Abstract
Purpose
Pediatric patients with inborn errors of immunity (IEI) undergoing umbilical cord blood transplantation (UCBT) are at risk of early mortality. Our aim was to develop and validate a prediction model for early mortality after UCBT in pediatric IEI patients based on pretransplant factors.
Methods
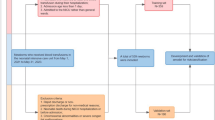
Data from 230 pediatric IEI patients who received their first UCBT between 2014 and 2021 at a single center were analyzed retrospectively. Data from 2014–2019 and 2020–2021 were used as training and validation sets, respectively. The primary outcome of interest was early mortality. Machine learning algorithms were used to identify risk factors associated with early mortality and to build predictive models. The model with the best performance was visualized using a nomogram. Discriminative ability was measured using the area under the curve (AUC) and decision curve analysis.
Results
Fifty days was determined as the cutoff for distinguishing early mortality in pediatric IEI patients undergoing UCBT. Of the 230 patients, 43 (18.7%) suffered early mortality. Multivariate logistic regression with pretransplant albumin, CD4 (absolute count), elevated C-reactive protein, and medical history of sepsis showed good discriminant AUC values of 0.7385 (95% CI, 0.5824–0.8945) and 0.827 (95% CI, 0.7409–0.9132) in predicting early mortality in the validation and training sets, respectively. The sensitivity and specificity were 0.5385 and 0.8154 for validation and 0.7667 and 0.7705 for training, respectively. The final model yielded net benefits across a reasonable range of risk thresholds.
Conclusion
The developed nomogram can predict early mortality in pediatric IEI patients undergoing UCBT.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human inborn errors of immunity (IEI), also known as primary immunodeficiencies, are a group of rare diseases caused by monogenic germline mutations. They can be life-threatening if appropriate medical treatment is not provided. Allogeneic hematopoietic stem cell transplantation (allo-HSCT) offers life-saving and curative treatment for the vast majority of IEI [1-5]. Although a human leukocyte antigen (HLA)-matched unaffected sibling is the preferred donor, it is available for only less than one-third of patients [6, 7]. Unrelated umbilical cord blood (UCB), as an important alternative stem cell source, is a suitable option for patients for whom a suitable donor is not available in due time [8]. However, transplant-related mortality (TRM) is a major obstacle in unrelated umbilical cord blood transplantation (UCBT).
Mortality after allo-HSCT can be classified into very early, early, intermediate, and late mortality, based on the time between HSCT and death [9]. The exact definitions of these classifications vary by transplant indication (e.g., malignancy vs. non-malignancy) and transplant characteristics (e.g., stem cell source). Early all-cause mortality is of great interest in patients undergoing UCBT for several reasons. First, mortality in patients with IEI occurs mainly in the early stage after UCBT, due to delayed engraftment, graft failure or poor graft function, acute graft-versus-host-disease (GvHD), and infection [3-5, 10]. Second, although the overall mortality after UCBT in children has decreased in recent years, the trend in early all-cause mortality has not been reported [7, 9]. Third, patients at high risk of early mortality must be identified for clinicians and patient guardians to make informed decisions.
Despite its importance, early mortality after UCBT in IEI patients has not been well-studied for several reasons. First, IEI are "very rare" and highly heterogeneous. As a result, it is difficult to obtain adequate sample size for IEI subgroup analysis. Second, analytic tools (e.g., machine learning algorithms) have rarely been used to reveal underlying patterns between risk factors and outcomes of interest (i.e., early all-cause mortality) in the landscape of hematology or transplantation. Lastly, the threshold for early mortality after UCBT, defined as a day post-transplant, when mortality transitions from a designation of "early" to “intermediate”, remains undetermined [9, 11-13].
Machine learning is a data-driven analytic approach [14] to establishing a model by recognizing the underlying patterns [15], including the thresholds for specific variables. Machine learning is likely to be increasingly integrated into the research and practice landscape of hematology [16]. However, machine learning is yet to be applied in identifying risk factors for early mortality after allo-HSCT and predicting early all-cause mortality in pediatric IEI patients.
Therefore, in this study, we first established a cutoff value for distinguishing different phases concerning mortality (e.g., early mortality) based on a data-driven approach and real-world evidence in pediatric IEI patients who underwent UCBT and developed and validated a prediction model for early all-cause mortality after UCBT based on pretransplant factors.
Materials and Methods
Patients
Patients included in this study were diagnosed with IEI according to the criteria proposed by the International Union of Immunological Societies, which include: typical clinical manifestations, monogenic mutations, and/or functional identifications [17]. All patients were between 0 and 14 years old and had undergone UCBT at the Children’s Hospital of Fudan University between February 2014 and December 2021. They all met the HSCT guidelines for IEI [18, 19] and had no matched-related donor. No exclusion criteria were set for this study.
This study was approved by the ethics board of the Children’s Hospital of Fudan University. Written informed consent was obtained from the guardians of all the patients.
