Abstract
Conflicting data exist in rheumatoid arthritis and the collagen-induced arthritis (CIA) murine model of autoimmune arthritis regarding the role of bacterial carnitine and choline metabolism into the inflammatory product trimethylamine (TMA), which is oxidized in the liver to trimethylamine-N-oxide (TMAO). Using two published inhibitors of bacterial TMA lyase, 3,3-dimethyl-1-butanol (DMB) and fluoromethylcholine (FMC), we tested if TMA/TMAO were relevant to inflammation in the development of CIA. Surprisingly, DMB-treated mice demonstrated > 50% reduction in arthritis severity compared to FMC and vehicle-treated mice, but amelioration of disease was independent of TMA/TMAO production. Given the apparent contradiction that DMB did not inhibit TMA, we then investigated the mechanism of protection by DMB. After verifying that DMB acted independently of the intestinal microbiome, we traced the metabolism of DMB within the host and identified a novel host-derived metabolite of DMB, 3,3-dimethyl-1-butyric acid (DMBut). In vivo studies of mice treated with DMB or DMBut demonstrated efficacy of both molecules in significantly reducing disease and proinflammatory cytokines in CIA, while in vitro studies suggest these molecules may act by modulating secretion of proinflammatory cytokines from macrophages. Altogether, our study suggests that DMB and/or its metabolites are protective in CIA through direct immunomodulatory effects rather than inhibition of bacterial TMA lyases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Rheumatoid arthritis (RA) is an autoimmune disease characterized by joint inflammation and destruction and exhibits a worldwide prevalence of 0.8–1.0% [1]. In spite of significant advancements for the treatment of RA, current therapies focus on managing symptoms and slowing disease progression [2]. Unfortunately, 43% of patients do not respond to first line therapy with methotrexate and require more aggressive therapy with biologic medications [3]. Overall, only 60–70% of patients with RA sufficiently respond to therapy within the first six months of administration [4]; after about two years ~ 50% discontinue therapy due to loss of efficacy (35%) or safety (20%), among other issues [5]. Thus, significant limitations in efficacy and safety remain, highlighting the need for improved therapeutics.
A novel therapeutic target for the treatment of RA stems from observations that the initial autoimmunity characteristic of RA may originate at mucosal surfaces [6]. Initial hypotheses for the development of this autoimmunity centered on a microbial-triggered process and later a microbial antigen cross-reactive with self. More recently, specific microbiota in the periodontium and intestine have been associated with RA, either by induction of autoantibody generation or by changes in microbial community composition [7, 8]. Further, animal models suggest that the presence of specific microbiota is necessary for the development of autoimmune arthritis. For example, utilizing the collagen-induced arthritis (CIA) model, mice given broad spectrum antibiotics to deplete their gut microbiomes have reduced intestinal IL-17 and IL-22, as well as reduced circulating IL-6, IL-1β and TNF-α [9]; similar dependencies of the gut microbiome on disease development are observed in the K/BxN and SKG models [10, 11]. These findings suggest that the composition of the gut microbiome and its effects can have an immunomodulatory role in RA pathogenesis. However, the mechanisms of how microbial dysbiosis can promote RA development, and how it may be manipulated for therapy, are not understood.
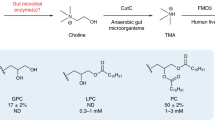
The microbiome can influence RA through the effect of bacterially-derived metabolites acting as potent immune modulators. Metabolic assessments of plasma from individuals at-risk for or with RA associate alterations in concentrations of numerous carnitines and choline with the future development or presence, respectively, of RA [12, 13]. Trimethylamine (TMA) is a bacterial product of dietary carnitine and choline. Some expanded populations of gut bacteria in RA, including Prevotella and Collinsella, express TMA lyases that are capable of metabolizing dietary choline and carnitine into cytotoxic TMA, which is subsequently absorbed into the host and oxidized primarily by hepatic flavin monooxygenase 3 (FMO3) into non-cytotoxic trimethylamine-N-oxide (TMAO) [7, 14, 15]. Although much of the TMAO produced is excreted in the urine within 24 h [16, 17] accumulation of TMAO in human plasma is associated with inflammation concomitant to kidney disease, heart failure, and cardiovascular disease via NLRP3 inflammasome activation and NF-kB signaling [18]. Notably, serum TMAO levels are significantly elevated in patients with psoriatic arthritis (PsA), and serum TMAO is significantly correlated with the number of swollen joints, disease severity scores, and serum CRP concentration [19]. Moreover, rat bone marrow mesenchymal stem cells cultured with TMAO showed attenuated osteogenesis and increased secretion of IL-1β, IL-6, TNF-α, and reactive oxygen species via activation of NF-kB in these cells [20]. In a rat model of osteoarthritis (OA), treatment with TMAO induced synovitis and increased IL-6 production via elevated expression of Piezo1 in chondrocytes, thereby making these cells susceptible to mechanical injury [21]. However, one study profiling urinary metabolites in RA found elevated TMAO, but statistical significance was achieved only after removal of outliers, and urinary TMAO correlated with a modest reduction in CRP, counterintuitive to the previous reports regarding the actions of TMAO [22]. In the same study, mice with CIA fed a 1% choline diet as a source of TMA/TMAO showed delayed disease onset and mildly reduced severity compared to mice given normal chow [22]. Critically, TMAO production from the increased choline consumption was not assessed in this study, and choline itself can act in an anti-inflammatory capacity in both RA and CIA through its involvement in the cholinergic anti-inflammatory pathway [23]. Therefore, herein we sought to resolve the discrepancy in the role of TMAO between these data in RA and CIA with that from PsA and OA.
Although FMO3 is not a viable drug target due to risk of drug toxicity, decreased TMAO generation has been reported to be achieved through inhibition of bacterial TMA lyase [24]. Previous studies using the apolipoprotein E (ApoE) knockout (ApoE−/−) model of atherosclerosis demonstrate that inhibition of TMA lyase via oral administration of the choline structural analogs 3,3-dimethyl-1-butanol (DMB) or fluoromethylcholine (FMC) is beneficial, such that mice given these inhibitors have reduced circulating TMAO levels coupled with reductions in endogenous macrophage foam cell and atherosclerotic lesion formations [24, 25]. Therefore, in this study, we utilized the CIA mouse model of autoimmune arthritis to test whether TMA/TMAO is a mediator in the development of inflammation and arthritis. After confirming our hypothesis was null (TMA/TMAO was not a mediator of CIA), but DMB was an effective therapeutic, we then set out to identify the mechanism of action for DMB.
RESULTS
DMB but not FMC Reduces CIA Severity Independent of TMA-Lyase Inhibition
Because TMA lyase inhibition using first (DMB) and second-generation (FMC) small molecule inhibitors (Fig. 1a) was effective in ameliorating the inflammatory disease processes underlying the ApoE−/− mouse model of atherosclerosis [24, 25], we questioned if TMA lyase inhibition using these same inhibitors would also be beneficial in another inflammatory disease model, CIA. Therefore, we induced CIA in male 6-week-old DBA/1j mice. At day 21, mice were treated with vehicle, 1% vol/vol DMB in drinking water, or 100 mg/kg FMC via oral gavage. These doses and routes of administration were chosen as they have been reported previously to effectively inhibit TMA generation in mice [24, 25]. Mice were monitored for arthritis until day 35 using the CIA scoring scale described in Methods. Mice given DMB exhibited significantly lower arthritis scores and a delayed incidence of disease compared to vehicle- and FMC-treated mice (Fig. 1b and c). Surprisingly, mice given FMC showed an increase in disease severity compared to vehicle-treated mice (Fig. 1b).
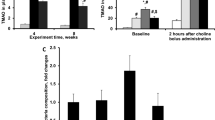
DMB, but not FMC, reduces CIA severity. a Chemical structures of the TMA lyase inhibitors used in this study. b CIA was induced in 6-week-old male DBA/1j mice. On day 21 post-initial immunization, mice were treated with either vehicle (CIA + Vehicle), 1% (v/v) DMB in drinking water (CIA + DMB), or with 100 mg/kg FMC via oral gavage (CIA + FMC). N = 14 (CIA), N = 14 (CIA + DMB), and N = 5 (CIA + FMC) per group from three pooled experiments. Data are reported as mean ± SEM. *, p < 0.05; **, p < 0.01; ns, non-significant as determined by two-way ANOVA with Bonferroni correction for multiple comparisons. c Arthritis incidence was calculated by dividing the number of mice showing clinical evidence of arthritis (CIA score ≥ 1) by the total number of mice per group. Differences between groups were non-significant as determined by chi-square test. d-e Cecal TMA d and serum TMAO e concentrations were measured via UHPLC-MS/MS using tissues harvested from mice at day 35 post-initial immunization. N = 14 (CIA + Vehicle), N = 14 (CIA + DMB), and N = 5 (CIA + FMC) per group and pooled from three independent experiments. Data are reported as fold change relative to TMA/TMAO median peak area in the CIA + Vehicle group (symbols) and group mean ± SEM (bars). *, p < 0.05; ****, p < 0.0001; ns, non-significant as determined by one-way ANOVA with Tukey’s correction for multiple comparisons.
