Abstract
Kale (Brassica oleracea. var. acephala) is a nutrient-rich green leafy vegetable consumed as food and used in traditional medicine worldwide. An essential step in describing the available genetic resources and ensuring their effective use in breeding programs is to characterize the genetic diversity of available germplasm. In this study, the genetic diversity and structure of 26 kale accessions from South-East Europe were examined using 26 agro-morphological traits collected in the field and 12 simple sequence repeat (SSR) markers. Considerable agro-morphological variability was found in most quantitative (CV = 17.26–42.42%) and qualitative (H' = 0.61–1.79) traits. Multifactorial analysis (MFA) showed that country of origin (33.01%) and morphotype (32.30%) significantly influenced kale diversification. Leaf blade shape (20.62%), leaf incision (19.43%), anthocyanin distribution (16.43%), and leaf colour (15.55%) were the traits that most clearly differentiated accessions. The three common commercial kale cultivars were identified as independent outliers that differed from the other kale accessions in both MFA and UPGMA clustering analysis. The SSR markers were highly informative with 108 alleles and polymorphic information content ranging from 0.395 to 0.856. Strong genetic diversity was detected at the accession level (H' = 0.58) while genetic differentiation was low (Fst = 0.05). Similar to UPGMA clustering, Bayesian clustering suggests that the kale collection can be divided into four clusters with a high degree of admixture and no geographic grouping pattern is apparent. Overall, the study showed that the kale collection studied represents a valuable reservoir of genetic and agro-morphological variability that could be used for future breeding initiatives.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The diploid species (2n = 18) Brassica oleracea L. (Brassicaceae), the most diverse species in the genus Brassica, includes several vegetable crops with a long history of cultivation and domestication worldwide, such as cauliflower, cabbage, broccoli, Brussels sprouts, kohlrabi, and kale (Golicz et al. 2016). In particular, kale leaves (B. oleracea var. acephala) has attracted great interest due to its excellent nutritional value compared to other crops. It is considered an ideal source of vitamins, essential minerals, and fatty acids (Pathirana et al. 2017). Therefore, it occupies a privileged position in the cuisine and diet of European, Asian and American populations (Gonçalves et al. 2012). Moreover, it is widely used in traditional medicine for the treatment of rheumatism, liver diseases, eye problems, bone weakness, anaemia, and obesity (Gonçalves et al. 2012; Kuerban et al. 2017; Thavarajah et al. 2016) and also as an animal feed (Cartea et al. 2003).
Brassica species are known as a highly diverse group of plants due to their ability of spontaneous cross-pollination and gene flow among sexually compatible relatives (Meglič and Pipan 2018; Pipan et al. 2013, 2011). B. oleracea var. acephala is classified into different morphotypes based on morphological characteristics. For example, kale (B. oleracea L. var. acephala DC.) has dark green and crinkled leaves, Scotch kale (B. oleracea L. var. acephala (DC.) Alef. var. sabellica L.) is characterized by grey-green and highly crinkled and wrinkled leaves, while Marrow stem kale (B. oleracea L. var. acephala (DC.) Alef. var. medullosa L.) is characterized by a soft and thick stem and different leaf types. In addition to variation among morphotypes, populations/cultivars within the same morphotype may also exhibit wide morphological variation due to a long history of domestication and adaptation to environmental conditions (Hahn et al. 2022). According to Hahn et al. (2022), three types of kale are distinguished according to their origin: curly kale (Scotch type, Brassica oleracea covar. acephala var. sabellica), Italian kale (Lacinato type, Brassica oleracea covar. acephala var. palmifolia), and collard (Brassica oleracea covar. acephala var. viridis). Indeed, in B. oleracea var. acephala, large morphological variation could occur both within populations as a result of cross-pollination and between populations due to extensive selection by farmers and/or adaptation to local environments (Cartea et al. 2003; Šamec et al. 2019a; Thavarajah et al. 2016).
Detailed agro-morphological characterization and assessment of patterns of genetic variability based on quantitative and qualitative traits is the first and crucial step in describing available genetic resources and their efficient use in breeding programs (Balkaya and Yanmaz 2005; Oumata et al. 2023). However, although agro-morphological traits can be controlled at the genetic level, they may be nonspecific and non-polymorphic, and their variation could be strongly influenced by environmental factors (Choudhury et al. 2022; Petit et al. 2020; Terlević et al. 2023). In contrast, DNA-based molecular markers are stable, intact, abundant in the genome, and free from environmental influences. Therefore, they are a powerful tool not only for comprehensive depiction the genetic diversity patterns, but also for other advanced analyses such as gene mapping and molecular marker-assisted breeding (Zhu et al. 2019). Although various molecular markers have been developed to date, simple requence Repeats (SSRs) or microsatellites remain the most attractive for genetic variability research and plant breeding because they are codominantly inherited, abundant, multi-allelic and highly polymorphic (Riangwong et al. 2020; Rivera et al. 2016), and are also easy to handle in the laboratory or automated by capillary sequencers (Schuelke 2000). Therefore, assessment of genetic variation using agro-morphological traits and SSR markers has proven to be a very powerful approach for genetic resources management, utilization, and conservation in many crops, including chickpea (Ghaffari et al. 2014), tomato (Mercati et al. 2015), pepper (Rivera et al. 2016), leafy mustard (Sharma et al. 2020), and cauliflower (Rakshita et al. 2021).
In the present study, a large set of quantitative and qualitative agro-morphological traits and SSRs markers were used to evaluate the genetic diversity and pattern of genetic structure of B. oleracea L. var. acephala collection from South-East Europe. The results of the study will be very useful for further use of the analysed accessions in ongoing and future breeding programs.
Materials and methods
Plant material
The plant material used in the study consists of 26 accessions of B. oleracea var. acephala, of which 23 accessions were B. oleracea L. var. acephala DC obtained from the two national plant gene banks, i.e. Croatia (six accessions) and Bosnia and Herzegovina (17 accessions). The remaining three accessions were commercial cultivars developed by three different seed producers. 'Kodrolistni ohrovt' and 'Krmni ohrovt' were two cultivars of Scotch kale and Marrow stem kale developed by Semenarna Ljubljana (Slovenia) and Semina Royal Seeds (Slovenia), respectively, while 'Ohrovt Nero di Toscana' was a cultivar of Lacinato kale, B. oleracea var. palmifolia developed by Franchi Sementi (Italy). All available passport data for the studied accessions are summarised in Supplementary Table S1.
Experimental design and agro-morphological characterization
In March 2019, four individuals from each accession were initially cultivated in seedling trays filled with fertilized peat. Two months after germination, seedlings were transplanted to open-field conditions and grown to maturity. Adjacent plants were spaced 50 cm apart and a black polyethylene cover was placed on the soil before planting. Each plant was phenotypically characterised using 11 quantitative and 15 qualitative non-destructive descriptors for Brassica spp. The whole plant, entire leaf, leaf blade and petiole were characterised according to the European Cooperative Programme for Plant Genetic Resources developed by the Community Plant Variety Office (CPVO 2011) and the International Board for Plant Genetic Resources (IBPGR 1990). Quantitative traits included plant height [PLANTHEIGHT], plant diameter [PLANTDIAM], plant height/diameter ratio [PLANTRATIO], leaf length [LEAFLENGTH], leaf area [LEAFAREA], leaf blade length [BLADELENGTH], leaf blade width [BLADEWIDTH], leaf blade width/leaf length ratio [BLADERATIO], petiole length [PETIOLELENGTH], petiole width [PETIOLEWIDTH], and petiole length/width ratio [PETIOLERATIO]. For qualitative traits, plant shape [PLANTSHAPE] and position of growing point relative to the top of the plant [PLANTPOSIT] were used to characterise plant shape. Leaf division (margin) [LEAFMARGIN], leaf division (incision) [LEAFINCISION], leaf apex shape [LEAFAPEX], leaf tip attitude [LEAFTIP], leaf lamina attitude [LEAFLAMINA], leaf anthocyanin coloration [LEAFANTHO], distribution of anthocyanin coloration on the leaf [LEAFDISTRIB], leaf colour [LEAFCOLOUR], and intensity of the colour of the fully developed leaf [LEAFINTENS] were evaluated to describe leaf morphology. The blade shape [BLADESHAPE] and blade blistering [BLADEBLISTER] were also used to characterise the leaf blade. Finally, the petiole by assessing petiole and/or midvein enlargement [PETIOLEMEDVEIN] and the colour of the petiole and/or midvein colour [PETIOLECOLOUR] were characterised. The full description of all analysed traits is given in the Supplementary Table S2. In addition to agro-morphological characterization, a young healthy leaf was taken from 2–3 plants per accession and stored at –80 °C for further molecular characterization. Non-healthy plants were excluded from agro-morphological and molecular characterization. The experiment was conducted at the experimental fields of the Agricultural Institute of Slovenia in Jablje (304 m a.s.l.; 46.151°N 14.562°E).
