Abstract
Concerns about food security worldwide have been raised due to the adverse impacts of climate change on agricultural productivity. Utilizing wild relatives of cultivated plants in breeding schemes is a potential solution to address the abovementioned challenges. Wild relatives refer to indigenous plant species that share genetic ancestry with recently cultivated plants. The capacity of plants to withstand abiotic stresses and enhance crop yield stability in cold and arid environments renders them a valuable prospective resource for breeding programs. Molecular markers are employed to assess the genetic variability of cultivated plants and their wild relatives under investigation. This study employed a set of five primers derived from ISSR (Inter Simple Sequence Repeats) and SCoT (Start Codon Targeted) to assess comparative research between cultivated species and their wild relatives from three families: Apiaceae (Apium gravolens L., A. leptophyllum (Pers.) F.Muell., Daucus carota L.& D. litoralis Sm.), Asteraceae (Lactuca sativa L. & L. serriola L.) and Brassicaceae (B. oleracea L., Brassica tourneforttii Gouan., Raphanus sativus L. & R. raphanistrum L.). The ISSR technique yields approximately 206 bands, whereas the SCoT primers generate approximately 207 bands. The ISSR primers yielded 169 polymorphic bands, exhibiting a polymorphism rate of 82.04%, whereas the SCoT primers produced 169 polymorphic bands with a polymorphism rate of 81.64%. The Brassicaceae has been found to exhibit the highest resolving power (RP) of 48.25, as determined by ISSR. The primary objective of DNA barcoding is to create a communal database of DNA sequences that can be utilized to identify organisms and clarify their taxonomic classification. The findings indicate that the internal transcribed spacer (ITS) regions exhibit high efficacy as a DNA barcoding tool. Overall, the results obtained from two molecular markers and DNA barcoding analyses suggest a significant degree of genetic diversity and relatedness among the cultivated taxa and their wild relatives examined in this study.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The forthcoming century presents a significant obstacle for humankind, namely the production of sustainable, just, and hygienic food amidst worldwide environmental transformations such as climate change, escalating environmental contamination, and the deterioration of soil quality. Furthermore, climatic condition alterations have adversely impacted worldwide agricultural output (Singh and Abhilash 2018; Mponya et al. 2021). Therefore, utilizing versatile and climate-resilient cultivars in cultivation is imperative to fulfill the growing population's increasing food requirements. The utilization of wild relatives of crop plants for arid and semi-arid climatic conditions is highly valuable due to their superior tolerance, adaptability, high nutritional value, wide range of ecosystem plasticity, and ability to cultivate with fewer external inputs such as pesticides and fertilizers and crop management than modern cultivars (Singh et al. 2018).
Moreover, they have the potential to enhance soil quality and promote soil carbon sequestration, all while preserving invaluable natural resources. Utilizing a significant gene pool of untamed relatives of domesticated species is imperative in developing novel cultivars and modifying existing agricultural plants to enhance their adaptability, tolerance, yield, and nutritional content in response to anticipated climatic conditions. According to Whitney et al. (2018), Crop wild relatives (CWRs) are wild relatives of crops that are closely related and possess a range of beneficial agronomic, nutritional, and breeding efficiency characteristics that can be utilized for crop improvement purposes (Mezghani et al. 2019; Perrino and Wagensommer 2021). Crop Wild Relatives (CWR) refer to plant species that are naturally occurring and possess remarkable adaptability to harsh climatic conditions, nutrient-deficient soil, and strong resistance to pests and diseases.
Additionally, research by Khoury et al. (2020) and Rocha et al. (2021) show that they confer advantageous characteristics for crop adaptation to current and projected agricultural challenges. Plant genetic resources for food and agriculture (PGRFA) are deemed significant in crop improvement, with their constituent elements regarded as highly valuable (Maxted and Vincent 2021). Plant Genetic Resources for Food and Agriculture (PGRFA) are the plant genetic resources primarily associated with human food production in agriculture. According to Ford-Lloyd et al. (2011) and Engels and Thormann (2020), plant genetic resources encompass the genetic variation present in domesticated plant species and their wild counterparts, which holds potential value for plant breeders.
The preservation of genetic diversity is widely recognized as a fundamental prerequisite for the survival of plant species in their natural habitats and for advancing crop breeding programs. Plant breeders can develop novel and enhanced cultivars with desirable traits, including those that are highly esteemed by farmers and breeders ( high yield potential, large seed), owing to the wide-ranging genetic resources available in plants, such as pest and disease resistance, photosensitivity, and other attributes. Since the inception of agriculture, crop species have been utilized to fulfill subsistence food needs by harnessing natural genetic variability (Bhandari et al. 2017; Kumar et al. 2023). In contemporary times, molecular markers have gained popularity due to their imperviousness to environmental influences (Hasan et al. 2021 and EL-Mansy et al. 2021).
Molecular markers have proven to be highly effective tools for investigating the genetic diversity of various plant species, particularly those that pose challenges to traditional classification methods (Payacan et al. 2017; Abd El-Moneim et al. 2021). Molecular markers are valuable in examining evolutionary relationships among various plant groups. According to Nadeem et al. (2018), molecular markers refer to genetic loci that can be conveniently monitored and measured within a population and may correlate with a particular gene or trait of significance. A genetic marker can manifest as a brief DNA sequence, such as a single nucleotide polymorphism (SNP) resulting from a single base-pair shift, or as a more extensive sequence, such as mini- and microsatellites. Molecular markers can categorize into three distinct groups: hybridization-based markers, which include restriction fragment length polymorphism (RFLP); PCR-based markers, which encompass random amplified polymorphic DNA (RAPD), amplified fragment length polymorphism (AFLP); and microsatellites or simple sequence repeats (SSR); and sequence-based markers, exemplified by single nucleotide polymorphism (SNP).
