Abstract
Soft rot Pectobacteriaceae (SRP) are the causative agents of soft rot and blackleg in potato. In this study, we investigated if potato seed lots of the same cultivar, but originating from different fields, inoculated with the same density of SRP and planted in the same field, showed differences in blackleg disease incidence. We tested if these differences were correlated with the microbial community composition in tuber, and the soil where the mother tubers were grown, as the microbiome is known to play a large role in plant disease resistance. We found that tubers from seed lots with a high disease incidence had a different microbial community composition than tubers from seed lots with a low disease incidence. Several taxa could be identified that were on average more abundant in seed lots with a low disease incidence. However, the taxa that differed in abundance were different between the two growing seasons. Most of the taxa that differed in abundance between seed lots with high and low disease incidence were also present in the soil of the fields from which the tubers originated. However, the taxa did not differ in abundance between the different fields. This raises the question as to how these taxa are recruited by the tuber.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Worldwide, soft rot Pectobacteriaceae (SRP) are major pathogens affecting potato (Czajkowski et al., 2011; Toth et al., 2011). By degrading the tuber cell wall, they cause tissue maceration in the tuber (soft rot) and in the stems (blackleg). In the Netherlands, the SRP species Dickeya solani and Pectobacterium brasiliense have emerged as the dominant causative agents of blackleg (van der Wolf et al., 2021a). Spread of the pathogen occurs mainly via seed tubers. Latently infected mother tubers can result in blackleg inflicted plants and infection of the daughter tubers, which can cause high disease incidences in later field generations (Pérombelon, 1992). In seed potato production even low disease incidences can lead to a rejection of a seed potato lot (Dupuis et al., 2021). There are currently no effective treatments against SRP. Control measures include the planting of disease-free minitubers from sterile culture, the testing of seed lots for the presence of SRP, and dry storage conditions (Czajkowski et al., 2011; Toth et al., 2011). In addition, some potato varieties appear to be less susceptible to SRP than others, although there are no varieties that are completely resistant. For all known SRP species, fundamental knowledge about origin, epidemiology and control measures are still scarce.
Since there are no direct control measures effective against SRP, such as chemical compounds, increasing attention is being paid to biological factors that influence disease resistance. A preliminary study found differences in disease incidence between different potato tuber lots from the same variety inoculated with the same density of SRP and planted in the same soil (unpublished results). Tubers from the same variety are genetically identical and therefore differences in resistance to blackleg between seed lots are not expected. It is still unknown what causes these differences, but it is likely that the soil at the location the tubers were grown plays a significant role in later disease suppressiveness. Both abiotic and biotic soil factors can alter plant growth and resistance to different kinds of stress (Amtmann et al., 2008; Choudhary et al., 2016). But especially biotic factors, have been demonstrated to influence plant growth and resistance.
The role of the microbiome in mitigating plant disease has been frequently reported. The soil-, rhizosphere-, and endophytic microbiome are known for containing taxa with potential beneficial effects on plant resistance (Berendsen et al., 2012). This protective effect can be caused by two main mechanisms: direct interactions between beneficial microbes and pathogens and induction of resistance in the plant. Direct interactions between microbes are for example competition for space and nutrients, direct antibiosis and disruption of quorum sensing (QS) communication (Compant et al., 2005). Induced resistance can be mediated via induced systemic resistance (ISR) (Kou et al., 2016) or systemic acquired resistance (SAR). ISR is triggered by beneficial microbes in either the rhizosphere or the endophytic compartment, and induces a primed state in the plant, leading to a faster and stronger defence response upon pathogen attack (Pieterse et al., 2014). It is mediated by jasmonate and ethylene. In contrast, SAR is defined as systemic resistance in response to a local pathogen infection and is mediated by salicylic acid. Both direct interactions and resistance induction have been found to play a role in SRP control. Liu et al. (2020b) obtained a collection of endophytic isolates from potato, several of which were able to produce at least one antibiotic compound and Padilla-Gálvez et al., (2021) reported a Streptomyces strain from potato with QS inhibition activity effective against SRP. An in vitro study found increased resistance against D. solani after treatment with salicylic acid (Czajkowski et al., 2015). In addition, bacteria isolated from rotting tubers that were shown to provide protection against D. solani under greenhouse conditions (Czajkowski et al., 2012).
The plant-associated microbiome has been found to be dependent on the soil microbial community as the major source of plant associated bacteria (Morales-Cedeño et al., 2021) (Compant et al., 2012). Consequentially, in potato, the endophytic microbial community of the mother tuber partly originates from the soil in which it was propagated (Buchholz et al., 2019) and therefore this soil may have an influence on pathogen resistance. However, it is still poorly understood how the soil in which the tubers were grown in the previous generation, affects the endophytic microbiome in potato. Moreover, the subsequent effect on resistance in the field is unknown.
The aim of the present study was to investigate possible causes of differences in disease resistance against blackleg disease between genetically identical potato lots grown in different soils. We tested if differences in disease incidences, found for tuber lots from the same cultivar but collected from different field crops, could be correlated with differences in the tuber microbiome. In addition, we assessed if these differentially abundant taxa might originate from the field soil of origin.
Material and methods
For a summary of the workflow see Fig. S1.