Transplantation-related definitions
Conditioning regimens were classified as myeloablative conditioning with busulfan (BU) >8 mg/kg in combination with cyclophosphamide (100 mg/kg) and/or fludarabine (150–175 mg/m2) and reduced-intensity conditioning with BU ≤8 mg/kg in combination with cyclophosphamide and/or fludarabine. The BU dosage for patients with severe combined immunodeficiency (SCID), very early onset inflammatory bowel disease (VEO-IBD), and chronic granulomatous disease (CGD) was (6.4–13.2), (8.0–14.4), and (12.0–19.2) mg/kg, respectively. GvHD prophylaxis mainly consisted of calcineurin inhibitors (cyclosporin A or tacrolimus) alone or in combination with mycophenolate mofetil. HLA compatibility was determined by high-resolution typing for HLA-A, -B, -C, -DR, and -DQ loci.
Characteristics and outcomes
Patient (e.g., demographics, clinical manifestations, medical history, laboratory results, and diagnosis), donor (stem cell count and HLA compatibility), and follow-up information (e.g., complications, survival status, and cause of death) were collected retrospectively using a standardized, computerized form.
Serum biochemical indices (albumin [Alb], alanine aminotransferase [ALT], total bilirubin, and immunoglobulin), inflammatory biomarkers (procalcitonin [PCT], ferritin, and Interleukin 6 [IL-6]), and lymphocyte subsets were measured approximately one week before the start of conditioning. C-reactive protein (CRP) was measured just before the initiation of conditioning. The elements in the medical history category (e.g., intestinal infection, severe pneumonia) are defined in detail in Table S1.
With the exception of the outcomes of interest (early mortality and causes of death), all the aforementioned analysis variables were pretransplant factors. A complete list of the variables is shown in Table S2.
We plotted the cumulative density plot of the interval between UCBT and the last follow-up or death. Early mortality was defined based on the pattern depicted in the plot using the ‘elbow’ method [20].
Modeling
When preparing the data for modeling, we first checked for missing patterns in the variables. Missing values were then handled with a multivariate imputation strategy to prepare the data matrix for modeling, and one of the five imputed data sets was randomly selected. Variables used in the analyses were converted to numeric or binary values (e.g., 1 indicates male, 0 indicates female). Using one-hot encoding method [21], we converted IEI disease diagnosis (categorical data) to multidimensional binary vectors. The primary outcome variable was converted to 0 (negative) or 1 (positive) indicating early mortality.
The cohort dataset was divided based on a marker time point. Data from patients who underwent UCBT before and after January 1, 2020, were assigned to a training or validation set, respectively, before feature selection and model development. We performed ensemble feature selection to alleviate and compensate for specific biases associated with single feature selection and to increase the stability of feature selection based on a different training set [22].
Machine learning algorithms were used to develop a prediction model. Multivariate logistic regression (LR), Lasso [23], random forest, and extreme gradient boosting (XGBoost) were used for the selected variable and all-variable sets, as appropriate. Subsequently, the optimal parameters of the machine learning algorithms were obtained through cross-validation using the training set. Finally, the performance of the model was compared using the validation set. Among all the models, the LR with the selected variables provided the highest area under the curve (AUC) using the validation set.
The resulting model was further validated using the bootstrap method (1000 times) after a performance assessment in the validation set [24]. Finally, a nomogram was developed to visualize the model.
Model evaluation
Performance of the best model (i.e., LR with the selected variables) was evaluated using the AUC. The accuracy of the optimal cutoff value was assessed using sensitivity, specificity, and positive and negative predictive values (PPV and NPV, respectively).
Patients in the temporal validation cohort were classified into two prognostic groups (i.e., high-risk group vs. low-risk group) based on their predicted probability of early mortality and the selected cutoff probability, and their survival curves were compared using the Kaplan–Meier method. In addition, a decision curve analysis (DCA) was performed to evaluate the potential clinical benefits of using the model to identify patients at high risk for early mortality. DCA [25] evaluates the value of a predictive model when making clinical decisions. Three strategies were compared: selecting all patients for intervention (i.e., treating all), selecting no patients (i.e., treating none), and selecting patients based on the nomogram.
Statistical analyses
Unpaired, two-tailed t-test and Wilcoxon test were used to compare the distribution of continuous variables. For variables that did not follow a normal distribution, the median and quantile values were compared. The chi-square test was used to quantify the relationship between categorical variables (e.g., label balance between the training and validation sets). The Multivariate Imputation by Chained Equations (MICE) package was used for missing value imputation. The roc.test() function in the ‘pROC’ package was used to compare the two Receiver Operator Characteristic (ROC) curves on the training and validation sets. Statistical significance was set at p = 0.05. Statistical analyses were performed using R version 4.0.1 (R Foundation for Statistical Computing, Vienna, Austria).
Results
Patient characteristics
Data from the 230 patients (181 male and 49 female) who underwent UCBT between February 2014 and December 2021 were included in this study. Patient demographics and UCBT characteristics are shown in Table 1. The median age at UCBT was 14.50 months (interquartile range [IQR], 8.71–29.18 months). Sixty-seven patients were diagnosed with CGD, 73 with VEO-IBD, 48 with SCID, and 42 with other types of IEI, including Wiskott–Aldrich syndrome (WAS), leukocyte adhesion deficiency (LAD), and hyper-IgM (HIgM) syndrome. The genetics of the diagnoses are detailed in Table S3. The engraftment rate was 81.7% (188/230). The median follow-up time was 30.17 months (IQR, 5.44–56.78 months) after UCBT. During follow up, 65 patients (28.3%) died after UCBT, resulting in an overall survival rate of 71.7% (Fig. S1). Early mortality was defined as death within 50 days after UCBT (Fig. 1). Of the 230 patients, 43 (18.7%) had early mortality due to infection (29/43), organ dysfunction (11/43), or acute GvHD (3/43).