According to Yang et al. (2020), these clusters of markers have the potential to identify dominance and co-dominance within a genome. The Inter Simple Sequence Repeat (ISSR) method is a widely utilized, cost-effective, stable, and reproducible technique for evaluating genetic diversity, cultivar identification, genetic mapping, evolution, and molecular ecology (Henareh et al. 2016). The Inter Simple Sequence Repeat (ISSR) technique has been used since 1994 (Zietkiewicz et al. 1994). ISSRs are as semi-arbitrary markers that undergo PCR amplification in the presence of a single primer complementary to a microsatellite target (Abd El-Moneim 2020). The methodology is utilized to enhance the inter-simple sequence repeat (SSR) regions of varying lengths primarily through utilizing microsatellites with a length of 16–25 base pairs as primers. The utilization of ISSRs in the botanical realm has been extended due to their provision of significant insights into inter-species connections (Arslan et al. 2020); Mesfer ALshamrani et al. 2022; Jabari et al. 2023).
Moreover, it is noteworthy that inter simple sequence repeats (ISSRs) typically comprise a substantial quantity of polymorphic bands, with the potential to detect a significant degree of genomic polymorphisms, reaching up to 97 (Monpara et al. 2017).
The Start Codon Targeted (SCoT) polymorphism is a gene-targeted marker system that utilizes polymerase chain reaction (PCR)-based DNA markers. It is widely utilized due to its simplicity and reliability. Satya et al. (2015) reported that this technique depends on identifying conserved regions within the genetic structure of plants that includes the ATG translation start codon. According to Etminan et al. (2016), this marker is a proficient methodology for conducting population research evaluating genetic diversity and structure, distinguishing cultivars, and performing DNA fingerprinting.
This method has advantages, including ease of use, affordability, high polymorphism level, and more information about genetic diversity (Chai et al. 2017). DNA barcoding is a valuable molecular identification tool for plants, which guarantees the safety and effectiveness of plant materials (Jamdade et al. 2022) in plant identification and the examination of their ecological and evolutionary interconnections while remaining conserved within a given species (Rayan 2019). Sequence-based specimen identification was proposed initially for land plants by Hollingsworth et al. (2009) and later extended to the animal kingdom by Taylor and Harris (2012). Hebert et al. (2003) standardized short DNA sequences to identify a broad sense of taxa. This method serves as a supplementary tool for morphological identification.
The technique in question relies on utilizing a limited number of loci, either singularly or in combination, as a cohesive unit, each characterized by a brief and distinctive DNA sequence (Mansour et al. 2020; Safhi et al. 2022). The DNA barcodes involve the use of particular chloroplast genomes, plastid DNA regions, or a combination of them, such as matK, trnH-psbA, rbcL, atpF-atphH, rpoB, psbK-psbIr, and rpoC1 (Hollingsworth et al. 2009). Nuclear ribosome DNA sequences, including the internal transcribed spacer (ITS), have been employed as DNA barcodes (Senapati et al. 2022).
The proliferation of DNA barcodes in public repositories can be attributed to many factors, including advancements in sequencing technology, the emergence of novel database management and computational software, and the proliferation of national and international consortia that facilitate DNA barcode sequencing. DeSalle and Goldstein (2019) emphasized the prevalence of DNA barcode implementations in phylogenetics and taxonomy. The primary objective of the current study was to i) inspect the genetic variation among cultivated plant species and their wild relatives from three families: Apiaceae (Apium gravolens L., A leptophyllum (Pers.) F.Muell., Daucus carota L.& D. litoralis Sm.), Asteraceae (Lactuca sativa L. & L.serriola L.) and Brassicaceae (Brassica oleracea L., B. tourneforttii Gouan., Raphanus sativus L. & R. raphanistrum L.) in Egypt using two molecular markers (ISSR and SCoT) ii) study the comparative analysis among the studied cultivated species and their wild relatives using DNA barcoding techniques and iii) assess the molecular relationships among the studied taxa using clustering heatmap, principle component analysis (PCA) based on molecular markers and DNA barcoding attributes.
Materials and methods
Plant materials
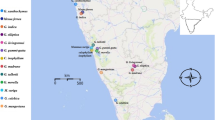
Viable seeds of cultivated species (A. gravolens, D. carota, L. sativa, B. oleracea and R. sativus) obtained from the National Gene Bank, Ministry of Agriculture and Land Reclamation, Egypt, and cultivated proliferated at the experimental field of the Faculty of Science, Mansoura University, Egypt, during two successive seasons, 2020/2021 and 2021/2022. The wild relatives were collected from their natural habitats in Egypt by the authority of the Faculty of Science, Mansoura University, Egypt. Wild taxa identified by Professor Ibrahim A. Mashaly, Professor of Plant Ecology and Flora, Botany Department, Faculty of Science, Mansoura University. Identification and nomenclature were according to Tackholm (1974) and Boulos (2009) as in Fig. 1. Table 1 illustrates the studied taxa's families, names, types, collection areas, and governorates.