To determine if DMB and FMC inhibited TMA lyase, ceca and sera were collected from mice at day 35 post-initial immunization and subjected to UPLC-MS/MS analysis for detection of TMA and TMAO. Only FMC treatment resulted in a significant reduction in cecal TMA levels compared to vehicle and DMB-treated mice, while serum TMAO was significantly reduced in FMC-treated mice compared to those given DMB. Moreover, mice given DMB displayed no reduction in TMA or TMAO relative to vehicle-treated mice (Fig. 1d and e).
Since DMB did not reduce TMA/TMAO production in vivo, we examined the ability of DMB to block the TMA lyase of Proteus mirabilis in vitro, which has been previously shown to block conversion of choline to TMA [24, 25]. Whole-cell P. mirabilis (ATCC 29906) were cultured with d9-choline and confirmed to produce d9-TMA in a dose-dependent manner (Supplemental Fig. 1a). Subsequently the cells were cultured with a dilution series of DMB (10−12 M–100 M) or FMC (10–12–10–3) for 2 h. LC–MS/MS analysis of the culture supernatants for production of d9-TMA showed a dose-dependent reduction in d9-TMA production only in the presence of concentrations of DMB greater than 10 mM (Supplemental Fig. 1b). Critically, the concentration of DMB administered to mice during in vivo experiments is about 83 μM. In line with previous reports, FMC reduced d9-TMA in a dose-dependent manner, with almost complete ablation of d9-TMA production at concentrations greater than 1 pM (Supplemental Fig. 1b). These data suggest that DMB does not universally inhibit TMA lyase.
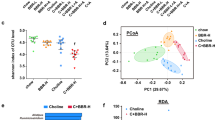
We next tested the possibility that DMB may impact the development of CIA via changes in the intestinal microbiome. Fecal pellets were harvested at day 35 and analyzed by 16S ribosomal RNA gene sequencing. Measures of alpha diversity indicated that DMB treatment significantly altered microbial evenness (ShannonE), but not richness (Chao1) or diversity (Simpson) compared to vehicle but not FMC-treated mice with CIA (Fig. 2a-c). Further, Bray–Curtis distance analysis of the bacterial communities revealed significant differences in beta diversity among all treatment groups; however, the microbial communities of each treatment group do not substantially separate (Fig. 2d). Non-FDR-corrected two-way ANOVA comparisons of bacterial taxa identified significant expansions of S24-7 and family Ruminococcaceae as well as significant depletions of Firmicutes, Bacilli, Lactobacialles, Lactobacillus, Clostridiales, and Lachnospiraceae in DMB-treated mice compared to vehicle-treated mice. Significant expansions of Clostridiales and Ruminococcaceae, and significant depletions of Bacilli, Lactobacialles,and Lactobacillus were also present in DMB-treated mice compared to FMC-treated mice (Supplemental Fig. 2a). Subsequent linear discriminant effect size analysis with FDR correction, however, identified only a significant expansion of specific taxa driven by S24-7 family bacteria in mice treated with DMB compared to vehicle (Fig. 2e) and no significantly altered bacterial taxa between DMB and FMC-treated mice (data not shown). Thus, because DMB and FMC-treated mice minimally differed in their microbiomes despite significant differences in disease activity, we concluded that the effect of DMB was likely independent of the microbiome.
DMB treatment does not confer substantial alterations in the host gut microbiome. Fecal pellets from DBA/1j mice were harvested at day 35 post-initial immunization and underwent genomic DNA extraction and16S ribosomal RNA gene sequencing to assess microbial diversity. N = 9 (CIA + vehicle), N = 9 (CIA + DMB), N = 5 (CIA + FMC) per group and pooled from two independent experiments. a-c Alpha diversity analyses of bacterial taxa. Data are reported as box and whisker plots with 5–95% confidence intervals of mean index values. *, p < 0.05; ns, non-significant as determined by one-way ANOVA with Tukey’s correction for multiple comparisons. d PCoA plot showing the beta diversity of microbiota from mice in each treatment group as determined by Bray–Curtis Index distance. Data are shown as individual mice (symbols) with 95% confidence intervals (ellipses). F-value = 4.2063; R.2 = 0.29608; p-value = 0.008 as determined by PERMANOVA between the three treatment groups. e Linear discriminant effect size analysis of OTU counts from mice treated with vehicle (CIA + vehicle) or DMB (CIA + DMB) using FDR < 0.1, LDA score > 2.0, and FDR-corrected p-value < 0.05.
DMB is Absorbed and Metabolized in Mice
Next, we examined the host metabolism of DMB. GC/MS analysis was used to examine the presence of DMB through identification of three ion peaks corresponding to DMB (Supplemental Fig. 3a). Indeed, mice given DMB had detectable and significantly greater concentrations of DMB in their serum compared to vehicle-treated mice, suggesting absorption into the host (Fig. 3a). Since DMB was administered orally, we hypothesized that DMB is subject to first-pass metabolism in the liver. DMB was detected at significantly higher concentrations in the livers of DMB-treated mice compared to untreated mice (Fig. 3b). Further, previous studies have shown that DMB is a substrate for alcohol dehydrogenase 1 (ADH1) [24]; however, the aldehyde product of DMB generated by ADH1 was undetectable by LC–MS in the livers of DMB-treated mice (data not shown). Instead, the downstream acylcarnitine metabolite, 3,3-dimethylbutyrylcarnitine (DMBC), was detected by LC–MS in the serum and liver (Fig. 3c and d), providing evidence for rapid metabolism of the DMB aldehyde to its carboxylic acid conjugate within the host.
DMB is absorbed into the host and metabolized to its fatty acid and acylcarnitine conjugates. a-d Serum (a, c) and liver (b, d) from male DBA1/j mice was harvested at day 35 post-initial immunization and subjected to GC–MS for detection of DMB (a-b) and LC–MS for detection of DMBC (c-d). N = 4 (CIA + vehicle) and N = 4 (CIA + DMB) per group from one representative experiment. Data are reported as mean ± SEM. *, p < 0.05 as determined by Mann–Whitney non-parametric t-test. e Hypothesized metabolism of DMB in the liver after it is absorbed into the host. f-g Serum (f) and liver (g) was subjected to GC–MS for detection of DMBut. N = 4 per group from one representative experiment. Data are reported as mean ± SEM. *, p < 0.05 as determined by Mann–Whitney non-parametric t-test.
In order to further define the metabolism of DMB by the murine host, we mapped the metabolic pathway from DMB to DMBC (Fig. 3e). In the pathway, we predicted the generation of the intermediate metabolite, 3–3-dimethyl-1-butyrate (DMBut). Therefore, we assayed the serum and liver of untreated and DMB-treated mice for the presence of DMBut, which was only present in the serum and liver of CIA mice given DMB but not vehicle-treated mice (Figs. 3f and g). We also assayed ceca from DMBut-treated mice for the presence of DMBut; however, no DMBut was detected in the ceca of mice given vehicle or DMBut (data not shown). The identification of two DMB metabolites within this pathway suggests that DMB is taken up into host tissues where it may be active in disease-modulating processes.
DMB and DMBut Administration Reduces Inflammatory Features of CIA
Like all acylcarnitines, we hypothesized that DMBC is an intermediate of DMB metabolism and formed via the esterification of carnitine with a fatty acid (DMBut) transferred from an acyl-CoA intermediate (DMBCoA). Acylcarnitines function to transport conjugated fatty acids of varying lengths across cell membranes, particularly from the cytosol into the mitochondrial matrix for energy production [26, 27]. Once the acylcarnitine translocates across the membrane and arrives at its cellular destination, the fatty acid is transferred to CoASH, reforming the acyl-CoA for subsequent utilization of the acyl groups in the fatty acid [28]. Hence, while DMBC is an intermediate downstream of DMBut, we hypothesized that DMBut is the biologically active molecule.
To test this hypothesis, we treated mice with CIA with either vehicle, 1% vol/vol DMB, or 1% vol/vol DMBut in the drinking water beginning on day 21 post-initial immunization. Mice were monitored for arthritis until day 35 using the CIA scoring scale described in Methods. Like mice given DMB, mice treated with DMBut had significantly reduced CIA scores compared to untreated mice (Fig. 4a). However, the incidence of arthritis only differed significantly between groups at 23 days post-immunization (Fig. 4b). Overall, these results suggest that both DMB and DMBut modulate the severity of inflammatory responses during CIA.
