Abstract
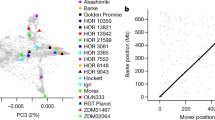
The discovery of Hordeum spontaneum C. Koch, a wild ancestor of cultivated barley, in Morocco in 1978 led to the proposal of a multicentric origin for this crop, as an alternative to the widely accepted theory of a single centre of domestication in the Fertile Crescent. Since this discovery, we have tested this hypothesis using the most advanced genetic techniques available at the time, from CM-proteins to RFLP and DNA-chloroplast markers. Nowadays, the availability of single nucleotide polymorphism (SNP) markers that are spread densely over the barley genome provides us with another powerful tool to give further support for the above. We have used 1,536 SNPs from the Barley Oligo Pool Assay 1 (BOPA1) of Illumina to characterize 107 wild and cultivated barley accessions from the Western Mediterranean, Fertile Crescent, Ethiopia, and Tibet. The results have confirmed that each location of the above-mentioned germplasm groups clusters separately. Analysis of molecular variance enabled us to focus on the chromosomal regions and loci that differentiated these groups of barley germplasm. Some of these regions contain vernalization and photoperiod response genes, some of the so-called domestication genes, as well as the most important quantitative trait locus for flowering time in the Mediterranean region. We have combined these results with other genetic evidence, and interpreted them in the framework of current theories on the onset of the Neolithic revolution in the Mediterranean region, to conclude that neither Ethiopia nor the Western Mediterranean can be ruled out as centres of barley domestication, together with the Fertile Crescent.
Access this article
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.



Similar content being viewed by others
References
Abbo S, Lev-Yadun S, Gopher A (2010) Agricultural origins: centers and noncenters; a Near Eastern reappraisal. Crit Rev Plant Sci 29:319–328
Allaby RG, Fuller DQ, Brown TA (2008) The genetic expectations of a protracted model for the origins of domesticated crops. Proc Natl Acad Sci USA 105:13892–13896
Azhaguvel P, Komatsuda T (2007) A phylogenetic analysis based on nucleotide sequence of a marker linked to the brittle rachis locus indicates a diphyletic origin of barley. Ann Bot 100:1009–1015
Baba T, Tanno K, Furusho M, Komatsuda T (2011) Allelic variation at the EF-G locus among northern Moroccan six-rowed barleys. Plant Genet Resour Charact Util 9:240–242
Badr A, Müller K, Schäfer-Pregl R, El Rabey H, Effgen S, Ibrahim HH, Pozzi C, Röhde W, Salamini F (2000) On the origin and domestication history of barley (Hordeum vulgare). Mol Biol Evol 17:499–510
Bjørnstad A, Abay F (2010) Multivariate patterns of diversity in Ethiopian barleys. Crop Sci 50:1579–1586
Blattner FR, Badani-Méndez A (2001) RAPD data do not support a second centre of barley domestication in Morocco. Genet Resour Crop Evol 48:13–19
Boyd WJR, Li CD, Grime CR, Cakir M, Potipibool S, Kaveeta L, Men S, Kamali MRJ, Barr AR, Moody DB, Lance RCM, Logue SJ, Raman H, Rea BJ (2003) Conventional and molecular genetic analysis of factors contributing to variation in the timing of heading among spring barley (Hordeum vulgare L.) genotypes grown over a mild winter growing season. Aust J Agric Res 54:1277–1301
Brown TA, Jones MK, Powell W, Allaby RG (2008) The complex origins of domesticated crops in the Fertile Crescent. Trends Ecol Evol 24:103–109
Casao MC, Igartua E, Karsai I, Bhat PR, Cuadrado N, Gracia MP, Lasa JM, Casas AM (2011) Introgression of an intermediate VRNH1 allele in barley (Hordeum vulgare L.) leads to reduced vernalization requirement without affecting freezing tolerance. Mol Breeding 28:475–484