Molecular analysis
The genomic DNA of each taxon under investigation was extracted from fresh leaves using the CTAB buffer methodology, as described by Cota-Sanchez et al. (2006) at the Plant Laboratory in Botany and Microbiology Department, Faculty of Science, Arish University, Egypt. The concentration of the extracted DNA was determined using nanodrop. Five primer pairs were used for ISSR and SCoT markers, as presented in Table 2. The process of DNA amplification was conducted using a 20 μL PCR reaction mixture comprising 10 μL of master (2X TOPsimpleTM DyeMIX-nTaq), 5 µL of each primer (0.1 µM), and 1 μL of genomic DNA (50 ng/μL); 4 μL from sterile distilled water was utilized to attain the final volume of 20 μL (Zietkiewicz et al. 1994); Collard and Mackill 2009). The PCR reaction conditions were conducted using the SimpliAmp™ Thermal Cycler. The protocol involved an initial pre-denaturation step at 95 °C for 5 min, followed by 45 cycles of denaturation at 94 °C for 30 s, annealing for 40 s, and extension at 72 °C for 1 min. The reaction was concluded with a final extension step at 72 °C for 5 min. Subsequently, the PCR products were subjected to electrophoresis on a 1.5% agarose gel. The gels were assigned a score of (1) to indicate presence and (0) to indicate absence. Genetic parameters assessed the efficacy of the markers in distinguishing the studied accessions. The polymorphism information content (PIC) calculates using the formula PIC = 1 − Σpi2, where pi represents the frequency of the ith allele(Anderson et al. 1993) stated. The effective multiplex ratio (EMR), that is the frequency of the total number of polymorphic loci (np) to the number of polymorphic loci per primer (n), was calculated by the formula of EMR = np (np/n). The marker index (MI) was estimated by Nagaraju et al. (2001) and Powell et al. (1996), as the follow of formula MI = PIC × EMR. The resolving power (RP) is the summation of band informativeness (Ib), which scale from 0 to 1. The Ib is the proportion of accessions containing the band represented by the formula \({\text{Ib}} = {1} - \left( {{2} - \left( {0.{5} - {\text{p}}} \right)} \right)\) (Prevost and Wilkinson 1999).
DNA barcoding
The DNA extraction was carried out on approximately 0.5 g of cultivated plants and their wild relatives using the DNeasy 96 Plant Mini Kit manufactured by QIAGEN in Hilden, Germany. The ITS1 (5′-TCTGTAGGTGAACCTGCGG-3′) and ITS2 (5′-TCCTCCGCTTATTGATATGC-3′), targeting the ITS region were used to barcode the studied taxa., targeting. Following purification with a QIAquick PCR purification kit from QIAGEN, Hilden, Germany, the DNA was subjected to DNA sequencing. The sequencing of the product PCR was carried out using the Big DyeTM Terminator Cycle Sequencing Kits in an automatic sequencer ABI PRISM 3730XL analyzer, as provided by the Microgen Company located in Moscow, Russia. The NCBI BLAST database was utilized for this purpose. The phylogenetic tree was constructed using MEGA 10 software and the statistical methodology of UPGMA (Unweighted Pair Group Method with Arithmetic Mean), as documented on the website https://www.megasoftware.net.
Statistical analysis
The construction of the heatmap analysis was carried out utilizing the TBtools software package. PAST (ver. 4, Past Software, University of Oslo, Oslo, Norway) was utilized to perform principal component analysis (PCA).
Results
Molecular analysis
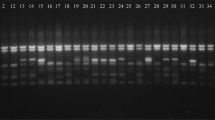
The use of molecular analysis is due to estimate the genetic diversity among the studied cultivated taxa and their wild relatives using five primers of ISSR and SCoT (Fig. 2 and 3). The molecular size of ISSR primers’ bands ranges from 178 to 1856 bp. About 206 bands are produced, including 169 polymorphic bands with polymorphism of 82.04%. The molecular size of bands produced by five SCoT primers ranges from 162 to 1825 bp. The SCoT profile produced 207 bands; the number of polymorphic bands produced 169, with a polymorphism of 81.64%.
The DNA profile generated from inter simple sequence repeat (ISSR) primers. M: Marker, 1: Apium graveolens, 2: Apium leptophyllum, 3: Daucus carota, 4: Daucus litoralis, 5: Lactuca sativa, 6: Lactuca serriola, 7: Brassica oleracea, 8: Brassica tournefortii, 9: Raphanus sativus, 10: Raphanus raphanistrum
For the Apiaceae, ISSR1 Produces the highest total number of bands (20 bands), while the lowest number is produced by ISSR3 primers (14 bands). The highest number of unique bands is produced by ISSR1 (11 bands). The highest value of resolving power (RP) is 10.13 and obtained by ISSR5, while the lowest value is 5.88 and obtained by ISSR3. ISSR1 shows the highest result of EMR and MI with values of 16.2 and 12.64, respectively, while ISSR5 records the lowest result of EMR and MI with values of 9.6 and 6.34. The highest value of polymorphism is 92.86%, recorded by ISSR3, while the lowest value is 80%, obtained by ISSR5 primer, as shown in Table 3. SCoT analysis shows that SCoT2 Produces the highest total number of bands (20 bands) while SCoT4 produces the lowest number of total bands (12 bands). The highest number of unique is 12 bands produced by SCoT2. Resolving power (RP) shows the highest value (12) obtained by SCoT5, while SCoT4 obtains the lowest value (6). SCoT2 shows the highest result of EMR and MI with values of 14.45 and 10.84, respectively, while SCoT4 records the lowest result of EMR and MI with values of 8.33 and 6.25. The value of polymorphism is 88.24% by SCoT1 and SCoT3 primers, while the lowest value is 77.78%, obtained by the SCoT5 primer, as shown in Table 3.
For the Asteraceae, ISSR4 Produces the highest number of bands (8 bands), while ISSR2 and ISSR3 primers (6 bands) produce the lowest number of bands. The highest value of resolving power (RP) is 7.5, obtained by ISSR 2 and ISSR3, while the lowest value is 5, obtained by ISSR5. The highest result of EMR and MI with values 5.14 and 3.29 is recorded in ISSR5, while the lowest is recorded in ISSR2 and ISSR3 as 1.5 and 0.57, respectively. The value of polymorphism becomes the maximum value (85.71) by ISSR5 while the minimum value is 50% and recorded by ISSR2 and ISSR3 primer as shown in Table 4. For SCoT analysis, the SCoT5 primer produces 11 bands as the highest total number of bands, while the SCoT3 primer produces the lowest number of bands (4). No unique bands are recorded. The highest value of resolving power (RP) is 11 and obtained by SCoT2, while the lowest value is (1) and obtained by SCoT4. The highest result of EMR and MI, with values of 5.82 and 3.2, respectively, is recorded by SCoT5, while SCoT1 records their lowest value (1.5 and 0.57, respectively). The maximum value of polymorphism is 80%, obtained by SCoT4 primer, and the minimum value is 50%, obtained by SCoT1, as shown in Table 4.