Molecular analysis
First, genomic DNA was extracted from 100 mg young and healthy leaves from 2 to 3 individuals per accession as described in (Pipan et al. 2017). DNA from all individuals was then checked for quality and quantity using a fluorimeter (Qubit 3.0; ThermoFisher Scientific, MA, USA) and then diluted to a uniform final concentration of 5.6 ng/μL and stored at –20 °C for further analysis.
DNAs from a total of 71 individuals were used separately (2–3 individuals per accession) and then amplified with a set of 12 genome-specific SSR markers previously used to analyse the genetic diversity of different B. oleracea crops (kale, cabbage, cauliflower, and Brussels sprouts) and found to be highly polymorphic (El-Esawi et al. 2016). PCR amplification was performed using fluorescently labelled universal primers according to the protocol described by (Schuelke 2000). First, for each SSR marker, the forward primer was labelled with the M13 sequence (5'-CACGACGTTGTAAAACGAC-3'). The PCR reaction was performed in a final volume of 11 μL containing 5.6 ng genomic DNA, 1 μL 10 × PCR buffer (Biotools, Spain), 0.5 μL 50 mM MgCl2 (Biotools, Spain), 0.2 μL each 10 mM dNTPs (Sigma-Aldrich, USA), 0.1 μL 10 μM forward primer (Sigma-Aldrich, USA), 0.25 μL 10 μM reverse primer (Sigma-Aldrich, USA), 0.183 μL 10 μM 5'-fluorescently labelled universal primer (with 6-FAM, NED or HEX; Omega, Slovenia), and 0.5 μL 5 U Taq DNA polymerase (Biotools, Spain). PCR amplification was performed in a thermal cycler (Veriti, ThermoFisher Scientific) using the following primer-specific touchdown profile: 94 °C for 4 min; 15 cycles at 94 °C for 1 min; temperature reduction from 60 (62) °C to 49.5 (51.5) °C at 0.7 °C per cycle for 30 s; 72 °C for 1 min; followed by 23 cycles at 94 °C for 30 s; 53 °C for 30 s; 72 °C for 1 min; and final extension for 5 min at 72 °C.
Ultimately, an Applied Biosystems ABI3130XL genetic analyser was used to resolve the DNA fragments. The allele sizes were then determined by comparison with an internal size standard (GeneScan-500 ROX; Applied Biosystems) using the GeneMapper 6.0 software (Applied Biosystems).
Agro-morphological data analysis
The obtained data were statistically analysed with a series of univariate and multivariate approaches using R statistical programming environment version 3.4.4 R Core Team (2021). First, the package "ggplot2" (Villanueva and Chen 2019) was used to visualise the frequency of distribution of all agro-morphological traits across accessions using histograms. To describe patterns of the agro-morphological variation, the package "summarytools"(Comtois and Comtois 2016) was used to compute basic descriptive statistics, including maximum and minimum values (max and min), standard deviation (SD), and coefficient of variation (CV) for the quantitative traits. Analysis of variance (ANOVA) was used to partition variation within and between accessions using the "aov" function of the "Stats" package (R Core Team 2021). For qualitative traits, the "vegan" package (Oksanen et al. 2019) was used to compute the Shannon–Weaver diversity index (Shannon and Weaver 1949) as a measure of phenotypic diversity. Patterns of correlations between pairwise of quantitative and qualitative traits was estimated separately via Pearson (r) and Cramer’s V (v) coefficients, with P < 0.05 set as significance level, using packages "corrplot" (Wei et al. 2017) and "vcd" (Meyer et al. 2021), respectively.
A multifactorial analysis (MFA) was conducted using the "FactoMineR" (Lê et al. 2008) and "factoextra" packages (Kassambara and Mundt 2020) to examine patterns of the agro-morphological differentiation and determine the factors and quantitative and qualitative agro-morphological traits that explain variability among accessions. Gower distance (Gower 1971) was calculated using the "daisy" function in the "cluster" package (Maechler et al. 2012) to create a dissimilarity matrix between accessions based on all traits, which was eventually used to visualise the relationship between accessions UPGMA clustering using the "hclust" function.
SSR data analysis
The data matrix resulting from allele sizing was first analysed using Power Marker version 3.25 with the default parameters (Liu and Muse 2005) to calculate several measures describing the genetic diversity at the loci and accession level, including number of different alleles (Na), effective number of alleles (Ne), major allele frequency (MAF), and polymorphic information content (PIC). We also used the default parameter settings in GenAlEx 6.5.2 software (Peakall and Smouse 2006) to calculate the Shannon information index (I), observed heterozygosity (Ho), and expected heterozygosity (He), as well as the fixation index (F) and the number of private alleles (No. PA). The same software was also used for the analysis of molecular variance (AMOVA) to estimate the proportion of variation explained within accessions, between accessions, and between geographic origins of accessions. In addition, to examine the genetic structure of the studied accessions using the data from the 12 SSR loci, Bayesian clustering was performed using the software STRUCTURE 2.3.4 (Pritchard et al. 2000). The analysis was performed using the Admixture model by adjusting the number of populations (K) from 1 to 10 based on 10 independent simulations for each K. Each simulation consisted of a burn-in period of 10.000 steps and 100.000 Markov Chain Monte Carlo (MCMC). The presumed number of clusters (optimal K) was then determined using the software STRUCTURE HARVESTER (Earl and VonHoldt 2012). Using the R package StructuRly 0.1.0 (Criscuolo and Angelini 2020), the Bayesian clusters of the studied accessions were visualised in the form of admixture bar plots. The genetic relationship between accessions was visualised via an unweighted pair group with arithmetic average (UPGMA) clustering using Poptree2 (Takezaki et al. 2010) based on the matrix of Nei genetic distance between accessions pairwise generated by GenAlEx 6.5.2 software (Peakall and Smouse 2006). Genetic relatedness among kale accessions was further analysed using UPGMA (Unweighted Pair Group Method Arithmetic Average) cluster analysis based on Nei genetic distance, and clustering accuracy was tested using 1000 bootstrap resamples with the R package "poppr" (Kamvar et al. 2014). Finally, the matrix of Nei’s genetic distance between accessions pairwise was used along with the dissimilarity matrix between accessions based on Gower distance to perform a Mantel test with the package "vegan" (Oksanen et al. 2019) to test the degree of association between agro-morphological and molecular variation of the studied kale accessions.
