Abstract
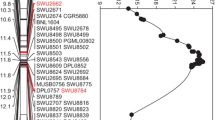
Upland cotton is an important economic crop that produces high-quality fiber for the textile industry. With the development of next-generation sequencing technology and improvements in human living standards, it has become possible to improve the fiber quality and yield of cotton with high-throughput molecular markers. Upland cotton 901-001 is an excellent, high-quality, non-transgenic cultivar, while the sGK156 strain shows high resistance to verticillium wilt. The phenotype of F1 plants, certified in 2008 as national variety CCRI70, shows positive transgressive characteristics such as high quality, high yield, and resistance to verticillium wilt. We developed a population of 250 recombination inbred lines from a cross between 901-001 and sGK156. The fiber strength trait of plants from nine environments was collected, and a genetic linkage map of Chr24 comprising 168 SNP marker loci covering a genetic distance of 107.46 cM and with an average distance of 0.64 cM was generated. QTLs were identified across the nine environments using the composite interval mapping method. A total of eight QTLs for FS were identified on Chr24, three of which were stably expressed in at least five environments. Some candidate genes located in qFS-c24-2 and qFS-c24-4 were functionally annotated as potentially playing important roles in fiber development, with homologous genes reported in Arabidopsis thaliana. These results suggest that QTLs identified in the present study could contribute to improving FS and may be applicable for marker-assisted selection.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/s0022-2836(05)80360-2
Ascencio-Ibanez JT, Sozzani R, Lee TJ, Chu TM, Wolfinger RD, Cella R, Hanley-Bowdoin L (2008) Global analysis of Arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection. Plant Physiol 148(1):436–454. https://doi.org/10.1104/pp.108.121038
Cai C, Zhu G, Zhang T, Guo W (2017) High-density 80K SNP array is a powerful tool for genotyping G. hirsutum accessions and genome analysis. BMC Genomics 18(1):654. https://doi.org/10.1186/s12864-017-4062-2
Cao Z, Zhu X, Chen H, Zhang T (2015) Fine mapping of clustered quantitative trait loci for fiber quality on chromosome 7 using a Gossypium barbadense introgressed line. Mol Breed 35(11):215. https://doi.org/10.1007/s11032-015-0393-3
Chen H, Qian N, Guo W, Song Q, Li B, Deng F, Dong C, Zhang T (2009a) Using three overlapped RILs to dissect genetically clustered QTL for fiber strength on Chro. D8 in Upland cotton. Theor Appl Genet 119(4):605–612. https://doi.org/10.1007/s00122-009-1070-x
Chen H, Qian N, Guo W, Song Q, Li B, Deng F, Dong C, Zhang T (2009b) Using three overlapped RILs to dissect genetically clustered QTL for fiber strength on Chro.D8 in Upland cotton: TAG. Theor Appl Genet (Theoretische und angewandte Genetik) 119(4):605–612. https://doi.org/10.1007/s00122-009-1070-x
Chothia C, Gough J, Vogel C, Teichmann SA (2003) Evolution of the protein repertoire. Science 300(5626):1701–1703. https://doi.org/10.1126/science.1085371
Fulton DC, Stettler M, Mettler T, Vaughan CK, Li J, Francisco P, Gil M, Reinhold H, Eicke S, Messerli G, Dorken G, Halliday K, Smith AM, Smith SM, Zeeman SC (2008) β-AMYLASE4, a noncatalytic protein required for starch breakdown, acts upstream of three active β-amylases in Arabidopsis Chloroplasts. Plant Cell 20(4):1040
Gardiner J, Overall R, Marc J (2011) Putative Arabidopsis homologues of metazoan coiled-coil cytoskeletal proteins. Cell Biol Int 35(8):767–774. https://doi.org/10.1042/cbi20100719