A comparison between the early mortality group and other patients group (died after day 50 or survived) is presented in Table 2. The serum albumin, CD4-positive lymphocytes (absolute count and ratio), CD19-positive lymphocytes (absolute count), and CD3-positive lymphocytes (absolute count) were significantly lower in the early mortality patients than in intermediate to late mortality patients and survivors. Patients with early mortality showed a significant positive correlation with elevated CRP (CRP >= 8 mg/L), higher IL-6, a medical history of sepsis, and intestinal infection. Early mortality was significantly lower in patients with CGD. Insignificant differences were found in other variables between the different mortality groups. A comparison of clinical variables between the training and validation sets is provided in Table S2.
Feature selection
Two of the 49 variables had missing values (missing rate IL6: 11/230 4.78%; ferritin: 26/230 11.3%). After missing value imputation, no significant outliers or perfect collinearity were found in the feature set, and the correlation matrix by heatmap is shown in Fig. S2. Next, we calculated the quantitative ensemble importance for each variable in the training set using the ensemble feature selection algorithm (Fig. S3) and all candidate variables were ranked based on their importance. In a step-wise approach, we added features from the top-ranked variables until Akaike information criterion (AIC) of the Linear Regression model fitting would not decrease (Table S4). Four variables (Alb, cd4_abs [pretransplant CD4 count], elevated CRP and sepsis) were selected. We did not select candidate variables based on univariate analysis.
Development and validation of the early mortality-predicting models
There was no statistical difference in early mortality incidence between the training and validation sets (19.74% vs. 16.67%, p = 0.699). The multivariate LR analysis with Alb, cd4_abs, elevatedCRP, and sepsis performed better than the other models in the validation set, with an AUC of 0.7385 (95% CI, 0.5824–0.8945) (Fig. 2A). The bootstrap-corrected AUC was 0.8052 (Table S4). In the validation cohort, the sensitivity, specificity, PPV, and NPV for differentiating early mortality were 0.5385, 0.8154, 0.3684, and 0.8983, respectively (Table 3). A comprehensive comparison of the model performance is presented in Table S5.
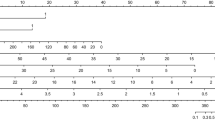
Nomogram for predicting early mortality and model evaluation. A. Receiver operating characteristic (ROC) curves for evaluating the model’s discrimination ability in validation. Panel A shows the ROC curve for the Four-Var LR model, the area under curve (AUC) is 0.7385 (95% confidence interval [CI], 0.5824–0.8945). B. Nomogram. The nomogram was developed in the derivation cohort with pretransplant albumin (Alb), pretransplant absolute CD4 (cd4_abs), elevated CRP, and medical history of sepsis (sepsis). Instructions on using nomogram: first, find the position of each variable on the corresponding axis; then, draw a line to the points axis for the number of points and add the points from all variables; and finally, draw a line from the total points axis to the lower line of the nomogram for determining the early mortality probabilities. C. Decision curve analysis (DCA) in the validation set. D. Comparison of survival curves (event: early mortality) in different risk groups by the risk estimation nomogram (p = 0.0041)
A nomogram was developed based on the best model (Fig. 2B). The DCA results for the model in the validation set are presented in Fig. 2C. The DCA results suggest that the machine learning-based prediction model had a net benefit compared with the “treat all” and “treat none” strategies for thresholds above 0.1.
Furthermore, a risk stratification based on the early mortality probability of each patient predicted by the nomogram was performed to divide all patients in the temporal validation cohort into two prognostic groups: the high-risk group (19/78, 7 early mortality, predicted probability >= 0.2) and the low-risk group (59/78, 6 early mortality, predicted probability < 0.2) and their survival curves were compared using the Kaplan-Meier method. The results showed that the difference was statistically significant (p = 0.0041) (Fig. 2D).
Discussion
Owing to the rarity of IEI and the difficulty in predicting the prognosis of transplantation, few studies have comprehensively investigated patients with IEI and early mortality after HSCT. Existing models were developed using patients with malignancy [13, 26, 27]. However, IEI patients are fundamentally different from patients with malignancy. The comorbidity index for hematopoietic cell transplantation has been validated and can distinguish between the risk of death after HSCT for patients with nonmalignant disease [28]. Nevertheless, it was not specific for predicting the prognosis of pediatric patients with IEI who underwent UCBT. Thus, a model that can predict early mortality after UCBT in children with IEI is urgently needed. To the best of our knowledge, this is the first study to develop such a model. Here, we included 230 pediatric patients with IEI who underwent UCBT to demonstrate the pattern of early mortality and proved that early mortality can be predicted using a machine learning approach.