Results
Morpho-agronomic variability of kales from South-East Europe
A total of 26 quantitative and qualitative morpho-agronomic descriptors related to plants, leaves, leaf blades, and petioles were evaluated for these 26 kale accessions originating from South-East Europe. Descriptive statistics revealed that the 11 quantitative traits had moderate (10% ≤ CV ≤ 20%) to very high (CV ≥ 30%) morpho-agronomic variation for all traits with an average coefficient of variation (CV) of 25.33% (Table 1). The frequencies of distribution of the 26 analysed accessions for the 11 quantitative traits are shown in Fig. 1. Plant height was ≤ 60 cm in 16 accessions with a mean value of 61.34 cm and a CV of 24.03%, plant diameter exceeded 60 cm in 18 accessions with an average of 57.97 cm and a CV of 19.46%, while height/diameter was above the average (1.09) in 11 accessions with a CV value of 17.38%. As for leaf characteristics, the length in most accessions (18) was 31–40 cm with a mean value of 36.39 cm and a CV of 19.22%, while the area in 17 accessions was 200–400 cm2 with a mean value of 261.94 cm2 and a CV of 42.42%. Phenotypic evaluation of leaf blade showed that leaf blade length ranged from 16 to 21 cm in 13 accessions, with an average of 19.19 cm and a CV of 19.07% for all collection. Leaf blade width was ≤ 15 cm in 8 accessions and 18–19 cm in 8 accessions with an average of 16.77 cm and a CV of 22.68%, while most accessions (16) had a width/length ratio above the average (0.46) with a CV of 17.26%. As for petiole, length in most accessions was between 14 and 19 cm with an average of 17.02 cm (CV = 32.07%), width in 15 accessions was above average (1.14) with CV of 25.19%, while length/width ratio in 14 accessions was above average (16.06) with CV of 39.86%. Correlations between pairwise quantitative traits (Fig. 3A) showed that, with the exception of the ratio between plant, leaf blade, and petiole, most of the remaining traits had strong positive associations with each other (r = 0.39, P < 0.05). The highest correlations were observed between leaf and petiole length (r = 0.86), leaf area and the blade width (r = 0.82), plant diameter and leaf length (r = 0.79), and plant diameter and leaf petiole length (r = 0.76), most likely reflecting the strong developmental associations among traits related to plant vigour. Interestingly, analysis of variance revealed a significant difference (P < 0.0001) among accessions for all quantitative traits. As shown in Fig. 4A, variation between accessions far exceeded variation within accessions, with a level of variation greater than 50% and an average of 73.98% for all traits. In addition, as indicated by the Shannon–Weaver index (H'), a wide variation was observed in all qualitative morphological descriptors. For instance, plant shape (H' = 0.69) was either inverted pyramid or a dome (13 accessions). In the latter, the position of the growing point (H' = 0.61) was deeply below the plant top for 10 accessions and slightly below in only 3 accessions (Fig. 2). Regarding leaf shape, the leaf margin (H' = 1.41) was mostly crenate and serrate (9 and 6 accessions, respectively), while the incision (H' = 1.02) was mostly lyrate and sinuate (14 and 9 accessions, respectively). Moreover, most accessions exhibited either a rounded (11) or broadly rounded (8) leaf apex (H' = 1.07), the tip was mostly dropping (17) (H' = 0.76), and the lamina was mostly either concave drooping (12) or straight (12) lamina (H' = 0.91). The blade shape (H' = 1.79) was mostly obovate (11) or oblong (5), while blade blistering (H' = 1.54) was mostly low (11) or intermediate (7) (Fig. 2). In addition, anthocyanin (H' = 0.68) was visible in the leaves of only 11 accessions, 10 of which had partial distribution in the leaf (H' = 0.49). In most accessions, leaf colour (H' = 1.03) was mostly green (13) or dark green (9) and intensity (H' = 1.01) was mostly medium (12) or dark (10). Petiole colour (H' = 1.21) was mostly light green (9), green (8), or purple (8) (Fig. 2), while petiole and/or midvein enlargement (H' = 0.82) was either intermediate (13) or narrow (12) in most accessions. Moreover, as revealed by the Cramer’s coefficient (Fig. 3B), plant shape and growing point position, as well as anthocyanin coloration and its distribution in the leaf, were completely interdependent (r = 1). Some other traits appeared to be strongly (r ≥ 0.75) interdependent, such as leaf colour and its intensity, as well as the blade shape and the distribution of anthocyanin coloration on the leaf. In contrast, no association was detected between leaf anthocyanin coloration and plant shape, and a weak association was found between leaf apex shape and anthocyanin coloration and its distribution on the leaf (r ≤ 0.15).
Frequency distribution of the 26 kale accessions for the 11 quantitative traits. a Plant height [PLANTHEIGHT]. b Plant diameter [PLANTDIAM]. c Plant height/diameter ratio [PLANTRATIO]. d Leaf length [LEAFLENGTH]. e Leaf area [LEAFAREA]. f Leaf blade length [BLADELENGTH] g Leaf blade width [BLADEWIDTH]. h Leaf blade width/leaf length ratio [BLADERATIO]. i Petiole length [PETIOLELENGTH]. j Petiole width [PETIOLEWIDTH]. k Petiole length/width ratio [PETIOLERATIO]
Frequency distribution of the 26 kale accessions for the 15 qualitative traits. a Plant shape [PLANTSHAPE]. b Position of growing point in relation to top of plant (only for those of dome shape) [PLANTPOSIT]. c Leaf division (margin) [LEAFMARGIN]. d Leaf division (incision) [LEAFINCISION]. (e) Leaf apex shape [LEAFAPEX]. f Leaf tip attitude [LEAFTIP]. g Leaf lamina attitude [LEAFLAMINA]. h Leaf blade shape [BLADESHAPE]. i Leaf blade blistering [BLADEBLISTER]. j Anthocyanin coloration of leaf [LEAFANTHO]. k Distribution of anthocyanin coloration on leaf [LEAFDISTRIB]. l Leaf colour [LEAFCOLOUR]. m Intensity of colour of fully developed leaf [LEAFINTENS]. n Petiole and/or midvein enlargement [PETIOLEMEDVEIN]. o Petiole and/or midvein colour [PETIOLECOLOUR]
Patterns of the agro-morphological differentiation among accessions
To examine patterns of the agro-morphological variation and determine the major sources of variation among kale accessions, data from both quantitative and qualitative traits as well as the morphotype and country of origin were analysed together using multifactorial analysis (MFA). The first three MFA components cumulatively explained 43.49% of the total variance. As shown in Fig. 4B, the traits that contributed most to the variation explained by the first three MFA dimensions, and therefore generally differed among the studied accessions, were the leaf margin (22.08%), leaf apex (20.85%), intensity of colour of fully developed leaf (19.72%), leaf blade blistering (16.38%) and leaf incision (15.53%). Among the quantitative traits, the petiole ratio (11.14%) and the leaf area (10.99%) and the petiole width (10.04%) were the main contributors to differentiation. Overall, the contribution of qualitative traits to the agro-morphological differentiation of the studied accessions slightly exceeded that of quantitative traits (11.28% and 10.88%, respectively). However, morphotype seems to be the main driver of agro-morphological diversification of kale accessions, contributing with 32.61% (Fig. 4B). This pattern was evidenced by the 2D plot of the first two MFA dimensions, which showed that the two Slovenian commercial cultivars of Scotch Kale (KIS19_R1) and Marrow stem Kale (KIS19_R3) and the Italian commercial cultivar of B. oleracea L. (KIS19_R2) formed three outgroups clearly separated from the other kale accessions (Fig. 5A). When only the accessions of B. oleracea L. var. acephala DC. were examined, the 2D MFA plot did not show a clear separation between the Croatian and Bosnian kales (Fig. 5B), suggesting that some variation may occur between geographically close sites within the same country.
