Abstract
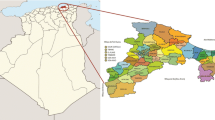
Unidentified olive plants naturally grow in the Golestan province of Iran, on different soils and under climates spanning from sub-temperate to desert conditions, represented by single trees or groups of few trees. We collected samples from representative sites and analyzed them by simple sequence repeat markers in order to determine their identity and their relationships to prominent Iranian and Mediterranean reference cultivars. Population structure analysis separated these ecotypes from Mediterranean and, surprisingly, from all Iranian cultivars, the parentage test excluded their direct contribution as candidate parents or offspring of cultivars, and they also showed a high level of admixture. Their differentiation from cultivated olives may be attributed to different factors: they could represent wild plants or could derive from natural dissemination of ancestral cultivated trees. Their survival up to now may be due to the fact that most of them are grown on sacred sites such as necropolis. Anyhow, the adaptation to strong environmental stresses, and their fruit size and oil content make the olive Golestan ecotypes a valuable source of genetic variation previously uncharacterized and currently threatened with extinction.



Similar content being viewed by others
References
Akhani H, Djamali M, Ghorbanalizadeh A, Ramezani E (2010) Plant biodiversity of Hyrcanian relict forests, N Iran: an overview of the flora, vegetation, palaeoecology and conservation. Pak J Bot 42:231–258
Ayoubi S, Khormali F, Sahrawat KL, Rodrigues de Lima AC (2011) Assessing impacts of land use change on soil quality indicators in a loessial soil in Golestan Province. Iran J Agr Sci Tech 13:727–742
Bagnoli F, Vendramin GG, Buonamici A, Doulis AG, Gonzalez-Martinez SC, Laporta N, Magri D, Raddi P, Sebastini F, Fineschi S (2009) Is Cupressus sempervirens native in Italy? An answer from genetic and palaeobotanical data. Mol Ecol 18:2276–2286
Baldoni L, Cultrera NGM, Mariotti R, Ricciolini C, Arcioni S, Vendramin GG (2009) A consensus list of microsatellite markers for olive genotyping. Mol Breed 24:213–231
Beghe D, Ferrarini A, Ganino T, Fabbri A (2011) Molecular characterization and identification of a group of local Olea europaea L. varieties. Tree Genet Genomes 7:1185–1198
Belaj A, Domingez-Garcia MC, Atienza SG, Urdiroz NM, De la Rosa R, Satovic Z, Martin A, Kilian A, Trujillo I, Valpuesta V (2012) Developing a core collection of olive (Olea europaea L.) based on molecular markers (DArTs, SSRs, SNPs) and agronomic traits. Tree Genet Genomes 8:365–378
Carriero F, Fontanazza G, Cellini F, Giorio G (2002) Identification of simple sequence repeats (SSRs) in olive (Olea europaea L.). Theor Appl Genet 104:301–307
Cipriani G, Marrazzo MT, Marconi R, Cimato A, Testolin R (2002) Microsatellite markers isolated in olive (Olea europaea L.) are suitable for individual fingerprinting and reveal polymorphism within ancient cultivars. Theor Appl Genet 104:223–228
D’Imperio M, Viscosi V, Scarano M, D’Andrea M, Zullo BA, Pilla F (2011) Integration between molecular and morphological markers for the exploitation of olive germplasm (Olea europaea). Sci Hort 130:229–240
De La Rosa R, James CM, Tobutt KR (2002) Isolation and characterization of polymorphic microsatellites in olive (Olea europaea L.) and their transferability to other genera in the Oleaceae. Mol Ecol Notes 2:265–267
Diez CM, Trujillo I, Barrio E, Belaj A, Barranco D, Rallo L (2011) Centennial olive trees as a reservoir of genetic diversity. Ann Bot 108:797–807
Doveri S, Gil FS, Diaz A, Reale S, Busconi M, Machado AC, Martin A, Fogher C, Donini P, Lee D (2008) Standardization of a set of microsatellite markers for use in cultivar identification studies in olive (Olea europaea L.). Sci Hort 116:367–373