Compared with bone marrow and peripheral blood stem cells, UCB has the advantages of rapid availability, less stringent HLA matching, lower risk of viral infection transmission, more versatile transplant planning, and no risk of donor refusal [29]. UCBT is a suitable option for patients without an available matching (related or unrelated) donor, especially for patients who need an urgent transplant (e.g., patients with SCID) and pediatric patients with low body weight. However, the disadvantages of UCBT, such as slower engraftment, graft failure, delay in immune reconstitution due to limited stem cell dose, and the associated significantly increased risk of infection, should not be underestimated [29]. Specifically, patients with IEI usually face the heavy burden of infection and inflammation. Given the limited cell dose of UCB and the underlying IEI disease, the risk of infection and organ dysfunction (the major cause of early mortality) after transplantation is of particular concern. Thus, there is a need to create and validate a model that can predict early mortality after UCBT in patients with IEI that can help clinicians make better-informed decisions considering the risk-benefit ratio.
In our study, the cumulative density plot of time from transplant to the last follow-up or death in all patients and in the subgroup of patients who died during the follow-up period showed that the majority (43/65, 66.2%) of mortality in pediatric patients with IEI after UCBT occurred within 50 days of transplantation. All-cause mortality is more heterogeneous (in terms of causes of death) than early mortality, which may explain the difficulty in predicting mortality after HSCT. Our results showed that the main causes of early mortality were infections, organ dysfunction, or acute GvHD. We further compared the interval between transplantation and the last follow-up among patients with different causes of death and found insignificant differences (Fig. S4). Given the dominance and relative homogeneity of early mortality, we chose “early mortality” as the outcome of interest.
To date, a consensus on the cutoff point for early mortality has not been reached. Some studies reported the cutoff point as 30–100 days in adult patients [9, 11, 12], and early mortality was defined as death during hospitalization for HSCT [13]. However, in our data, the cumulative density plot of the interval between UCBT and the last follow-up demonstrated a sudden change in the slope at the 50-day cutoff point. Thus, we defined 50 days as a cutoff point to distinguish patients with early mortality from patients who died after 50 days or survivors.
To account for the heterogeneity of this IEI cohort with multiple disease subtypes, we adopted a one-hot encoding approach, i.e., the presence or absence of a particular IEI subtype considered during modeling. After the modeling process was completed, we attempted to add the disease information (one-hot encoded disease variable) back to the best-performing model and observed that the model performance did not improve in either the training or validation set (Table S6). This indicated that the selected variables were informative and that disease heterogeneity was represented by these selected variables (cd4_abs, Alb, elevatedCRP, and sepsis). To further verify the disease heterogeneity represented by the selected variables, we used the selected variables to depict different disease subtypes; the risk profile in the radar plot indicated that the selected variables constitute a good representation of disease heterogeneity (Fig. 3). Overall, these findings highlight that the prediction model could estimate the risk of early mortality independently of the underlying disease.
Risk profile for different IEI subtypes represented by the four selected variables. The mean value of each variable was first calculated across patients in each IEI subtype (for binary variables [i.e., sepsis, elevated CRP], mean values equal to its positive rate); each variable has three disease-specific mean values, these summaries are then rescaled to the [0, 1] interval
In the risk estimation nomogram, pretransplant serum albumin was negatively correlated with the risk of early mortality after UCBT. Serum albumin is a biomarker of nutrition, inflammation, and hepatic synthetic function [30]. Several studies have shown that pretransplant serum albumin is an important prognostic marker of autologous stem cell transplantation and allo-HSCT [31-33]. Patients with IEI are at risk of recurrent infections, inflammatory diseases, growth retardation and failure to thrive resulting in hypoalbuminemia. Furthermore, hypoalbuminemia is associated with poor nutritional status, severe disease burden, repeated exposure to therapeutic drugs, and decreased tolerance to conditioning regimens, which could contribute to early mortality after UCBT. Our study highlights the predictive value of pretransplant serum albumin level in patients with IEI who underwent UCBT. Moreover, serum albumin detection is inexpensive and practical, making it an ideal biomarker for clinical decision-making, such as aggressive nutritional intervention and effective infection control before UCBT.
Infection and inflammation at transplant are critical risk factors that affect HSCT outcomes [34]. Despite optimal intervention (e.g., protective isolation, antibiotics, immunoglobulin replacement, and targeted agents) before transplantation, infection and inflammation cannot be completely relieved at transplantation due to the nature of IEI. In this study, acute phase reactant proteins (e.g., CRP, ferritin) and cytokine (e.g., IL-6), as objective inflammatory biomarkers, were used to reflect the status of infection and inflammation and we found that elevated CRP was a significant risk factor for post-UCBT early mortality in patients with IEI. Since elevated CRP is associated with active inflammation and previous studies showed that control of inflammation activity is the key to successful outcome [34], patients with elevated CRP prior to transplant should proceed with caution or be under intensive monitoring after UCBT.
In our study, a low absolute count of CD4-positive lymphocytes was associated with a high risk of early mortality. CD4-positive lymphocytes play an important role in adaptive immunity. The deficiency of immune defense and surveillance caused by CD4-positive lymphocytopenia can provoke fatal opportunistic infections and lead to malignancy [35, 36]. In patients with IEI, a decrease in CD4-positive lymphocytes is often closely associated with combined immunodeficiency diseases, especially SCID [37-40]. Even when we excluded SCID patients (N = 48) characterized by low CD4 levels from our cohort, the absolute count of CD4-positive lymphocytes in the early mortality group remained significantly lower than that in the other patients group (p = 0.03). Thus, our study highlights the association between lower CD4 levels and early mortality, and clinicians should be alert to patients whose CD4-positive lymphocyte count decrease before UCBT.