For the Brassicaceae, the highest number of bands is 22 bands obtained by ISSR4, but the lowest number (16 bands) is produced by ISSR2, ISSR3 and ISSR5 primers. ISSR4 produces the highest number of unique bands (12 bands). The highest value of resolving power (RP) is 11.5 and obtained by ISSR1, while the lowest value is 8.25 and obtained by ISSR3. ISSR4 shows the highest result of EMR (26.41), while the lowest result of EMR (10.56) is recorded by ISSR2 and ISSR5 primers. ISSR4 records the highest result of MI (12.31), while the lowest result is 7.40 and recorded by ISSR1. The maximum value of polymorphism is 87.5%, recorded in ISSR3, while the minimum value is 77.78, recorded in ISSR1 primer, as shown in Table 5. Using SCoT primer, SCoT1 and SCoT2 produce the highest number of bands (19 bands), while SCoT3 produces the lowest number of total bands (15 bands). The highest number of unique bands is 11, produced by SCoT1 and SCoT5 (Table 6). Resolving power (RP) shows the highest value (12.5) obtained by SCoT2, while the lowest value is 7.88, obtained by SCoT5 primer. SCoT5 shows the highest result of EMR is 14.22, but the lowest value is 11.27, produced by SCoT3 primer. The marker index (MI) shows the highest value scored 11.09 for SCoT5, while the lowest result of MI recorded 7.93) for SCoT2 primer. The maximum value of polymorphism is 88.89% and produced by SCoT5 primer, but the minimum value is 78.95%, produced by SCoT2, as shown in Table 5.
DNA barcoding
The accession numbers with sequences of the studied taxa are presented in Table 7. The sequence length, AT% and GC % content for studied taxa are recorded in Table 8. In the Apiaceae, the sequence length ranged from 536 bp in Daucus litoralis to 649 bp in D. carota. AT% content also ranged from 45.3% in Apium leptophyllum to 48% in D. carota, while GC% content ranged from 52% in D. carota to 54.7% in A. leptophyllum. The region of ITS (rDNA) used as the identity in the BLAST search (https://blast.ncbi.nlm.nih.gov/Blast.cgi)shows a pairwise identity (PI) of 99.83% for A. graveolens and 99.66% for A. leptophyllum with A. graveolens (FJ986043) (China). For Daucus species, the pairwise identity (PI) of 99.85% for D. carota with D. carota (MH711237) (China) while the pairwise identity (PI) of 99.44% for D. litoralis with D. virgatus (KT347710) (Poland) as shown in Table 8 and Fig. 4.
The phylogenetic analysis in the Apiaceae is presented in Fig. 4. The phylogenetic tree showed two main clades. The large clade is divided into two main subclades. The largest one includes the studied Apium graveolens, A. leptophyllum and D. carota with different Apium and Daucus sp. from different countries with bootstrap support (100% BS). The small sub-clade includes A. garveolens (Netherlands) and D. carota (China) with bootstrap support BS of 34%. The small clade includes the studied D. litoralis with A. graveolens (United Kingdom) and D. carota (Republic of Korea) with bootstrap support BS of 38%.
Table 8 shows that the sequence length in the Asteraceae is 638 bp in Lactuca sativa and 640 bp in L. serriola. Lactuca sativa has 45.5% AT and 54.5% GC content, while L. serriola contains 45.3% AT and 54.7% GC content. The BLAST search showed a pairwise identity (PI) of 100% for studied L. sativa and L. serriola with each other and with different Lactuca sp. from different countries.
The phylogenetic analysis in the Asteraceae is presented in Fig. 5. The phylogenetic tree showed two main clades. The small clade includes Lactuca sativa from different countries. The large clade is divided into two main subclades with bootstrap support (47%). The largest one includes the studied L. sativa, L. serriola, with different Lactuca species from different countries with bootstrap support (45% BS). The studied L. sativa and L. serriola have bootstrap support BS of 99%.
In the Brassicaceae, the sequence length ranged from 443 bp in B. tournefortii to 611 bp in R. sativus. GC% content ranged from 52.4% in B. tournefortii to 53.7% in R. sativus. AT% content ranged from 46.3% in R. sativus to 47.6% in B. tournefortii. The BLAST search showed a pairwise identity (PI) of 99.84% for B. oleracea with B. oleracea (MN401733) (Republic of Korea) and 99.77% for B. tournefortii with B. tournefortii (LC089997) (Japan). The pairwise identity (PI) of 99.82% for R. sativus and R. raphanistrum with R. sativus (AY722480) (Canada) is shown in Table 8.
The phylogenetic analysis in the Brassicaceae is presented in Fig. 6. The phylogenetic tree showed two main clades with bootstrap support BS of 28%. The small clade includes Brassica and Raphanus species from different countries. The large clade is divided into two main subclades with bootstrap support (33% BS). The small sub-clade includes B. tournefortii (LC089997) (Japan). The largest sub-clade includes all studied taxa (B. tournefortii, B. oleracea, R. sativus and R. raphanistrum). Raphanus sativus and R. raphanistrum (Egypt) with bootstrap support BS of 46%, while B. tournefortii and B. oleracea are with bootstrap support BS of 99%.
Statistical analysis
The findings obtained from the hierarchical cluster analysis of the heatmap indicated the presence of two distinct types of dendrograms. Specifically, the studied taxa dendrogram was observed vertically, while the molecular markers (ISSR and SCoT) were arranged horizontally (Fig. 7). The dendrogram of the cultivated and its relative wild plants reveals the presence of two main clusters. The first contains four Apiaceae taxa (A. gravolens L., A. leptophyllum (Pers.) F.Muell., D. carota L. and D. litoralis Sm.), while the second cluster comprises two subgroups. The initial subgroup comprises two Asteraceae taxa (L. sativa L. and L. serriola L.); the second comprises four Brassicaceae taxa. The observed red color denoted a significant resemblance among the examined taxa, whereas the blue signified a comparatively lower degree of similarity.