DMB and its metabolite DMBut reduce arthritis severity and proinflammatory cytokines in CIA. a CIA was induced in 6-week-old male DBA/1j mice. On day 21 post-initial immunization, mice were left treated with vehicle (CIA + Vehicle), 1% (v/v) DMB in drinking water (CIA + DMB), or 1% (v/v) DMBut in drinking water (CIA + DMBut). N = 24 (CIA + Vehicle), N = 23–25 (CIA + DMB), and N = 10 (CIA + DMBut) per group pooled from 5 independent experiments. Data are reported as mean ± SEM. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns, non-significant as determined by two-way ANOVA with Bonferroni correction for multiple comparisons. b Arthritis incidence was calculated by dividing the number of mice showing clinical evidence of arthritis (CIA score \(\ge\) 1) by the total number of mice per group. N = 24 (CIA + Vehicle), N = 23–25 (CIA + DMB), and N = 10 (CIA + DMBut) per group pooled from 5 independent experiments. *, p < 0.05; ns, non-significant as determined by chi-square test. c-d Serum was harvested from male DBA/1j mice at day 35 post-initial immunization and analyzed for proinflammatory cytokines by a 6-plex immunoassay (Mesoscale). N = 28 (CIA + Vehicle), N = 23 (CIA + DMB), and N = 14 (CIA + DMBut) per group pooled from 5 independent experiments. Data are reported as fold change normalized to the cytokine concentrations in the CIA + Vehicle group (symbols) and group mean ± SEM (bars). **, p < 0.01; ***, p < 0.001; ****, p < 0.0001 as determined by one-way ANOVA with Tukey’s correction for multiple comparisons.
To investigate how DMB and DMBut administration modulate the inflammatory response in CIA, we harvested serum from vehicle-treated, DMB-treated, or DMBut-treated mice 35 days post-initial immunization and analyzed circulating proinflammatory cytokines and collagen-type II (CII)-specific antibodies. The cytokines IFN-γ, IL-17A, IL-1β, IL-23, IL-6, and TNF were chosen for their relevance to the pathogenesis of CIA and RA [29,30,31,32]. DMB and DMBut-treated mice both had significantly reduced circulating concentrations of IL-1β and IL-6 compared to untreated controls (Figs. 4c and d). Individually, DMBut significantly reduced TNF-α and IL-23 compared to untreated mice (Supplemental Fig. 4a and b); whereas DMB treatment significantly reduced IFN-γ (Supplemental Fig. 4c). Interestingly, neither compound significantly reduced serum IL-17A compared to vehicle-treated mice (Supplemental Fig. 4d). Total anti-CII IgG was not affected by DMB or DMBut treatment, and only treatment with DMB significantly reduced the concentration of pathogenic isotype IgG2b anti-CII antibodies compared to vehicle-treated mice (Supplemental Fig. 4e and f). Thus, DMB and DMBut appear to have the most profound effect on the cytokines IL-1β and IL-6.
Butyrate is associated with the expansion of regulatory T cell subsets and modulating the balance of helper T cells and regulatory T cells, particularly the Th17/Treg ratio [33,34,35], whereas Tfh-derived IL-1β is critical for plasma cell differentiation and IL-6 induces terminal B cell differentiation via stimulation of IgD production [36, 37]. Therefore, we next sought to investigate changes in T and B cell differentiation during CIA in response to DMB treatment. Flow cytometric analysis of splenocytes for T lymphocyte populations (Supplemental Fig. 5a-e) showed that mice treated with DMB or DMBut had no changes in the percent of regulatory, Th1, follicular helper, or Th17 T lymphocytes compared to vehicle controls (Supplemental Fig. 5f-i). Within the splenic B cell compartment (Supplemental Fig. 6), flow cytometric analysis showed that treatment with DMB significantly reduced the frequency of B1, B2, and immature B cells compared to vehicle-treated controls (Fig. 5a-c). However, frequencies of memory B cell and plasma cell populations were unaffected by DMB treatment (Fig. 5d and e). Finally, since IL-1β and IL-6 are derived by monocytes, we examined this population. Mice treated with DMB similarly showed a significant reduction in total monocytes, though it is unclear from this analysis how macrophage or dendritic cell populations are affected by DMB treatment (Fig. 5f). Altogether, these data suggest that DMB is likely to affect innate responses as indicated by reductions in cytokines IL-1β and IL-6 rather than T cell responses.
DMB alters B cell and monocyte populations in CIA. Splenocytes were harvested from male DBA/1j mice 35 days post-initial immunization and underwent flow cytometry analysis to assess altered populations of B cell subsets and monocytes in untreated (CIA + Vehicle) and DMB-treated (CIA + DMB) mice. a-e Frequency of B cell subsets. N = 9 (CIA + Vehicle) and N = 7 (CIA + DMB) pooled from 3 independent experiments. Data are reported as individual mice (symbols) and the group mean ± SEM (bars). *, p < 0.05; ** p < 0.01; ns, non-significant as determined by unpaired t-test. f Frequency of monocytes. N = 9 (CIA + Vehicle) and N = 7 (CIA + DMB) pooled from 3 independent experiments. Data are reported as individual mice (symbols) and the group mean ± SEM (bars). *, p < 0.05 as determined by unpaired t-test.
DMB and DMBut Reduce Macrophage Secretion of Pro-inflammatory Cytokines
We next hypothesized that DMB and DMBut interact with monocytes to reduce secretion of IL-1β and IL-6. To investigate this, macrophages derived from the bone marrow of 6—10-week-old DBA/1j mice were cultured with DMB or DMBut in the presence or absence of E. coli K12 lipopolysaccharide (LPS). Cells without LPS stimulation showed no IL-1β or IL-6 secretion in response to DMB or DMBut at any concentration in the culture media (Fig. 6a-d). Further, both DMB and DMBut reduced secretion of IL-1β and IL-6 from LPS-stimulated macrophages in a dose-dependent manner (Fig. 6a-d), Since mice treated with DMBut, but not DMB, showed reduced serum concentrations of TNF-α, we next investigated the effect of DMBut and DMB on secretion of this cytokine from cultured monocytes. Again, cells without LPS stimulation show no TNF-α productionin response to DMB or DMBut at any concentration (Fig. 6e-f). Surprisingly, both DMB and DMBut reduced secretion of TNF-α from LPS-stimulated macrophages in a dose-dependent manner (Fig. 6e and f). Overall, neither DMB nor DMBut treatment of LPS-stimulated or non-stimulated macrophages significantly reduced viability of these cells at the highest concentration used (25 µM, Supplemental Fig. 7a-b). Taken together, these data suggest that DMB and DMBut may modulate the innate immune system by reducing secretion of pro-inflammatory cytokines from macrophages.
DMB and DMBut reduce production of proinflammatory cytokines from murine BMDMs. Bone marrow-derived macrophages differentiated from hematopoietic cells isolated from DBA/1j mice were cultured in the presence of 0, 25 μM, or 10–4-10–12 M DMB a, c, e or DMBut b, d, f with or without stimulation by E. coli K12 LPS for 24 h. Supernatants were collected and concentrations of IL-1β (a, b), IL-6 (c, d), and TNF-α (e, f), were analyzed by ELISA. Data are reported as the mean ± SEM of two technical replicates pooled from 5 independent experiments (symbols). N = 3 (0 DMB ± LPS, 0 DMBut ± LPS), N = 2 (25 μM DMB ± LPS, 25 μM DMBut ± LPS), N = 4 (1 μM DMB ± LPS, DMBut ± LPS), N = 4 (10 nM DMB ± LPS, 10 nM DMBut ± LPS), N = 4 (100 pM DMB ± LPS, 100 pM DMBut ± LPS), and N = 4 (1 pM DMB ± LPS, 1 pM DMBut ± LPS). *, p < 0.05; ****, p < 0.0001; ns, non-significant as determined by two-way ANOVA with Dunnett’s correction for multiple comparisons.
DISCUSSION
The initial aim of our study was to elucidate the role of TMA lyase activity during inflammatory arthritis in order to propose a novel druggable target for the treatment of RA. To test this, we administered previously characterized TMA lyase inhibitors, DMB and FMC, to mice with CIA and monitored their disease progression. Unexpectedly, blockade of TMA lyase with FMC had no effect on mice with CIA while the significant anti-inflammatory properties of DMB during CIA were found to be independent of TMA lyase activity based on both metabolomic and microbiome analyses. This led us to further investigate the possibility that DMB is absorbed systemically by the murine host. In doing so, we identified novel metabolites, such as DMBut, stemming from the host’s metabolism of DMB. Previous studies have shown that butyrate itself is beneficial in RA and CIA [33, 34, 38,39,40,41,42]; therefore, we hypothesized that DMBut is an immunomodulatory metabolite of DMB. Indeed, the administration of DMB or DMBut both ameliorated disease in our CIA mouse model, and our data showing significant reductions in serum IL-1β and IL-6 without significant effects on autoantibody production or T lymphocyte subsets suggested that DMB alters innate immune responses.
