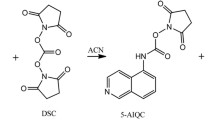
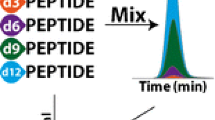
Abstract
Amino acid analysis is a powerful bioanalytical technique for many biomedical research endeavors, including cancer, emergency medicine, nutrition and neuroscience research. In the present study, we present a 3 min analytical method for underivatized amino acid analysis that employs ultra high-performance liquid chromatography and high-resolution quadrupole orbitrap mass spectrometry. This method has demonstrated linearity (mM to nM range), reproducibility (intra-day <5 %, inter-day <20 %), sensitivity (low fmol) and selectivity. Here, we illustrate the rapidity and accuracy of the method through comparison with conventional liquid chromatography–mass spectrometry methods. We further demonstrate the robustness and sensitivity of this method on a diverse range of biological matrices. Using this method we were able to selectively discriminate murine pancreatic cancer cells with and without knocked down expression of hypoxia-inducible factor 1α; plasma, lymph and bronchioalveolar lavage fluid samples from control versus hemorrhaged rats; and muscle tissue samples harvested from rats subjected to both low-fat and high-fat diets. Furthermore, we were able to exploit the sensitivity of the method to detect and quantify the release of glutamate from sparsely isolated murine taste buds. Spiked in light or heavy standards (13C6-arginine, 13C6-lysine, 13C 155 N2-glutamine) or xenometabolites (5-fluorouracil) were used to determine coefficients of variation, confirm linearity of relative quantitation in four different matrices, and overcome matrix effects for absolute quantitation. The presented method enables high-throughput analysis of low-abundance samples requiring only one percent of the material extracted from 100,000 cells, 10 µl of biological fluid, or 2 mg of muscle tissue.




Similar content being viewed by others
Abbreviations
- 3AA:
-
Three-minute method for amino acid analysis
- BALF:
-
Bronchioalveolar lavage fluid
- CV:
-
Coefficient of variation
- GC/MS:
-
Gas chromatography/mass spectrometry
- HFD:
-
High-fat diet
- HIF1α:
-
Hypoxia-inducible factor 1α
- HILIC:
-
Hydrophilic interaction liquid chromatography
- HPLC:
-
High-performance liquid chromatography
- LFD:
-
Low-fat diet
- LOD:
-
Limit of detection
- LOQ:
-
Limit of quantification
- MAP:
-
Mean arterial pressure
- MLD:
-
Mesenteric lymph diversion
- MRM:
-
Multiple reaction monitoring
- MS:
-
Mass spectrometry
- m/z :
-
Mass-to-charge ratio
- PCA:
-
Principal component analysis
- PLS-DA:
-
Partial least-square discriminant analysis
- SD:
-
Standard dilution
- S/N:
-
Signal to noise ratio
- T/HS:
-
Trauma/hemorrhagic shock
- T/SS:
-
Trauma/sham shock
- TOF:
-
Time of flight
- UHPLC:
-
Ultra high-performance liquid chromatography
- XIC:
-
Extracted ion chromatogram
References
Amelio I, Cutruzzolá F, Antonov A et al (2014a) Serine and glycine metabolism in cancer. Trends Biochem Sci 39:191–198
Amelio I, Markert EK, Rufini A et al (2014b) p73 regulates serine biosynthesis in cancer. Oncogene 33(42):5039–5046
Armstrong M, Jonscher K, Reisdorph NA (2007) Analysis of 25 underivatized amino acids in human plasma using ion-pairing reversed-phase liquid chromatography/time-of-flight mass spectrometry. Rapid Commun Mass Spectrom RCM 21:2717–2726
Badawy AA-B (2012) The EZ: faast family of amino acid analysis kits: application of the GC-FID kit for rapid determination of plasma tryptophan and other amino acids. Methods Mol Biol Clifton NJ 828:153–164
Benson JR, Hare PE (1975) O-phthalaldehyde: fluorogenic detection of primary amines in the picomole range. Comparison with fluorescamine and ninhydrin. Proc Natl Acad Sci U S A 72:619–622
Bidlingmeyer BA, Cohen SA, Tarvin TL (1984) Rapid analysis of amino acids using pre-column derivatization. J Chromatogr B Biomed Sci App 336:93–104
Buiarelli F, Gallo V, Di Filippo P et al (2013) Development of a method for the analysis of underivatized amino acids by liquid chromatography/tandem mass spectrometry: application on Standard Reference Material 1649a (urban dust). Talanta 115:966–972