Principal Component Analysis (PCA) was estimated for both markers to demonstrate the clustering and correlation among the studied taxa under investigation. PCA1 accounts for a significant portion of the variance (39.9%), and PCA2 accounts for 29.5% (Fig. 8). The Principal Component Analysis (PCA) yielded a distinct separation of the taxa from Brassicaceae in a separate group which the molecular marker ISSR 1 and SCoT 3 can separate this group. At the same time, the taxa from Apiaceae and Asteraceae obtained were superimposed by the effect of molecular markers ISSR (2, 3 & 4) and SCoT (1, 2 & 5).
Discussion
For numerous years, crop wild relatives (CWR), commonly referred to as wild counterparts of cultivated plant species, have served as sources of genetic diversity (Tirnaz et al. 2022). They can confer advantageous traits to cultivated plant species. The progenitors of cultivated plants originate from their indigenous habitats, where they have adapted to survive in diverse abiotic and biotic stressors (Dempewolf et al. 2017). As a result, it is recommended that breeding programs incorporate these genetic lineages into cultivated plant species. The utilization of SCoT and ISSR markers in assessing genetic diversity and relatedness is of great significance in the enhancement of crops. Utilizing multilocus DNA markers, including ISSRs and SCoTs, is common among numerous plant species and genera (Soliman et al. 2023; Safhi et al. 2022). However, the efficacy of these markers may be subject to considerable variation contingent upon the specific plant species employed (Abd El-Moneim et al. 2020; Abd El-Moneim et al.2021; EL-Mansy et al. 2021).
The Apiaceae exhibits a polymorphism percentage of 84.52% with 71 polymorphic bands when analyzed using the SCoT marker. Similarly, ISSR analysis of the family revealed a polymorphism percentage of 86.72% with a total of 73 polymorphic bands. The percentage of polymorphism resulting from the ISSR marker was observed to be greater than the percentage of polymorphism resulting from the three ISSR primers utilized in the investigation of genetic diversity across four species of the Apiaceae (Altameme and Ibraheam 2018). The profile generated Approximately 50 polymorphic bands with a polymorphism percentage of 59.53%.
The Asteraceae exhibits a notable polymorphism, with the SCoT marker yielding the highest percentage at 67.55%. The marker analysis produced a total band of 36, and polymorphic bands of 24. The ISSR molecular marker exhibited a polymorphism of 66.43%, with a total band of 34, and polymorphic bands of 23. The findings presented here are in contrast to the research on genetic diversity among certain Indian lettuce (Asteraceae), wherein ISSR analysis revealed a collective count of 60 bands, of which 22 bands exhibited polymorphism, with a percentage of polymorphism amounting to 36.32% (Pham et al. 2022).
The Brassicaceae exhibits a polymorphism percentage of 82.83% through ISSR, resulting in 73 polymorphic bands. Using SCoT primers produced 74 polymorphic bands a polymorphism of 85.24%. Khalil and EL-Zayat (2019) conducted a study wherein they performed ISSR analysis on various Brassica species. The results indicated that 326 total bands were generated using 19 ISSR primers, out of which 36 were monomorphic and 290 were polymorphic bands; the level of polymorphism was 89%. The study by Jelvehgar et al. (2021) showcases the potential of SCoT markers in assessing genetic diversity within the Brassicaceae. The researchers discovered a substantial degree of genetic variation and postulated that certain species of Brassicaceae could be employed in conservation initiatives and breeding programs to satisfy the market's needs for vegetables and medicinal plants.
Polymorphism information content (PIC) refers to the ability to detect polymorphism within a given population. The Apiaceae exhibits a maximum PIC value of 0.75 when utilizing the ISSR marker, whereas the Asteraceae demonstrates a minimum PIC value of 0.5 when utilizing the ISSR primer. Several investigations have reported a Polymorphic Information Content (PIC) value greater than 0.5 for dominant markers (Olatunji and Afolayan 2019; EL-Mansy et al. 2021). The study indicated variation in the Polymorphism Information Content (PIC) ranging from 0.73, as observed in SCoT to 0.74, as demonstrated by ISSR. The polymorphic information content (PIC) is a metric that varies between 0 and 1, with values closer to 1 denoting greater levels of polymorphism. Furthermore, it is noteworthy that PIC can be categorized into three groups based on its level of informativeness: high informative (PIC > 0.5), moderate informative (0.25 < PIC < 0.5), and low informative (PIC < 0.25) (Abd El-Moneim et al. 2021; Botstein et al. 1980).
The scientific technique known as "DNA barcoding" pertains to a contemporary approach utilized for identifying plants, which involves the analysis of a concise and uniform DNA segment (Hebert et al. 2003). The fundamental principle of the discriminatory system is predicated on the notion that to differentiate between species; there must be a substantial degree of variation in the DNA barcode sequence that distinguishes them from one another. A DNA sequence derived from a standardized DNA region can be generated using a small tissue sample collected from an unidentified organism and subsequently matched against a repository of reference sequences sourced from documented species. Identifying an unknown organism can be expedited and dependable by conducting a comparative analysis between the organism's sequence and a reference sequence, as outlined by Casiraghi et al. (2010).