A Variation decomposition within and among accessions for the 11 quantitative traits and B the contribution of morphotype, the 15 qualitative (filled bar plots) and the 11 quantitative (non-filled bar plots) agro-morphological traits to the total variance explained by the first three dimensions of the multifactorial analysis
Two-dimensional multifactorial analysis plots showing the relationship patterns between all 26 kale accessions (A) and only the B. oleracea var. acephala accession based on the data of the 26 agro-morphological traits(B). C The optimal number of clusters and D the UPGMA dendrogram depicting the relationship between the 26 kale accessions based on the Gower distance matrix between pairwise accessions
To further investigate the agro-morphological differentiation among kale accessions, the degree of dissimilarity was estimated using Gower distance (GD) based on phenotypic data of all traits. The results showed that KIS19_R1 and KIS19_R3 were the most divergent accessions (GD = 0.87), while the two Bosnian accessions KIS19_R16 and KIS19_R26 (GD = 0.22) were the most phenotypically similar, with an average GD of 0.45 between all kale accessions. Cluster analysis based on Gower distance divided the 26 accessions into 8 groups (Fig. 5C). Consistent with the MFA results, accessions KIS19_R1, KIS19_R2, and KIS19_R3 diverged from the other accessions in three different groups (C6, C1, and C2). Cluster C2 included only Bosnian accessions, while the four remaining clusters were a mixture of Croatian and Bosnian accessions (Fig. 5D).
Microsatellite polymorphism and genetic diversity level
The twelve SSR markers used to characterise the genetic diversity of the 26 kale accessions yielded a total of 108 alleles, with the observed number of alleles (Na) ranging from three (Na14-C12) to 14 (Ol10-A03a and Ol13-C12), with an average of nine alleles per locus. The effective number of alleles (Ne) ranged from 2.01 to 3.16 in Na14-C12 and Ol10-A03a, respectively, with a mean of 2.62. The 12 SSR markers had a high polymorphism rate, with a polymorphism information content (PIC) of 0.4 (Na14-C12) to 0.86 (Ol12-F02) and a mean PIC of 0.68. Major allele frequencies were lowest in Ol12-F02 (MAF = 0.27) and highest in Na12-C08 (MAF = 0.54), with a mean frequency of 0.39. In addition, observed heterozygosity (Ho) varied from 0.76 (Ol13-C12) to 1 (Ra2-E11, Ol12-F11, and Ra2-E03) with a mean of 0.95. Importantly, high gene diversity expressed by expected heterozygosity (He) was observed in all SSR markers, with values ranging from 0.52 in Na14-C12 to 0.87 in Ol12-F02 and He of 0.73 in all markers. This pattern is reflected in the negative Fis values obtained for all markers, indicating a general excess of heterozygosity (Table 2).
Genetic diversity estimates obtained at the accession level are shown in Table 3. The lowest number of observed (Na) and effective (Ne) alleles was observed in accessions KIS19_R21 and KIS19_R27 (Na = 1.750; Ne = 1,750), whereas the highest allelic richness was observed in accessions KIS19_R1 (Na = 3.667; Ne = 3.264), KIS19_R5 (Na = 3.5; Ne = 3.160), and KIS19_R11 (Na = 3.667; Ne = 3.124). Across all analysed accessions, the mean values for Na and Ne were 2.881 and 2.616, respectively. Moreover, the Shannon diversity index (I) and expected heterozygosity (He), with mean values of 0.953 and 0.578 across accessions, respectively, indicated that accessions KIS19_R11 were the most genetically diverse (I = 1.887 and He = 0.662), whereas accessions KIS19_R21 and KIS19_R27 were the least diverse (I = 0.587 and He = 0.417). Remarkably, the observed heterozygosity was higher than the expected heterozygosity in all accessions, with an average value of 0.939, resulting in negative inbreeding coefficients (Fis) that averaged -0.671, confirming the high level of heterozygosity in the entire collection.
Genetic differentiation and population structure
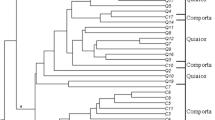
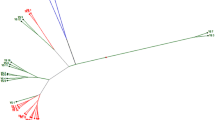
Analysis of molecular variance (AMOVA) revealed that 94% of the genetic variation occurred within accessions. Genetic differentiation among accessions (Fst) ranged from 0.01 to 0.294, with an average of 0.06 and a level of gene flow of 3.55 migrants per generation. However, no genetic differentiation was observed between countries of origin (Fst = 0). Bayesian clustering using STRUCTURE revealed that ΔK reached its maximum (ΔK = 3.84) at K = 4, indicating that the accessions could be divided into four main clusters (Fig. 6A) without a geographic clustering pattern. With the exception of the Italian commercial cultivar (KIS19_R2), the Croatian accession KIS19_R9, and the three Bosnian accessions KIS19_R20, KIS19_R25, and KIS19_R27, which were assigned to cluster 3 with a membership coefficient Q > 75%, the remaining accessions exhibited mixed clustering with partial membership in multiple clusters (Fig. 6B). Remarkably, very little divergence in allele frequency (DAF) was detected among the four clusters (< 5%), with completely identical genetic composition in Cluster 2 and Cluster 4 (DAF = 0), indicating little genetic differentiation among kale accessions. Similar to the STRUCTURE based clustering, UPGMA clustering based on Nei’s genetic distance did not reveal any geographic grouping patterns. As shown in Fig. 7, the 26 kale accessions could be divided into two main groups with 100% bootstrap support. The first group included six Bosnian accessions, while the 20 remaining accessions were mixed in a second major group that was divided into several subgroups differentiated with low bootstraps support. Within the second group, the three commercial cultivars were separated from the remaining accessions in a single group.
Association between agro-morphological and molecular variation
The Mantel test revealed a very low and non-significant association between the Nei’s genetic distance matrix generated from the molecular data and the Gower distance matrix among accessions, with a cophenetic coefficient (r) of 0.06 (P = 0.213). When only the qualitative trait data were used, r was 0.06 (P = 0.249) and dropped to 0.03 (P = 0.520) when only the quantitative traits were used. This result suggests that molecular differentiation is not correlated with agro-morphological differentiation among kale accessions, as clearly evidenced by the different patterns of clustering in our study.
Discussion
Detailed genetic characterization of a particular germplasm is instrumental for establishing appropriate management, preservation and breeding practices. Within the Brassicaceae family, B. oleracea var. acephala L. (leafy kale) has attracted great interest worldwide as a 'superfood' and ornamental plant (Šamec et al. 2018). Therefore, several investigations have already been conducted to characterize the genetic diversity of local accessions in different countries based on agro-morphological or nutritional traits, e.g. in Portugal (Dias et al. 1994), Spain (Cartea et al. 2003; Padilla et al. 2007), Turkey (Balkaya and Yanmaz 2005), Croatia (Šamec et al. 2019a) and Italy (Lotti et al. 2018). Some other investigations have been conducted using molecular markers such as random amplified polymorphic DNA (RAPD) (Farnham 1996; Okumus and Balkaya 2007), amplified fragment length polymorphism (AFLP) (Christensen et al. 2011), single nucleotide polymorphism (SNP) (Hahn et al. 2022) and SSR markers (Šutković et al. 2021). In addition to these valuable reports, detailed characterization of kale genetic resources will be very useful to breed superior accessions, promote their valorisation and management to halt genetic erosion and economic losses due to climate change.