Hölscher C, Lutterbey M-C, Lansing H, Meyer T, Fischer K, von Schaewen A (2016) Defects in peroxisomal 6-phosphogluconate dehydrogenase isoform PGD2 prevent gametophytic interaction in Arabidopsis thaliana. Plant Physiol 171(1):192–205
Huang X, Zhao Y, Wei X, Li C, Wang A, Zhao Q, Li W, Guo Y, Deng L, Zhu C, Fan D, Lu Y, Weng Q, Liu K, Zhou T, Jing Y, Si L, Dong G, Huang T, Lu T, Feng Q, Qian Q, Li J, Han B (2011) Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat Genet 44(1):32–39. https://doi.org/10.1038/ng.1018
Islam MS, Zeng L, Delhom CD, Song X, Kim HJ, Li P, Fang DD (2014) Identification of cotton fiber quality quantitative trait loci using intraspecific crosses derived from two near-isogenic lines differing in fiber bundle strength. Mol Breed 34(2):373–384
Jamshed M, Jia F, Gong J, Palanga KK, Shi Y, Li J, Shang H, Liu A, Chen T, Zhang Z (2016) Identification of stable quantitative trait loci (QTLs) for fiber quality traits across multiple environments in Gossypium hirsutum recombinant inbred line population. BMC Genomics 17(1):197
John ME (1995) Characterization of a cotton (Gossypium hirsutum L.) fiber mRNA (Fb-B6). Plant Physiol 107(4):1477–1478
Kosambi DD (2016) The estimation of map distances from recombination values. In: Ramaswamy R (ed) D.D. Kosambi: selected works in mathematics and statistics. Springer India, New Delhi, pp 125–130. https://doi.org/10.1007/978-81-322-3676-4_16
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):R25. https://doi.org/10.1186/gb-2009-10-3-r25
Li H, Ye G, Wang J (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175(1):361–374. https://doi.org/10.1534/genetics.106.066811
Li W, Shang H, Ge Q, Zou C, Cai J, Wang D, Fan S, Zhang Z, Deng X, Tan Y, Song W, Li P, Koffi PK, Jamshed M, Lu Q, Gong W, Li J, Shi Y, Chen T, Gong J, Liu A, Yuan Y (2016) Genome-wide identification, phylogeny, and expression analysis of pectin methylesterases reveal their major role in cotton fiber development. BMC Genomics 17:1000. https://doi.org/10.1186/s12864-016-3365-z
Liu Q, Talbot M, Llewellyn DJ (2013) Pectin methylesterase and pectin remodelling differ in the fibre walls of two gossypium species with very different fibre properties. PLoS ONE 8(6):e65131. https://doi.org/10.1371/journal.pone.0065131
Liu D, Ma C, Hong W, Huang L, Liu M, Liu H, Zeng H, Deng D, Xin H, Song J, Xu C, Sun X, Hou X, Wang X, Zheng H (2014) Construction and analysis of high-density linkage map using high-throughput sequencing data. PLoS ONE 9(6):e98855. https://doi.org/10.1371/journal.pone.0098855
Liu D, Zhang J, Liu X, Wang W, Liu D, Teng Z, Fang X, Tan Z, Tang S, Yang J, Zhong J, Zhang Z (2016) Fine mapping and RNA-Seq unravels candidate genes for a major QTL controlling multiple fiber quality traits at the T1 region in upland cotton. BMC Genomics 17:295. https://doi.org/10.1186/s12864-016-2605-6
Ma X, Ding Y, Zhou B, Guo W, Lv Y, Zhu X, Zhang T (2008) QTL mapping in A-genome diploid Asiatic cotton and their congruence analysis with AD-genome tetraploid cotton in genus Gossypium. J Genet Genomics 35(12):751–762. https://doi.org/10.1016/S1673-8527(08)60231-3
Meng L, Li H, Zhang L, Wang J (2015) QTL IciMapping: integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J 3(3):269–283
Menges M, Hennig L, Gruissem W, Murray JAH (2002) Cell cycle-regulated gene expression in Arabidopsis. J Biol Chem 277(44):41987–42002. https://doi.org/10.1074/jbc.M207570200
Ohno S, Wolf U, Atkin NB (1968) Evolution from fish to mammals by gene duplication. Hereditas 59(1):169–187
Percival AE (1999) Taxonomy and germplasm resources. In: Smith WC, Cothren JT (eds) Cotton; origin, history, technology, and production. Wiley, New York, pp 33–63
