Abstract
Plant architecture-related traits are important in breeding maize (Zea mays L.). Suwan maize germplasm has been widely used in tropical/subtropical regions because of its abundant genetic diversity compared to temperate germplasm. To investigate the influence of the genetic base on the detection of quantitative trait loci (QTLs) controlling plant architecture-related traits, a population of 150 F2:3 progeny derived from ZHL908 and HCL645 were developed and genotyped using the genotyping by sequencing (GBS) method. Through inclusive composite interval mapping using 5636 SNP markers, a total of 25 QTL-controlling plant architecture-related traits were identified. Seventeen QTLs had been previously reported, which may be important in future MAS breeding. Eight new QTLs were reported for the first time in this study. Of the 25 QTLs, 7 QTLs were related to plant height (PH), 4 QTLs were related to ear height (EH), 4 QTLs were related to leaf number (LN), 5 QTLs were related to tassel length (TL), and 5 QTLs were related to tassel primary branch number (TPBN). Four plant architectural trait-related genes, sparse inflorescence1, ramosa3, dwarf1, and zea floricaula/leafy1, were found to be located in qLN4, qTL5, qTPBN1 and qTPBN5, respectively. These results not only provide new insights into the genetic research investigating maize plant architecture variation but also provide molecular evidence that may enable the improvement of Suwan germplasm in modern maize breeding.







Similar content being viewed by others
References
Bajgain P, Rouse MN, Tsilo TJ, Macharia GK, Bhavani S, Jin Y et al (2016) Nested association mapping of stem rust resistance in wheat using genotyping by sequencing. PLoS ONE 11(5):e0155760
Bouchet S, Servin B, Bertin P, Madur D, Combes V, Dumas F et al (2013) Adaptation of maize to temperate climates: mid-density genome-wide association genetics and diversity patterns reveal key genomic regions, with a major contribution of the Vgt2 (ZCN8) locus. PLoS ONE 8(8):e71377
Bouchet S, Bertin P, Presterl T, Jamin P, Coubriche D, Gouesnard B et al (2017) Association mapping for phenology and plant architecture in maize shows higher power for developmental traits compared with growth influenced traits. Heredity 118(3):249–259
Brown PJ, Upadyayula N, Mahone GS, Tian F, Bradbury PJ, Myles S et al (2011) Distinct genetic architectures for male and female inflorescence traits of maize. PLoS Genet 7(11):e1002383
Cao S, Loladze A, Yuan Y, Wu Y, Zhang A, Chen J et al (2017) Genome-wide analysis of tar apot complex resistance in maize using genotyping-by-sequencing SNPs and whole-genome prediction. Plant Genome 10(2):1–14
Chen ZL, Wang BB, Dong XM, Liu H, Ren LH, Chen J et al (2014) An ultra-high density bin-map for rapid QTL mapping for tassel and ear architecture in a large F2 maize population. BMC Genom 15:1–10
David HT, Paul HH, Christopher RD (1988) Development and spread of improvement maize varities and hybrids in developing countries. Metrotec, Inc., Washinton, DC, pp 1–26
Ding X, Wu X, Lin C, Li C, Shi Y, Song Y et al (2017) Both major and minor QTL associated with plant height can be identified using near-isogenic lines in maize. Euphytica 213:21
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES et al (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6(5):e19379
Frascaroli E, Cane MA, Landi P, Pea G, Gianfranceschi L, Villa M et al (2007) Classical genetic and quantitative trait loci analyses of heterosis in a maize hybrid between two elite inbred lines. Genetics 176(1):625–644
Gong HJ, Zhu XY, Chen KM et al (2005) Silicon alleviates oxidative damage of wheat plants in pots under drought. Plant Sci 169:313–321
Hallauer A, Miranda J (1988) Quantitative genetics in maize breeding. Iowa State University Press, Ames. pp 1–26
Hill WG, Weir BS (1994) Maximum-likelihood estimation of gene location by linkage disequilibrium. Am J Hum Genet 54(4):705–714
Lai J, Li R, Xu X, Jin W, Xu M, Zhao H et al (2010) Genome-wide patterns of genetic variation among elite maize inbred lines. Nat Genet 42(11):1027–1030