DNA barcoding has emerged as a viable alternative to traditional phyto-morphological techniques for species identification across all taxa due to the inherent stability of DNA sequences, which remain unaffected by environmental factors and remain constant throughout developmental stages (Liu et al. 2011). The present investigation utilized phylogenetic analysis of ITS regions to demonstrate the utility of nuclear regions in identifying and estimating molecular variations among the taxa under study. Within the Apiaceae, Daucus carota exhibits the highest pairwise identity at 99.85%, whereas Daucus litoralis displays the lowest value at 99.44%. Several investigations have documented the phylogenetic associations among Apiaceae constituents utilizing the ITS regions of 45S nrDNA, as evidenced by Downie et al. (2010) and Liu et al. (2014). The results of the phylogenetic analysis utilizing ITS regions in Asteraceae indicate that both Lactuca sativa and Lactuca serriola exhibit a pairwise identity of 100%, as per the investigation conducted. According to Gao et al. (2010) and Safhi et al. (2022), the utilization of DNA barcoding, specifically through the employment of the ITS region, is a valuable mechanism for categorizing and recognizing distinct species within the Asteraceae family. The application of DNA barcoding technology at the genera and species level within the family Asteraceae has been recommended to resolve classification issues (Safhi et al. 2023). The results of the phylogenetic analysis conducted on the Brassicaceae indicate that Brassica oleraceae exhibits the highest pairwise identity at 99.84%, whereas Brassica tournefortii displays the lowest value at 99.77%. Sun et al. (2015) employed DNA barcoding to identify species of significant economic value within the Brassicaceae. The Internal Transcribed Spacer 2 (ITS2) could serve as a viable plant barcode due to its ease of amplification and sequencing.
A heatmap was employed to investigate the congruencies and incongruities within wild and cultivated taxa. A hierarchical cluster heatmap is a viable approach to elucidate the interrelationships among various sample parameters, including molecular markers and the studied plant taxa. On the other hand, principal component analysis (PCA) is an unsupervised learning technique that effectively reduces the dataset's dimensionality and extracts the most significant information for analysis, (Khan et al. 2019). The study utilized heatmap and multivariate principal component analysis (PCA) techniques to visually represent the interrelationships, similarities, and dissimilarities among different parameters concerning studied cultivated genotypes and wild taxa based on their respective distances. The principal component analysis (PCA) outcome exhibited a segregation of taxa based on their respective molecular parameters (Mesfer ALshamrani et al. 2022; Safhi et al. 2022).
Conclusion
The anticipated global population growth of over 9 billion by 2050 necessitates the presence of genetic diversity in crops to facilitate their continued advancement. This diversity provides breeders alternative options for generating cultivars and hybrids. The attainment of diversity could favor using various molecular characterization techniques. The current investigation demonstrates that utilizing molecular markers such as ISSR and SCoT and molecular identification via DNA barcoding can be highly advantageous in discerning inter-species diversity, even with a limited number of primers. The findings indicate a significant of genetic diversity and relatedness among the cultivated taxa under investigation and their wild counterparts. The findings of this study offer genetic insights into specific taxa, providing valuable information for enhancing crop resistance to pathogens and improving their ability to thrive in adverse environmental conditions.
Data availability
The ITS sequences of the studied taxa were deposited at NCBI GenBank. All data generated or analyzed during this study are included in this published article.
References
Abd El-Moneim D (2020) Characterization of ISSR and SCoT markers and TaWRKY gene expression in some Egyptian wheat genotypes under drought stress. J Plant Prod Sci 8:31–46
Abd El-Moneim D, ELsarag EIS, Aloufi S, El-Azraq AM, ALshamrani SM, Safhi FAA, Ibrahim AA (2021) Quinoa (Chenopodium quinoa Willd.): genetic diversity according to ISSR and SCoT markers, relative gene expression, and morpho-physiological variation under salinity stress. Plants 10:2802. https://doi.org/10.3390/plants10122802
Altameme HJM, Ibraheam IA (2018) RAPD and ISSR analysis of the genetic relationship among some species in Rutaceae in and Apiceae in Iraq. Iraqi J Agric Sci 50(2):608–616
Anderson JA, Churchill GA, Autrique JE, Tanksley SD, Sorrells ME (1993) Optimizing parental selection for genetic linkage maps. Genome 36:181–186
Arslan E, Mutlu EG, Dursun O, Bagci SA (2020) Comparative analysis of agronomic traits and ISSR method among some soybeans [Glycine Max (L.) Merr.] genotypes. J Agric Nat 23(3):687–696. https://doi.org/10.18016/ksutarimdoga.v23i53104.631286
Bhandari HR, Bhanu AN, Srivastava K, Singh MN, Shreya HA (2017) Assessment of genetic diversity in crop plants—an overview. Adv Plants Agric Res 7(3):279–286. https://doi.org/10.15406/apar.2017.07.00255
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Boulos L (2009) Flora of Egypt checklist. Al-Hadara Publishing, Cairo, pp 198–201
Casiraghi M, Labra M, Ferri E, Galimberti A, De Mattia F (2010) DNA barcoding: a six-question tour to improve users’ awareness about the method. Brief Bioinform 11(4):440–453