Our result also showed that a history of sepsis was positively associated with early mortality after UCBT in patients with IEI. Few studies have shown the association between a history of sepsis and HSCT prognosis [41]. Sepsis is a life-threatening condition caused by a dysregulated host response to infection, and it remains a leading cause of morbidity and mortality in pediatric patients worldwide [42, 43]. Virtually all tissues and organs can undergo dysfunction following sepsis [44]. Although sepsis was under control in our cohort before transplantation, patients with a history of sepsis were more likely to experience early mortality. We speculate that the long-term consequences of sepsis had a negative impact on UCBT prognosis. Our results highlight the need to identify the history of sepsis as a risk factor for an unfavorable prognosis after UCBT in patients with IEI since patients who undergo UCBT before the onset of sepsis may have better outcomes.
CD34-positive cell dose and HLA compatibility are important markers for HSCT outcomes. A sufficient number of CD34-positive stem cells is the key to rapid and durable engraftment of donor cells [45-49]. HLA-locus mismatch increases the risk of graft failure, GvHD, and mortality [50]. However, in the present study, none of these factors was selected by feature selection, and therefore, were omitted from the final model. Furthermore, no significant difference in CD34-positive stem cell dose and HLA compatibility was observed between patients with early mortality and other patients (Table 2). Since all patients included in this study were pediatric, the CD34 counts of this cohort were rather high in nearly all the patients (225/230), thus its impact may not be obvious. Similarly, due to the availability of high-resolution typing and stringent donor selection, HLA typing was a relatively good match (8–9/10) for UCBT, and its impact will also be less important in this setting. Furthermore, the reduced impact can also be attributed to the biological characteristics of UCB - high proliferative capacity and high immune plasticity, which allowed engraftment despite a 1-log lower cell number and a wider HLA disparity between donor and recipient [51, 52]. Moreover, with the improvement of HSCT technology, such as the intensity of the conditioning regimen and GvHD prophylaxis, a limited stem cell count or an HLA-related barrier no longer has an independent impact on the outcomes of patients with IEI.
The findings of the present study have several clinical implications. First, our model yielded a good AUC, indicating a good discriminatory ability. The DCA of the validation set revealed that the model provided the highest overall net benefit when the probability threshold was approximately 0.2. At this threshold probability, sensitivity and specificity were 0.5385 and 0.8154, respectively. Second, the risk estimate provided a net benefit across reasonable probability thresholds (0.1–0.5) using DCA. Clinicians have the flexibility to adjust the threshold for better sensitivity (i.e., a lower risk threshold) or better specificity (i.e., a higher risk threshold) in a case-by-case manner. Third, the nomogram developed in this study can be used to facilitate clinical decision-making and determine the predicted risk to optimize patient treatment or determine the optimal time frame for transplantation. Clinicians can use this model to modify treatment options (e.g., gene therapy and clinical trials). Furthermore, patients and their families can make informed decisions regarding predicted risk, in cases where the risk level of early mortality cannot be altered.
Limitations
This study had several limitations. First, the analysis was based on patient data from a single center, which may be influenced by selection bias. The selection bias may also induce lack of heterogeneity in some variables, including BU-based conditioning, degree of HLA match, and CD34 count, which are important for transplant outcomes. The lack of heterogeneity limits the utility of these variables in the early mortality risk model. A multicenter study and external validation are needed to confirm our results. Second, due to the long-term nature of the study, we could not find or access the information needed to determine ‘performance status’, so we had to exclude it from the list of potential predictors when designing the study. However, by including variables that can be objectively measured, we ensured the reliability and accuracy of the variables included. Third, the present study was retrospective in nature. A prospective study is required to confirm the usefulness of the nomogram developed in this study. Nonetheless, the large dataset in the analysis, the temporal validation approach, the ensemble feature selection, and the bootstrap validation greatly improved the validity and robustness of the final model.
Conclusions
Using real-world data from pediatric IEI patients and data-driven analysis, we developed a nomogram that can predict early mortality after UCBT based on four pretransplant risk factors. The nomogram performed well in training and validation sets. The personalized prediction model based on machine learning can improve decision-making and outcomes for patients with IEI who have undergone UCBT.
Data Availability
The datasets generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Laberko A, Gennery AR. Clinical considerations in the hematopoietic stem cell transplant management of primary immunodeficiencies. Expert Rev Clin Immunol. 2018;14(4):297–306. https://doi.org/10.1080/1744666x.2018.1459189.
Castagnoli R, Delmonte OM, Calzoni E, Notarangelo LD. Hematopoietic Stem Cell Transplantation in Primary Immunodeficiency Diseases: Current Status and Future Perspectives. Front Pediatr. 2019;7:295. https://doi.org/10.3389/fped.2019.00295.
Pai S-Y, Logan BR, Griffith LM, Buckley RH, Parrott RE, Dvorak CC, et al. Transplantation Outcomes for Severe Combined Immunodeficiency, 2000-2009. N Engl J Med. 2014;371(5):434–46. https://doi.org/10.1056/NEJMoa1401177.