Field experiment set-up 2018
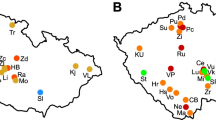
In 2018, 384 cultivar Kondor tubers each from 14 different seed lots, originating from different fields throughout the Netherlands in 2017 (Table S1), were used. We define a seed lot as a batch of seed potatoes from the same cultivar (thus with identical genetic background) grown in the same field. The cultivar Kondor was used as it is known to be susceptible to SRP (van der Wolf et al., 2021b). The 14 seed lots were vacuum inoculated with D. solani (IPO2222) or P. brasiliense (IPO3649) at a concentration of 106 cells/ml at the Dutch seed potato trading company HZPC (Metslawier, The Netherlands) (Fig. 1a). This concentration is known to lead to a high disease incidence (van der Wolf et al., 2017), which is necessary to assess potential differences between seed lots. As a control, tubers from the same seed lots were inoculated with only demineralized water. After inoculation, 10 tubers from 3 lots were used to assess inoculation efficiency. Two pieces of potato peel from each tuber were transferred to an extraction bag with a synthetic intermediate layer (Bioreba AG, Switzerland) with the addition of Ringer’s ¼ strength solution in a 1:1 w/v ratio and crushed with a mallet. Fifty µl of the undiluted, 10 × or 100 × diluted extracts were plated on double-layer crystal violet pectate (DL-CVP) medium (first layer: 5.5 g/l CaCl2 × 2H2O, 1 g/l tryptone, 1.5 ml 0.1% Crystal violet, 1.6 g/l NaNO3, 15 g/l agar, 1 ml cycloheximide (200 mg/ml stock); second layer: 5.5 ml EDTA (5.5%, pH = 8), 6 ml NaOH 5 M, 25 g dipecta pectin) (Hélias et al., 2012). Cavity forming colonies were counted.
For assessment of disease incidence per lot, 4 replicates of 16 tubers were planted in both a sandy soil in Driezum (The Netherlands) and a clay soil in Munnekezijl (The Netherlands). In addition, to examine microbiome composition, from each lot 16 tubers were sampled. From each tuber, one third was removed from the stolon-end and a thick peel was taken, finely chopped, flash-frozen in liquid nitrogen and stored at -80 °C for later analysis of the microbiome.
Field experiment set-up 2019
In 2019, 576 tubers per lot of 20 lots of cultivar Kondor harvested in 2018 (Table S1) were vacuum inoculated with D. solani or P. brasiliense at a concentration of 106 cells/ml and planted in both a sandy soil at Kolummerzwaag (The Netherlands) and in a clay soil in Munnekezijl (The Netherlands) in four replicates of 24 tubers per lot (Fig. 1b). In addition, 24 tubers per lot were sampled and frozen for microbiome analysis as in 2018. Like in 2018, 10 tubers from 3 lots were tested for inoculation efficiency. It needs to be noted that each lot is unique and that lots with the same number from 2018 and 2019 respectively are independent from each other.
Disease assessment
Tubers were planted at the end of April in a randomized design and disease-incidence was assessed once per week until mid-June separately for whole and cut tubers. Based on these results, each year three lots of cultivar Kondor with a high disease incidence (in 2018: K9-18, K13-18, K14-18; in 2019: K18-19, K19-19, K20-19) and three with a low disease incidence (in 2018: K6-18, K8-18, K10-18; in 2019: K13-19, K14-19, K17-19) were selected for further analysis (Table 1).
Soil sampling
In 2018, directly after harvesting of the seed lots that were used in 2019 for the field experiment, soil samples were taken from each field. The fields were separated into eight blocks. From each block, twenty samples were taken randomly at a depth of 5–25 cm, pooled and frozen at -80 °C. For each of the six selected seed lots this yielded 8 soil samples and therefore 46 soil samples in the year 2018. From each soil sample, 250 mg frozen soil was extracted using the MagAttract PowerSoil DNA KF kit (Qiagen) according to manufacturer’s instructions.
Sequencing of the tuber and soil microbiome
For each of the six selected lots, ten tuber peel samples that had been frozen previously were analysed. The frozen samples were transferred to 15 ml bead beating tubes filled each with two ceramic beads (2.8 mm) and two metal beads (2.8 mm) (Precellys, Bertin, Montigny-le-Bretonneux, France). Each sample was beaten 2–3 times for 10 s at 6000 rpm and 0 °C in a Precellys Evolution bead beater with a Cryolys cooling system. Between rounds, samples were cooled in liquid nitrogen for 1–2 s to prevent thawing. Then 250 mg of the resulting frozen powder was used for DNA extraction with the MagAttract PowerSoil DNA KF kit (Qiagen) according to manufacturer’s instructions.
Amplification of the bacterial 16S rDNA region from tuber and soil DNA was carried out using the primers MSAf-B-515f (5’-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGGTGCCAGCMGCCGCGGTAA-3’) and MSAr-B-806r (5’-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGGACTACHVGGGTWTCTAAT-3’) including MiSeq-adapters, a pPNA clamp (5- GGCTCAACCCTGGACAG-3’) and a mPNA clamp (5’- GGCAAGTGTTCTTCGGA-3’) (Lundberg et al., 2013). The following protocol was used: 5.75 µl water, 1 µl dNTPs (5 mM), 5 µl 5xQ5 reaction buffer, 1.25 µl of each primer, 4 µl each of each PNA, 0.25 µl Q5 HF DNA polymerase and 2.5 µl DNA resulting in a final volume of 25 µl. The samples were amplified with a starting temperature of 98 °C for 30 s, followed by 30 cycles of 98 °C for 10 s, 75 °C for 10 s, 50 °C for 30 s, 72 °C for 30 s and a final 2 min at 72 °C.
The amplification of the ITS region from tuber and soil DNA was carried out using the primers MSAf-F-gITS7 (5’- TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGGTGARTCATCGARTCTTTG-3’) and MSAr-F-ITS4 (5’- GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGTCCTCCGCTTATTGATATGC-3’) using the following protocol: 16 µl water, 1 µl dNTPs (5 mM), 5 µl 5xQ5 reaction buffer, 0.125 µl of each primer, 0.25 µl Q5 HF DNA polymerase and 2.5 µl DNA. The samples were amplified with a starting temperature of 98 °C for 30 s, followed by 30 cycles of 10 s of 98 °C, 30 s of 50 °C, 30 s of 72 °C and a final 2 min at 72 °C. Illumina MiSeq sequencing was carried out at the Bioscience group of WUR in 2018 and at Baseclear (Leiden, The Netherlands) in 2019, with 2 × 250nt paired end reading. All sequencing data has been deposited at the NCBI sequence read archive (SRA) under BioProject accession number PRJNA903099.
Microbiome data analysis
All analyses were done in R version 3.6.1.
