Abstract
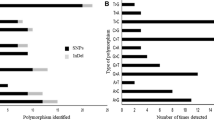
This study reports the implementation of three strategies for the development of genetic markers and their evaluation in both progenitors of an F2 population used for the construction of a genetic map of Coffea arabica. The strategies were Cleaved Amplified Polymorphic Sequences (CAPS), Single Strand Conformational Polymorphism (SSCP), and sequence analysis predicted Single Nucleotide Polymorphism (SNP). The methodologies were developed from different sequence sources: For CAPS, we used 25 COS sequences derived from Hedyotis spp. and 29 COSII sequences derived from Solanaceae and Rubiaceae species; for SSCP, we used 111 coffee EST sequences, 50 COSII sequences, and 10 C. arabica BAC end sequences. A low polymorphism was identified with the CAPS and SSCP methodologies. A total of 61 SNPs were identified in silico from 5,371 ESTs of coffee and from amplified, cloned, and sequenced COSII markers. Sixteen of these SNPs were validated with Luminex technology and 2 of them were polymorphic in C. arabica genotypes. This study highlights the difficulties of finding polymorphism in the species C. arabica where SNP identification seems to be the best strategy to search for polymorphic markers for this low diversity plant.



Similar content being viewed by others
References
Baruah A, Naik V, Hendre PS, Rajkumar R, Rajendrakumar P, Aggarwal RK (2003) Isolation and characterization of nine microsatellite markers from Coffea arabica L., showing wide cross-species amplifications. Mol Ecol Notes 3:647–650
Bernatzky R, Tanksley SD (1986) Toward a saturated linkage map in tomato based on isozymes and random cDNA sequences. Genetics 112(4):887–898
Bertin I, Zhu JH, Gale MD (2005) SSCP-SNP in pearl millet—a new marker system for comparative genetics. Theor Appl Genet 110:1467–1472
Carvalho A (1988) Principles and practices of coffea plant breeding for productivity and quality factors: Coffea arabica. In: Clarke RJ, Macrae R (eds) Coffea, vol 4. Agronomy. Elsevier Applied Science, London, pp 129–165
Chaparro AP, Cristancho M, Cortina H, Gaitán A (2004) Genetic variability of Coffea arabica L. accessions from Ethiopia evaluated with RAPDs. Genet Res Crop Evol 51:291–297
Charrier A, Berthaud J (1985) Botanical classification of coffee. In: Clifford MN, Willson KC (eds) Coffee: botany biochemistry and production of beans and beverage. Croom Helm Ltd, Sydney, pp 13–47
Ching A, Caldwell KS, Jung M, Dolan M, Smith OS, Tingey S, Morgante M, Rafalski Aj (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19
Diniz LEC, Ruas CF, Carvalho VP, Torres FM, Ruas EA, Santos MO, Sera T, Ruas PM (2005) Genetic diversity among forty coffee varieties assessed by RAPD markers associated with restriction digestion. Braz Arch Biol Technol 48:511–521
Ferwerda FP (1976) Coffee. In: Simpsons NW (ed) Evolution of crop plants. Longman, London, pp 257–260
Fujimura N, Yoshinobu K, Kazunori O, Masafumi Y, Hideki K (2002) Rapid multiplex single nucleotide polymorphism genotyping based on single base extension reactions and color-code beads. J Biosci Bioeng 4:368–370
Fulton RE, Salasek ML, DuTeau NM, WC Black IV (2001) SSCP analysis of cDNA markers provides a dense linkage map of the Aedesaegypti genome. Genetics 158:715–726
Fulton TM, Vand Der Hoeven R, Eannetta NT, Tanksley SD (2002) Identification, analysis and utilization of conserved ortholog set markers for comparative genomics in higher plants. Plant Cell 14:1457–1467