Rong J, Abbey C, Bowers JE, Brubaker CL, Chang C, Chee PW, Delmonte TA, Ding X, Garza JJ, Marler BS, C-h Park, Pierce GJ, Rainey KM, Rastogi VK, Schulze SR, Trolinder NL, Wendel JF, Wilkins TA, Williams-Coplin TD, Wing RA, Wright RJ, Zhao X, Zhu L, Paterson AH (2004) A 3347-locus genetic recombination map of sequence-tagged sites reveals features of genome organization, transmission and evolution of cotton (Gossypium). Genetics 166(1):389–417. https://doi.org/10.1534/genetics.166.1.389
Rong J, Feltus FA, Waghmare VN, Pierce GJ, Chee PW, Draye X, Saranga Y, Wright RJ, Wilkins TA, May OL, Smith CW, Gannaway JR, Wendel JF, Paterson AH (2007) Meta-analysis of polyploid cotton QTL shows unequal contributions of subgenomes to a complex network of genes and gene clusters implicated in lint fiber development. Genetics 176(4):2577–2588. https://doi.org/10.1534/genetics.107.074518
Rotmistrovsky K, Jang W, Schuler GD (2004) A web server for performing electronic PCR. Nucleic Acids Res 32(Web Server issue):W108–W112. https://doi.org/10.1093/nar/gkh450
Said JI, Knapka JA, Song M, Zhang J (2015a) Cotton QTLdb: a cotton QTL database for QTL analysis, visualization, and comparison between Gossypium hirsutum and G. hirsutum × G. barbadense populations. Mol Genet Genomics 290(4):1615–1625. https://doi.org/10.1007/s00438-015-1021-y
Said JI, Song M, Wang H, Lin Z, Zhang X, Fang DD, Zhang J (2015b) A comparative meta-analysis of QTL between intraspecific Gossypium hirsutum and interspecific G. hirsutum × G. barbadense populations. Mol Genet Genomics MGG 290(3):1003–1025. https://doi.org/10.1007/s00438-014-0963-9
Sasidharan R, Chinnappa CC, Staal M, Elzenga JT, Yokoyama R, Nishitani K, Voesenek LA, Pierik R (2010) Light quality-mediated petiole elongation in Arabidopsis during shade avoidance involves cell wall modification by xyloglucan endotransglucosylase/hydrolases. Plant Physiol 154(2):978–990. https://doi.org/10.1104/pp.110.162057
Shappley ZW, Jenkins JN, Meredith WR, McCarty JC Jr (1998) An RFLP linkage map of Upland cotton, Gossypium hirsutum L. Theor Appl Genet 97(5):756–761. https://doi.org/10.1007/s001220050952
Shen X, Guo W, Lu Q, Zhu X, Yuan Y, Zhang T (2007) Genetic mapping of quantitative trait loci for fiber quality and yield trait by RIL approach in Upland cotton. Euphytica 155(3):371–380. https://doi.org/10.1007/s10681-006-9338-6
Sparla F, Costa A, Lo Schiavo F, Pupillo P, Trost P (2006) Redox regulation of a novel plastid-targeted β -Amylase of Arabidopsis. Plant Physiol 141(3):840
Takahashi N, Nakazawa M, Shibata K, Yokota T, Ishikawa A, Suzuki K, Kawashima M, Ichikawa T, Shimada H, Matsui M (2005) shk1-D, a dwarf Arabidopsis mutant caused by activation of the CYP72C1 gene, has altered brassinosteroid levels. Plant J Cell Mol Biol 42(1):13–22. https://doi.org/10.1111/j.1365-313X.2005.02357.x
Tan Z, Fang X, Tang S, Zhang J, Liu D, Teng Z, Li L, Ni H, Zheng F, Liu D, Zhang T, Paterson AH, Zhang Z (2015) Genetic map and QTL controlling fiber quality traits in upland cotton (Gossypium hirsutum L.). Euphytica 203(3):615–628. https://doi.org/10.1007/s10681-014-1288-9
Tang S, Teng Z, Zhai T, Fang X, Liu F, Liu D, Zhang J, Liu D, Wang S, Zhang K, Shao Q, Tan Z, Paterson AH, Zhang Z (2015) Construction of genetic map and QTL analysis of fiber quality traits for Upland cotton (Gossypium hirsutum L.). Euphytica 201(2):195–213. https://doi.org/10.1007/s10681-014-1189-y
Taylor DR, Ingvarsson PK (2003) Common features of segregation distortion in plants and animals. Genetica 117(1):27–35. https://doi.org/10.1023/A:1022308414864