Chiesa R, Wang J, Blok H-J, Hazelaar S, Neven B, Moshous D, et al. Hematopoietic cell transplantation in chronic granulomatous disease: a study of 712 children and adults. Blood. 2020;136(10):1201–11. https://doi.org/10.1182/blood.2020005590.
Burroughs LM, Petrovic A, Brazauskas R, Liu X, Griffith LM, Ochs HD, et al. Excellent outcomes following hematopoietic cell transplantation for Wiskott-Aldrich syndrome: a PIDTC report. Blood. 2020;135(23):2094–105. https://doi.org/10.1182/blood.2019002939.
Gragert L, Eapen M, Williams E, Freeman J, Spellman S, Baitty R. HLA match likelihoods for unrelated donor grafts in the US registry. N Engl J Med. 2014;371(4):339–48. https://doi.org/10.1056/NEJMsa1311707.
Spees LP, Martin PL, Kurtzberg J, Stokhuyzen A, McGill L, Prasad VK, et al. Reduction in mortality after umbilical cord blood transplantation in children over a 20-year period (1995-2014). Biol Blood Marrow Transplant. 2019;25(4):756–63. https://doi.org/10.1016/j.bbmt.2018.11.018.
Ballen KK, Gluckman E, Broxmeyer HE. Umbilical cord blood transplantation: the first 25 years and beyond. Blood. 2013;122(4):491–8. https://doi.org/10.1182/blood-2013-02-453175.
Styczynski J, Tridello G, Koster L, Iacobelli S, van Biezen A, van der Werf S, et al. Death after hematopoietic stem cell transplantation: changes over calendar year time, infections and associated factors. Bone Marrow Transplant. 2020;55(1):126–136. https://doi.org/10.1038/s41409-019-0624-z.
Bohannon LM, Page KM, Ren Y, Jung S-H, Giri VK, Lew MV, et al. Decreased mortality after the first year of allogeneic hematopoietic stem cell transplant in recipients of umbilical cord blood vs. matched related or matched unrelated donors. Blood. 2019;134:4613. https://doi.org/10.1182/blood-2019-126906.
Kong SG, Jeong S, Lee S, Jeong J-Y, Kim DJ, Lee HS. Early transplantation-related mortality after allogeneic hematopoietic cell transplantation in patients with acute leukemia. BMC Cancer. 2021;21(1):177. https://doi.org/10.1186/s12885-021-07897-3.
Iioka F, Maesako Y, Nakamura F, Hayashi T, Ohno H. Outcome of second/third allogeneic hematopoietic stem cell transplantations for a relapse of acute leukemia after the first transplantation. Tenri Medical Bulletin. 2013;16(1):17–24. https://doi.org/10.12936/tenrikiyo.16-001.
Godara A, Siddiqui NS, Holtzman NG, Munigala S, Kansagra AJ, Rapoport AP, et al. Factors Associated with Early Mortality in Patients Undergoing Allogeneic Stem Cell Transplantation. Biol Blood Marrow Transplant. 2020;26(3):S44. https://doi.org/10.1016/j.bbmt.2019.12.113.
Arai Y, Kondo T, Fuse K, Shibasaki Y, Masuko M, Sugita J, et al. Using a machine learning algorithm to predict acute graft-versus-host disease following allogeneic transplantation. Blood. Advances. 2019;3(22):3626–34. https://doi.org/10.1182/bloodadvances.2019000934.
Shouval R, Labopin M, Unger R, Giebel S, Ciceri F, Schmid C, et al. Prediction of Hematopoietic Stem Cell Transplantation Related Mortality-Lessons Learned from the In-Silico Approach: A European Society for Blood and Marrow Transplantation Acute Leukemia Working Party Data Mining Study. Plos One. 2016;11(3):e0150637. https://doi.org/10.1371/journal.pone.0150637.
Radakovich N, Nagy M, Nazha A. Machine learning in haematological malignancies. Lancet Haematol. 2020;7(7):E541–E50. https://doi.org/10.1016/S2352-3026(20)30121-6.
Bousfiha A, Jeddane L, Picard C, Ailal F, Gaspar HB, Al-Herz W, et al. The 2017 IUIS Phenotypic Classification for Primary Immunodeficiencies. J Clin Immunol. 2018;38(1):129–43. https://doi.org/10.1007/s10875-017-0465-8.
Lankester AC, Albert MH, Booth C, Gennery AR, Guengoer T, Hoenig M, et al. EBMT/ESID inborn errors working party guidelines for hematopoietic stem cell transplantation for inborn errors of immunity. Bone Marrow Transplant. 2021;56(9):2052–62. https://doi.org/10.1038/s41409-021-01378-8.
Enric Carreras, Carlo Dufour, Mohamad Mohty, Nicolaus Kröger. The EBMT Handbook: Hematopoietic Stem Cell Transplantation and Cellular Therapies[M]. 2019. https://doi.org/10.1007/978-3-030-02278-5
Robert LT. Who belongs in the family? Psychometrika. 1953;18(4):267–76. https://doi.org/10.1007/BF02289263.
Nelson RJ, Huffman DA. The synthesis of sequential switching circuits. J Franklin Inst. 1955;257:161–90. https://doi.org/10.1016/0016-0032(54)90618-3.
Neumann U, Genze N, Heider D. EFS: an ensemble feature selection tool implemented as R-package and web-application. BioData Mining. 2017;10(1):1–9. https://doi.org/10.1186/s13040-017-0142-8.