Chai XT, Dong R, Liu WX, Wang YR, Liu ZP (2017) Optimizing sample size to assess the genetic diversity in common vetch (Vicia sativa L.) populations using Start Codon Targeted (SCoT) markers. Molecules 22:2–10
Collard BCY, Mackill DJ (2009) Start Codon Targeted (SCoT) polymorphism: a simple novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol Bio 27:86–93
Cota-Sanchez H, Remarchuk K, Ubayasenaj K (2006) Ready-to-use DNA extracted with a CTAB method adapted for herbarium specimens and mucilaginous plant tissue. Plant Mol Biol Report 24:161–167
Dempewolf H, Baute G, Anderson J, Kilian B, Smith C, Guarino L (2017) Past and future use of wild relatives in crop breeding. Crop Sci 57(3):1070–1082
DeSalle R, Goldstein P (2019) Review and interpretation of trends in DNA barcoding. Front Ecol Evol 7:302
Downie SR, Spalik K, Katz-Downie DS, Reduron JP (2010) Major clades within Apiaceae subfamily Apioideae as inferred by phylogenetic analysis of nrDNA ITS sequences. Plant Div Evol 128:111–136
EL-Mansy AB, Abd El-Moneim D, ALshamrani SM, Alsafhi FA, Abdein MA, Ibrahim AA (2021) Genetic diversity analysis of tomato (Solanum lycopersicum L.) with morphological, cytological, and molecular markers under heat stress. Horticulturae 7:65
Engels JM, Thormann I (2020) Main challenges and actions needed to improve conservation and sustainable use of our crop wild relatives. Plants 9(8):968
Etminan A, Pour-Aboughadareh AR, Mohammadi R, Ahmadi-Rad A, Noori A, Mahdavian Z, Moradi Z (2016) Applicability of start codon targeted (SCoT) and inter-simple sequence repeat (ISSR) markers for genetic diversity analysis in durum wheat genotypes. Biotechnol Biotechnol Equip 30(6):1075–1081
Ford-Lloyd BV, Schmidt M, Armstrong SJ, Barazani O, Engels J, Hadas R, Hammer K, Kell SP, Kang D, Khoshbakht K, Li Y, Long C, Lu B-R, Ma K, Nguyen VT, Qiu L, Ge S, Wei W, Zhang Z, Maxted N (2011) Crop wild relatives—undervalued, underutilized and under threat? Bioscience 61(7):559–565
Gao T, Yao H, Song J, Zhu Y, Liu C, Chen S (2010) Evaluating the feasibility of using candidate DNA barcodes in discriminating species of the large Asteraceae family. BMC Evol Biol 10:1–7
Hasan N, Choudhary S, Naaz N, Sharma N, Laskar RA (2021) Recent advancements in molecular marker-assisted selection and applications in plant breeding programmes. Genet Eng Biotechnol 19(1):2–26
Hebert PDN, Cywinska A, Ball SL, DeWaard JR (2003) Biological identification through DNA barcodes. Proc R Soc Lond [biol] 270:313–321
Henareh M, Dursun A, Abdollahi-Mandoulakani B, Haliloğlu K (2016) Assessment of genetic diversity in tomato landraces using ISSR markers. Genetika 48:25–35
Hollingsworth ML, Clark ALEX, Forrest LL, Richardson J, Pennington RT, Long DG, Hollingsworth PM (2009) Selecting barcoding loci for plants: evaluation of seven candidate loci with species-level sampling in three divergent groups of land plants. Mol Ecol Resour 9(2):439–457
Jabari M, Golparvar A, Sorkhilalehloo B, Shams M (2023) Investigation of genetic diversity of Iranian wild relatives of bread wheat using ISSR and SSR markers. J Genet Eng Biotechnol 21(1):1–16. https://doi.org/10.1186/s43141-023-00526-5
Jamdade R, Mosa KA, El-Keblawy A, Al Shaer K, Al Harthi E, Al Sallani M, Al Jasmi M, Gairola S, Shabana H, Mahmoud T (2022) DNA barcodes for accurate identification of selected medicinal plants (Caryophyllales): toward barcoding flowering plants of the United Arab Emirates. Diversity 14(4):262
Jelvehgar N, Miri SM, Mostafavi K, Mohammadi A (2021) Assessing the suitability of SCoT markers for studying genetic variation and genetic structure of Lepidium species. Iran J Genet Plant Breed 10(1):91–100
Khalil RMA, El-Zayat MAS (2019) Molecular characterization of some Brassica species. Adv Plants Agric Res 1(3):112–119
Khan N, Bano A, Babar MA (2019) Metabolic and physiological changes induced by plant growth regulators and plant growth promoting rhizobacteria and their impact on drought tolerance in Cicer arietinum L.. PLoS ONE 14(3):e0213040. https://doi.org/10.1371/journal.pone.0213040
Khoury CK, Carver D, Kates HR, Achicanoy HA, van Zonneveld M, Thomas E, Heinitz C, Jarret R, Labate JA, Reitsma K, Nabhan GP, Greene SL (2020) Distributions, conservation status, and abiotic stress tolerance potential of wild cucurbits (Cucurbita L.). Plant People Planet 2(3):269–283
Kumar GP, Pathania P, Goyal N, Gupta N, Parimalan R, Radhamani J, Gomashe SS, Kadirvel P, Rajkumar S (2023) Genetic diversity and population structure analysis to construct a core collection from safflower (Carthamus tinctorius L.) germplasm through SSR markers. Agriculture 13(4):836. https://doi.org/10.3390/agriculture13040836
Liu C, Liang D, Gao T, Pang X, Song J, Yao H, Chen S (2011) PTIGS-IdIt, a system for species identification by DNA sequences of the psbA-trnH intergenic spacer region. BMC Bioinform 12:1–7
Liu J, Shi L, Han J, Li G, Lu H, Hou J, Zhou X, Meng F, Downie SR (2014) Identification of species in the angiosperm family Apiaceae using DNA barcodes. Mol Ecol Res 14:1231–1238
Mansour HM, Younis RAA, Eldomiati FM, Rashed MA, Hassanein SH (2020) Phylogenetic analyses of some Egyptian genus of Lamiaceae family using rbcL sequences. Arab Univ J Agric Sci 28(1):229–237. https://doi.org/10.21608/AJS.2020.26513.1184
Maxted N, Vincent H (2021) Review of congruence between global crop wild relative hotspots and centers of crop origin/diversity. Genet Resour Crop Evol 68:1283–1297
Mesfer ALshamrani S, Safhi FA, Alshaya DS, Ibrahim AA, Mansour H, Abd El Moneim D (2022) Genetic diversity using biochemical, physiological, karyological and molecular markers of Sesamum indicum L. Front Genet 13:1035977. https://doi.org/10.3389/fgene.2022.1035977