Filtering, error removal, dereplication, merging of paired end reads and chimera removal were done using the package DADA2 (Callahan et al., 2016). Data from 2018 and 2019 were merged after chimera detection and removal. For taxonomic identification of 16S rDNA sequences, the silva train set version 128 was used and for fungal ITS sequences the sh general release dataset from 02.02.2019 was used. Operational taxonomic unit (OTU) contingency tables, sample data and taxonomic trees were stored in phyloseq objects (McMurdie and Holmes, 2013).
Mitochondrial and chloroplast sequences were removed from the 16S rDNA data. Subsequently, OTUs that showed an overall abundance below twenty were removed and the datasets were split into tuber and soil data. Then OTUs that were present in less than 1% of all samples were also removed from the tuber dataset and OTUs that were present in less than 5% of all samples were removed from the soil dataset. This was due to the higher amount of OTUs found in soil samples, which had to be reduced for computational reasons. Sample counts were transformed to relative abundance and the package microeco. was used for calculating the relative abundance of phyla in the different lots (Liu et al., 2020a). For comparison of tuber and soil microbiome composition, the datasets were not split and OTUs with a presence in less than 5% of all samples were removed.
Non-metric multidimensional scaling (NMDS) was applied using the built-in ordinate function with weighted unifrac dissimilarity in order to reduce the dimensions of the microbiome community data to only two dimensions. The vegan package (Oksanen et al., 2007) was used to assess differences in community composition between different lots and lots with high or low disease incidence by means of a permutational multivariate analysis of variance using the adonis function. In addition, the package DESeq2 (Love et al., 2014) was used to detect OTUs with differing abundances between seed lots with a high and a low disease incidence. For 16S rDNA data, filtered with an abundance threshold of 0.1, a random forest analysis was performed with incidence for classification. This analysis was carried out with the package randomForest (Liaw & Wiener, 2002).
Statistical analysis
Disease incidence in the field was analysed with a generalized linear model (binomial distribution), using the number of diseased plants per plot as the response variable and seed lot, bacterial species and location as the explanatory variable. This was followed by an analysis of variance (anova) of the model using the function Anova() from the package car (Fox & Weisberg, 2011). The effect of seed lot on disease incidence alone was analysed with a generalized linear mixed model (binomial distribution) with the number of diseased plants per plot as the response variable, the seed lot as the explanatory variable and bacterial species and location as random variables. The effect of bacterial strain on disease incidence was analysed in the same way. Pairwise differences were assessed with the emmeans function from the package emmeans (Lenth et al., 2018).
Results
Disease incidence in the field
Each lot is unique and lots with the same numbers from 2018 and 2019 respectively are independent from each other.
In 2018, disease incidence in sandy soil was on average 36.06% and thus significantly higher than in clay soil with an average of 23.33% (χ2 = 138.90, p < 0.01). In addition, disease incidence was significantly higher in plants inoculated with D. solani (46.40%) compared to P. brasiliense (40.34%; z-ratio = 3.85, p < 0.01) and to the water control (2.34%; z-ratio = 22.39, p < 0.01). Disease incidence with. P. brasiliense was also significantly different from the water control (z-ratio = 20.77, p < 0.01). There was also an interaction between seed lot, soil type and pathogen strain (χ2 = 50.87, p < 0.01), showing that disease incidence for each seed lot varied with soil type and pathogen strain (Fig. S2a). In spite of the high variation, significant differences in disease incidence between lots were detected (χ2 = 223.55, p < 0.01) (Fig. 2a). Selection of lots for microbiome analysis was subsequently based on disease incidence averaged over both soil types and pathogen strains. The lots K6-18, K8-18 and K10-18 with average disease incidences of 21.01%, 12.83% and 23.49% respectively, were selected as representatives of low disease incidence lots and the lots K9-18, K13-18 and K14-18 with disease incidences of 46.08%, 35.63% and 41.15% respectively, were selected as representatives of high disease incidence lots.
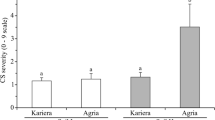
Bar chart of average disease incidence (%) in a) the 14 seed lots in 2018 after inoculation averaged over all bacterial treatments and over location and b) the 20 seed lots in 2019 after inoculation averaged over all bacterial treatments and over location. Error bars represent the standard error. Different letters represent statistically significant differences
In 2019, average disease incidence in sandy soil was again higher than in clay soil with 44.40% disease incidence compared to 26.58% (χ2 = 551.5, p < 0.01). There was no significant difference in disease incidence between plants infected with D. solani and P. brasiliense with 51.78% and 52.92% disease incidence respectively. Both pathogen treatments had a significantly higher disease incidence than the water control with 1.78% (z-ratio = 32.99, p < 0.01 for D. solani and z-ratio = 33.35, p < 0.01 for P. brasiliense). There were significant interactions between seed lot and pathogen strain (χ2 = 52.10, p < 0.01) (Fig. S2b). Still, disease incidence differed significantly between seed lots (χ2 = 201.20, p < 0.01) (Fig. 2b).
There was also a significant interaction between seed lot and soil type (χ2 = 302.90, p < 0.01). However, there were no significant differences in pairwise interactions, with disease incidence being higher in sandy soil for all lots.
The lots K13-19, K14-19 and K17-19 were chosen as representatives of seed lots with a low disease incidence. Average disease incidences in lot K13-19, K14-19 and K17-19 were 19.31%, 35.57% and 33.53% respectively. The lots K18-19, K19-19 and K20-19 with disease incidences of 42.20%, 46.51% and 38.21% respectively, were chosen as representatives of lots with a high disease incidence.
Tuber microbiome
After filtering, the dataset from 2018 and 2019 combined consisted of 4383 bacterial and 269 fungal OTUs.
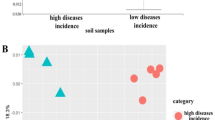
The NMDS showed substantial overlap between both the bacterial and fungal communities in tubers with a high and a low disease incidence (Fig. 3a, b). Still, a permanova analysis indicated significant differences between the two groups for both bacterial and fungal communities (bacteria: F = 7.88, p < 0.01, fungi: F = 3.69, p = 0.013). In addition, seed lots differed from each other (bacteria: F = 20.77, p < 0.01; fungi: F = 9.37, p < 0.01).