Casas AM, Yahiaoui S, Ciudad F, Igartua E (2005) Distribution of MWG699 polymorphism in Spanish European barleys. Genome 48:41–45
Castro AJ, Hayes P, Viega L, Vales I (2008) Transgressive segregation for phenological traits in barley explained by two major QTL alleles with additivity. Plant Breeding 127:561–568
Close TJ, Bhat PR, Lonardi S, Wu Y, Wanamaker S, Rostoks N, Ramsay L, Stein N, Svensson JT, Bozdag S, Moscou M, Varshney R, Sato K, DeYoung J, Chao S, Waugh R, Marshall D, Graner A, Roose ML, Muehlbauer G, Matthews D, Madishetty K, Fenton RD, Condamine P (2009) Development and implementation of high-throughput SNP genotyping in barley. BMC Genomics 10:582
Cockram J, White J, Zuluaga DL, Smith D, Comadran J, Macaulay M, Luo Z, Kearsey MJ, Werner P, Harrap D, Tapsell C, Liu H, Hedley PE, Stein N, Schulte D, Steuernagel B, Marshall DF, Thomas WTB, Ramsay L, Mackay I, Balding D, The AGOUEB Consortium, Waugh R, O’Sullivan DM (2011) Genome-wide association mapping to candidate polymorphism resolution in the unsequenced barley genome. Proc Natl Acad Sci USA 107:21611–21616
Comadran J, Ramsay L, MacKenzie K, Hayes P, Close TJ, Muehlbauer G, Stein N, Waugh R (2011a) Patterns of polymorphism and linkage disequilibrium in cultivated barley. Theor Appl Genet 122:523–531
Comadran J, Russell JR, Booth A, Pswarayi A, Ceccarelli S, Grando S, Stanca AM, Pecchioni N, Akar T, Al-Yassin A, Benbelkacem A, Ouabbou H, Bort J, van Eeuwijk FA, Thomas WTB, Romagosa I (2011b) Mixed model association scans of multi-environmental trial data reveal major loci controlling yield and yield related traits in Hordeum vulgare in Mediterranean environments. Theor Appl Genet 122:1363–1373
Cuesta-Marcos A, Igartua E, Ciudad FJ, Codesal P, Russell JR, Molina-Cano JL, Moralejo M, Szűcs P, Gracia MP, Lasa JM, Casas AM (2008) Heading date QTL in a spring x winter barley cross evaluated in Mediterranean environments. Mol Breeding 21:455–471
Cuesta-Marcos A, Szűcs P, Close TJ, Filichkin T, Muehlbauer GJ, Smith KP, Hayes PM (2010) Genome-wide SNPs and re-sequencing of growth habit and inflorescence genes in barley: implications for association mapping in germplasm arrays varying in size and structure. BMC Genomics 11:707
Druka A, Franckowiak J, Lundqvist U, Bonar N, Alexander J, Houston K, Radovic S, Shahinnia F, Vendramin V, Morgante M, Stein N, Waugh R (2011) Genetic dissection of barley morphology and development. Plant Physiol 155:617–627
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Excoffier L, Hofer T, Foll M (2009) Detecting loci under selection in a hierarchically structured population. Heredity 103:285–298
Gross BL, Olsen KM (2010) Genetic perspectives on crop domestication. Trends Plant Sci 15:529–537
Hammer K, Lehmann CO, Perrino P (1985) Character variability and evolutionary trends in a barley hybrid swarm – a case study. Biol Zentralblatt 104:511–517
Harlan JR (1992) Crops & Man. ASA, CSSA, Madison
Hübner S, Günther T, Flavell A, Fridman E, Graner A, Korol A, Schmid KJ (2012) Islands and streams: clusters and gene flow in wild barley populations from the Levant. Mol Ecol 21:1115–1129
Komatsuda T, Maxim P, Senthil N, Mano Y (2004) High-density AFLP map of nonbrittle rachis 1 (btr1) and 2 (btr2) genes in barley (Hordeum vulgare L.). Theor Appl Genet 109:986–995
Komatsuda T, Pourkheirandish M, He C, Azhaguvel P, Kanamori H, Perovic D, Stein N, Graner A, Wicker T, Tagiri A, Lundqvist U, Fujimura T, Matsuoka M, Matsumoto T, Yano M (2007) Six-rowed barley originated from a mutation in a homeodomain-leucine zipper I-class homeobox gene. Proc Natl Acad Sci USA 104:1424–1429