Hsu C, An C, Saha S, Ma D, Jenkins J, Scheffler B, Stelly D (2008) Molecular and SNP characterization of two genome specific transcription factor genes GhMyb8 and GhMyblO in cotton species. Euphytica 159:259–273
Huang X, Madan A (1999) CAP3: a DNA sequence assembly program. Genome Res 9:868–877
Humphries S, Gudnadson V, Whitall R, Day I (1997) Single strand conformation polymorphism analysis with high throughput modifications and its use in mutation detection in familial hypercholesterolemia. Clin Chem 43:427–435
IDEAS Iniciativas de Economía Alternativa y Solidaria, Observatorio de Corporaciones Transnacionales (2006) El mercado internacional del café. Boletín 14. Córdoba, Spain. 48p. (http://www.ideas.coop/acerca-de-ideas/memorias-y-documentos/doc_details/13-el-mercado-internacional-del-cafe-actualizado.HTML)
Keller I, Bensasson D, Nichols RA (2007) Transition-transversion bias is not universal: a counter example from grasshopper pseudogenes. PLoS Genet 3(2):e22
Kent JW (2002) BLAT—the BLAST-like alignment tool. Genome Res 12(4):656–664
Konieczny A, Ausubel MF (1993) A procedure for mapping Arabidopsis mutations using co-dominant ecotype-specific PCR based markers. Plant J 4(2):403–410
Krug CA, Carvalho A (1951) The genetics of coffee. Adv Genet 4:127–158
Kunihisa M, Fukino N, Matsumoto S (2003) Development of cleavage amplified polymorphic sequence (CAPS) markers for identification of strawberry cultivars. Euphytica 134:209–215
Lai Z, Livingstone K, Zou Y, Church SA, Knapp SJ, Andrews J, Rieseberg LH (2005) Identification and mapping of SNPs from ESTs in sunflower. Theor Appl Genet 111(8):1532–1544
Lashermes F, Combes M C, Cros J, Trouslot P, Anthony F, Charrier A (1995) Origin and genetic diversity of Coffea arabica L. based on DNA molecular markers. 16th conference of ASIC, Kyoto, Japan
Lashermes P, Trouslot P, Anthony F, Combes MC, Charrier A (1996) Genetic diversity for RAPD markers between cultivated and wild accessions of Coffea arabica. Euphytica 87:59–64
Lashermes P, Combes MC, Robert J, Trouslot P, D’hont A, Anthony F, Charrier A (1999) Molecular characterization and origin of the Coffea arabica L. genome. Mol Gen Genet 26:259–266
Le Dantec L, Chagne D, Pot D, Cantin O, Garnier-Gere P, Bedon F, Frigerio J-M, Chaumeil P, Leger P, Garcia V, Laigret F, de Daruvar A, Plomion C (2004) Automated SNP detection in expressed sequence tags: statistical considerations and application to maritime pine sequences. Plant Mol Biol 54:461–470
Li WH, Graur D (1991) Fundamentals of molecular evolution. Sinauer Associates, Sunderland, MA, 284p
Loïck DL, Chagné D, Pot D, Cantin O, Garnier-Géré P, Bedon F, Frigerio JM, Chaumeil P, Léger P, Garcia V, Laigret F, Daruvar A, Plomion C (2004) Automated SNP detection in expressed sequence tags: statistical considerations and application to maritime pine sequences. Plant Mol Biol 54:461–470
Lopez C, Piegu B, Cooke R, Delseny M, Tohme J, Verdier V (2005) Using cDNA and genomic sequences as tools to develop SNP strategies in cassava (ManihotesculentaCrantz). Theor Appl Genet 110:425–431
Marth GT, Korf I, Yandell MD, Yeh RT, Gu Z (1999) A general approach to single-nucleotide polymorphism discovery. Nat Genet 23:452–456
Minamiyama Y, Kinoshita S, Inaba K, Inoue M (2005) Development of a cleaved amplified polymorphic sequence (CAPS) marker linked to pungency in pepper. Plant Breed 124:288–291
Möhring S, Horstmann V, Esch E (2005) Development of molecular CAPS marker for the self-incompatibility locus in Brassicanapus and identification of different S alleles. Plant Breed 124:105–110