The two ordination axes of non-metric multidimensional scaling (NMDS) with weighted unifrac distances for a) bacterial and b) fungal communities analysed in the six selected seed lots per year with a high and low disease incidence (n = 12), with ten sampled tubers per lot; colour indicates disease incidence in the respective seed lot
For bacteria, on phylum level, a higher abundance of Proteobacteria and of Bacteroidetes was observed in tubers with a high disease incidence (Proteobacteria High: 22.1%, Low: 16.03%, Bacteroidetes: High: 4.01%, Low: 2.41%), whereas a higher abundance of Firmicutes and Actinobacteria was observed in tubers with a low disease incidence (Firmicutes: High: 7.42%, Low: 12.65%; Actinobacteria: High: 24.17%, Low = 27.45%) (Fig. 4). For fungi, Ascomycota was the most abundant phylum. No consistent differences could be observed between seed lots with a high and a low disease incidence.
The differential abundance analysis in DESeq2 showed that 310 bacterial OTUs differed in abundance between lots with a high and a low disease incidence. Of those, 60 OTUs, belonging to 47 genera, had a higher abundance in tubers with a low disease incidence. Among the most abundant were the genera Staphylococcus, Rhodococcus, Pseudomonas, Pantoea, Curtobacterium and Arthrobacter (Fig. 5a). However, several low abundance taxa showed the highest log2-fold change, such as Stenotrophomonas and Massilia (Fig, 5b). Arthrobacter was both abundant and showed a high log2-fold change. The random forest analysis also identified the genera Pantoea, Arthrobacter and Rhodococcus among the 30 OTUs that best explained the differentiation between low and high disease incidence lots (Fig. S3, Table S2).
Results of a differential abundance analysis with DESeq2 depicting genera containing OTUs with a significantly higher abundance in tubers of the selected seed lots with low disease incidence in comparison to selected seed lots with high disease incidence in 2018 and 2019; a) Relative abundance of bacterial genera in seed tuber lots with a low and high disease incidence; colour indicates disease incidence b) log2-fold change of bacterial genera in seed tuber lots with a low compared to a high disease incidence; colour indicates phylum, c) relative abundance of fungal genera with a significantly higher abundance in seed tuber lots with low and high disease incidence; colour indicates disease incidence and d) log2-fold changes of fungal genera in seed lots of low compared to high disease incidence; colour indicates phylum
Looking at the abundances of the respective OTUs in the individual seed lots showed that for example the genera Pantoea and Staphylococcus were especially abundant in only one or two of the seed lots with a low disease incidence. On the other hand, Rhodococcus, Pseudomonas and Arthrobacter were more abundant in several or even all seed lots with a low disease incidence (Figure S4).
For fungi, 41 OTUs differed in abundance between seed lots with a high and low disease incidence, of which 20 OTUs, belonging to 16 genera, had a higher abundance in tubers with a low disease incidence (Fig. 5c, d). Similar to bacterial taxa, most taxa were particularly highly abundant in only one of the seed lots. Only the genera Vishniacozyma, Pyrenochaeta and Acremonium were more abundant in several lots with a low disease incidence compared to a high disease incidence (Figure S5).
Soil microbiome
The filtered soil microbiome dataset sampled in 2018 from the soil of origin of tubers planted in 2019 included 3746 bacterial and 992 fungal OTUs. Separation between bacterial communities from soil with high and low disease incidences occurred mostly along the first axis of the NMDS, which was due to seed lot 13 and 14 differing strongly from all other seed lots (Fig. 6). Still, differences in communities between lots with a high and a low disease incidence were significant (F = 9.73, p < 0.01). For fungal communities, on the other hand, there were little differences on the first two axes of the NMDS, although communities differed significantly between high and low disease incidence according to permanova (F = 12.83, p < 0.01).
The two ordination axes of non-metric multidimensional scaling (NMDS) with weighted unifrac distances for a) bacterial and b) fungal communities in soil sampled in 2018, which were the soils of origin of the six selected seed lots in 2019 with a high and low disease incidence (n = 6), with eight samples per lot; colour indicates disease incidence in the respective seed lot
Similar to tubers, higher abundances of Bacteroidetes were found in soil samples with a high disease incidence (High: 8.54%, Low: 2.87%), while more Actinobacteria and Firmicutes were found with a low disease incidence (Actinobacteria: High:2.69%, Low:3.31%; Firmicutes: High: 0.45%, Low: 2.00%) (Fig. 7). For fungi, Ascomycota were again the most abundant phylum.
In total, 826 bacterial OTUs were significantly differentially abundant, 336 of which showed a higher abundance at low disease incidence. Of these 336 the 100 most abundant OTUs contained the genera Terrabacter, Sphingomonas, Candidatus Udaeobacter, Bacillus and Bradyrhizobium (Fig. 8a). In most cases, these genera were highly abundant in the seed lots 13 and 14, such as Candidatus Udaeobacter and Bacillus, while Bradyrhizobium had a higher abundance in all lots with a low disease incidence. For fungi, 277 OTUs were differentially abundant, 221 of which were more abundant at low disease incidence. The taxa that were more abundant at low disease incidence included the genera Vishniacozyma, Fusarium, Cladosporium, Trichoderma, Pyrenochaeta and Acremonium (Fig. 8b).