Konishi T (2001) Genetic diversity in Hordeum agriocrithon E. Åberg, six-rowed barley with brittle rachis, from Tibet. Genet Res Crop Evol 48:27–34
Massman J, Cooper B, Horsley R, Neate S, Dill-Macky R, Chao S, Dong Y, Schwarz P, Muehlbauer GJ, Smith KP (2011) Genome-wide association mapping of Fusarium head blight resistance in contemporary barley breeding germplasm. Mol Breeding 27:439–454
Molina-Cano JL, Conde J (1980) Hordeum spontaneum C. Koch emend Bacht. collected in Southern Morocco. Barley Genet Newsl 10:44–47
Molina-Cano JL, Gómez Campo C, Conde J (1982) Hordeum spontaneum C. Koch as a weed of barley fields in Morocco. Z Pflanzenzücht 88:161–167
Molina-Cano JL, Fra-Mon P, Salcedo G, Aragoncillo C, Roca de Togores F, García Olmedo F (1987) Morocco as a possible domestication center for barley. Biochemical and agromorphological evidence. Theor Appl Genet 73:531–536
Molina-Cano JL, Moralejo M, Igartua E, Romagosa I (1999) Further evidence supporting Morocco as a center of origin of barley. Theor Appl Genet 98:913–918
Molina-Cano JL, Igartua E, Casas AM, Moralejo MA (2002) New views on the origin of cultivated barley. In: Slafer GA, Molina-Cano JL, Savin R, Araus JL, Romagosa I (eds) Barley science: recent advances from molecular biology to agronomy of yield and quality. The Haworth Press, Binghamton
Molina-Cano JL, Russell J, Moralejo MA, Escacena JL, Arias G, Powell W (2005) Chloroplast-DNA microsatellite analysis supports a polyphyletic origin for barley. Theor Appl Genet 110:613–619
Moragues M, Comadran R, Waugh R, Milne I, Flavell AJ, Russell JR (2010) Effects of ascertainment bias and marker number on estimations of barley diversity from high-throughput SNP genotype data. Theor Appl Genet 120:1525–1534
Morrell PL, Clegg MT (2007) Genetic evidence for a second domestication of barley (Hordeum vulgare) east of the Fertile Crescent. Proc Natl Acad Sci USA 104:3289–3294
Muzzolini A (1989) La “Néolithisation” du Nord de l’Afrique et ses causes. In: Aurenche O, Cauvin J (eds) Néolithisations: Proche et Moyen Orient, Méditerranée orientale, Nord de l’Afrique, Europe méridionale, Chine, Amérique du Sud. Bar International Series 516, pp 145–186
Orabi J, Backes G, Wolday A, Yahyaoui A, Jahoor A (2007) The horn of Africa as a center of barley diversification and a potential domestication site. Theor Appl Genet 114:1117–1127
Orabi J, Jahoor A, Backes G (2009) Genetic diversity and population structure of wild and cultivated barley from West Asia and North Africa. Plant Breeding 128:606–614
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/darwin
Pourkheirandish M, Komatsuda T (2007) The importance of barley genetics and domestication in a global perspective. Ann Bot 100:999–1008
Purugganan MD, Fuller DQ (2009) The nature of selection during plant domestication. Nature 457:843–848
Ramsay L, Comadran J, Druka A, Marshall DF, Thomas WTB, Macaulay M, MacKenzie K, Simpson CG, Fuller J, Hayes PM, Lundqvist U, Franckowiak JD, Close TJ, Muehlbauer G, Waugh R (2011) INTERMEDIUM-C, a modifier of lateral spikelet fertility in barley, is an ortholog of the maize domestication gene TEOSINTE BRANCHED 1. Nat Genet 43:169–173
Rostoks N, Mudie S, Cardle L, Russell J, Ramsay L, Booth A, Svensson JT, Wanamaker SI, Walia H, Rodriguez EM, Hedley PE, Liu H, Morris J, Close TJ, Marshall DF, Waugh R (2005) Genome-wide SNP discovery and linkage analysis in barley based on genes responsive to abiotic stress. Mol Genet Genomics 274:515–527