Our initial finding that DMB did not inhibit TMA lyase has been corroborated by the findings of other studies published since DMB was first characterized as a putative TMA lyase inhibitor. In vitro incubation of live bacteria from human fecal samples, recombinant Desulfovibrio alaskensis G20 choline trimethylamine-lyase (CutC/D), live P. mirabilis ATCC 29906, or P. mirabilis ATCC 29906 protein lysate with 10 nM—10 mM DMB revealed that DMB did not reduce relative CutC activity or production of TMA from choline [43,44,45]. Moreover, using 1% (v/v) DMB as an inhibitor of in vivo TMAO production in numerous disease models has resulted in conflicting efficacy outcomes, such that administration of 1% (v/v) DMB has been shown to either reduce or have no effect on circulating TMAO levels [46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63]. The results of our study in conjunction with the findings of these other studies strongly indicate that the amelioration of disease in these models cannot be attributed solely to the suppression of TMAO generation in vivo. Rather, the mechanism of DMB action should be considered independent of TMA/TMAO production. In addition, directly modulating TMAO production with a specific TMA lyase inhibitor, FMC, does not reduce disease severity in CIA. Altogether, our study strongly suggests that the generation of TMAO from the gut microbiome is not a crucial mediator in the development of CIA or RA.
Next, we sought to characterize the metabolism of DMB in mice. DMB is a substrate of ADH in vitro and the acylcarnitine conjugate of DMB, DMBC, has been previously identified in the urine of mice given DMB [24]. Our findings present further insights into the metabolism of DMB in vivo. The predicted product of ADH activity on DMB is 3,3-dimethylbutric aldehyde [24]; however, we and others were unable to detect this metabolite in host tissues, presumably due to the rapid metabolism of cytotoxic aldehydes. Instead, we were successful in detecting the downstream acylcarnitine, DMBC, which corroborates previous findings. From these, we predicted the generation of the novel molecules originating from the hepatic metabolism of DMB since DMB is administered orally and would be subject to first-pass metabolism. One of these predicted novel DMB metabolites, the carboxylic acid DMBut, was also detectable only in mice treated with DMB. Notably, DMBut was only present in the serum and liver of CIA mice given DMB, but not in the cecum, suggesting it is not microbially derived and lending evidence to support the absorption and metabolism of DMB within the host.
DMB and DMBut together elicited the most profound immunologic effects on circulating IL-6 and IL-1β. Both IL-6 and IL-1β are key mediators in the pathogenesis of both RA and CIA [64, 65]. Blockade of IL-6 signaling in CIA using a monoclonal antibody targeting IL-6 or its receptor starting at the time of CIA initiation, but not at day 21, reduces CIA severity [66] and serum anti-CII antibodies and TNF concentration [67]. Similarly, blockade of TNF or IL-1 receptor during the induction phase as well as the established arthritis phase of CIA resulted in reduced disease severity, serum amyloid P, and TNF and IL-1β expression in the arthritic joints [68]. IL-6 and IL-1β are important mediators of joint damage in RA: IL-1 and TNF induce IL-6 in RA synoviocytes [69] and IL-6 enhances IL-1β-induced matrix metalloproteinases in synovial fibroblasts [70]. Our BMDM experiments show DMB and DMBut inhibit LPS-induced IL-1β, IL-6, and TNF-α release in M1-like differentiated macrophages without substantially affecting viability of these cells. The activity of DMB and DMBut on monocyte populations in vivo was not assessed in our study, which is a limitation. Ideally, synovial macrophages from mice with CIA treated with vehicle, DMB, or DMBut would be examined for their expression of key disease-mediating cytokines such as IL-1β, IL-6 and TNF. However, the isolation of synovial macrophages from mice and yield of RNA is a significant limitation. In other disease models and mouse strains, DMB reduces circulating and tissue-specific expression of IL-1β, TNF-α, and IL-6 [53, 56, 59,60,61,62,63, 71,72,73,74,75]. Another study in male C57BL/6 J mice fed a high-choline diet suggested that treatment with 1.3% (v/v) DMB reduced IL-1β protein expression in heart tissue through acting on cGAS-STING upstream of NLRP3 expression in macrophages, though this effect was attributed to the presumed inhibition of TMAO production by DMB [76]. Thus, our data and that of others supports the hypothesis that DMB and DMBut are likely to act on macrophage secretion of key inflammatory cytokines.
A significant limitation of our study is that the specific mechanism of DMB and DMBut remains unclear. We presume that the active metabolite DMBut may work similarly to butyrate. In addition to being a histone deacetylase (HDAC) inhibitor, butyrate is also known to activate the GPCRs GPR43 (FFA2), GPR41 (FFA3), and GPR109A, which are expressed in RA-relevant immune cells and mediate anti-inflammatory responses [77]. Therefore, it is possible that DMBut signals through the same receptors to drive amelioration of arthritis severity in CIA. DMB is also reported to inhibit choline dehydrogenase, causing accumulation of choline in liver and kidney tissue, inhibition of choline phosphorylation, and reduction of available choline to cross into the circulation [78]. The choline transporter CTL1 is highly expressed in macrophage-like and fibroblast-like synoviocytes in the synovia and cartilage of patients with RA, and deficient choline uptake through CTL1 in macrophages reduces IL-1β production via attenuation of mitochondrial ATP synthesis, thereby driving activation of AMPK-mediated mitophagy and termination of NLRP3 inflammasome activation [79, 80]. Thus, DMB and DMBut have multiple potential mechanistic pathways by which they reduce arthritis severity during CIA.
In summary, we identify the small molecules DMB and DMBut as potent agents to decrease the severity of CIA, and strikingly, provide protection after the initiation of disease. Although not an effective TMA lyase inhibitor as previously published, DMB and DMBut have multiple potential mechanisms by which they reduce circulating IL-1β and IL-6, possibly through histone deacetylase, GPCR, or NLRP3 pathways in innate immune cells. DMB has been shown to be a viable therapy in the context of atherosclerosis, which is a known co-morbidity of RA, partly through inhibiting endogenous foam cell formation. Though it is unclear if DMB is functioning as an inhibitor of TMA lyases, it may function as an inhibitor of the inflammatory mechanisms contributing to both foam cell formation and autoimmune arthritis. Further studies should focus on the effects of DMB/DMBut administration during CIA and the mechanisms by which they influence the inflammatory pathways underlying both RA and atherosclerosis that are independent of TMA/TMAO production.
METHODS
Collagen-induced Arthritis
Male 6-week-old DBA/1j mice were injected intradermally at the base of the tail on days 0 and 21 with 100 µl of an emulsion containing 200 µg nasal bovine type II collagen (CII, Elastin) in 0.01 M glacial acetic acid and an equal volume of complete Freund’s adjuvant (Millipore Sigma). On day 21, mice were treated with either 1% (vol/vol) DMB (TCI Chemicals), 1% (vol/vol) DMBut (Sigma-Aldrich), or no additional additive (vehicle), in drinking water with 100 mg/ml grape-flavored sugar-sweetened Kool-Aid (Kraft Foods) added to encourage consumption, or 100 mg/kg FMC (Jubilant Biosys Limited) gavaged orally every other day. Mice were monitored for onset of arthritis and severity of disease until 35 days after the initial immunization. Disease for each of the four paws was scored on a scale from 0–4 according to established metrics: 0—no erythema or joint swelling; 1—erythema and one swollen digit; 2—erythema and two swollen digits; 3—erythema and three swollen digits; 4—ankylosis. The score for each paw was summed to generate a total score per mouse. The incidence of arthritis was defined as a non-zero score. Therefore, the rate of incidence, as a percentage, indicates how many mice in the treatment group had a non-zero score on a given day. At euthanasia on day 35, blood was collected by cardiac puncture and the serum stored at -20 °C for future analysis. Spleens and inguinal lymph nodes were harvested and processed. Feces were collected at day 35 for 16S ribosomal RNA sequencing analysis. Terminal euthanasia of animals involved intraperitoneal injection of a mixture of ketamine (100 mg/kg) and xylazine (10 mg/kg), with subsequent cardiac puncture as a secondary method of euthanasia.
Cytokine Quantification
Serum taken from mice with CIA on day 35 post-immunization was analyzed for cytokine concentrations using a Meso Scale Discovery U-Plex assay platform according to manufacturer instructions. Assay plates were imaged on a MESO QuickPlex SQ 120 at the University of Colorado, Anschutz Medical Campus Human Immune Monitoring Shared Resource (HIMSR).