Moncada P, McCouch S (2004) Simple sequence repeat diversity in diploid and tetraploid Coffea species. Genome 47:501–509
Orita M, Iwahana H, Kanazana H, Hayashi K, Sekiya T (1989) Detection of polymorphisms of human DNA by electrophoresis as single strand conformation polymorphisms. Proc Natl Acad Sci 86:2766–2770
Picoult-Newberg L, Ideker L, Pohl TE, Taylor MG, Donaldson SL, Nickerson MA, Boyce-Jacino M (1999) Mining SNPs from EST databases. Genome Res 9:167–174
Plomion C, Hurme P, Frigerio JM, Ridolfi M, Pot D, Pionneau C, Avila C, Gallardo F, David H, Neutelings G, Campbell M, Canovas FM, Savolainen O, Bodénès C, Kremer A (1999) Developing SSCP markers in two Pinus species. Mol Breed 5:21–31
Ragoussis J, Elvidge GP, Kaur K, Colella S (2006) Matrix-assisted laser desorption/ionisation, time of flight mass spectrometry in genomics research. PLoS Genet 2(7):e100
Ruas PM, Ruas CF, Rampim L, Carvalho VP, Ruas EA, Sera T (2003) Genetic relationship in Coffea species and parentage determination of interspecific hybrids using ISSR (inter-simple sequence repeat) markers. Genet Mol Biol 26:319–327
Sato Y, Nishio T (2003) Mutation detection in rice waxy mutants by PCR-RF-SSCP. Theor Appl Genet 107(3):560–567
Schneider K, Kulosa D, Soerensen TR, Möhring S, Heine M, Durstewitz G, Polley A, Weber E, Jamsari LJ, Hohmann U, Tahiro E, Weisshaar B, Schulz B, Koch G, Jung C, Ganal M (2007) Analysis of DNA polymorphisms in sugar beet (Beta vulgaris L.) and development of an SNP-based map of expressed genes. Theor Appl Genet 115:601–615
Snelling WM, Casas E, Stone RT, Keele JW, Harhay GP, Bennett GL, Smith TP (2005) Linkage mapping bovine EST-based SNP. BMC Genomics 6:74
Somers DJ, Kirkpatrick R, Moniwa M, Walsh A (2003) Mining single-nucleotide polymorphisms from hexaploid wheat ESTs. Genome 49:431–437
Taylor J, Provart NJ (2006) CapsID: a web-based tool for developing parsimonious sets of CAPS molecular markers for genotyping. BMC Genet 7:27
Tsui C, Coleman LE, Griffith JL, Bennett AE, Goodson SG, Scott JD, Pittard SW, Devine SE (2003) Single nucleotide polymorphisms (SNPs) that map to gaps in the human SNP map. Nucleic Acids Res 31(16):4910–4916
Van der Vossen HAM (1985) Coffee selection and breeding. In: Clifford MN, Willson KC (eds) Coffee: botany, biochemistry and production of beans and beverage. Croom Helm, London, pp 48–96
Wrigley G (1995) Coffee-coffea spp. (Rubiaceae). In: Smartt J, Simmonds NW (eds) Evolution of crop plants. Longman Scientific and Technical, Harlow, England, pp 438–443
Wu F, Mueller LA, Crouzillat D, Pétiard V, Tanksley SD (2006) Combining bioinformatics and phylogenetics to identify large sets of single-copy orthologous genes (COSII) for comparative, evolutionarity and systematic studies: a test case in the Euasterid plant clade. Genetics 174:1407–1420
Zhu YL, Song QJ, Hyten DL, Tassell Van CP, Matukumalli LK, Grimm DR, Hyatt SM, Fickus EW, Young ND, Cregan PB (2003) Single-nucleotide polymorphisms in soybean. Genetics 163:1123–1134
Acknowledgments
The authors gratefully acknowledge The Ministry of Agriculture and Rural Development of Colombia for financial support. We also thank Luis Fernando Rivera for sequence analysis and bioinformatics support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zarate, LA., Cristancho, MA. & Moncada, P. Strategies to develop polymorphic markers for Coffea arabica L.. Euphytica 173, 243–253 (2010). https://doi.org/10.1007/s10681-009-0102-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-009-0102-6