In the current study, a large number of qualitative and quantitative agro-morphological traits and SSR markers was combined to characterise the genetic variation of 23 Croatian and Bosnian leafy kale accessions for inclusion in future breeding programmes. Similar to previous research on Spanish (Cartea et al. 2003) and Indian (Gorka et al. 2018; Singh et al. 2017) accessions, our study showed considerable agro-morphological variation in quantitative traits among the studied kale accessions. Despite the similarity in leaf morphology in terms of length and width, the Indian kale accessions had shorter plant height (average 47.71 cm versus 61.34 cm) compared to the accessions analysed in our study (Singh et al. 2017). In contrast, the Spanish accessions were characterised by outstanding plant vigour with plant height exceeding 160 cm. Interestingly, most of the quantitative traits analysed in our study showed strong positive correlation among themselves, indicating polygenic control through the pleiotropic effect of genes on different traits or linkage disequilibrium among different loci (Chebib and Guillaume 2021). Similar results were also observed in B. oleracea L. var. botrytis (Kumar et al. 2019), B. rapa var. rapa L. (Khadivi et al. 2022) and B. carinata A. Braun (Yimer et al. 2021) accessions. Moreover, our results revealed that most of the phenotypic variation in quantitative traits resided among accessions. Overall, the strong correlations among traits and the large phenotypic diversity among accessions provide a great opportunity to use both traits and accessions analysed in our study as a source of genetic diversity in future breeding programmes. In addition, a large phenotypic diversity was found in qualitative traits (H' = 0.49 – 1.54). Similar to the Indian accessions analysed by (Gorka et al. 2018), most of our accessions were either an inverted pyramid or a dome shaped with green leaves. In contrast to the low variation in leaf blade shape among Indian accessions, which were mainly characterised by elliptic blade shape, this trait was most variable among the accessions analysed in our study (H' = 1.54).
Previous research has demonstrated that leaf shape is strongly influenced by environmental factors in addition to its genetic control (Dkhar and Pareek 2014) and that its variation may be the result of strong adaptation to environmental stresses (Kidner and Umbreen 2010). Šamec et al. (2019a) reported that kale exhibits broad tolerance to different temperatures. On the other hand, variation in leaf shape in kale is thought to be the result of selection by the breeder (Arias et al. 2021). Thus, the variation in leaf shape observed in our study could be due to the adaptation of the studied accessions to their local environment, since they originated from different locations. Cartea et al. (2003) also considered that the agro-morphological variation among Spanish kale accessions could be due to both adaptation to local ecological conditions and selection by farmers. In addition, the MFA analysis revealed that qualitative rather than quantitative traits contributed most to agro-morphological diversification among the studied kale accessions. The MFA analysis and UPGMA clustering based on the Gower distance also revealed that the three commercial cultivars were classified as independent outliers that deviated from the other kale accessions, with geographic origin being an important contributor to diversification. A similar pattern of agro-morphological differentiation was observed in cabbage and Galega kale landraces from Portugal (Dias et al. 1994) and Spanish kale populations (Cartea et al. 2003). The considerable effect of geographic origin on differentiation and the evidence of adaptation to the local environment suggest that the influence of genotype by environment on several agro-morphological traits is important. Moreover, the divergence among the three commercial cultivars with each other and with the kale accessions was expected, as they belong to different morphotypes and indicate that they were selected for different agro-morphological traits. Interestingly, in agreement with our results, previous research has shown that qualitative traits such as leaf colour and division, leaf blade blistering, anthocyanin coloration, and anthocyanin coloration distribution were the main drivers of variation in Croatian accessions of B. oleracea var. acephala with different seed source (Batelja et al. 2009; Šamec et al. 2019a).
In addition to agro-morphological characterization, molecular characterization, particularly via codominant markers such as microsatellites (SSRs), is required to provide a robust appraisal of the genetic diversity of kale accessions. The joint analysis maximizes the comprehensive assessment of the diversity of available genetic material, facilitates its organization, and enables the identification of desired traits and distinctive alleles, allowing the accurate selection of promising genetic resources for breeding programs (Li et al. 2020). In our study, the twelve SSR markers used for genetic characterization of kale accessions, previously shown to be highly informative (El-Esawi et al. 2016), were found to be highly polymorphic with a mean PIC value (0.68) and an average of nine alleles. These values were higher than those reported for the same SSR set in Irish accessions (El-Esawi et al. 2016) and for other SSR markers in different commercial cultivars of B. oleracea L. (Izzah et al. 2013; Tonguç and Griffiths 2004). This result indicates the high reliability of these SSR markers for the analysis of the genetic structure of kale accessions.
Our results revealed that the accessions analysed in this study had high genetic diversity (He = 0.578), which exceeded the level of Irish kale (He = 0.459), cabbage (He = 0.548), cauliflower (He = 0.446) and sprouts (He = 0.536) using SSR markers (El-Esawi et al. 2016). Moreover, similar to the above study, a significant excess of heterozygosity was observed in our study as reflected by the high observed heterozygosity and negative inbreeding coefficient (Fis). This was to be expected considering the small number of individuals within the studied accessions (2 to 3 individuals) and the outcrossing breeding system of B. oleracea L.
On the other hand, in contrast to the results we obtained for quantitative traits, the SSR data revealed that most of the genetic variation occurred at the level within accessions (95%), indicating low genetic differentiation and weak genetic structure in the studied accessions (Fst = 0.05). Although genetic differentiation was greater, a similar pattern was observed in Irish kale (Fst = 11%), cauliflower (Fst = 11%), and Brussels sprouts (Fst = 9%) (El-Esawi et al. 2016), as well as in closely related cultivars such as B. oleracea L. var. italica (Fst = 13%) (Shen et al. 2021) and B. rapa subsp. rapa L (Fst = 10%) (Soengas et al. 2011). The low genetic differentiation observed in our study was supported by the relatively high gene flow observed between accessions (Nm = 3.55), which was higher than the level observed for B. rapa subsp. rapa (Nm = 2.25) (Soengas et al. 2011) and B. oleracea L. var. italica (Nm = 1.74) (Shen et al. 2021). Such excellent gene flow can occur naturally or via seed exchange between farmers given the open pollination in B. oleracea L. In agreement with the above research, as well as with the study of Shen et al. (2021) on B. oleracea L. var. italica, the weak genetic structure of the studied accessions was confirmed by UPGMA clustering and Bayesian analysis, which did not reveal a clear relationship between clustering of accessions and geographical origin. A similar pattern was also observed in wild populations of B. oleracea L. (Christensen et al. 2011; Lanner-Herrera et al. 1996; Mittell et al. 2020) and B. oleracea L. var. italica (Shen et al. 2021). In contrast, in a large collection of Spanish B. oleracea L. wild accessions, restricted gene flow (Nm = 0.26) and large genetic differentiation (Fst = 55.1%) were observed, mainly promoted by geographic isolation (Tortosa et al. 2017). However, in our study, UPGMA clustering showed that the three commercial standard cultivars, although belonging to three different subspecies, were clearly differentiated from kale accessions in a single group, suggesting that their genomes harbour common genomic regions and could be bred for similar agronomic traits. Moreover, our results revealed that the grouping of accessions based on the agro-morphological data did not match the grouping determined from the SSR data. This was confirmed by the weak and non-significant association between the dissimilarity matrices between the accessions determined by the Mantel test for both pairs of data. The weak association between the phenotypic and genotypic matrices was also reported in broccoli (Li et al. 2019) and cabbage (Kang et al. 2011) and several other crops (Hartings et al. 2008; Roldan-Ruiz et al. 2000; Soriano et al. 2016). Previous research indicated that such incongruence highlights the complementarity and non-overlap between the agro-morphological and genotypic data to investigate the extent and nature of genetic variability in crops (Da Silva et al. 2017; Darkwa et al. 2020; Geleta et al. 2006). Importantly, this pattern indicates that the polymorphism detected by SSR markers does not fully contribute to the agro-morphological variability among kale accessions, suggesting that only a few genes are involved in the desired agro-morphological traits. Another plausible explanation could be the massive influence of genotype-environment interaction on the quantitatively inherited complex agro-morphological traits controlled by several genomic regions, including SSR loci which were not used in the present study.