Overnight bone marrow-derived monocyte (BMDM) cell cultures treated with vehicle, DMB, and DMBut with or without the presence of stimulating agents were pelleted at 1660 rpm for 5 min to collect cell-free supernatants for quantification of secreted cytokine concentrations via enzyme-linked immunosorbent assay (ELISA). Concentrations of secreted murine IL-6, IL-1β, and TNFα were determined using DuoSet ELISA kits (R&D Systems) according to manufacturer instructions at room temperature. Briefly, assay plates were coated with capture antibody diluted to the appropriate working concentrations in PBS overnight and subsequently blocked with 1% bovine serum albumin (BSA) in PBS for 2 h. Unknown supernatant samples were either undiluted (IL-1β), or diluted 1:10 (IL-6 and TNFα) in 1% BSA/PBS. Samples incubated on the assay plate for 2 h. Assay plates were developed in 100 µl TMB Substrate Solution (Thermofisher Scientific) for 20 min and subsequently stopped with 50 µl of 2N H2SO4 stop solution. Assay plates were imaged immediately after addition of stop solution on a SpectraMax iD5 plate reader at 450 nm with correction at 540 nm.
Anti-collagen Type II-IgG Antibody Quantification
Serum from day 35 post-immunization was evaluated for anti-collagen type II antibody concentrations via ELISA. All steps were performed on ice. ELISA-grade CII (Chondrex) was diluted 1:10 in 1 × collagen dilution buffer (Chondrex) and incubated on the assay plate at 4 °C overnight with gentle rocking while covered with aluminum foil. The assay plate was blocked with 0.5% BSA (Sigma-Aldrich) in PBS for 4 h at 4 °C with gentle rocking. A relative standard was generated using serum from a mouse with robust CIA, not otherwise treated, diluted 1:1,000 in 0.5% BSA/PBS and serially diluted 1:4. Unknown serum samples were diluted 1:10,000 in 0.5% BSA/PBS and incubated on the assay plate overnight at 4 °C with gentle rocking while covered in aluminum foil. Goat anti-mouse IgG Fab-HRP, IgG1-HRP, IgG2a-HRP, and IgG2b-HRP antibodies (Southern Biotech) were diluted 1:10,000 in PBS and incubated on the assay plate at room temperature for 2 h with gentle shaking. The assay was developed with 100 µl of 1:1 BD OptEIA TMB reagents (BD Bioscience) at room temperature for 20 min, and subsequently stopped with 100 µl 2N H2SO4 stop solution. Assay plates were imaged immediately after addition of stop solution on a SpectraMax iD5 plate reader at 450 nm with correction at 570 nm.
Flow Cytometry
Splenocytes were strained through 70 µm cell strainers (Fisher Scientific) and washed with serum-free RPMI 1640. The cell suspensions were pelleted at 4 °C and 300xg for 5 min and the supernatant was discarded. Lymphocytes were resuspended in 1 ml 5% fetal bovine serum (FBS) in PBS. Splenocytes were resuspended in 1 ml 1 × red blood cell lysis buffer (Invitrogen) and incubated on ice for 5 min. Lysis was stopped with 10 ml PBS and the cell suspension was pelleted at 4 °C and 300xg for 5 min. Splenocytes were resuspended in 1 ml 5% FBS in PBS. 100 µl of each cell suspension was added to a 5 ml polystyrene round-bottom tube (Corning) and incubated in 1 µl Human TruStain FcX (Biolegend) for 5 min at 4 °C. 10 µl Brilliant Stain Buffer Plus (BD Biosciences) was added to each tube and cells were stained for viability and surface markers as noted in Supplemental Table 1. Stained cells incubated at 4 °C for 30 min. Cells were washed with 1 ml 5% FBS in PBS at 4 °C and 300xg for 5 min, then fixed and made permeable in 1 ml Foxp3/Transcription Factor Staining Buffer (Tonbo Biosciences). Cells incubated at 4 °C for 30 min. Fixed cells were washed twice with 1 ml 1 × Flow Cytometry Perm Buffer (Tonbo Biosciences) at 4 °C and 300xg for 5 min and stained for intracellular markers. Stained cells incubated at 4 °C for 45 min, then washed with 1 ml 1 × Flow Cytometry Perm Buffer at 4 °C and 300xg for 5 min and resuspended in 300 µl 5% FBS in PBS for analysis. Analysis of data was performed using FlowJo (version 10.8.1). Supplemental Table 2 lists the definitions of the T and B lymphocyte populations presented.
TMA Lyase Inhibition Assay
Proteus mirabilis (ATCC 29906) was cultured in 5 ml Difco Nutrient Broth (BD Biosciences) overnight at 37 °C and 215 rpm without antibiotic selection. Overnight cultures were sub-cultured at a dilution of 1:20 in fresh nutrient broth and grown overnight at 37 °C and 215 rpm to serve as the starting material for downstream assays.
Inhibition of the P. mirabilis TMA lyase enzyme complex CutC/CutD by DMB or FMC was assessed as previously described with some modifications [24, 25, 81]. Briefly, overnight P. mirabilis cultures were pelleted by centrifugation at 3000 rpm for 30 min and the broth supernatant was discarded. Cells were resuspended in 10 ml PBS and 400 µl of cell suspension was allocated to 13 × 100 mm screw cap culture tubes (Pyrex) with gas-tight 13 mm-425 Mininert valve caps (Supelco). To determine functionality of endogenous P. mirabilis CutC/CutD, bacteria were incubated at 37 °C in the presence of 0 µM, 25 µM, 50 µM, 75 µM, and 100 µM D9-choline (Cambridge Isotope Laboratories) for 2, 4, 6, and 24 h. To determine the inhibition of endogenous P. mirabilis CutC/CutD by DMB, bacteria were incubated in the presence of 1 M, 10 mM, 100 µM, 1 µM, 10 nM, 100 pM, and 1 pM DMB or 1 mM, 100 µM, 1 µM, 10 nM, 100 pM, and 1 pM FMC for 15 min, then 100 µM D9-choline was added to the reaction vials. Reactions were performed at 37 °C for 2 h. Reactions were quenched with 200 µl of cold 1 M NaOH and submerged in a liquid nitrogen bath. 2 ml hexanes, 1 ml butanol, and 200 µl 1N NaOH were added to the reaction vials, vortexed for 1 min, and centrifuged for 15 min at 4 °C and 2500 rpm. The upper phase was transferred to a new 13 × 100 mm screw cap culture tube with PTFE-lined caps and 200 µl of 0.2N formic acid was added. Vials were vortexed for 1 min and centrifuged for 15 min at 4 °C and 2500 rpm. The lower aqueous phase was collected and stored at -80 °C until analysis by stable isotope dilution LC–MS/MS.
Bone Marrow-derived Macrophage Differentiation
Bone marrow-derived macrophages were isolated from 6–10-week-old DBA/1j mice as previously described [82]. Briefly, bone marrow was flushed and processed to a single cell suspension from the femur and tibia using 1 ml cold PBS. The single cell suspension was cultured in 9 ml of complete RPMI 1640 supplemented with 10% fetal bovine serum, 2% HEPES, 0.6% penicillin/streptomycin, 0.1% 2-mercaptoethanol, and 20 ng/ml recombinant mouse GM-CSF (Peprotech). After 72 h, the cell culture media was replaced with fresh differentiation media to remove non-adherent cells. After 6 days of culture, adherent cells were washed with cold PBS and resuspended in RPMI 1640 without recombinant mouse GM-CSF. Cells were stimulated with 10 µg/µl ultrapure E. coli K12 lipopolysaccharide (InvivoGen) and treated with 25 μM, 1 μM, 10 nM, 100 pM, or 1 pM DMB or DMBut overnight.
Gas Chromatography
Serum samples were thawed to room temperature. After a brief vortex, 50μL of each serum sample was transferred to a glass vial. 3N HCl (20 μL) was added followed by hexane (50 μL) and vortexed thoroughly. The material was then transferred to respective vial inserts (150 μL). All the samples were then centrifuged at 4 °C, 10 min, 3000 rpm. The upper hexane layer was then taken out and transferred into respective new vial inserts and capped immediately for chromatography.
Frozen liver samples were ground in a mortar with pestle in liquid N2. The suspension of ground liver in liquid N2 was quickly poured into a pre-weighed glass vial, allowing the liquid N2 to evaporate prior to capping and weighing the sample. To each sample, hexane was added in a weight (mg) to volume (μL) ratio of 1:2 and stored at -20 °C. 3N HCl (20 μL) was added to each hexane-suspension, sonicated for 15 min, and vortexed vigorously for 5 min. The 100 μL top liquid layer was then transferred to respective vial inserts and capped. All the samples were then centrifuged at 4 °C for 10 min at 3000 rpm. The upper hexane layer (50 μL) was then transferred into a new vial insert and capped immediately for chromatography.
Hexane extracts (1 μL) were injected into a Trace 1310 GC coupled to a Thermo ISQ-LT MS, at splitless mode. The inlet was held at 250 °C. Peak separation was achieved on a 30 m DB-WAXUI column (J&W, 0.25 mm ID, 0.25 μm film thickness). Oven temperature was held at 80 °C for 2 min, ramped at 20 °C/min to 125 °C, then ramped at 40 °C/min to 175 °C and then to 240 °C at 20 °C/min with a final hold for 20 min. Helium carrier gas flow was held at 1.2 mL/min. Temperatures of transfer line and ion source were both held at 250 °C. SIM mode was used to scan ions m/z 57, 69, 87 for DMB and m/z 59, 57, 101 for DMBut with scan time of 0.1 s/ion under electron impact mode. Peak integration was completed using Chromeleon software (ThermoFisher) (Supplemental Fig. 3b and c).