Cardaci S, Ciriolo MR (2012) Deprive to kill: glutamine closes the gate to anticancer monocarboxylic drugs. Autophagy 8:1830–1832
Castorena CM, Arias EB, Sharma N, Cartee GD (2014) Postexercise improvement in insulin-stimulated glucose uptake occurs concomitant with greater AS160 phosphorylation in muscle from normal and insulin-resistant rats. Diabetes 63:2297–2308
Chen G, Wang J (2014) Threonine metabolism and embryonic stem cell self-renewal. Curr Opin Clin Nutr Metab Care 17:80–85
Clasquin MF, Melamud E, Rabinowitz JD (2012) LC-MS data processing with MAVEN: a metabolomic analysis and visualization engine. Curr Protoc Bioinforma Ed Board Andreas Baxevanis Al Chapter 14:Unit14.11
Cubbon S, Antonio C, Wilson J, Thomas-Oates J (2010) Metabolomic applications of HILIC-LC-MS. Mass Spectrom Rev 29:671–684
D’Alessandro A, Zolla L (2013) Proteomics and metabolomics in cancer drug development. Expert Rev Proteomics 10:473–488
D’Alessandro A, Gevi F, Zolla L (2011) A robust high resolution reversed-phase HPLC strategy to investigate various metabolic species in different biological models. Mol BioSyst 7:1024–1032
D’Alessandro A, Gevi F, Palini S et al (2012a) A mass spectrometry-based targeted metabolomics strategy of human blastocoele fluid: a promising tool in fertility research. Mol BioSyst 8:953–958
D’Alessandro A, Giardina B, Gevi F et al (2012b) Clinical metabolomics: the next stage of clinical biochemistry. Blood Transfus Trasfus Sangue 10(Suppl 2):s19–s24
D’Alessandro A, Amelio I, Berkers CR et al (2014a) Metabolic effect of TAp63α: enhanced glycolysis and pentose phosphate pathway, resulting in increased antioxidant defense. 5(17):7722–7733
D’Alessandro A, Cervia D, Catalani E et al (2014b) Protective effects of the neuropeptides PACAP, substance P and the somatostatin analogue octreotide in retinal ischemia: a metabolomic analysis. Mol BioSyst 10(6):1290–1304
D’Alessandro A, Nemkov T, Kelher M et al (2014c) Routine storage of red blood cell (RBC) units in additive solution-3: a comprehensive investigation of the RBC metabolome. Transfusion. doi:10.1111/trf.12975
D’Alessandro A, Kriebardis AG, Rinalducci S et al (2015a) An update on red blood cell storage lesions, as gleaned through biochemistry and omics technologies. Transfusion 55(1):205–219
D’Alessandro A, Moore HB, Moore EE et al (2015b) Early hemorrhage triggers metabolic responses that build up during prolonged shock. Am J Physiol Regul Integr Comp Physiol. doi:10.1152/ajpregu.00030.2015
Dorresteijn RC, Berwald LG, Zomer G et al (1996) Determination of amino acids using o-phthalaldehyde-2-mercaptoethanol derivatization effect of reaction conditions. J Chromatogr A 724:159–167
Droge W (2005) Oxidative stress and ageing: is ageing a cysteine deficiency syndrome? Philos Trans R Soc B Biol Sci 360:2355–2372
Dunn WB, Lin W, Broadhurst D et al (2014) Molecular phenotyping of a UK population: defining the human serum metabolome. Metabolomics 11:9–26
Fonville JM, Richards SE, Barton RH et al (2010) The evolution of partial least squares models and related chemometric approaches in metabolomics and metabolic phenotyping. J Chemom 24:636–649
Fürst P, Pollack L, Graser TA et al (1990) Appraisal of four pre-column derivatization methods for the high-performance liquid chromatographic determination of free amino acids in biological materials. J Chromatogr 499:557–569
Heinrikson RL, Meredith SC (1984) Amino acid analysis by reverse-phase high-performance liquid chromatography: precolumn derivatization with phenylisothiocyanate. Anal Biochem 136:65–74
Hirayama A, Soga T (2012) Amino acid analysis by capillary electrophoresis-mass spectrometry. Methods Mol Biol Clifton NJ 828:77–82
Hiscock N, Pedersen BK (2002) Exercise-induced immunodepression– plasma glutamine is not the link. J Appl Physiol 93:813–822
Husain FA, Martin MJ, Mullenix PS et al (2003) Serum lactate and base deficit as predictors of mortality and morbidity. Am J Surg 185:485–491
Kaspar H, Dettmer K, Gronwald W, Oefner PJ (2008) Automated GC–MS analysis of free amino acids in biological fluids. J Chromatogr B 870:222–232