Ulloa M, Meredith WR Jr, Shappley ZW, Kahler AL (2002) RFLP genetic linkage maps from four F2.3 populations and a joinmap of Gossypium hirsutum L. Theor Appl Genet 104(2):200–208. https://doi.org/10.1007/s001220100739
Ulloa M, Saha S, Jenkins JN, Meredith WR Jr, McCarty JC Jr, Stelly DM (2005) Chromosomal assignment of RFLP linkage groups harboring important QTLs on an intraspecific cotton (Gossypium hirsutum L.) Joinmap. J Hered 96(2):132–144. https://doi.org/10.1093/jhered/esi020
Van Ooijen JW (2006) JoinMap®4.0 software for the calculation of genetic linkage maps in experimental populations. Kyazma B.V., Wageningen, Netherlands
van Os H, Stam P, Visser RG, van Eck HJ (2005) SMOOTH: a statistical method for successful removal of genotyping errors from high-density genetic linkage data. TAG Theor Appl Genet (Theoretische und angewandte Genetik) 112(1):187–194. https://doi.org/10.1007/s00122-005-0124-y
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93(1):77–78
Wang S (2007) Windows QTL cartographer 2.5. http://statgen.ncsu.edu/qtlcart/WQTLCart.htm
Wang Y, Li J, Shi Y, Liu A, Shang H, Gong J, Wang T, Gong W, Yuan Y (2010) Molecular marker of QTL for fibre quality traits of upland cotton with elite fibre quality. Cotton Sci 22(6):533–538
Wang K, Wang Z, Li F, Ye W, Wang J, Song G, Yue Z, Cong L, Shang H, Zhu S, Zou C, Li Q, Yuan Y, Lu C, Wei H, Gou C, Zheng Z, Yin Y, Zhang X, Liu K, Wang B, Song C, Shi N, Kohel RJ, Percy RG, Yu JZ, Zhu YX, Wang J, Yu S (2012) The draft genome of a diploid cotton Gossypium raimondii. Nat Genet 44(10):1098–1103. https://doi.org/10.1038/ng.2371
Wang C, Ulloa M, Shi X, Yuan X, Saski C, Yu JZ, Roberts PA (2015) Sequence composition of BAC clones and SSR markers mapped to Upland cotton chromosomes 11 and 21 targeting resistance to soil-borne pathogens. Front Plant Sci 6:791. https://doi.org/10.3389/fpls.2015.00791
Willats WG, McCartney L, Mackie W, Knox JP (2001) Pectin: cell biology and prospects for functional analysis. Plant Mol Biol 47(1–2):9–27
Wu J, Mao X, Cai T, Luo J, Wei L (2006) KOBAS server: a web-based platform for automated annotation and pathway identification. Nucleic Acids Res 34(Web Server issue):W720–724. https://doi.org/10.1093/nar/gkl167
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39(Web Server issue):W316–322. https://doi.org/10.1093/nar/gkr483
Xu Y, Zhu L, Xiao J, Huang N, McCouch SR (1997) Chromosomal regions associated with segregation distortion of molecular markers in F2 , backcross, doubled haploid, and recombinant inbred populations in rice (Oryza sativa L.). Mol Gen Genet MGG 253(5):535–545. https://doi.org/10.1007/s004380050355
Yang X, Wang Y, Zhang G, Wang X, Wu L, Ke H, Liu H, Ma Z (2016) Detection and validation of one stable fiber strength QTL on c9 in tetraploid cotton. Mol Genet Genomics 291(4):1625–1638. https://doi.org/10.1007/s00438-016-1206-z
Zeng ZB (1994) Precision mapping of quantitative trait loci. Genetics 136(4):1457–1468
Zhang T, Yuan Y, Yu J, Guo W, Kohel RJ (2003) Molecular tagging of a major QTL for fiber strength in Upland cotton and its marker-assisted selection. TAG Theor Appl Genet (Theoretische und angewandte Genetik) 106(2):262–268. https://doi.org/10.1007/s00122-002-1101-3
Zhang Z-S, Hu M-C, Zhang J, Liu D-J, Zheng J, Zhang K, Wang W, Wan Q (2009) Construction of a comprehensive PCR-based marker linkage map and QTL mapping for fiber quality traits in upland cotton (Gossypium hirsutum L.). Mol Breed 24(1):49–61. https://doi.org/10.1007/s11032-009-9271-1
Zhang K, Zhang J, Ma J, Tang S, Liu D, Teng Z, Liu D, Zhang Z (2012) Genetic mapping and quantitative trait locus analysis of fiber quality traits using a three-parent composite population in upland cotton (Gossypium hirsutum L.). Mol Breed 29(2):335–348. https://doi.org/10.1007/s11032-011-9549-y
