017) Nucleic acid aptamers: emerging applications in medical imaging, nanotechnology, neurosciences, and drug delivery. Int J Mol Sci 18:2430
Khan S, Hussain A, Fahimi H et al (2022) A review on the therapeutic applications of aptamers and aptamer-conjugated nanoparticles in cancer, inflammatory and viral diseases. Arab J Chem 15:103626
Walia S, Chandrasekaran AR, Chakraborty B et al (2021) Aptamer-programmed DNA nanodevices for advanced, targeted cancer theranostics. ACS Appl Bio Mater 4:5392–5404
Kovacevic KD, Greisenegger S, Langer A et al (2021) The aptamer BT200 blocks von Willebrand factor and platelet function in blood of stroke patients. Sci Rep-UK 11:3092
Yu A-M, Tu M-J (2022) Deliver the promise: RNAs as a new class of molecular entities for therapy and vaccination. Pharmacol Ther 230:107967
Zhou L-Y, Qin Z, Zhu Y-H et al (2019) Current RNA-based therapeutics in clinical trials. Curr Gene Ther 19:172–196
Shigdar S, Schrand B, Giangrande PH et al (2021) Aptamers: cutting edge of cancer therapies. Mol Ther 29:2396–2411
Morita Y, Leslie M, Kameyama H et al (2018) Aptamer therapeutics in cancer: current and future. Cancers 10:80
Vinores SA (2006) Pegaptanib in the treatment of wet, age-related macular degeneration. Int J Nanomed 1:263–268
Yoon S, Rossi JJ (2018) Targeted molecular imaging using aptamers in cancer. Pharmaceuticals-Base 11:71
Xiang Z, Zhao J, Yi D et al (2021) Peptide nucleic acid (PNA)-guided peptide engineering of an aptamer sensor for protease-triggered molecular imaging. Angew Chem Int Edit 60:22659–22663
Yin J, Chen S, Song Y et al (2020) Fluorescent imaging of cytoplasmic nucleolin in live cells by a functionalized-engineered aptamer. Chem Commun 56:14171–14174
Zhu L, Wang L, Liu Y et al (2018) CAIX aptamer-functionalized targeted nanobubbles for ultrasound molecular imaging of various tumors. Int J Nanomed 13:6481–6495
Fang K, Wang L, Huang H et al (2020) Construction of nucleolin-targeted lipid nanobubbles and contrast-enhanced ultrasound molecular imaging in triple-negative breast cancer. Pharm Res-Dordr 37:145
Chen Z, Peng Y, Li Y et al (2021) Aptamer-dendrimer functionalized magnetic nano-octahedrons: theranostic drug/gene delivery platform for near-infrared/magnetic resonance imaging-guided magnetochemotherapy. ACS Nano 15:16683–16696
Koudrina A, McConnell EM, Zurakowski JA et al (2021) Exploring the unique contrast properties of aptamer-gadolinium conjugates in magnetic resonance imaging for targeted imaging of thrombi. ACS Appl Mater Inter 13:9412–9424
Song W, Song Y, Li Q et al (2022) Advances in aptamer-based nuclear imaging. Eur J Nucl Med Mol I 49:2544–2559
Filippi L, Bagni O, Nervi C (2020) Aptamer-based technology for radionuclide targeted imaging and therapy: a promising weapon against cancer. Expert Rev Med Devic 17:751–758
Cheng S, Jacobson O, Zhu G et al (2019) PET imaging of EGFR expression using an 18F-labeled RNA aptamer. Eur J Nucl Med Mol I 46:948–956
Li J, Zheng H, Bates PJ et al (2014) Aptamer imaging with Cu-64 labeled AS1411: preliminary assessment in lung cancer. Nucl Med Biol 41:179–185
Ireson CR, Kelland LR (2006) Discovery and development of anticancer aptamers. Mol Cancer Ther 5:2957–2962
Funding
National Natural Science Foundation of China (No. 82060324) and "Youth Science and Technology Talents in Colleges and Universities" Support Program Project of Inner Mongolia Education Department (NJYT22016).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, P., Wang, C., Wang, W. et al. Preliminary evaluation of a 64Cu-labeled DNA aptamer for PET imaging of glioblastoma. J Radioanal Nucl Chem 332, 2279–2284 (2023). https://doi.org/10.1007/s10967-023-08835-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10967-023-08835-2