Floor W (2005) Olive tree. Encyclopaedia Iranica
Ghavami S, Mohammad AD, Saeid S, Ga Saeid, Saeb J (2007) An investigation of spider fauna of olive orchards in northern part of Iran. Pak J Biol Sci 10(15):2562–2568
Hakim IR, Kammoun NG, Makhloufi E, Rebai A (2010) Discovery and potential of SNP markers in characterization of Tunisian olive germplasm. Diversity 2:17–27
Hannachi H, Breton C, Monji M, Ben EI, Hadj S, Gazzah M, Berville A (2008) Differences between native and introduced olive cultivars as revealed by morphology of drups, oil composition and SSR polymorphisms: a case study in Tunisia. Sci Hort 116:280–290
Haouane H, El Bakkali A, Moukhli A, Tollon C, Santoni S, Oukabli A, El Modafar C, Khadari B (2011) Genetic structure and core collection of the world olive Germplasm Bank of marrakech: towards the optimized management and use of Mediterranean olive genetic resources. Genetica 139:1083–1094
Heshmati GA (2013) Indigenous plant species from the drylands of Iran, distribution and potential for habitat maintenance and repair. In: Heshmati GA, Squires VR (eds) Combating Desertification in Asia, Africa and the Middle East. Springer, NY
IOC (2000) World olive cultivars catalogue
Khoshbakht K, Hammer K (2006) Savadkouh (Iran): an evolutionary centre for fruit trees and shrubs. Genet Resour Crop Evol 53:641–651
Marshall TC, Salte J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655
Muzzalupo I, Chiappetta A, Benincasa C, Perri E (2010) Intra-cultivar variability of three major olive cultivars grown in different areas of central-southern Italy and studied microsatellite markers. Sci Hort 126:324–329
Noormohammadi Z, Hosseini-Mazinani M, Trujillo I, Belaj A (2009) Study of intracultivar variation among main Iranian olive cultivars using SSR markers. Acta Biologica Szegediensis 53:27–32
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/darwin
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rechinger KH (1968) Trib. Magnoliopsida, Lamiales (Oleaceae) In: Rechinger KH (ed.) Flora Iranica, vol 52. Graz
Rehman AU, Mailer RJ, Belaj A, de la Rosa R, Raman H (2012) Microsatellite marker-based identification of mother plants for the reliable propagation of olive (Olea europaea L.) cultivars in Australia. J Hortic Sci Biotech 87:647–653
Sarri V, Baldoni L, Porceddu A, Cultrera NGM, Contento A, Frediani M, Belaj A, Trujillo I, Cionini PG (2006) Microsatellite markers are powerful tools for discriminating among olive cultivars and assigning them to geographically defined populations. Genome 49:1606–1615
Sefc KM, Lopes MS, Mendonca D, Rodrigues dos Santos M, da Camara Laimer, Machado M, da Camara Machado A (2000) Identification of microsatellite loci in olive (Olea europaea) and their characterization in Italian and Iberian olive trees. Mol Ecol 9:1171–1173
Tatjana K, De la Rosa R, Satovic Z, Leon L, Belaj A (2013) Utility of wild germplasm in olive breeding. Sci Hort 152:92–101
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Acknowledgments
The research was performed within the international collaboration between NIGEB (Iran) and CNR-IGV (Italy) with the contribution of the Iranian Ministry of Agriculture (Olive Research Office), who contributed to collect samples from different parts of the Golestan province. We would like to thank Tarbiat Modares University (TMU) for providing facilities and financial support.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mousavi, S., Hosseini Mazinani, M., Arzani, K. et al. Molecular and morphological characterization of Golestan (Iran) olive ecotypes provides evidence for the presence of promising genotypes. Genet Resour Crop Evol 61, 775–785 (2014). https://doi.org/10.1007/s10722-013-0071-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-013-0071-4