Le A, Ng A, Kwan T et al (2014) A rapid, sensitive method for quantitative analysis of underivatized amino acids by liquid chromatography-tandem mass spectrometry (LC-MS/MS). J Chromatogr B Analyt Technol Biomed Life Sci 944:166–174
Maddocks ODK, Berkers CR, Mason SM et al (2013) Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature 493(542–546):3
Manning JM (1993) The contributions of Stein and Moore to protein science. Protein Sci Publ Protein Soc 2:1188–1191
Moore S, Spackman DH, Stein WH (1958) Chromatography of Amino Acids on Sulfonated Polystyrene Resins. An Improved System. Anal Chem 30:1185–1190
Morrison AB, Middleton EJ, McLaughlan JM (1961) Blood amino acid studies: ii. effects of dietary lysine concentration, sex, and growth rate on plasma free lysine and threonine levels in the rat. Can J Biochem Physiol 39:1675–1680
Pan Z, Gu H, Talaty N et al (2007) Principal component analysis of urine metabolites detected by NMR and DESI-MS in patients with inborn errors of metabolism. Anal Bioanal Chem 387:539–549
Panopoulos AD, Yanes O, Ruiz S et al (2012) The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Res 22:168–177
Peltz E, D’Alessandro A, Moore E et al (2015) Pathologic metabolism: an exploratory study of the plasma metabolome of critical injury. J Trauma Acute Care Surg. doi:10.1097/TA.0000000000000589
Piraud M, Vianey-Saban C, Petritis K et al (2005) Ion-pairing reversed-phase liquid chromatography/electrospray ionization mass spectrometric analysis of 76 underivatized amino acids of biological interest: a new tool for the diagnosis of inherited disorders of amino acid metabolism. Rapid Commun Mass Spectrom 19:1587–1602
Platten M, Wick W, Van den Eynde BJ (2012) Tryptophan catabolism in cancer: beyond IDO and tryptophan depletion. Cancer Res 72:5435–5440
Prada PO, Hirabara SM, de Souza CT et al (2007) l-glutamine supplementation induces insulin resistance in adipose tissue and improves insulin signalling in liver and muscle of rats with diet-induced obesity. Diabetologia 50:1949–1959
Rutherfurd SM, Gilani GS (2009) Amino acid analysis. Curr Protoc Protein Sci Editor Board John E Coligan Al Chapter 11:Unit 11.9
Salazar C, Armenta JM, Cortés DF, Shulaev V (2012) Combination of an AccQ·Tag-ultra performance liquid chromatographic method with tandem mass spectrometry for the analysis of amino acids. Methods Mol Biol Clifton NJ 828:13–28
Shimbo K, Oonuki T, Yahashi A et al (2009) Precolumn derivatization reagents for high-speed analysis of amines and amino acids in biological fluid using liquid chromatography/electrospray ionization tandem mass spectrometry. Rapid Commun Mass Spectrom 23:1483–1492
Stein S, Böhlen P, Stone J et al (1973) Amino acid analysis with fluorescamine at the picomole level. Arch Biochem Biophys 155:203–212
Stringham JR, Moore EE, Gamboni F et al (2014) Mesenteric lymph diversion abrogates 5-lipoxygenase activation in the kidney following trauma and hemorrhagic shock. J Trauma Acute Care Surg 76:1214–1221
Thiele B, Stein N, Oldiges M, Hofmann D (2012) Direct analysis of underivatized amino acids in plant extracts by LC-MS-MS. Methods Mol Biol Clifton NJ 828:317–328
Vandenbeuch A, Tizzano M, Anderson CB et al (2010) Evidence for a role of glutamate as an efferent transmitter in taste buds. BMC Neurosci 11:77
Yamaguchi S, Ninomiya K (2000) Umami and Food Palatability. J Nutr 130:921S–926S
Yang X-D, Ma JYC, Barger MW, Ma JKH (2002) Transport and utilization of arginine and arginine-containing peptides by rat alveolar macrophages. Pharm Res 19:825–831
Yang W-C, Mirzaei H, Liu X, Regnier FE (2006) Enhancement of amino acid detection and quantification by electrospray ionization mass spectrometry. Anal Chem 78:4702–4708
Yao X, Zhou G, Tang Y et al (2013) Direct determination of underivatized amino acids from Ginkgo biloba leaves by using hydrophilic interaction ultra high performance liquid chromatography coupled with triple quadrupole mass spectrometry. J Sep Sci 36:2878–2887
Yuan M, Breitkopf SB, Yang X, Asara JM (2012) A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat Protoc 7:872–881