Zhang D, Hua Y, Wang X, Zhao H, Shi L, Xu F (2014) A high-density genetic map identifies a novel major QTL for boron efficiency in oilseed rape (Brassica napus L.). PLoS ONE 9(11):e112089. https://doi.org/10.1371/journal.pone.0112089
Zhang T, Hu Y, Jiang W, Fang L, Guan X, Chen J, Zhang J (2015a) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol. https://doi.org/10.1038/nbt.3207
Zhang T, Hu Y, Jiang W, Fang L, Guan X, Chen J, Zhang J, Saski CA, Scheffler BE, Stelly DM, Hulse-Kemp AM, Wan Q, Liu B, Liu C, Wang S, Pan M, Wang Y, Wang D, Ye W, Chang L, Zhang W, Song Q, Kirkbride RC, Chen X, Dennis E, Llewellyn DJ, Peterson DG, Thaxton P, Jones DC, Wang Q, Xu X, Zhang H, Wu H, Zhou L, Mei G, Chen S, Tian Y, Xiang D, Li X, Ding J, Zuo Q, Tao L, Liu Y, Li J, Lin Y, Hui Y, Cao Z, Cai C, Zhu X, Jiang Z, Zhou B, Guo W, Li R, Chen ZJ (2015b) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol 33(5):531–537. https://doi.org/10.1038/nbt.3207
Zhang Z, Li J, Muhammad J, Cai J, Jia F, Shi Y, Gong J, Shang H, Liu A, Chen T, Ge Q, Palanga KK, Lu Q, Deng X, Tan Y, Li W, Sun L, Gong W, Yuan Y (2015c) High resolution consensus mapping of quantitative trait loci for fiber strength, length and micronaire on Chromosome 25 of the Upland Cotton (Gossypium hirsutum L.). PLoS ONE 10(8):e0135430. https://doi.org/10.1371/journal.pone.0135430
Zhang Z, Shang H, Shi Y, Huang L, Li J, Ge Q, Gong J, Liu A, Chen T, Wang D, Wang Y, Palanga KK, Muhammad J, Li W, Lu Q, Deng X, Tan Y, Song W, Cai J, Li P, Rashid H, Gong W, Yuan Y (2016) Construction of a high-density genetic map by specific locus amplified fragment sequencing (SLAF-seq) and its application to Quantitative Trait Loci (QTL) analysis for boll weight in upland cotton (Gossypium hirsutum.). BMC Plant Biol 16:79. https://doi.org/10.1186/s12870-016-0741-4
Zhang Z, Ge Q, Liu A, Li J, Gong J, Shang H, Shi Y, Chen T, Wang Y, Palanga KK, Muhammad J, Lu Q, Deng X, Tan Y, Liu R, Zou X, Rashid H, Iqbal MS, Gong W, Yuan Y (2017) Construction of a high-density genetic map and its application to QTL identification for fiber strength in Upland Cotton. Crop Sci 57(2):774. https://doi.org/10.2135/cropsci2016.06.0544
Zhiyuan N, Chen H, Mei H, Zhang T (2014) Molecular tagging of QTLs for fiber quality and yield in the upland cotton cultivar Acala-Prema. Euphytica 195(1):143–156
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31471538, 31621005) and the National Agricultural Science and Technology Innovation project for CAAS, and Fundamental Research Funds for Central research institute (Y2017JC48). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
YLY and HHS designed the experiments and provided the resources for all experiments. XYZ performed the experiments, data analysis, and wrote the manuscript. JWG, XJ, and ZZ constructed the RIL population. LD, SMF, and QG investigated phenotypes of the RILs. AYL, WKG, JWL, and YZS prepared the materials and performed the DNA extractions. YLW, LQF, and RXL screened markers and constructed the linkage genetic map and analyzed collinearity. KL and QZ performed QTL identification and compared these results with previous studies. All authors have read, edited, and approved the current version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zou, X., Gong, J., Duan, L. et al. High-density genetic map construction and QTL mapping for fiber strength on Chr24 across multiple environments in a CCRI70 recombinant inbred lines population. Euphytica 214, 102 (2018). https://doi.org/10.1007/s10681-018-2177-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-018-2177-4