Results of differential abundance analysis with DESeq2 depicting genera containing OTUs with significantly higher abundance in soils from the selected seed lots with a low disease incidence in comparison to the selected seed lots with a high disease incidence; a) Relative abundance of bacterial genera in soils from lots with low and high disease incidences, colour indicates disease incidence b) log2-fold change of bacterial genera in soils from lots with a low compared to a high disease incidence; colour indicates phylum, c) relative abundance of fungal genera with a significantly higher abundance in soils from lots with low disease incidence; colour indicates disease incidence and d) log2-fold changes of fungal genera in soils from lots with low compared to high disease incidence; colour indicates phylum
Comparison between soil and tuber
For the comparison between tubers and soil, only tubers harvested in 2019 were included. The soil used was sampled in 2018 and represents the soil the tubers used in 2019 originated from. In total, 1639 OTUs were shared between tuber and soil for bacteria and of these 29 OTUs were significantly increased in both tubers and soil from seed lots with a low disease incidence compared to a high disease incidence (Table S3). Similar to the compiled results from tubers from both years, the taxa significantly more abundant in tubers included the genera Staphylococcus, and Pseudarthrobacter. However, also other taxa, such as the genus Glutamicibacter, was significantly more abundant in tubers with a low disease incidence. This genus was not detected as differentially abundant in the data of both years compiled. Subsequently, we retrieved the abundance of these taxa in the sequencing data from soil. Broadly, three scenarios were observed. 1) The taxon was more abundant in seed lots with a low disease incidence in both soil and tubers, 2) the taxon was observed in both soil and tubers, but only differentially abundant in tubers, 3) the taxon was observed mostly in tubers. In the first scenario for example, an unidentified taxon of the family Bacillaceae and the genus Sphingomonas were significantly more abundant in both tuber and soil in lots with a low disease incidence (Fig. S6a, b). The second scenario was observed for the genera Pseudarthrobacter and the third for Glutamicibacter, which did not occur in most soil samples but was present in all tubers (Fig. S6c, d). For fungal OTUs, 234 were shared between soil and tuber, but only 3 OTUs had a higher abundance in both tuber and soil from lots with a low disease incidence in 2019 (Table S4), belonging to the genera Pyrenochaeta, Fusarium, and Acremonium. All were more abundant in several low disease incidence seed lots both in soil and in tubers (Fig. S7).
Discussion
This work demonstrated that seed lots of the same cultivar of potato, Kondor, originating from different locations, could differ in susceptibility to D. solani and P. brasiliense. These differences indicated that the origin of the mother tuber has an influence on of the next field generation resistance of cultivar Kondor against D. solani and P. brasilience. Furthermore, it was observed that disease incidence was generally higher in sandy soils than in clay soils. This was expected as sandy soils generally increase in temperature more quickly than clay soils during daytime, which is advantageous for pathogen growth and consequently for plant colonization and disease development (Du Raan et al., 2016). We also detected differences in disease incidence of the different lots inoculated with the two pathogens. Nevertheless, differences were sufficiently large to select lots with a generally higher and lower disease incidence in both years.
Differences in microbial communities were observed between seed lots. Small but significant differences in microbial community composition between seed lots with a high and low disease incidence, for both bacteria and fungi, suggested that the microbial community in the tuber was involved in disease resistance. Variability between the individual seed lots indicated that other factors, such as chemical composition and dry matter content of the tubers, may affect disease resistance. It is also possible that different taxa in the individual seed lots were associated with resistance against SRP. Various bacterial and fungal taxa have been identified as being involved in plant protection, for example Bacillus sp., Pseudomonas sp., Bradyrhizobium sp., and Trichoderma sp. (Diallo et al., 2011; Raaijmakers et al., 2002). It has also been indicated that different combinations of taxa and the interactions between those taxa may be effective in combating plant pathogens and in the induction of resistance in the plant (Czajkowski et al., 2020).
The phyla Firmicutes and Actinobacteria were more abundant in tubers and soils from lots with a low disease incidence. These two phyla have commonly been observed in soils and rhizosphere in correlation with suppressiveness against several different diseases (Lee et al., 2021; Mendes et al., 2011; Trivedi et al., 2017). In the current study, Actinobacteria, such as Arthrobacter sp., Curtobacterium sp. and Rhodococcus sp., Kocuria sp., and Glutamicibacter sp. and a Firmicute belonging to the family of Bacillacae, were more abundant in tubers and in soil with a low disease incidence. Most of these taxa are well described as including biocontrol bacteria. Several members of the genus Arthrobacter have been found to exhibit antagonism against a range of phytopathogenic fungi, but also against Pectobacterium carotovorum (Giannelli et al., 2022, López et al., 2021, Mahmoudi et al., 2011; Velázquez-Becerra et al., 2013). Biocontrol mechanisms of this genus include siderophore production, degradation of acetyl homoserine lactones (AHL) involved in quorum sensing and the production of volatile organic compounds. Glutamicibacter as well as Rhodoccus and Bacillus isolates were shown to reduce soft rot in potato caused by Pectobacterium carotovorum through AHL-degradation (Garge & Nerurkar, 2017; Vesuna & Nerurkar, 2020). Curtobacterium strains, on the other hand, have been suggested to act through the production of antibiotics against Agrobacterium vitis in grapevine (Ferrigo et al., 2017). Members of other phyla, such as the genus Pantoea, and especially the species P. agglomerans are known for their biocontrol activity through the production of antibiotic compounds and were found to be effective among others against Erwinia amylovora in apple and Botrytis cinerea in grapevine (Pusey et al., 2011, Trotel-Aziz et al., 2008). The genus Pseudomonas is widely known for its ability to produce antimicrobial compounds and its antagonistic activity against a variety of bacterial and fungal pathogens. Several strains of Pseudomonas sp. were found to reduce SRP growth in vitro and in plant assays, likely through the production of antibiotics and siderophores (Krzyzanowska et al., 2012, Krzyzanowska et al., 2019).
In addition, in this study we found several fungal taxa to be correlated with a low disease incidence of potato blackleg. Antagonistic activity of fungi against bacterial pathogens has been poorly investigated. However, several taxa have been previously reported for their antagonistic activity against fungal pathogens. Vishniacozyma victoriae has shown antagonism against e.g. Penicillium expansum and Botrytis cinerea (Gramisci et al., 2018), Plectosphaerella was an important predictor of plant health of sugar beets (Kusstatscher et al., 2019), and Acremonium sp. shown to affect different fungal pathogens by the production of antibiotics (Lo Piccolo et al., 2015, Wicklow et al., 2005). Endophytic fungi belonging to the genera Fusarium and Penicillium are known for their ability to induce resistance (Fontana et al., 2021). For example, Fusarium oxysporum strains reduced disease caused by Verticillium dahliae, Pythium ultimum or pathogenic Fusarium strains by induced resistance that did not seem to be associated with ISR nor SAR (de Lamo & Takken, 2020). Nevertheless, in most cases the mechanisms of fungal antagonistic activity have not yet been investigated. Given their frequently reported effectivity, more effort should be exerted to characterize potentially antagonistic fungal species.