Tibshirani R. Regression shrinkage and selection via the lasso. J R Stat Soc Series B Stat Methodol. 1996;58(1):267–88. https://doi.org/10.2307/2346178.
Steyerberg EW, Vergouwe Y. Towards better clinical prediction models: seven steps for development and an ABCD for validation. Eur Heart J. 2014;35(29):1925–31. https://doi.org/10.1093/eurheartj/ehu207.
Grant RC, Moineddin R, Yao Z, Powis M, Kukreti V, Krzyzanowska MK. Development and validation of a score to predict acute care use after initiation of systemic therapy for cancer. JAMA Netw Open. 2019;2(10):e1912823-e. https://doi.org/10.1001/jamanetworkopen.2019.12823.
Shouval R, Labopin M, Bondi O, Mishan Shamay H, Shimoni A, Ciceri F, et al. Prediction of allogeneic hematopoietic stem-cell transplantation mortality 100 days after transplantation using a machine learning algorithm: a European Group for Blood and Marrow Transplantation Acute Leukemia Working Party retrospective data mining study. J Clin Oncol. 2015;33:3144–51. https://doi.org/10.1200/JCO.2014.59.1339.
Parimon T, Au DH, Martin PJ, Chien JW. A risk score for mortality after allogeneic hematopoietic cell transplantation. Ann Intern Med. 2006;144(6):407–14. https://doi.org/10.7326/0003-4819-144-6-200603210-00007.
Thakar MS, Broglie L, Logan B, Artz A, Bunin N, Burroughs LM, et al. The Hematopoietic Cell Transplant Comorbidity Index predicts survival after allogeneic transplant for nonmalignant diseases. Blood. 2019;133(7):754–62. https://doi.org/10.1182/blood-2018-09-876284.
Zhu X, Tang B, Sun Z. Umbilical cord blood transplantation: Still growing and improving. Stem Cells Transl Med. 2021;10(S2):S62–74. https://doi.org/10.1002/sctm.20-0495.
Fanali G, Di Masi A, Trezza V, Marino M, Fasano M, Ascenzi P. Human serum albumin: from bench to bedside. Mol Aspects Med. 2012;33(3):209–90. https://doi.org/10.1016/j.mam.2011.12.002.
Artz AS, Logan B, Zhu X, Akpek G, Bufarull RM, Gupta V, et al. The prognostic value of serum C-reactive protein, ferritin, and albumin prior to allogeneic transplantation for acute myeloid leukemia and myelodysplastic syndromes. Haematologica. 2016;101(11):1426. https://doi.org/10.3324/haematol.2016.145847.
Luo C, Li Q, Li X, Wu G, Huang X, Zhang Y, et al. Prognostic Role of Serum Albumin Level in Patients with Lymphoma Undergoing Autologous Stem Cell Transplantation. Ann Transplant. 2021;26:e933365–1. https://doi.org/10.12659/AOT.933365.
Vaughn JE, Storer BE, Armand P, Raimondi R, Gibson C, Rambaldi A, et al. Design and validation of an augmented hematopoietic cell transplantation-comorbidity index comprising pretransplant ferritin, albumin, and platelet count for prediction of outcomes after allogeneic transplantation. Biol Blood Marrow Transplant. 2015;21(8):1418–24. https://doi.org/10.1016/j.bbmt.2015.04.002.
Shamriz O, Chandrakasan S. Update on advances in hematopoietic cell transplantation for primary immunodeficiency disorders. Immunol Allergy Clin North Am. 2019;39(1):113–28. https://doi.org/10.1016/j.iac.2018.08.003.
Zonios DI, Falloon J, Bennett JE, Shaw PA, Chaitt D, Baseler MW, et al. Idiopathic CD4+ lymphocytopenia: natural history and prognostic factors. Blood. 2008;112(2):287–94. https://doi.org/10.1182/blood-2007-12-127878.
Ahmad DS, Esmadi M, Steinmann WC. Idiopathic CD4 Lymphocytopenia: Spectrum of opportunistic infections, malignancies, and autoimmune diseases. Avicenna J Med. 2013;3(02):37–47. https://doi.org/10.4103/2231-0770.114121.
Aranda CS, Guimaraes RR, Pimentel MG-P. Combined immunodeficiencies. Jornal De. Pediatria. 2021;97:S39–48. https://doi.org/10.1016/j.jped.2020.10.014.
Mandola AB, Sharfe N, Nagdi Z, Dadi H, Vong L, Merico D, et al. Combined immunodeficiency caused by a novel homozygous NFKB1 mutation. J Allergy Clin Immunol. 2021;147(2):727. https://doi.org/10.1016/j.jaci.2020.08.040.
Ban SA, Salzer E, Eibl MM, Linder A, Geier CB, Santos-Valente E, et al. Combined Immunodeficiency Evolving into Predominant CD4+Lymphopenia Caused by Somatic Chimerism in JAK3. J Clin Immunol. 2014;34(8):941–53. https://doi.org/10.1007/s10875-014-0088-2.
Chinn IK, Shearer WT. Severe Combined Immunodeficiency Disorders. Immunol Allergy Clin North Am. 2015;35(4):671–94. https://doi.org/10.1016/j.iac.2015.07.002.