Leipner J, Jompuk C, Camp KH, Stamp P, Fracheboud Y (2008) QTL studies reveal little relevance of chilling-related seedling traits for yield in maize. Theor Appl Genet 116(4):555–562
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25(14):1754–1760
Li H, Ribaut JM, Li Z, Wang J (2008) Inclusive composite interval mapping (ICIM) for digenic epistasis of quantitative traits in biparental populations. Theor Appl Genet 116(2):243–260
Liu R, Meng Q, Zheng F, Kong L, Yuan J, Lubberstedt T (2017) Genetic mapping of QTL for maize leaf width combining RIL and IF2 populations. PLoS ONE 12(12):e0189441
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al (2010) The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303
Nathan AB (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3(10):e3376
Pan Q, Xu Y, Li K, Peng Y, Zhan W, Li W et al (2017) The genetic basis of plant architecture in 10 maize recombinant inbred line populations. Plant Physiol 175(2):858–873
Peiffer JA, Romay MC, Gore MA, Flint-Garcia SA, Zhang Z, Millard MJ et al (2014) The genetic architecture of maize height. Genetics 196(4):1337–1356
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci 81(24):8014–8018
Salvi S, Corneti S, Bellotti M, Carraro N, Sanguineti MC, Castelletti S, Tuberosa R (2011) Genetic dissection of maize phenology using an intraspecific introgression library. BMC Plant Biol. https://doi.org/10.1186/1471-2229-11-4
Teng F, Zhai L, Liu R, Bai W, Wang L, Huo D et al (2013) ZmGA3ox2, a candidate gene for a major QTL, qPH3.1, for plant height in maize. Plant J 73(3):405–416
Tian F, Bradbury PJ, Brown PJ, Hung H, Sun Q, Flint-Garcia S et al (2011) Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat Genet 43(2):159-U113
Upadyayula N, da Silva HS, Bohn MO, Rocheford TR (2006) Genetic and QTL analysis of maize tassel and ear inflorescence architecture. Theor Appl Genet 112(4):592–606
van Heerwaarden J, Hufford MB, Ross-Ibarra J (2012) Historical genomics of North American maize. Proc Natl Acad Sci 109(31):12420–12425
Wu X, Li Y, Li X, Li C, Shi Y, Song Y et al (2015) Analysis of genetic differentiation and genomic variation to reveal potential regions of importance during maize improvement. BMC Plant Biol 15:256
Wu X, Li Y, Shi Y, Song Y, Zhang D, Li C et al (2016) Joint-linkage mapping and GWAS reveal extensive genetic loci that regulate male inflorescence size in maize. Plant Biotechnol J 14(7):1551–1562
Yan J, Shah T, Warburton ML, Buckler ES, McMullen MD, Crouch J (2009) Genetic characterization and linkage disequilibrium estimation of a global maize collection using SNP markers. PLoS ONE 4(12):e8451
Yang X, Xu Y, Shah T, Li H, Han Z, Li J et al (2011) Comparison of SSRs and SNPs in assessment of genetic relatedness in maize. Genetica 139(8):1045–1054
Zhang L, Li H, Wang J (2012) The statistical power of inclusive composite interval mapping in detecting digenic epistasis showing common F2 segregation ratios. J Integr Plant Biol 54(4):270–279
Acknowledgements
This research was supported by the National Key Research and Development Program of China (2018YFD0100104), the National Natural Science Foundation of China (31760387), the Guizhou Academy of Agricultural Science Innovation Program ([2014] 006), the Guizhou Natural Science Foundation ([2017]1413), the Guizhou Major Special Projects ([2013]6022), the Guizhou Science and Technology Support Program ([2017]2507, [2018]2296, and [2017] 2504-1), the Qiankehe Talent Platform ([2018]5629), and government subsidies to local platform construction projects (QKZYD[2018]4003).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
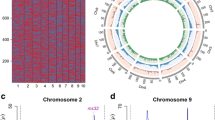
Fig. S1
Distribution of SNPs across the genome. Light color indicates a low density of markers, and deep color indicates a high density of markers. (DOCX 871 kb)
Rights and permissions
About this article
Cite this article
Wu, X., Guo, X., Wang, A. et al. Quantitative trait loci mapping of plant architecture-related traits using the high-throughput genotyping by sequencing method. Euphytica 215, 212 (2019). https://doi.org/10.1007/s10681-019-2535-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2535-x