It appears likely that the soil in which the mother tuber was grown had an influence on the tuber microbiome in our study. Our results are supported by previous findings that the soil has a larger effect on the tuber microbiome than the cultivar (Buchholz et al., 2019), although in our study we did not compare different cultivars. In support of this idea, we found that in 2019 many taxa occurring in the tubers could also be found in the soil of origin, sampled in 2018. However, only in a few cases the abundances of the taxa involved in resistance were likewise increased in soils that produced tubers with a low disease incidence. Both for fungi and for bacteria, the presence or a high abundance of specific taxa in tubers was not always accompanied by a higher abundance of the same taxa in soil. Rather, it appeared that these taxa were either differentially recruited to the tuber or, upon recruitment, were able to reach higher abundance in some seed lots, but not the others. Recruitment can be affected by a number of factors, as for example the field location, soil type and cultivation practices (Edwards et al., 2015; Liu et al., 2017). Interestingly, a number of taxa in the tubers, such as Glutamicibacter and Staphylococcus, were not found in soil, indicating that they originated either from the mother tuber or entered the above-ground part of the plant (Frank et al., 2017). It is also possible that the respective taxa were below detection limit in the soil. In addition, there could have been spatial and temporal differences in microbial dispersal i.e., the taxa only detected in tubers could have been present in soil at the time and/or specific location of plant growth, but not the time and/or location of soil sampling.
Conclusion
We conclude that potato seed lots of the same variety can differ in their resistance to disease caused by D. solani and P. brasiliense and that this difference in disease resistance is associated with differences in the bacterial and fungal community composition. A majority of differentially abundant taxa might have been recruited from the soil of the previous field generation. However, abundance in the soil was not always correlated with abundance in the tuber and so the soil microbiome cannot be used as an indicator of tuber resistance. It cannot be excluded that other, yet unknown factors are involved in the differences in blackleg resistance between seed lots. Our results also strongly suggest that for the testing of resistance of cultivars, more than one seed lot should be used to account for differences within one variety.
Data Availability
All data is available at doi: https://doi.org/10.4121/19237146.v1.
References
Amtmann, A., Troufflard, S., & Armengaud, P. (2008). The effect of potassium nutrition on pest and disease resistance in plants. Physiologia Plantarum, 133(4), 682–691. https://doi.org/10.1111/j.1399-3054.2008.01075.x
Berendsen, R. L., Pieterse, C. M. J., & Bakker, P. A. H. M. (2012). The rhizosphere microbiome and plant health. Trends in Plant Science, 17(8), 478–486. https://doi.org/10.1016/j.tplants.2012.04.001
Buchholz, F., Antonielli, L., Kostić, T., Sessitsch, A., & Mitter, B. (2019). The bacterial community in potato is recruited from soil and partly inherited across generations. PLoS ONE, 14(11), e0223691. https://doi.org/10.1371/journal.pone.0223691
Callahan, B., McMurdie, P., Rosen, M., Han, A., Johnson, A., & Holmes, S. (2016). DADA2: High-resolution sample inference from Illumina amplicon data. Nature Methods, 13, 581–583. https://doi.org/10.1038/nmeth.3869
Choudhary, D. K., et al. (2016). Bacterial-mediated tolerance and resistance to plants under abiotic and biotic stresses. Journal of Plant Growth Regulation, 35(1), 276–300. https://doi.org/10.1007/s00344-015-9521-x
Compant, S., Duffy, B., Nowak, J., Clément, C., & Barka, E. A. (2005). Use of plant growth-promoting bacteria for biocontrol of plant diseases: Principles, mechanisms of action, and future prospects. Applied and Environment Microbiology, 71(9), 4951–4959.
Compant, S., Sessitsch, A., & Mathieu, F. (2012). The 125th anniversary of the first postulation of the soil origin of endophytic bacteria – a tribute to M.L.V. Galippe. Plant and Soil, 356(1), 299–301. https://doi.org/10.1007/s11104-012-1204-9
Czajkowski, R., Pérombelon, M. C. M., van Veen, J. A., & van der Wolf, J. M. (2011). Control of blackleg and tuber soft rot of potato caused by Pectobacterium and Dickeya species: A review. Plant Pathology, 60(6), 999–1013. https://doi.org/10.1111/j.1365-3059.2011.02470.x
Czajkowski, R., De Boer, W., Van Veen, J., & Van der Wolf, J. (2012). Characterization of bacterial isolates from rotting potato tuber tissue showing antagonism to Dickeya sp. biovar 3 in vitro and in planta. Plant Pathology, 61(1), 169–182.
Czajkowski, R., et al. (2015). Salicylic acid can reduce infection symptoms caused by Dickeya solani in tissue culture grown potato (Solanum tuberosum L.) plants. European Journal of Plant Pathology, 141(3), 545–558. https://doi.org/10.1007/s10658-014-0561-z
Czajkowski, R., Maciag, T., Krzyzanowska, D. M., & Jafra, S. (2020). Biological control based on microbial consortia–from theory to commercial products. How research can stimulate the development of commercial biological control against plant diseases (pp. 183–202).
de Lamo, F. J., & Takken, F. L. W. (2020). Biocontrol by Fusarium oxysporum using endophyte-mediated resistance. Frontiers in Plant Science, 11. https://doi.org/10.3389/fpls.2020.00037
Diallo, S., Crépin, A., Barbey, C., Orange, N., Burini, J.-F., & Latour, X. (2011). Mechanisms and recent advances in biological control mediated through the potato rhizosphere. FEMS Microbiology Letters, 75(3), 351–364. https://doi.org/10.1111/j.1574-6941.2010.01023.x
Du Raan, S., Coutinho, T. A., & Van der Waals, J. E. (2016). Cardinal temperature differences, determined in vitro, between closely related species and subspecies of pectinolytic bacteria responsible for blackleg and soft rot on potatoes. European Journal of Plant Pathology, 144(2), 361–369.