Zhang Y, Dai Y, Wen J et al (2011) Detrimental effects of adenosine signaling in sickle cell disease. Nat Med 17:79–86
Zhou G, Pang H, Tang Y et al (2013) Hydrophilic interaction ultra-performance liquid chromatography coupled with triple-quadrupole tandem mass spectrometry for highly rapid and sensitive analysis of underivatized amino acids in functional foods. Amino Acids 44:1293–1305
Acknowledgments
The Authors are grateful to Drs. Anthony W. Bacon, Aurelie Vandenbeuch, Bryan Bergman, Craig Jordan, Carlos M. Castorena, Gregory D. Cartee, Jeniann Yi, Hunter Moore, Sean Newsom, Agnieszka Kendrick for providing the biological samples, as detailed in the paper. The authors would also like to thank Dr. Robert Hodges for helpful discussions. Research reported in this publication was supported by the National Institute of General Medical Sciences of the National Institutes of Health under Award Numbers P50GM049222, R33CA183685 and T32GM008315, and Grant #P50 GM049222 from NIGMS, NIH (CS). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Conflict of interest
All the authors disclose no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: D. Tsikas.
A. D’Alessandro and T. Nemkov contributed equally and share the first authorship.
Electronic supplementary material
Below is the link to the electronic supplementary material.
726_2015_2019_MOESM3_ESM.tif
Supplementary material 3 (TIFF 16061 kb). Supplementary Fig. 1 – Extracted Ion Chromatograms of amino acids. Extracted Ion Chromatograms (XIC) for 32 representative amino acids assayed in this study have been overlaid to demonstrate chromatographic separation (top). Amino acid categories are color coded as follows: aromatic (pink), acidic (red), basic (blue), neutral (yellow), imino (light blue), or other (green) amino acids. All the single XICs are presented to illustrate peak shape (bottom). Parts per million (ppm) error value windows on the intact mass needed to detect each amino acid are provided for each XIC
726_2015_2019_MOESM4_ESM.tif
Supplementary material 4 (TIFF 25866 kb). Supplementary Fig. 2 – Quantitative measurements of 15 representative basic, acidic, neutral and aromatic amino acid standards and relative quadratic correlations (RSQ). Extracted Ion Chromatograms for 15 representative amino acids are depicted in each panel. The inset depicts peak areas for each injection amount, with the RSQ value indicated beneath the bar graph. A mass-to-charge window used for detection is indicated next to the amino acid name to highlight high mass accuracy of the platform
726_2015_2019_MOESM5_ESM.tif
Supplementary material 5 (TIFF 3195 kb). Supplementary Fig. 3 – Comparability of the results obtained through the alternative methods for amino acid analysis using underivatized UHPLC/MS. Pancreatic cancer cell extracts were compared using the three min C18 method and two conventional HILIC or Amide column 15 min methods. Raw samples were either not-diluted prior to injection (ND) or injected upon a fivefold dilution. Raw values were Z-score normalized for each amino acid (intra-row) and plotted as heat maps (blue to red = one- to fivefold increase in amino acid level intra-row) using the software GENE E (Broad Institute). Consistently, undiluted samples were detected to have higher levels of amino acids through all three methods, but the C18 three min method was the one showing the average of the medians of amino acid fold changes across replicates closer to five (red columns for ND samples)
726_2015_2019_MOESM6_ESM.tif
Supplementary material 6 (TIFF 2867 kb). Supplementary Fig. 4 – Extracted ion chromatograms for heavy labeled 13C6-arginine and 5-fluorouracil in thirty-five different rat plasma samples. Coefficients of variation (CVs) were calculated (standard deviation/mean) to test technical reproducibility of the approach
Rights and permissions
About this article
Cite this article
Nemkov, T., D’Alessandro, A. & Hansen, K.C. Three-minute method for amino acid analysis by UHPLC and high-resolution quadrupole orbitrap mass spectrometry. Amino Acids 47, 2345–2357 (2015). https://doi.org/10.1007/s00726-015-2019-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-015-2019-9