UHPLC-tandem Mass Spectrometry
Frozen P. mirabilis supernatants and mouse serum samples were thawed on ice and extracted with ice cold methanol, acetonitrile, and water (5:3:2, respectively) at a 1:25 ratio. Frozen cecum and liver samples were weighed to the nearest 0.1 mg and extracted at 15 mg/ml in the same extraction buffer.
Extractions were vortexed for 30 min at 4 °C and then insoluble materials were pelleted by centrifugation at 18,000xg for 10 min at 4 °C. Supernatants were analyzed using a Thermo Vanquish UHPLC coupled to a Thermo Q Exactive MS and run in positive and negative ion modes (separate runs). Injection volumes were 20 µl for serum and broth extracts and 10 µl for tissue extracts. UHPLC phases were water (A) and acetonitrile (B) supplemented with 0.1% formic acid for positive mode runs and 1 mM ammonium acetate for negative mode runs. Metabolites were separated on a Kinetex C18 column (2.1 × 150 mm, 1.7 µm, Phenomenex) equipped with a guard column using a 5-min gradient method with the following conditions: Flow rate 0.45 ml/min; column temperature 45 °C; sample compartment temperature 7 oC; solvent gradient: 0–0.5 min 5% B, 0.5–1.1 min 5–95% B, 1.1–2.75 min hold at 95% B, 2.75–3 min 95–5% B, 3–5 min hold at 5% B. The mass spectrometer was operated in full MS mode at a resolution of 70,000, maximum injection time of 200 ms, microscans 2, automatic gain control (AGC) ions, electrospray source voltage 4.0 kV, capillary temperature 320 °C, and sheath gas 45, auxiliary gas 25, and sweep gas 0 (all nitrogen). Instrument stability and quality control were assessed using replicate injections of a technical mixture every 15 runs as previously described [83, 84]. Raw data files were converted to mzXML using RawConverter and metabolites were annotated and peaks integrated using Maven [85,86,87] in conjunction with the Kyoto Encyclopedia of Genes and Genomes (KEGG) database.
For D9-trimethylamine (TMA), TMA, and TMAO measurements, the mass spectrometer was operated as above with a scan range of 50–750 m/z. For 3,3-diemethylbutyrylcarnitine measurements, the mass spectrometer was operated as above with a scan range of 65–900 m/z.
Microbiome Analysis
Bacterial profiles were determined by broad-range amplification and sequence analysis of 16S rRNA genes following our previously described methods [8, 88, 89]. In brief, amplicons were generated using primers that target approximately 400 base pairs of the V3V4 variable region of the 16S rRNA gene. PCR products were normalized using a SequalPrepTM kit (Invitrogen, Carlsbad, CA), pooled, lyophilized, purified and concentrated using a DNA Clean and Concentrator Kit (Zymo, Irvine, CA). Pooled amplicons were quantified using Qubit Fluorometer 2.0 (Invitrogen, Carlsbad, CA). The pool was diluted to 4 nM and denatured with 0.2 N NaOH at room temperature. The denatured DNA was diluted to 15 pM and spiked with 25% of the Illumina PhiX control DNA prior to loading the sequencer. Illumina paired-end sequencing was performed on the Miseq platform with versions v2.4 of the Miseq Control Software and of MiSeq Reporter, using a 600-cycle version 3 reagent kit.
Illumina Miseq paired-end reads were aligned to human reference genome hg19 with bowtie2 and matching sequences discarded [90, 91]. As previously described, the remaining non-human paired-end sequences were sorted by sample via barcodes in the paired reads with a python script [92]. Sorted paired end sequence data were deposited in the NCBI Short Read Archive under accession number PRJNA1006768. The sorted paired reads were assembled using phrap [93, 94]. Pairs that did not assemble were discarded. Assembled sequence ends were trimmed over a moving window of 5 nucleotides until average quality met or exceeded 20. Trimmed sequences with more than 1 ambiguity or shorter than 350 nucleotides were discarded. Potential chimeras identified with Uchime (usearch6.0.203_i86linux32) using the Schloss Silva reference sequences were removed from subsequent analyses [95, 96]. Assembled sequences were aligned and classified with SINA (1.3.0-r23838) using the 418,497 bacterial sequences in Silva 115NR99 as reference configured to yield the Silva taxonomy [97, 98]. Operational taxonomic units (OTUs) were produced by clustering sequences with identical taxonomic assignments. This process generated 4136760 sequences for 23 samples (median sample size: 169668 sequences/sample; IQR: 113008 to 253290 sequences/sample). The median Goods coverage score was ≥ 99.97%. The software package Explicet (v2.10.5, www.explicet.org) was used for data organization and alpha-diversity calculations [99].
Statistics
Unless specified otherwise, data was analyzed using GraphPad Prism software version 9; specific statistical tests for comparisons are referenced in the figure legends.
Ethics Declarations
All animal studies and methods were approved by the University of Colorado School of Medicine Institutional Animal Care and Use Committee (protocol #173). All animal studies and methods were performed in accordance with the ethics guidelines and regulations put forth by the University of Colorado School of Medicine Institutional Animal Care and Use Committee. All studies are reported in accordance with the ARRIVE 2.0 guidelines [100, 101].
Data Availability
16S sequencing data are publicly deposited under BioProject accession: PRJNA1006768 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1006768). All other data are available upon request to the corresponding author.
References
Almutairi, K., et al. 2021. The global prevalence of rheumatoid arthritis: A meta-analysis based on a systematic review. Rheumatology International 41 (5): 863–877.
Aletaha, D., and J. S. Smolen. 2018. Diagnosis and management of rheumatoid arthritis. JAMA 320 (13): 1360.
Sergeant, J. C., et al. 2018. Prediction of primary non-response to methotrexate therapy using demographic, clinical and psychosocial variables: Results from the UK Rheumatoid Arthritis Medication Study (RAMS). Arthritis Research & Therapy 20 (1): 147.
Nam, J. L., et al. 2017. Efficacy of biological disease-modifying antirheumatic drugs: A systematic literature review informing the 2016 update of the EULAR recommendations for the management of rheumatoid arthritis. Annals of the Rheumatic Diseases 76 (6): 1113–1136.
Strand, V., et al. 2017. Discontinuation of biologic therapy in rheumatoid arthritis: Analysis from the Corrona RA registry. Rheumatol Ther 4 (2): 489–502.
Holers, V. M., et al. 2018. Rheumatoid arthritis and the mucosal origins hypothesis: Protection turns to destruction. Nature Reviews Rheumatology 14 (9): 542–557.
Maeda, Y., and K. Takeda. 2019. Host–microbiota interactions in rheumatoid arthritis. Experimental & Molecular Medicine 51 (12): 1–6.
Chriswell, M. E., et al. 2022. Clonal IgA and IgG autoantibodies from individuals at risk for rheumatoid arthritis identify an arthritogenic strain of Subdoligranulum. Science Translation Medicine 14 (668): eabn5166. https://doi.org/10.1126/scitranslmed.abn5166.
Jubair, W. K., et al. 2018. Modulation of inflammatory arthritis in mice by gut microbiota through mucosal inflammation and autoantibody generation. Arthritis & Rheumatology 70 (8): 1220–1233.
Maeda, Y., et al. 2016. Dysbiosis contributes to arthritis development via activation of autoreactive T cells in the intestine. Arthritis & Rheumatology 68 (11): 2646–2661.
Wu, H. - J., et al. 2010. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32 (6): 815–827.
Chu, S. H., et al. 2020. Circulating plasma metabolites and risk of rheumatoid arthritis in the Nurses’ Health Study. Rheumatology 59 (11): 3369–3379.
Costenbader, K. H., et al. 2021. Circulating blood metabolite trajectories and risk of rheumatoid arthritis among military personnel in the Department of Defense Biorepository. Annals of the Rheumatic Diseases 80 (8): 989.
Martínez-del Campo, A., et al., Characterization and Detection of a Widely Distributed Gene Cluster That Predicts Anaerobic Choline Utilization by Human Gut Bacteria. mBio, 2015. 6(2): p. e00042–15.
Janeiro, M., et al. 2018. Implication of Trimethylamine N-Oxide (TMAO) in disease: Potential biomarker or new therapeutic target. Nutrients 10 (10): 1398.
Al-Waiz, M., et al. 1987. The metabolism of 14C-labelled trimethylamine and its N-oxide in man. Xenobiotica 17 (5): 551–558.
Zeisel, S. H., et al. 1989. Conversion of dietary choline to trimethylamine and dimethylamine in rats: Dose-response relationship. Journal of Nutrition 119 (5): 800–804.