Dupuis, B., Nkuriyingoma P., & Van Gijsegem F. (2021). Economic impact of Pectobacterium and Dickeya species on potato crops: A review and case study. In (Ed.), Plant diseases caused by dickeya and pectobacterium species (pp. 263–282)
Edwards, J., et al. (2015). Structure, variation, and assembly of the root-associated microbiomes of rice. PNAS, 112(8), E911–E920. https://doi.org/10.1073/pnas.1414592112
Ferrigo, D., Causin, R., & Raiola, A. (2017). Effect of potential biocontrol agents selected among grapevine endophytes and commercial products on crown gall disease. BioControl, 62(6), 821–833. https://doi.org/10.1007/s10526-017-9847-3
Fontana, D. C., et al. (2021). Endophytic fungi: Biological control and induced resistance to phytopathogens and abiotic stresses. Pathogens, 10(5), 570.
Fox. J., & Weisberg, S. (2011). An {R} companion to applied regression (trans. Thousand Oaks CA: Sage
Frank, A. C., Saldierna Guzmán, J. P., & Shay, J. E. (2017). Transmission of bacterial endophytes. Microorganisms, 5(4), 70. https://doi.org/10.3390/microorganisms5040070
Garge, S. S., & Nerurkar, A. S. (2017). Evaluation of quorum quenching Bacillus spp. for their biocontrol traits against Pectobacterium carotovorum subsp. carotovorum causing soft rot. Biocatalysis and Agricultural Biotechnology, 9, 48–57. https://doi.org/10.1016/j.bcab.2016.11.004
Giannelli G., et al. (2022). Phyto-beneficial traits of rhizosphere bacteria: in vitro exploration of plant growth promoting and phytopathogen biocontrol ability of selected strains isolated from harsh environments. Plants, 11(2), 230. https://www.mdpi.com/2223-7747/11/2/230
Gramisci, B. R., Lutz, M. C., Lopes, C. A., & Sangorrín, M. P. (2018). Enhancing the efficacy of yeast biocontrol agents against postharvest pathogens through nutrient profiling and the use of other additives. Biological Control, 121, 151–158. https://doi.org/10.1016/j.biocontrol.2018.03.001
Hélias, V., Hamon, P., Huchet, E., Wolf, J. V. D., & Andrivon, D. (2012). Two new effective semiselective crystal violet pectate media for isolation of Pectobacterium and Dickeya. Plant Pathology, 61(2), 339–345. https://doi.org/10.1111/j.1365-3059.2011.02508.x
Kou, R., et al. (2016). Benefits and challenges with applying unique molecular identifiers in next generation sequencing to detect low frequency mutations. PLoS ONE, 11(1), e0146638. https://doi.org/10.1371/journal.pone.0146638
Krzyzanowska, D., et al. (2012). Rhizosphere bacteria as potential biocontrol agents against soft rot caused by various Pectobacterium and Dickeya spp. strains. Journal of Plant Pathology, 94(2), 367–378.
Krzyzanowska, D. M., Maciag, T., Siwinska, J., Krychowiak, M., Jafra, S., & Czajkowski, R. (2019). Compatible mixture of bacterial antagonists developed to protect potato tubers from soft rot caused by Pectobacterium spp and Dickeya spp. Plant Disease, 103(6), 1374–1382. https://doi.org/10.1094/pdis-10-18-1866-re
Kusstatscher, P., et al. (2019). Disease incidence in sugar beet fields is correlated with microbial diversity and distinct biological markers. Phytobiomes Journal, 3(1), 22–30. https://doi.org/10.1094/pbiomes-01-19-0008-r
Lee, S.-M., Kong, H. G., Song, G. C., & Ryu, C.-M. (2021). Disruption of Firmicutes and Actinobacteria abundance in tomato rhizosphere causes the incidence of bacterial wilt disease. ISME Journal, 15(1), 330–347. https://doi.org/10.1038/s41396-020-00785-x
Lenth, R., Singmann, H., Love, J., Buerkner, P., Herve, M. (2018) 'emmeans: estimated marginal means, aka least-square means' 1.2.3. p.^pp. Available at: https://github.com/rvlenth/emmeans (Accessed
Liaw, A., Wiener, M. (2002). Classification and regression by randomForest. R News, 2(3), 18–22. https://CRAN.R-project.org/doc/Rnews/
Liu, H., et al. (2017). Inner plant values: diversity, colonization and benefits from endophytic bacteria. Frontiers in Microbiology, 8(2552). https://doi.org/10.3389/fmicb.2017.02552
Liu, C., Cui, Y., Li, X., & Yao, M. (2020a). microeco: an R package for data mining in microbial community ecology. FEMS Microbiology Letters, 97(2). https://doi.org/10.1093/femsec/fiaa255
Liu, J.-M., et al. (2020b). Antimicrobial activity against phytopathogens and inhibitory activity on solanine in potatoes of the endophytic bacteria isolated from potato tubers. Frontiers in Microbiology, 11
Lo Piccolo, S., Alfonzo, A., Giambra, S., Conigliaro, G., Lopez-Llorca, L. V., & Burruano, S. (2015). Identification of Acremonium isolates from grapevines and evaluation of their antagonism towards Plasmopara viticola. Annales De Microbiologie, 65(4), 2393–2403. https://doi.org/10.1007/s13213-015-1082-5
López, S. M. Y., Pastorino, G. N., & Balatti, P. A. (2021). Volatile organic compounds profile synthesized and released by endophytes of tomato (Solanum lycopersici L.) and their antagonistic role. Archives of Microbiology, 203(4), 1383–1397. https://doi.org/10.1007/s00203-020-02136-y
Love, M. I., Huber, W., & Anders, S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology, 15(12), 550. https://doi.org/10.1186/s13059-014-0550-8
Lundberg, D. S., Yourstone, S., Mieczkowski, P., Jones, C. D., & Dangl, J. L. (2013). Practical innovations for high-throughput amplicon sequencing. Nature Methods, 10(10), 999–1002. https://doi.org/10.1038/nmeth.2634
Mahmoudi, E., Tabatabaei, B. E. S., & Venturi, V. (2011). Virulence attenuation of Pectobacterium carotovorum using N-acyl-homoserine lactone degrading bacteria isolated from potato rhizosphere. Plant Pathology Journal, 27(3), 242–248.
