Abstract
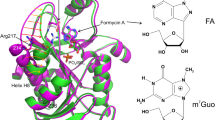
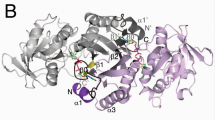
Uridine–cytidine kinase catalyzes phosphorylation of the pyrimidine nucleosides uridine and cytidine and plays an important role in nucleotide metabolism. However, the detailed molecular mechanism of these reactions remains to be elucidated. Here, we determined the structure of the ternary complex of Uridine–cytidine kinase from Thermus thermophilus HB8 with both cytidine and β,γ-methyleneadenosine 5′-triphosphate, a non-hydrolysable ATP analogue. Substrate binding is accompanied by substantial domain movement that allows the substrate-binding cleft to close. The terminal phosphodiester bond of the ATP analogue is in an ideal location for an inline attack of the 5′-hydroxyl group of cytidine. Asp40 is located near the 5′-hydroxyl group of cytidine. Mutation of this conserved residue to Asn or Ala resulted in a complete loss of enzyme activity, which is consistent with the notion that Asp40 acts as a general base that activates the 5′-hydroxyl group of cytidine. The pH profile of the activity showed an apparent pK a value of 7.4. Based on this structure, a likely mechanism of the catalytic step is discussed.









Similar content being viewed by others
Abbreviations
- AMPPCP:
-
β,γ-methyleneadenosine 5′-triphosphate
- CMP:
-
Cytidine monophosphate
- cv:
-
Column volume
- NMP:
-
Nucleoside monophosphate
- ttUCK:
-
Uridine–cytidine kinase in T. thermophilus HB8
- UCK:
-
Uridine–cytidine kinase
- WT:
-
Wild type
References
Anderson EP (1973) Nucleoside and nucleotide kinases. In: Boyer PD (ed) The enzymes, vol Vol IX, 3rd edn. Academic Press, New York, pp 49–96
O’Donovan GA, Neuhard J (1970) Pyrimidine metabolism in microorganisms. Bacteriol Rev 34:278–283
Anderson EP, Brockman RW (1964) Feedback inhibition of uridine kinase by cytidine triphosphate and uridine triphosphate. Biochim Biophys Acta 91:380–386
Murata D, Endo Y, Obata T, Sakamoto K, Syouji Y, Kadohira M, Matsuda A, Sasaki T (2004) A crucial role of uridine/cytidine kinase 2 in antitumor activity of 3′-ethynyl nucleosides. Drug Metab Dispos 32:1178–1182
Orengo A (1969) Regulation of enzymic activity by metabolites. I. Uridine–cytidine kinase of Novikoff ascites rat tumor. J Biol Chem 244:2204–2209
Shimamoto Y, Koizumi K, Okabe H, Kazuno H, Murakami Y, Nakagawa F, Matsuda A, Sasaki T, Fukushima M (2002) Sensitivity of human cancer cells to the new anticancer ribo-nucleoside TAS-106 is correlated with expression of Uridine–cytidine kinase 2. Jpn J Cancer Res 93:825–833
Canellakis ES (1957) Pyrimidine metabolism. II. Enzymatic pathways of uracil anabolism. J Biol Chem 227:329–338
Sköld O (1960) Uridine kinase from Ehrlich ascites tumor: purification and properties. J Biol Chem 235:3273–3279
Liacouras AS, Garvey TQ III, Millar FK, Anderson EP (1975) Uridine cytidine kinase: kinetic studies and reaction mechanism. Arch Biochem Biophys 168:74–80
Yan H, Tsai M (1999) Nucleoside monophosphate kinases: structure, mechanism, and substrate specificity. Adv Enzymol Relat Areas Mol Biol 73:103–134
Suzuki NN, Koizumi K, Fukushima M, Matsuda A, Inagaki F (2004) Structural basis for the specificity, catalysis, and regulation of human Uridine–cytidine kinase. Structure 12:751–764
Tomoike F, Nakagawa N, Kuramitsu S, Masui R (2011) A single amino acid limits the substrate specificity of Thermus thermophilus uridine–cytidine kinase to cytidine. Biochemistry 50:4597–4607
Smith AJ, Li Y, Houk KN (2009) Quantum mechanics/molecular mechanics investigation of the mechanism of phosphate transfer in human uridine–cytidine kinase 2. Org Biomol Chem 7:2716–2724
Kuramitsu S, Hiromi K, Hayashi H, Morino Y, Kagamiyama H (1990) Pre-steady state kinetics of Escherichia coli aspartate aminotransferase catalized reactions and thermodynamic aspects of its substrate specificity. Biochemistry 29:5469–5476