Ilyas, A., et al. 2022. Implications of trimethylamine N-oxide (TMAO) and betaine in human health: Beyond being osmoprotective compounds. Frontiers in Molecular Biosciences 9: 964624.
Coras, R., et al. 2019. Choline metabolite, trimethylamine N-oxide (TMAO), is associated with inflammation in psoriatic arthritis. Clinical and Experimental Rheumatology 37 (3): 481–484.
Lin, H., et al. 2020. The role of gut microbiota metabolite trimethylamine N-oxide in functional impairment of bone marrow mesenchymal stem cells in osteoporosis disease. Ann Transl Med 8 (16): 1009.
Zhuang, H., et al. 2023. Trimethylamine-N-oxide sensitizes chondrocytes to mechanical loading through the upregulation of Piezo1. Food and Chemical Toxicology 175: 113726.
Koh, J. H., et al. 2019. Distinct urinary metabolic profile in rheumatoid arthritis patients: A possible link between diet and arthritis phenotype. Journal of Rheumatic Diseases 26 (1): 46.
Van Maanen, M. A., M. J. Vervoordeldonk, and P. P. Tak. 2009. The cholinergic anti-inflammatory pathway: Towards innovative treatment of rheumatoid arthritis. Nature Reviews Rheumatology 5 (4): 229–232.
Wang, Z., et al. 2015. Non-lethal Inhibition of Gut Microbial Trimethylamine Production for the Treatment of Atherosclerosis. Cell 163 (7): 1585–1595.
Roberts, A. B., et al. 2018. Development of a gut microbe–targeted nonlethal therapeutic to inhibit thrombosis potential. Nature Medicine 24 (9): 1407–1417.
Dambrova, M., et al. 2022. Acylcarnitines: Nomenclature, biomarkers, therapeutic potential, drug targets, and clinical trials. Pharmacological Reviews 74 (3): 506–551.
McCoin, C. S., T. A. Knotts, and S. H. Adams. 2015. Acylcarnitines–old actors auditioning for new roles in metabolic physiology. Nature Reviews Endocrinology 11 (10): 617–625.
McGarry, J. D., and N. F. Brown. 1997. The mitochondrial carnitine palmitoyltransferase system. From concept to molecular analysis. European Journal of Biochemistry 244 (1): 1–14.
McInnes, I. B., and G. Schett. 2011. The pathogenesis of rheumatoid arthritis. New England Journal of Medicine 365 (23): 2205–2219.
Luross, J. A., and N. A. Williams. 2001. The genetic and immunopathological processes underlying collagen-induced arthritis. Immunology 103 (4): 407–416.
Taams, L. S. 2020. Interleukin-17 in rheumatoid arthritis: Trials and tribulations. Journal of Experimental Medicine 217 (3): e20192048.
Langrish, C. L., et al. 2005. IL-23 drives a pathogenic T cell population that induces autoimmune inflammation. Journal of Experimental Medicine 201 (2): 233–240.
Hui, W., et al. 2019. Butyrate inhibit collagen-induced arthritis via Treg/IL-10/Th17 axis. International Immunopharmacology 68: 226–233. https://doi.org/10.1016/j.intimp.2019.01.018.
Takahashi, D., et al. 2020. Microbiota-derived butyrate limits the autoimmune response by promoting the differentiation of follicular regulatory T cells. eBioMedicine 58: 102913.
He, J., et al. 2022. Intestinal butyrate-metabolizing species contribute to autoantibody production and bone erosion in rheumatoid arthritis. Sci Adv 8 (6): eabm1511.
Crotty, S. 2014. T follicular helper cell differentiation, function, and roles in disease. Immunity 41 (4): 529–542.
Maeda, K., et al. 2010. IL-6 increases B-cell IgG production in a feed-forward proinflammatory mechanism to skew hematopoiesis and elevate myeloid production. Blood 115 (23): 4699–4706.
Yao, Y., et al. 2022. Short-chain fatty acids regulate B cells differentiation via the FFA2 receptor to alleviate rheumatoid arthritis. British Journal of Pharmacology 179 (17): 4315–4329.
Rosser, E. C., et al. 2020. Microbiota-derived metabolites suppress arthritis by amplifying aryl-hydrocarbon receptor activation in regulatory B cells. Cell Metabolism 31 (4): 837-851.e10.
Kim, D. S., et al. 2018. Attenuation of rheumatoid inflammation by sodium butyrate through reciprocal targeting of HDAC2 in Osteoclasts and HDAC8 in T cells. Frontiers in Immunology 9: 1525.
Segain, J. P., et al. 2000. Butyrate inhibits inflammatory responses through NFkappaB inhibition: Implications for Crohn’s disease. Gut 47 (3): 397–403.
Lee, C., et al. 2017. Sodium butyrate inhibits the NF-kappa B signaling pathway and histone deacetylation, and attenuates experimental colitis in an IL-10 independent manner. International Immunopharmacology 51: 47–56.
Orman, M., et al. 2019. Structure-guided identification of a small molecule that inhibits anaerobic choline metabolism by human gut bacteria. Journal of the American Chemical Society 141 (1): 33–37.
Day-Walsh, P., et al. 2021. The use of an in-vitro batch fermentation (human colon) model for investigating mechanisms of TMA production from choline, L-carnitine and related precursors by the human gut microbiota. European Journal of Nutrition 60 (7): 3987–3999.
Iglesias-Carres, L., et al. 2021. Development of a high-throughput method to study the inhibitory effect of phytochemicals on trimethylamine formation. Nutrients 13 (5): 1466.
Casso, A. G., et al. 2022. Initiation of 3,3-dimethyl-1-butanol at midlife prevents endothelial dysfunction and attenuates in vivo aortic stiffening with ageing in mice. The Journal of Physiology 600: 4633–4651.
Yang, Y., et al. 2022. High-circulating gut microbiota-dependent metabolite trimethylamine N-oxide is associated with poor prognosis in pulmonary arterial hypertension. European Heart Journal Open 2 (5): oeac021.
Brunt, V. E., et al. 1985. 2022 Suppression of trimethylamine N-oxide with DMB mitigates vascular dysfunction, exercise intolerance, and frailty associated with a Western-style diet in mice. Journal of Applied Physiology 133 (4): 798–813.
González-Correa, C., et al. 2021. Trimethylamine N-oxide promotes autoimmunity and a loss of vascular function in toll-like receptor 7-driven lupus mice. Antioxidants (Basel) 11 (1): 84.
Li, J., et al. 2022. Trimethylamine N-oxide induces osteogenic responses in human aortic valve interstitial cells in vitro and aggravates aortic valve lesions in mice. Cardiovascular Research 118 (8): 2018–2030.
Hsu, C. N., et al. 2021. Maternal 3,3-dimethyl-1-butanol therapy protects adult male rat offspring against hypertension programmed by perinatal TCDD exposure. Nutrients 13 (9): 3041.
Mao, J., et al. 2021. Repeated 3,3-Dimethyl-1-butanol exposure alters social dominance in adult mice. Neuroscience Letters 758: 136006.
Chen, L., et al. 2020. Trimethylamine N-oxide impairs perfusion recovery after hindlimb ischemia. Biochemical and Biophysical Research Communications 530 (1): 95–99.
Brunt, V. E., et al. 2020. Trimethylamine-N-Oxide promotes age-related vascular oxidative stress and endothelial dysfunction in mice and healthy humans. Hypertension 76 (1): 101–112.
Wu, K., et al. 2020. The gut microbial metabolite trimethylamine N-oxide aggravates GVHD by inducing M1 macrophage polarization in mice. Blood 136 (4): 501–515.
Gao, Q., et al. 2019. Decreased levels of circulating trimethylamine N-oxide alleviate cognitive and pathological deterioration in transgenic mice: A potential therapeutic approach for Alzheimer’s disease. Aging (Albany NY) 11 (19): 8642–8663.
Hsu, C. N., et al. 2019. Targeting on gut microbial metabolite Trimethylamine-N-Oxide and short-chain fatty acid to prevent maternal high-fructose-diet-induced developmental programming of hypertension in adult male offspring. Molecular Nutrition & Food Research 63 (18): e1900073.
Li, X., et al. 2019. Reductions in gut microbiota-derived metabolite trimethylamine N-oxide in the circulation may ameliorate myocardial infarction-induced heart failure in rats, possibly by inhibiting interleukin-8 secretion. Molecular Medicine Reports 20 (1): 779–786.
Chen, H., et al. 2019. Increased circulating trimethylamine N-oxide plays a contributory role in the development of endothelial dysfunction and hypertension in the RUPP rat model of preeclampsia. Hypertension in Pregnancy 38 (2): 96–104.
Li, T., et al. 2018. Increased circulating trimethylamine N-oxide contributes to endothelial dysfunction in a rat model of chronic kidney disease. Biochemical and Biophysical Research Communications 495 (2): 2071–2077.