Mezghani N, Khoury CK, Carver D, Achicanoy HA, Simon P, Flores FM, Spooner D (2019) Distributions and conservation status of carrot wild relatives in Tunisia: a case study in the Western Mediterranean Basin. Crop Sci 59:2317–2328
Monpara J, Kiran C, Vrinda T (2017) ISSR studies on small and large seed varieties of Glycine max. J Pharmacogn Phytochem 6:1652–1656
Mponya NK, Chanyenga T, Brehm JM, Maxted N (2021) In situ and ex situ conservation gap analyses of crop wild relatives from Malawi. Genet Resour Crop Evol 68:759–771
Nadeem MA, Nawaz MA, Shahid MQ, Doğan Y, Comertpay G, Yıldız M, Hatipoğlu R, Ahmad F, Alsaleh A, Labhane N, Özkan H (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32(2):261–285
Nagaraju J, Reddy KD, Nagaraja GM, Sethuraman BN (2001) Comparison of multilocus RFLPs and PCR-based marker systems for genetic analysis of the silkworm, Bombyx mori. Heredity (edinb) 86:588–597
Olatunji TL, Afolayan AJ (2019) Evaluation of genetic relationship among varieties of Capsicum annuum L. and Capsicum frutescens L. in West Africa using ISSR markers. Heliyon 5:e01700
Payacan C, Moncada X, Rojas G, Clarke A, Chung KF, Allaby R, Seelenfreund D, Seelenfreund A (2017) Phylogeography of herbarium specimens of asexually propagated paper mulberry [Broussonetia papyrifera (L.) L’Hér. Ex vent. (Moraceae)] reveals genetic diversity across the Pacific. Ann Bot 120(3):1–18
Perrino EV, Wagensommer RP (2021) Crop Wild Relatives (CWR) priority in Italy: distribution, ecology, in situ and ex situ conservation and expected actions. Sustainability 13:1–27
Pham TTT, Tran HTT, Cao PB, Ninh PT, Do NH, Dinh ST (2022) High genetic diversity of 16 Indian lettuce (Lactuca indica L.) accessions from Vietnam. Pak J Biol Sci 25:201–209
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112
Rayan WA (2019) Evaluating the efficiency of DNA Barcode rbcl for detection of genetic relationships between four Moringa spp. Genotypes. Egypt J Exp Biol (bot) 15(2):333–340
Rocha V, Duarte MC, Catarino S, Duarte I, Romeiras MM (2021) Cabo Verde’s Poaceae Flora: a reservoir of crop wild relatives diversity for crop improvement. Front Plant Sci 12:1–17
Safhi FA, Alshamrani SM, Bogmaza AFM, El-Moneim DA (2023) DNA barcoding of wild plants with potential medicinal properties from Faifa Mountains in Saudi Arabia. Genes 14:469. https://doi.org/10.3390/genes14020469
Safhi FA, Ashamrani SM, Jalal AS, El-Moneim DA, Alyamani AA, Ibrahim AA (2022) Genetic characterization of some Saudi Arabia’s accessions from Commiphora gileadensis using physio-biochemical parameters, molecular markers, DNA barcoding analysis and relative gene expression. Genes 13:2099. https://doi.org/10.3390/genes13112099
Satya P, Karan M, Jana S, Mitra S, Sharma A, Karmakar PG, Ray DP (2015) Start codon targeted (SCoT) polymorphism reveals genetic diversity in wild and domesticated populations of ramie (Boehmeria nivea L. Gaudich.), a premium textile fiber producing species. Meta Gene 3:62–70
Senapati A, Basak S, Rangan L (2022) A review on application of DNA barcoding technology for rapid molecular diagnostics of adulterants in herbal medicine. Drug Saf 24:193–213
Singh A, Abhilash PC (2018) Agricultural biodiversity for sustainable food production. J Clean Prod 172:1368–1369
Singh A, Dubey PK, Chaurasia R, Mathur N, Kumar G, Bharati S, Abhilash PC (2018) Indian spinach: an underutilized perennial leafy vegetable for nutritional security in developing world. Energy Ecol Environ 3:195–205
Soliman MI, Mohammed NS, Safhi FA, ALshamrani SM, Alyamani AA, Alharthi B, Qahl SH, Al Kashgry NA, Ibrahim AA (2023) Antibacterial, antioxidant activities, GC-mass characterization, and cyto/genotoxicity effect of green synthesis of silver nanoparticles using latex of Cynanchum acutum L. Plants 12(1):172. https://doi.org/10.3390/plants12010172
Sun XQ, Qu YQ, Yao H, Zhang YM, Yan QQ, Hang YY (2015) Applying DNA barcodes for identification of economically important species in Brassicaceae. Genet Mol Res 14(4):15050–15061
Tackholm V (1974) Students’ flora of Egypt. Cairo University Egypt Publishing, Beirut, p 888
Taylor HR, Harris WE (2012) An emergent science on the brink of irrelevance: a review of the past 8 years of DNA barcoding. Mol Ecol Resour 12(3):377–388
Tirnaz S, Zandberg J, Thomas WJ, Marsh J, Edwards D, Batley J (2022) Application of crop wild relatives in modern breeding: an overview of resources, experimental and computational methodologies. Front Plant Sci 13:1008904. https://doi.org/10.3389/fpls.2022.1008904
Whitney CW, Luedeling E, Tabuti JR, Nyamukuru A, Hensel O, Gebauer J, Kehlenbeck K (2018) Crop diversity in home-gardens of southwest Uganda and its importance for rural livelihoods. Agric Hum Values 35:399–424
Yang M, Abdalrahman H, Sonia U, Mohammed A, Vestine U, Wang M, Ebadi A, Toughani M (2020) The application of DNA molecular markers in the study of Codonopsis species genetic variation, a review. Cell Mol Biol 66(2):23–30
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This research received no external funding.
Author information
Authors and Affiliations
Contributions
MIS and AAI conceived and designed the experiments; AAI analyzed the data; MIS, AAI and NME summarized the data; AAI, MIS and NME wrote the manuscript; MIS and AAI revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
All authors declare they have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Essa, N.M., Ibrahim, A.A. & Soliman, M.I. Association study between some cultivated species and their wild relatives from Apiaceae, Asteraceae and Brassicaceae families based on molecular and DNA barcoding in Egypt. Genet Resour Crop Evol 71, 1125–1143 (2024). https://doi.org/10.1007/s10722-023-01681-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01681-x