Iino H, Naitow H, Nakamura Y, Nakagawa N, Agari Y, Kanagawa M, Ebihara A, Shinkai A, Sugahara M, Miyano M, Kamiya N, Yokoyama S, Hirotsu K, Kuramitsu S (2008) Crystallization screening test for the whole-cell project on Thermus thermophilus HB8. Acta Cryst F64:487–491
Ueno G, Kanda H, Hirose R, Ida K, Kumasaka T, Yamamoto M (2006) RIKEN structural genomics beamlines at the SPring-8; high throughput protein crystallography with automated beamline operation. J Struct Funct Genomics 7:15–22
Otwinowski Z, Minor W (1997) Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol 276:307–326
Vagin A, Teplyakov A (1997) MOLREP: an automated program for molecular replacement. J Appl Cryst 30:1022–1025
Duncan EM (1992) A visual protein crystallographic software system for X11/Xview. J Mol Graph 10:44–46
Brünger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, Jiang J-S, Kuszewski J, Nilges M, Pannu NS, Read RJ, Rice LM, Simonson T, Warren GL (1998) Crystallography & NMR System: a new software suite for macromolecular structure determination. Acta Crystallogr D Biol Cryst 54:905–921
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structreus. J Appl Crystallogr 26:283–291
Kleywegt GJ (2007) Crystallographic refinement of ligand complexes. Acta Crys D Biol Crystallogr 63:94–100
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Iwai T, Kuramitsu S, Masui R (2004) The Nudix hydrolase Ndx1 from Thermus thermophilus HB8 is a diadenosine hexaphosphate hydrolase with a novel activity. J Biol Chem 279:21732–21739
Fersht AR (1999) Structure and mechanism in protein science. Freeman and Company, New York
Bellinzoni M, Haouz A, Graña M, Munier-Lehmann H, Shepard W, Alzari PM (2006) The crystal structure of Mycobacterium tuberculosis adenylate kinase in complex with two molecules of ADP and Mg2+ supports an associative mechanism for phosphoryl transfer. Protein Sci 15:1489–1493
Sekulic N, Shuvalova L, Spangenberg O, Konrad M, Lavie A (2002) Structural characterization of the closed conformation of mouse guanylate kinase. J Biol Chem 277:30236–30243
Antosiewicz J, McCammon JA (1996) The determinants of pKas in proteins. Biochemistry 35:7819–7833
Cowan JA (1995) The biological chemistry of magnesium. Wiley, New York
Picard M, Toyoshima C, Champeil P (2005) The average conformation at micromolar [Ca2+] of Ca2+-atpase with bound nucleotide differs from that adopted with the transition state analog ADP.AlFx or with AMPPCP under crystallization conditions at millimolar [Ca2+]. J Biol Chem 280:18745–18754
Taira K, Uebayasi M, Maeda H, Furukawa K (1990) Energetics of RNA cleavage: implications for the mechanism of action of ribozymes. Protein Eng 3:691–702
Pörtner HO, Heisler N, Grieshaber MK (1984) Anaerobiosis and acid-base status in marine invertebrates: a theoretical analysis of proton generation by anaerobic metabolism. J Comp Physiol B 155:1–12
Kim YG, Yoo JS, Kim JH, Kim CM, Oh JW (2007) Biochemical characterization of a recombinant Japanese encephalitis virus RNA-dependent RNA polymerase. BMC Mol Biol 8:59
Karp DA, Stahley MR, Garcia-Moreno B (2010) Conformational consequences of ionization of Lys, Asp, and Glu buried at position 66 in staphylococcal nuclease. Biochemistry 49:4138–4146
Acknowledgments
This work was partially supported by the Ministry of Education, Science, Sports and Culture, Grant-in-Aid for Challenging Exploratory Research 25650008 (to RM) and the Japan Society for the Promotion of Science for Young Scientists (10J0180400-00, FT).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tomoike, F., Nakagawa, N., Kuramitsu, S. et al. Structural and Biochemical Studies on the Reaction Mechanism of Uridine–Cytidine Kinase. Protein J 34, 411–420 (2015). https://doi.org/10.1007/s10930-015-9636-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-015-9636-8