Conclusion
In conclusion, the present study presented a detailed assessment of the genetic diversity and structure of kale accessions using a large set of agro-morphological traits and SSR markers. The results revealed that the kale accessions exhibit wide variation in several economically important agro-morphological traits. The agro-morphological characterization revealed that qualitative traits contributed more than quantitative traits to phenotypic differentiation among accessions. The diversity among accessions in some particular traits such as leaf shape, leaf colour and distribution of anthocyanin coloration on the leaf is useful to select a subset of accessions for breeding programs. The SSR markers used in our study, which proved to be very useful for molecular charcterization, showed great genetic diversity at the molecular level. As expected, the SSR data showed a large intra-accession variation and a weak genetic structure non-associated with the geographic origin of the kale accessions. Importantly, the weak and nonsignificant association between dissmilarity matrices among accessions generated by agro-morphological and SSR data indicates the complementarity of the multidisciplianry approaches used in our study. Overall, the results of our study pointed to the importance of the kale collection studied as a valuable reservoir of agro-morphological and genetic variability that could be exploited in future breeding programs.
Data availability
The authors confirm that all the experimental data are available and accessible via the main text and/or the supplementary information. Raw data will be made available on request.
References
Arias T, Niederhuth CE, McSteen P, Pires JC (2021) The molecular basis of kale domestication: transcriptional profiling of developing leaves provides new insights into the evolution of a Brassica oleracea vegetative morphotype. Front Plant Sci 12:637115. https://doi.org/10.3389/fpls.2021.637115
Balkaya A, Yanmaz R (2005) Promising kale (Brassica oleracea var. acephala) populations from Black Sea region, Turkey. New Zeal J Crop Hort Sci 33:1–7. https://doi.org/10.1080/01140671.2005.9514324
Batelja K, Goreta Ban S, Žanić K, Miloš B, Dumičić G, Matotan Z (2009) Autochtonous kale populations (Brassica oleracea L. var. acephala) in Croatian coastal region. Poljoprivreda 15:8–14
Cartea ME, Picoaga A, Soengas P, Ordás A (2003) Morphological characterization of kale populations from northwestern Spain. Euphytica 129:25–32. https://doi.org/10.1023/A:1021576005211
Chebib J, Guillaume F (2021) Pleiotropy or linkage? Their relative contributions to the genetic correlation of quantitative traits and detection by multitrait GWA studies. Genetics 219:iyab159. https://doi.org/10.1093/genetics/iyab159
Choudhury A, Deb S, Kharbyngar B, Rajpal VR, Rao SR (2022) Dissecting the plant genome: through new generation molecular markers. Genet Resour Crop Evol 69:2661–2698. https://doi.org/10.1007/s10722-022-01441-3
Christensen S, von Bothmer R, Poulsen G, Maggioni L, Phillip M, Andersen BA, Jørgensen RB (2011) AFLP analysis of genetic diversity in leafy kale (Brassica oleracea L. convar. acephala (DC.) Alef.) landraces, cultivars and wild populations in Europe. Genet Resour Crop Evol 58:657–666. https://doi.org/10.1007/s10722-010-9607-z
Comtois D, Comtois MD (2016) Package ‘summarytools.’
CPVO (2011) Protocol for Distinctness, Uniformity and Stability tests: Brassica oleracea L. var. sabellica L. Curly kale: 1–15. Available online: https://cpvo.europa.eu/sites/default/files/documents/phaseolus_vulgaris .pdf
Criscuolo NG, Angelini C (2020) StructuRly: A novel shiny app to produce comprehensive, detailed and interactive plots for population genetic analysis. PLoS ONE 15:e0229330. https://doi.org/10.1371/journal.pone.0229330
Da Silva AR, Malafaia G, Menezes IPP (2017) Biotools: an R function to predict spatial gene diversity via an individual-based approach. Genet Mol Res 16:1–6. https://doi.org/10.4238/gmr16029655
Darkwa K, Agre P, Olasanmi B, Iseki K, Matsumoto R, Powell A, Bauchet G, De Koeyer D, Muranaka S, Adebola P, Asiedu R, Terauchi R, Asfaw A (2020) Comparative assessment of genetic diversity matrices and clustering methods in white Guinea yam (Dioscorea rotundata) based on morphological and molecular markers. Sci Rep 10:13191. https://doi.org/10.1038/s41598-020-69925-9
Dias JS, Monteiro AA, Kresovich S (1994) Genetic diversity and taxonomy of Portuguese Tronchuda cabbage and Galega kale landraces using isozyme analysis. Euphytica 75:221–230. https://doi.org/10.1007/BF00025607
Dkhar J, Pareek A (2014) What determines a leaf’s shape? EvoDevo 5:1–19. https://doi.org/10.1186/2041-9139-5-47
Earl DA, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
El-Esawi MA, Germaine K, Bourke P, Malone R (2016) Genetic diversity and population structure of Brassica oleracea germplasm in Ireland using SSR markers. CR Biol 339:133–140. https://doi.org/10.1016/j.crvi.2016.02.002
Farnham MW (1996) Genetic variation among and within United States collard cultivars and landraces as determined by randomly amplified polymorphic DNA markers. J Am Soc Hortic Sci 121:374–379. https://doi.org/10.21273/JASHS.121.3.374
Geleta N, Labuschagne MT, Viljoen CD (2006) Genetic diversity analysis in sorghum germplasm as estimated by AFLP, SSR and morpho-agronomical markers. Biodivers Conserv 15:3251–3265. https://doi.org/10.1007/s10531-005-0313-7
Ghaffari P, Talebi R, Keshavarzi F (2014) Genetic diversity and geographical differentiation of Iranian landrace, cultivars, and exotic chickpea lines as revealed by morphological and microsatellite markers. Physiol Mol Biol Plants 20:225–233. https://doi.org/10.1007/s12298-014-0223-9
Golicz AA, Bayer PE, Barker GC, Edger PP, Kim H, Martinez PA, Chan CKK, Severn-Ellis A, McCombie WR, Parkin IA (2016) The pangenome of an agronomically important crop plant Brassica oleracea. Nat Commun 7:13390. https://doi.org/10.1038/ncomms13390
Gonçalves ÁLM, Lemos M, Niero R, de Andrade SF, Maistro EL (2012) Evaluation of the genotoxic and antigenotoxic potential of Brassica oleracea L. var. acephala D.C. in different cells of mice. J Ethnopharmacol 143:740–745. https://doi.org/10.1016/j.jep.2012.07.044
Gorka S, Samnotra RK, Kumar S, Chopra S, Gupta M (2018) Analysis of genetic diversity in kale (Brassica oleracea L. var. acephala) genotypes of jammu and kashmir region based on morphological descriptors. Int J Curr Microbiol Appl Sci 7:2176–2181. https://doi.org/10.20546/ijcmas.2018.702.261
Gower JC (1971) A general coefficient of similarity and some of its properties. Biometrics 27:857–871. https://doi.org/10.2307/2528823
Hahn C, Howard NP, Albach DC (2022) Different Shades of Kale - Approaches to analyze kale variety interrelations. Genes 13:232. https://doi.org/10.3390/genes13020232
Hartings H, Berardo N, Mazzinelli GF, Valoti P, Verderio A, Motto M (2008) Assessment of genetic diversity and relationships among maize (Zea mays L.) Italian landraces by morphological traits and AFLP profiling. Theor Appl Genet 117:831–842. https://doi.org/10.1007/s00122-008-0823-2
IBPGR (1990). Descriptors for Brassica and Raphanus. International Board for Plant Genetic Resources, Rome, Italy: 1–58. https://cropgenebank.sgrp.cgiar.org/images/file/learning_space/descriptors_brassica_raphanus .pdf
Izzah NK, Lee J, Perumal S, Park JY, Ahn K, Fu D, Kim GB, Nam YW, Yang TJ (2013) Microsatellite-based analysis of genetic diversity in 91 commercial Brassica oleracea L. cultivars belonging to six varietal groups. Genet Resour Crop Evol 60:1967–1986. https://doi.org/10.1007/s10722-013-9966-3
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281
Kang J, Fang Z, Wang X, Xu D, Liu Y, Yang L, Zhuang M, Zhang Y (2011) Genetic diversity and relationships among cabbage (Brassica oleracea var. capitata) landraces in China revealed by AFLP markers. Afr J Biotechnol 10:5940–5949
Kassambara A, Mundt F (2020a) Extrac and visualize the results of multivariate data analyses. R Package Version 1.0. 3. R package version.