Gennery AR, Slatter MA, Grandin L, Taupin P, Cant AJ, Veys P, et al. Transplantation of hematopoietic stem cells and long-term survival for primary immunodeficiencies in Europe: entering a new century, do we do better? J Allergy Clin Immunol. 2010;126(3):602–10. e11. https://doi.org/10.1016/j.jaci.2010.06.015.
Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, et al. The third international consensus definitions for sepsis and septic shock (Sepsis-3). Jama. 2016;315(8):801–10. https://doi.org/10.1001/jama.2016.0287.
Goldstein B, Giroir B, Randolph A. International pediatric sepsis consensus conference: definitions for sepsis and organ dysfunction in pediatrics. Pediatr Crit Care Med. 2005;6(1):2–8. https://doi.org/10.1097/01.PCC.0000149131.72248.E6.
Coopersmith CM, De Backer D, Deutschman CS, Ferrer R, Lat I, Machado FR, et al. Surviving sepsis campaign: research priorities for sepsis and septic shock. Intensive Care Med. 2018;44(9):1400–26. https://doi.org/10.1007/s00134-018-5175-z.
Kamel AM, El-Sharkawy N, Mahmoud HK, Khalaf MR, El Haddad A, Fahmy O, et al. Impact of CD34 subsets on engraftment kinetics in allogeneic peripheral blood stem cell transplantation. Bone Marrow Transplant. 2005;35(2):129–36. https://doi.org/10.1038/sj.bmt.1704755.
Gupta AO, Wagner JE. Umbilical Cord Blood Transplants: Current Status and Evolving Therapies. Front Pediatr. 2020;8:570282. https://doi.org/10.3389/fped.2020.570282.
Maffini E, Labopin M, Blaise D, Ciceri F, Gulbas Z, Deconinck E, et al. CD34+cell dose effects on clinical outcomes after T-cell replete haploidentical allogeneic hematopoietic stem cell transplantation for acute myeloid leukemia using peripheral blood stem cells. A study from the acute leukemia working Party of the European Society for blood and marrow transplantation (EBMT). Am J Hematol. 2020;95(8):892–9. https://doi.org/10.1002/ajh.25826.
Yokoyama Y, Maie K, Fukuda T, Uchida N, Mukae J, Sawa M, et al. A high CD34(+) cell dose is associated with better disease-free survival in patients with low-risk diseases undergoing peripheral blood stem cell transplantation from HLA-matched related donors. Bone Marrow Transplant. 2020;55(9):1726–35. https://doi.org/10.1038/s41409-020-0817-5.
Remberger M, Torlen J, Ringden E, Engstrom M, Watz E, Uhlin M, et al. Effect of Total Nucleated and CD34(+) Cell Dose on Outcome after Allogeneic Hematopoietic Stem Cell Transplantation. Biol Blood Marrow Transplant. 2015;21(5):889–93. https://doi.org/10.1016/j.bbmt.2015.01.025.
Petersdorf EW, Optimal HLA. Matching in Haematopoietic Cell Transplantation. Curr Opin Immunol. 2008;20(5):588–93. https://doi.org/10.1016/j.coi.2008.06.014.
Goldstein G, Toren A, Nagler A. Human umbilical cord blood biology, transplantation and plasticity. Curr Med Chem. 2006;13(11):1249–59. https://doi.org/10.2174/092986706776872998.
Stanevsky A, Goldstein G, Nagler A. Umbilical cord blood transplantation: pros, cons and beyond. Blood Rev. 2009;23(5):199–204. https://doi.org/10.1016/j.blre.2009.02.001.
Acknowledgments
We thank our patients and their families for their participation in this study.
Funding
This study was supported by the Shanghai Municipal Committee of Science and Technology (21Y31900302) and National Multidisciplinary Cooperative Diagnosis and Treatment Capability Construction Project for Major Diseases.
Author information
Authors and Affiliations
Contributions
Xiaowen Zhai, Xiaowen Qian, Xiaochuan Wang, Xinjun He, Ping Wang and Chao Liu designed this study, reviewed and revised the manuscript. Xiaowen Qian, Xiaochuan Wang, Ying Huang, Jinqiao Sun, Jia Hou, Yuhuan Wang, Hua Sun, Wenjin Jiang and Ping Wang participated in patients’ management. Ping Wang, Zhongling Wei, Wenjin Jiang collected clinical data and followed the patients. Statistical analysis was performed by Chao Liu and Yao Wang. Ping Wang and Chao Liu interpreted data and drafted the manuscript. All authors contributed to the article and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki and approved by the ethics board of the Children’s Hospital of Fudan University (Shanghai, China). Written informed consent was obtained from the guardians of each patient.
Consent to participate
Written informed consent was obtained from patient’s guardians.
Consent to publish
Written informed consent for publication of the study was obtained from patient’s guardians. All authors approved the final manuscript.
Competing Interests
The authors have no relevant financial or non-financial interests to disclose
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 1314 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, P., Liu, C., Wei, Z. et al. Nomogram for Predicting Early Mortality after Umbilical Cord Blood Transplantation in Children with Inborn Errors of Immunity. J Clin Immunol 43, 1379–1392 (2023). https://doi.org/10.1007/s10875-023-01505-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-023-01505-8