Russell JR, Dawson IK, Flavell AJ, Steffenson B, Weltzien E, Booth A, Ceccarelli S, Grando S, Waugh R (2011) Analysis of >1000 single nucleotide polymorphism in geographically matched samples of landrace and wild barley indicates secondary contact and chromosome-level difference in diversity around domestication genes. New Phytol 191:564–578
Saisho D, Purugganan MD (2007) Molecular phylogeography of domesticated barley traces expansion of agriculture in the Old World. Genetics 177:1765–1776
Sakuma S, Salomon B, Komatsuda T (2011) The domestication syndrome genes responsible for the major changes in plant form in the Triticeae crops. Plant Cell Physiol 52:738–749
Salamini F, Özkan H, Brandolini A, Schäffer-Pregl R, Martin W (2002) Genetics and geography of wild cereal domestication in the Near East. Nat Rev Genet 3:429–441
Sameri M, Takeda K, Komatsuda T (2006) Quantitative trait loci controlling agronomic traits in recombinant inbred lines from a cross of oriental- and occidental-type barley cultivars. Breeding Sci 56:243–252
Sneath PHA, Sokal RR (1973) Numerical taxonomy. WH Freeman and Company, San Francisco
Takahashi R (1955) The origin of cultivated barley. Adv Genet 7:227–266
Taketa S, Yao T, Sakurai Y, Miyake S, Ichii M (2011) Molecular mapping of the short awn 2 (lks2) and dense spike 1 (dsp1) genes on barley chromosome 7H. Breeding Sci 61:80–85
Tanno K, Takeda K (2004) On the origin of six-rowed barley with brittle rachis, agriocrithon [Hordeum vulgare ssp. f. agriocrithon (Åberg) Bowd.], based on a DNA marker closely linked to the vrs1 (six-row gene) locus. Theor Appl Genet 110:145–150
Tanno K, Taketa S, Takeda K, Komatsuda T (2002) A DNA marker closely linked to the vrs1 locus (row-type gene) indicates multiple origins of six-rowed cultivated barley (Hordeum vulgare L.). Theor Appl Genet 104:54–60
Trevaskis B (2010) The central role of the VERNALIZATION1 gene in the vernalization response of cereals. Funct Plant Biol 37:479–487
Weiss E, Kislev ME, Hartmann A (2006) Autonomous cultivation before domestication. Science 312:1608–1610
Willcox G (2005) The distribution, natural habitats and availability of wild cereals in relation to their domestication in the Near East: multiple events, multiple centres. Veget Hist Archaeobot 14:534–541
Zohary D (1999) Monophyletic vs. polyphyletic origin of crops on which agriculture was founded in the Near East. Gen Res Crop Evol 46:133–142
Acknowledgments
We want to thank INIA (MICINN) for partially funding this work through different grants. The Centre UdL-IRTA forms part of the Centre CONSOLIDER on Agrigenomics funded by the Spanish Ministry of Education and Science and acknowledges partial funding from grant AGL2005-07195-C02-02. Genotyping of the RIL population with BOPA1 was funded by the Spanish Ministry of Science and Innovation; project GEN2006-28560-E. We thank Prof. Nicolás Jouve Universidad de Alcalá de Henares, Spain) for supplying us with interesting information regarding mutation frequencies. We also thank Dr. Helmut Knüpffer (IPK, Gatersleben, Germany), who kindly supplied us with the Hordeum agriochrithon accessions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Lluís Torres: Deceased.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Igartua, E., Moralejo, M., Casas, A.M. et al. Whole-genome analysis with SNPs from BOPA1 shows clearly defined groupings of Western Mediterranean, Ethiopian, and Fertile Crescent barleys. Genet Resour Crop Evol 60, 251–264 (2013). https://doi.org/10.1007/s10722-012-9831-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-012-9831-9