Khadivi A, Mirheidari F, Moradi Y (2022) Selection of superior accessions of turnip (Brassica rapa var. rapa L.) based on tuber quality-related characters. Food Sci Nutr 10:2667–2680. https://doi.org/10.1002/fsn3.2871
Kidner CA, Umbreen S (2010) Why is leaf shape so variable. Int J Plant Dev Biol 4:64–75
Kuerban A, Yaghmoor SS, Almulaiky YQ, Mohamed YA, Razvi SSI, Hasan MN, Moselhy SS, Al-Ghafari AB, Alsufiani HM, Kumosani TA (2017) Therapeutic effects of phytochemicals of brassicaceae for management of obesity. J Pharm Res Int 19:1–11. https://doi.org/10.9734/JPRI/2017/37617
Kumar V, Thakur RK, Kumar P (2019) Assessment of heavy metals uptake by cauliflower (Brassica oleracea var botrytis) grown in integrated industrial effluent irrigated soils: A prediction modeling study. Sci Hortic 257:108682. https://doi.org/10.1016/j.scienta.2019.108682
Lanner-Herrera C, Gustafeson M, Filt AS, Bryngelsson T (1996) Diversity in natural populations of wild Brassica oleracea as estimated by isozyme and RAPD analysis. Genet Resour Crop Evol 43:13–23. https://doi.org/10.1007/BF00126936
Li Z, Mei Y, Liu Y, Fang Z, Yang L, Zhuang M, Zhang Y, Lv H (2019) The evolution of genetic diversity of broccoli cultivars in China since 1980. Sci Hortic 250:69–80. https://doi.org/10.1016/j.scienta.2019.02.034
Li D, Long C, Pang X, Ning D, Wu T, Dong M, Han X, Guo H (2020) The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions. Plant Biol 8:e8573. https://doi.org/10.7717/peerj.8573
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Lotti C, Iovieno P, Centomani I, Marcotrigiano AR, Fanelli V, Mimiola G, Summo C, Pavan S, Ricciardi L (2018) Genetic, bio-agronomic, and nutritional characterization of kale (Brassica oleracea L. var. acephala) diversity in Apulia Southern Italy. Diversity 10:25. https://doi.org/10.3390/d10020025
Maechler M, Rousseeuw P, Struyf A, Hubert M, Hornik K (2012) Cluster: cluster analysis basics and extensions.
Meglič V, Pipan B (2018) Spatial and temporal assessment of Brassica napus L. maintaining genetic diversity and gene flow potential: An empirical evaluation. In: El-Esawi MA (ed) Brassica Germplasm - Characterization, Breeding and Utilization. InTech
Mercati F, Longo C, Poma D, Araniti F, Lupini A, Mammano MM, Fiore MC, Abenavoli MR, Sunseri F (2015) Genetic variation of an Italian long shelf-life tomato (Solanum lycopersicon L.) collection by using SSR and morphological fruit traits. Genet Resour Crop Evol 62:721–732. https://doi.org/10.1007/s10722-014-0191-5
Meyer D, Zeileis A, Hornik K, Gerber F, Friendly M, Meyer MD (2021) Package ‘vcd.’ Visualizing Categorical Data. R Package Version 1–4.
Mittell EA, Cobbold CA, Ijaz UZ, Kilbride EA, Moore KA, Mable BK (2020) Feral populations of Brassica oleracea along Atlantic coasts in western Europe. Ecol Evol 10:11810–11825. https://doi.org/10.1002/ece3.6821
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Imports MASS (2019). Package ‘vegan’. Community ecology package, version, 2(9)
Okumus A, Balkaya A (2007) Estimation of genetic diversity among Turkish kale populations (Brassica oleracea var. acephala L.) using RAPD markers. Russ J Genet 43:411–415. https://doi.org/10.1134/S1022795407040096
Oumata S, Monneveux P, Zaharieva M, Mekliche-Hanifi L, David J (2023) Variation of morphological traits among wheat (Triticum aestivum L.) landraces from two regions of the Algerian Sahara. Potential interest for wheat breeding. Genet Resour Crop Evol 70:235–250. https://doi.org/10.1007/s10722-022-01429-z
Padilla G, Cartea ME, Velasco P, de Haro A, Ordás A (2007) Variation of glucosinolates in vegetable crops of Brassica rapa. Phytochemistry 68:536–545. https://doi.org/10.1016/j.phytochem.2006.11.017
Pathirana I, Thavarajah P, Siva N, Wickramasinghe AN, Smith P (2017) Moisture deficit effects on kale (Brassica oleracea L. var. acephala) biomass, mineral, and low molecular weight carbohydrate concentrations. Sci Hortic 226:216–222. https://doi.org/10.1016/j.scienta.2017.08.050
Peakall ROD, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Petit J, Salentijn EMJ, Paulo MJ, Thouminot C, van Dinter BJ, Magagnini G, Gusovius HJ, Tang K, Amaducci S, Wang S, Uhrlaub B, Müssig J, Trindade LM (2020b) Genetic variability of morphological, flowering, and biomass quality traits in hemp (Cannabis sativa L.). Front Plant Sci 11:102. https://doi.org/10.3389/fpls.2020.00102
Pipan B, Šuštar-Vozlič J, Meglič V (2011) Cultivation, varietal structure and possibilities for cross-pollination of Brassica napus L. in Slovenia. Acta Agric Slov 97:247. https://doi.org/10.2478/v10014-011-0019-3
Pipan B, Šuštar-Vozlič J, Meglič V (2013) Genetic differentiation among sexually compatible relatives of Brassica napus L. Genetika 45:309–327. https://doi.org/10.2298/GENSR1302309P
Pipan B, Žnidarčič D, Meglič V (2017) Evaluation of genetic diversity of sweet potato [Ipomoea batatas (L.) Lam.] on different ploidy levels applying two capillary platforms. J Hortic Sci Biotechnol 92:192–198. https://doi.org/10.1080/14620316.2016.1249963
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
R Core Team (2021). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/.