McMurdie, P. J., & Holmes, S. (2013). phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLOS ONE, 8(4), e61217. https://doi.org/10.1371/journal.pone.0061217
Mendes, R., et al. (2011). Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science, 332(6033), 1097–1100. https://doi.org/10.1126/science.1203980
Morales-Cedeño, L. R., Orozco-Mosqueda, M. D. C., Loeza-Lara, P. D., Parra-Cota, F. I., de los Santos-Villalobos, S., & Santoyo, G. (2021). Plant growth-promoting bacterial endophytes as biocontrol agents of pre- and post-harvest diseases: Fundamentals, methods of application and future perspectives. Microbiological Research, 242, 126612. https://doi.org/10.1016/j.micres.2020.126612
Oksanen, J., et al. (2007). The Vegan Package. Community Ecology Package, 10(631–637), 719.
Padilla-Gálvez, N., et al. (2021). Antagonistic activity of endophytic actinobacteria from native potatoes (Solanum tuberosum subsp. tuberosum L) against Pectobacterium carotovorum subsp. carotovorum and Pectobacterium atrosepticum. BMC Microbiology, 21(1), 1–17.
Pérombelon, M. C. M. (1992). Potato blackleg: Epidemiology, host-pathogen interaction and control. Netherlands Journal of Plant Pathology, 98(2), 135–146. https://doi.org/10.1007/bf01974480
Pieterse, C. M. J., Zamioudis, C., Berendsen, R. L., Weller, D. M., Van Wees, S. C. M., & Bakker, P. A. H. M. (2014). Induced systemic resistance by beneficial microbes. Annual Review of Phytopathology, 52(1), 347–375. https://doi.org/10.1146/annurev-phyto-082712-102340
Pusey, P., Stockwell, V., Reardon, C., Smits, T., & Duffy, B. (2011). Antibiosis activity of Pantoea agglomerans biocontrol strain E325 against Erwinia amylovora on apple flower stigmas. Phytopathology, 101(10), 1234–1241.
Raaijmakers, J. M., Vlami, M., & de Souza, J. T. (2002). Antibiotic production by bacterial biocontrol agents. Antonie Van Leeuwenhoek, 81(1), 537. https://doi.org/10.1023/a:1020501420831
Toth, I. K., et al. (2011). Dickeya species: An emerging problem for potato production in Europe. Plant Pathology, 60(3), 385–399. https://doi.org/10.1111/j.1365-3059.2011.02427.x
Trivedi, P., Delgado-Baquerizo, M., Trivedi, C., Hamonts, K., Anderson, I. C., & Singh, B. K. (2017). Keystone microbial taxa regulate the invasion of a fungal pathogen in agro-ecosystems. Soil Biology & Biochemistry, 111, 10–14.
Trotel-Aziz, P., Couderchet, M., Biagianti, S., & Aziz, A. (2008). Characterization of new bacterial biocontrol agents Acinetobacter, Bacillus, Pantoea and Pseudomonas spp. mediating grapevine resistance against Botrytis cinerea. Environmental and Experimental Botany, 64(1), 21–32. https://doi.org/10.1016/j.envexpbot.2007.12.009
van der Wolf, J. M., et al. (2017). Virulence of Pectobacterium carotovorum subsp brasiliense on potato compared with that of other Pectobacterium and Dickeya species under climatic conditions prevailing in the Netherlands. Plant Pathology, 66(4), 571–583. https://doi.org/10.1111/ppa.12600
van der Wolf, J. M. et al. (2021a). Diseases caused by Pectobacterium and Dickeya species around the world. In (Ed.), Plant Diseases Caused by Dickeya and Pectobacterium Species (pp. 215–261). Springer
van der Wolf, J. M., et al. (2021b). Management of diseases caused by Pectobacterium and Dickeya species. Plant diseases caused by Dickeya and Pectobacterium species, 175–214
Velázquez-Becerra, C., et al. (2013). The rhizobacterium Arthrobacter agilis produces dimethylhexadecylamine, a compound that inhibits growth of phytopathogenic fungi in vitro. Protoplasma, 250(6), 1251–1262. https://doi.org/10.1007/s00709-013-0506-y
Vesuna, A. P., & Nerurkar, A. S. (2020). Biocontrol impact of AHL degrading actinobacteria on quorum sensing regulated virulence of phytopathogen Pectobacterium carotovorum subsp. carotovorum BR1. Plant and Soil, 453(1), 371–388. https://doi.org/10.1007/s11104-020-04623-z
Wicklow, D. T., Roth, S., Deyrup, S. T., & Gloer, J. B. (2005). A protective endophyte of maize: Acremonium zeae antibiotics inhibitory to Aspergillus flavus and Fusarium verticillioides, Dedicated to John Webster on the occasion of his 80th birthday. Mycological Research, 109(5), 610–618. https://doi.org/10.1017/S0953756205002820
Funding
This research was funded by the Dutch Ministry of Agriculture, Nature and Food Quality (LNV) in the Topsector Program Agriculture & Food (project title: “Effect van de bodem op weerbaarheid van aardappelknollen tegen biotische stress”; project number: AF-17003).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kurm, V., Mendes, O., Gros, J. et al. Potato tuber origin and microbial composition determines resistance against soft rot Pectobacteriaceae. Eur J Plant Pathol 168, 383–399 (2024). https://doi.org/10.1007/s10658-023-02763-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-023-02763-3