Sun, G., et al. 2017. Gut microbial metabolite TMAO contributes to renal dysfunction in a mouse model of diet-induced obesity. Biochemical and Biophysical Research Communications 493 (2): 964–970.
Li, T., et al. 2017. Elevated circulating Trimethylamine N-Oxide levels contribute to endothelial dysfunction in aged rats through vascular inflammation and oxidative stress. Frontiers in Physiology 8: 350.
Chen, K., et al. 2017. Gut microbiota-dependent metabolite Trimethylamine N-Oxide contributes to cardiac dysfunction in western diet-induced obese mice. Frontiers in Physiology 8: 139.
Chung, Y., et al. 2009. Critical regulation of early Th17 cell differentiation by interleukin-1 signaling. Immunity 30 (4): 576–587.
Sarkar, S., L. A. Cooney, and D. A. Fox. 2010. The role of T helper type 17 cells in inflammatory arthritis. Clinical and Experimental Immunology 159 (3): 225–237.
Takagi, N., et al. 1998. Blockage of interleukin-6 receptor ameliorates joint disease in murine collagen-induced arthritis. Arthritis and Rheumatism 41 (12): 2117–2121.
Liang, B., et al. 2009. Evaluation of anti-IL-6 monoclonal antibody therapy using murine type II collagen-induced arthritis. Journal of Inflammation (London) 6: 10.
Williams, R. O., et al. 2000. Evaluation of TNF-alpha and IL-1 blockade in collagen-induced arthritis and comparison with combined anti-TNF-alpha/anti-CD4 therapy. The Journal of Immunology 165 (12): 7240–7245.
Guerne, P. A., et al. 1989. Synovium as a source of interleukin 6 in vitro Contribution to local and systemic manifestations of arthritis. Journal of Clinical Investigation 83 (2): 585–92.
Ito, A., et al. 1992. Effects of interleukin-6 on the metabolism of connective tissue components in rheumatoid synovial fibroblasts. Arthritis and Rheumatism 35 (10): 1197–1201.
Jiang, W. Y., et al. 2022. Trimethylamine N-oxide facilitates the progression of atrial fibrillation in rats with type 2 diabetes by aggravating cardiac inflammation and connexin remodeling. Journal of Physiology and Biochemistry 78: 855–867.
Liu, G., et al. 2022. Inhibition of microbiota-dependent Trimethylamine N-Oxide production ameliorates high salt diet-induced sympathetic excitation and hypertension in rats by attenuating central neuroinflammation and oxidative stress. Frontiers in Pharmacology 13: 856914.
Zou, D., Y. Li, and G. Sun. 2021. Attenuation of circulating Trimethylamine N-Oxide prevents the progression of cardiac and renal dysfunction in a rat model of chronic cardiorenal syndrome. Frontiers in Pharmacology 12: 751380.
Shuai, W., et al. 2020. High-choline diet exacerbates cardiac dysfunction, fibrosis, and inflammation in a mouse model of heart failure with preserved ejection fraction. Journal of Cardiac Failure 26 (8): 694–702.
Wang, G., et al. 2020. 3,3-Dimethyl-1-butanol attenuates cardiac remodeling in pressure-overload-induced heart failure mice. Journal of Nutritional Biochemistry 78: 108341.
Bai, L., et al. 2022. Hydrogen sulfide ameliorated high choline-induced cardiac dysfunction by inhibiting cGAS-STING-NLRP3 inflammasome pathway. Oxidative Medicine and Cellular Longevity 2022: 1392896.
Parada Venegas, D., et al. 2019. Short Chain Fatty Acids (SCFAs)-mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Frontiers in Immunology 10: 277.
Barlow, P., and R. M. Marchbanks. 1985. The effects of inhibiting choline dehydrogenase on choline metabolism in mice. Biochemical Pharmacology 34 (17): 3117–3122.
Beckmann, J., et al. 2015. Expression of choline and acetylcholine transporters in synovial tissue and cartilage of patients with rheumatoid arthritis and osteoarthritis. Cell and Tissue Research 359 (2): 465–477.
Sanchez-Lopez, E., et al. 2019. Choline uptake and metabolism modulate macrophage IL-1β and IL-18 production. Cell Metabolism 29 (6): 1350-1362.e7.
Jia, X., L. J. Osborn, and Z. Wang. 2020. Simultaneous Measurement of Urinary Trimethylamine (TMA) and Trimethylamine N-Oxide (TMAO) by liquid chromatography-mass spectrometry. Molecules 25 (8): 1862.
Zhang, X., R. Goncalves, and D.M. Mosser. 2008. The isolation and characterization of murine macrophages. Current Protocols in Immunology 83 (1): 14.1.1-14.1.14.
Nemkov, T., A. D’Alessandro, and K. C. Hansen. 2015. Three-minute method for amino acid analysis by UHPLC and high-resolution quadrupole orbitrap mass spectrometry. Amino Acids 47 (11): 2345–2357.
Nemkov, T., K. C. Hansen, and A. D’Alessandro. 2017. A three-minute method for high-throughput quantitative metabolomics and quantitative tracing experiments of central carbon and nitrogen pathways. Rapid Communications in Mass Spectrometry 31 (8): 663–673.
He, L., et al. 2015. Extracting accurate precursor information for tandem mass spectra by RawConverter. Analytical Chemistry 87 (22): 11361–11367.
Melamud, E., L. Vastag, and J. D. Rabinowitz. 2010. Metabolomic analysis and visualization engine for LC−MS data. Analytical Chemistry 82 (23): 9818–9826.
Clasquin, M. F., E. Melamud, and J. D. Rabinowitz. 2012. LC-MS data processing with MAVEN: A metabolomic analysis and visualization engine. Current Protocols in Bioinformatics 14: Unit14.11.
Frank, D. N., et al. 2022. A dysbiotic microbiome promotes head and neck squamous cell carcinoma. Oncogene 41 (9): 1269–1280.
Soderborg, T. K., et al. 2018. The gut microbiota in infants of obese mothers increases inflammation and susceptibility to NAFLD. Nature Communications 9 (1): 4462.
Homo Sapiens UCSC Hg19 Human Genome Sequence from iGenome.
Langmead, B., and S. L. Salzberg. 2012. Fast gapped-read alignment with Bowtie 2. Nature Methods 9 (4): 357–359.
Markle, J. G., et al. 2013. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339 (6123): 1084–1088.
Ewing, B., and P. Green. 1998. Base-calling of automated sequencer traces using Phred. II. error probabilities. Genome Research 8 (3): 186–194.
Ewing, B., et al. 1998. Base-calling of automated sequencer traces using Phred. I. accuracy assessment. Genome Research 8 (3): 175–185.
Edgar, R. C., et al. 2011. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27 (16): 2194–2200.
Schloss, P. D., and S. L. Westcott. 2011. Assessing and improving methods used in operational taxonomic unit-based approaches for 16S rRNA gene sequence analysis. Applied and Environmental Microbiology 77 (10): 3219–3226.
Pruesse, E., J. Peplies, and F. O. Glöckner. 2012. SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28 (14): 1823–1829.
Quast, C., et al. 2013. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research 41 (Database issue): D590-6.
Robertson, C. E., et al. 2013. Explicet: Graphical user interface software for metadata-driven management, analysis and visualization of microbiome data. Bioinformatics 29 (23): 3100–3101.
Percie Du Sert, N., et al. 2020. The ARRIVE guidelines 2.0: Updated guidelines for reporting animal research. PLOS Biology 18 (7): e3000410.
Percie Du Sert, N., et al. 2020. Reporting animal research: Explanation and elaboration for the ARRIVE guidelines 2.0. PLOS Biology 18 (7): e3000411.
Acknowledgements
This study was supported by U01HL152405, T32AI074491, State of Colorado Office of Economic Development and International Trade Advanced Industries Accelerator Program. The funders had no role in the study design, data collection and analysis, decision to publish or preparation of the manuscript.
The authors thank the following for their aid: Francesca Cendali and Angelo D’Alessandro (University of Colorado School of Medicine Metabolomics Core); Linxing Yao and Prithwiraj De (Colorado State University, Analytical Resources Core—Bioanalysis and Omics); and Scott Beard (Barbara Davis Center Flow Cytometry Core).
Author information
Authors and Affiliations
Contributions
SF and BEA contributed equally.
SF, BEA, VMH, and KAK designed the study, analyzed and interpreted the data, and wrote the manuscript.
SF, BEA, MEC, WKJ, CER, JNK, and DNF generated data.
All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fechtner, S., Allen, B.E., Chriswell, M.E. et al. 3,3-Dimethyl-1-Butanol and its Metabolite 3,3-Dimethylbutyrate Ameliorate Collagen-induced Arthritis Independent of Choline Trimethylamine Lyase Activity. Inflammation (2024). https://doi.org/10.1007/s10753-024-02126-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10753-024-02126-y