Rakshita KN, Singh S, Verma VK, Sharma BB, Saini N, Iquebal MA, Sharma A, Dey SS, Behera TK (2021) Agro-morphological and molecular diversity in different maturity groups of Indian cauliflower (Brassica oleracea var. botrytis L.). PLoS ONE 16:e0260246. https://doi.org/10.1371/journal.pone.0260246
Riangwong K, Wanchana S, Aesomnuk W, Saensuk C, Nubankoh P, Ruanjaichon V, Kraithong T, Toojinda T, Vanavichit A, Arikit S (2020) Mining and validation of novel genotyping-by-sequencing (GBS)-based simple sequence repeats (SSRs) and their application for the estimation of the genetic diversity and population structure of coconuts (Cocos nucifera L.) in Thailand. Hortic Res 7:156. https://doi.org/10.1038/s41438-020-00374-1
Rivera A, Monteagudo AB, Igartua E, Taboada A, García-Ulloa A, Pomar F, Riveiro-Leira M, Silvar C (2016) Assessing genetic and phenotypic diversity in pepper (Capsicum annuum L.) landraces from North-West Spain. Sci Hortic 203:1–11. https://doi.org/10.1016/j.scienta.2016.03.006
Roldan-Ruiz I, Dendauw J, Van Bockstaele E, Depicker A, De Loose M (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Mol Breed 6:125–134. https://doi.org/10.1023/A:1009680614564
Šamec D, Pavlović I, Radojčić Redovniković I, Salopek-Sondi B (2018) Comparative analysis of phytochemicals and activity of endogenous enzymes associated with their stability, bioavailability and food quality in five Brassicaceae sprouts. Food Chem 269:96–102. https://doi.org/10.1016/j.foodchem.2018.06.133
Šamec D, Kruk V, Ivanišević P (2019a) Influence of seed origin on morphological characteristics and phytochemicals levels in Brassica oleracea var. acephala. Agronomy 9:502. https://doi.org/10.3390/agronomy9090502
Šamec D, Urlić B, Salopek-Sondi B (2019b) Kale (Brassica oleracea var. acephala) as a superfood: Review of the scientific evidence behind the statement. Crit Rev Food Sci Nutr 59:2411–2422. https://doi.org/10.1080/10408398.2018.1454400
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234. https://doi.org/10.1038/72708
Sebastien L, Julie J, Francois H (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25(1):1–18. https://doi.org/10.18637/jss.v025.i01
Shannon CE, Weaver W (1949) The mathematical theory of communication. Bell Sys Tech J 27:379–423. https://doi.org/10.1002/j.1538-7305.1948.tb01338.x
Sharma D, Nanjundan J, Singh L, Singh SP, Parmar N, Sujith Kumar MS, Singh KH, Mishra AK, Singh R, Verma KS (2020) Genetic diversity in leafy mustard (Brassica juncea var. rugosa) as revealed by agro-morphological traits and SSR markers. Physiol Mol Biol Plants 26:2005–2018. https://doi.org/10.1007/s12298-020-00883-2
Shen Y, Wang J, Shaw RK, Yu H, Sheng X, Zhao Z, Li S, Gu H (2021) Development of GBTS and KASP panels for genetic diversity, population structure, and fingerprinting of a large collection of broccoli (Brassica oleracea L. var. italica) in China. Front Plant Sci 12:655254. https://doi.org/10.3389/fpls.2021.655254
Singh SR, Ahamed N, Kumar D, Srivatsava KK, Yousuf S, Mir A (2017) Genetic divergence assessment in kale (Brassica oleracea L. var. acephala (DC.) Alef) by using the multivariate analysis. J Hortic Sci 12:42–48
Soengas P, Cartea ME, Francisco M, Lema M, Velasco P (2011) Genetic structure and diversity of a collection of Brassica rapa subsp. rapa L. revealed by simple sequence repeat markers. J Agric Sci 149:617–624. https://doi.org/10.1017/S002185961100013X
Soriano JM, Villegas D, Aranzana MJ, García del Moral LF, Royo C (2016) Genetic structure of modern durum wheat cultivars and Mediterranean landraces matches with their agronomic performance. PLoS ONE 11:e0160983. https://doi.org/10.1371/journal.pone.0160983
Šutković J, Glamočija P, Karić L, Yildirim A (2021) Genetic characterization of Brassica oleracea L. var. acephala DC cultivars from Bosnia and Herzegovina. Genet Appl. 5:40–50. https://doi.org/10.31383/ga.vol5iss1pp40-50
Takezaki N, Nei M, Tamura K (2010) POPTREE2: Software for constructing population trees from allele frequency data and computing other population statistics with Windows interface. Mol Biol Evol 27:747–752. https://doi.org/10.1093/molbev/msp312
Terlević A, Temunović M, Bogdanović S, Grgurev M, Ljubičić I, Rešetnik I (2023) Morphological and environmental variability of Dianthus sylvestris (Caryophyllaceae) in the Balkan Peninsula. Bot J Linn Soc 201:377–389. https://doi.org/10.1093/botlinnean/boac058
Thavarajah P, Abare A, Basnagala S, Lacher C, Smith P, Combs GF Jr (2016) Mineral micronutrient and prebiotic carbohydrate profiles of USA-grown kale (Brassica oleracea L. var. acephala). J Food Compos Anal 52:9–15. https://doi.org/10.1016/j.jfca.2016.07.003
Tonguç M, Griffiths PD (2004) Genetic relationships of Brassica vegetables determined using database derived simple sequence repeats. Euphytica 137:193–201. https://doi.org/10.1023/B:EUPH.0000041577.84388.43
Tortosa M, Velasco P, Afonso D, Padilla G, Ríos D, Soengas P (2017) Characterization of a Spanish Brassica oleracea collection by using molecular and biochemical markers. Sci Hortic 219:344–350. https://doi.org/10.1016/j.scienta.2017.03.021
Villanueva RAM, Chen ZJ (2019) ggplot2: elegant graphics for data analysis
Wei T, Simko V, Levy M, Xie Y, Jin Y, Zemla J (2017) Package ‘corrplot.’ Statistician 56:e24
Yimer O, Mohammed W, Teju E (2021) Correlation and path coefficient analysis studies in Ethiopian Mustard (Brassica carinata A. Braun). Afr J Plant Sci 15:250–256. https://doi.org/10.5897/AJPS2021.2156
Zhu S, Niu E, Shi A, Mou B (2019) Genetic diversity analysis of olive germplasm (Olea europaea L.) with genotyping-by-sequencing technology. Front Genet 10:755. https://doi.org/10.3389/fgene.2019.00755
Acknowledgements
This study was part of the research programme “Agrobiodiversity” (P4-0072) financially supported by the Slovenian Research Agency, Ljubljana, Slovenia; part of the EUBRASWILD project funded by the international ECPGR project (Third Call – Phase X 2019-2023); and part of the bilateral projects funded by ARRS: Slovenia-Serbia (BI-RS/20-21-015) and Slovenia-Bosnia and Herzegovina (BI-BA/19-20-009 and BI-BA/21-23-011). The authors thank Boštjan Ogorevc, Mojca Polak, Neža Kermavnar, Ana Špilak, and Teja Krpan for technical assistance, evaluations, and analyses. The authors thank the Genetic Resources Institute, University of Banja Luka (Bosnia and Herzegovina) and the Institute for Adriatic Crops and Karst Reclamation, Split (Croatia) for providing accessions from their gene banks.
Funding
This study was part of the research programme “Agrobiodiversity” (P4-0072) financially supported by the Slovenian Research Agency, Ljubljana, Slovenia; part of the EUBRASWILD project funded by the international ECPGR project (Third Call – Phase X 2019-2023); and part of the bilateral projects funded by ARRS: Slovenia-Serbia (BI-RS/20–21-015) and Slovenia-Bosnia and Herzegovina (BI-BA/19-20-009 and BI-BA/21-23-011).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by BP, LS and MN. The first draft of the manuscript was written by LS and all authors commented on previous versions of the manuscript. VM obtained the funding. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest. All the authors read and approved the manuscript in its final form.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pipan, B., Neji, M., Meglič, V. et al. Genetic diversity of kale (Brassica oleracea L. var acephala) using agro-morphological and simple sequence repeat (SSR) markers. Genet Resour Crop Evol 71, 1221–1239 (2024). https://doi.org/10.1007/s10722-023-01686-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01686-6