Abstract
Viral diseases in grapevine cause large economic losses due to decreased irregular yield and unbalanced ripening, and can even lead to plant mortality. There is a large number of grapevine viral agents, and a few of them have a prominent impact due to their worldwide distribution, virulence, and incidence. Although previous research has evaluated variations in viral load between organs and time since infection, there is still a lack of knowledge on how the viruses are transported toward developing tissues. In this work, we present the results of two experiments that contribute to understanding the spread dynamics of four major grapevine viruses (GFLV, GFkV, GLRaV-1, and GLRaV -3). Bud and leaf tissues were sampled from shoots obtained from cv. ‘Garnacha’ cuttings known to be infected with one of these viruses. Bud samples taken at early development stages were used to understand short-distance transport, while leaves taken from young shoots represented long-distance transport, driven mainly through the phloem. Our results show that all viruses were able to invade tissues from the beginning of development. The dissemination ability of GFLV was considerable, as the viral load detected in young organs was as high as in the dormant shoot. Furthermore, for GFLV and GFkV, it was shown that the viral load in young shoots does not follow the general assumption of older tissues accumulating a higher viral load but, conversely, a higher viral load closer to the shoot tip might be driven by the sink strength.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The grape growing industry is concerned about viral diseases since they result in significant economic losses due to decreased yields, irregular and unbalanced ripening, and even plant mortality (Andret-Link et al., 2004; Mannini & Digiaro, 2017; Martínez et al., 2016; Moutinho-Pereira et al., 2012; Naidu et al., 2014; Vega et al., 2011; Vigne et al., 2015; Wu et al., 2020). Viral diseases are provoked by many diverse viral agents, and in the case of cultivated grapevine (Vitis vinifera L.), it is known to be the host of over 80 distinct virus species (Fuchs, 2020). Some of these viruses have a prominent impact due to their worldwide distribution, virulence, and incidence, and specific control regulations are implemented to contain and reduce their dissemination. For instance, at the European level, regulations specify the obligation to test for the presence of Grapevine fanleaf virus (GFLV) and Arabis mosaic virus (ArMV) of the genus Nepovirus, Grapevine leafroll-associated virus 1 and 3 (GLRaV-1 and -3) of the genus Ampelovirus, while testing for Grapevine fleck virus (GFkV) of the genus Maculavirus is only mandatory in rootstock plant material (Directive, 1968; OEPP/EPPO, 2008).
Since the viral distribution along the plant is not uniform, previous research has been conducted on viral detection and quantification in different plant tissues with two main aims. First, identify organs and time points in the season with the highest viral load for each virus in order to improve diagnosis, trying to prevent false negatives (Bouyahia et al., 2003; Čepin et al., 2010; Chooi et al., 2016; Fiore et al., 2009; Gasparro et al., 2019; Komínek et al., 2009; Krebelj et al., 2015; Pacifico et al., 2011; Rowhani et al., 1992; Shabanian et al., 2020; Walter & Etienne, 1987). Secondly, identify virus-free tissues, with the objective to generate virus-free plants (sanitation) through in vitro culture. As in other horticultural crops, viral concentration decreases in the apical meristem, and sequential in vitro propagation has been used for grapevine sanitation since the 1950s (Esau, 1967; Gautheret, 1983; Limasset & Cornuet, 1949; Mori & Hosokawa, 1977; Youssef et al., 2009).
Different reasons are proposed for the inhibition of virus replication in meristematic tips (Waigmann & Heinlein, 2007), mainly (i) the yet scarce development of conductive tissues, which can hinder particularly the transportation of phloem-limited viruses (Gilbertson & Lucas, 1996); (ii) the potentially inhibitory role of plant hormones such as auxins, whose content is higher in developing tissues (Espinoza et al., 2007; Padmanabhan et al., 2005), or (iii) viral RNA silencing defence from the plant (Schwach et al., 2005).
In the current study, we designed an experiment to understand the spread dynamics of GFLV, GFkV, GLRaV-1, and GLRaV -3 viruses towards developing tissues. Our work comprises a two-sided approach, each linked to a viral agent transport mechanism (Carrington et al., 1996). On the one hand, we studied developing bud tissues to understand short-distance transport, which comprehends the transport of viruses through the plasmodesmata of neighbouring cells. On the other hand, we analysed leaves of young developing shoots to study the long-distance transport system, in which viral agents use the phloem to spread throughout the plant.
Material and methods
Vitis vinifera L. cv. Garnacha (syn. Grenache) plants from a public germplasm collection of regional Garnacha biotypes located in the Public University of Navarre (Pamplona, Navarre, Spain) were used as starting material. Since all the plants in the collection had been historically tested for the presence of GFLV, GFkV, GLRaV-1, and GLRaV-3 by enzyme-linked immunosorbent assay (ELISA), it was possible to select five plants that were positive for just one of the four viral infections considered. At the beginning of the experiment, those results were confirmed by real time RT-PCR. Nonetheless, it must be considered that other viruses, which were not tested, could be present. The dormant shoots of these plants were sampled in winter and kept in a cold chamber (4ºC) until potting. One week later, dormant shoots were cut into three winter buds-cuttings. To prevent desiccation, the distal end of each dormant cutting was sealed with paraffin, while the basal end was gently crushed and immersed in a rooting promoter (Hormon, Productos Flower, Tàrrega, Spain) to increase rooting success. These cuttings were planted in 2L pots, using a mixture of 75% peat—25% perlite substrate, and placed in a phytotron under controlled conditions (T = 20 °C, 16 h light photoperiod), and watered every second day. After 10 days, the start of winter bud development was evident.
As detailed above, two types of samples were used based on the virus transport distance considered. Thus, to represent the short-distance transport, buds of four development stages, defined according to the BBCH scale (Biologische Bundesanstalt, Bundessortenamt und Chemische Industrie scale) (Meier, 1997), were used: BBCH01 (beginning of bud swelling), BBCH03 (end of bud swelling), BBCH05 (wool stage), and BBCH07 (beginning of bud burst) (Fig. 1A). Three replicates were obtained for every bud stage. The long-distance transport system was evaluated using leaves placed at different positions in the shoot. For that reason, all the buds left were allowed to develop into a shoot with its developing leaves. When shoots had reached ca. 30 cm- in length, three shoots were selected and single leaf samples (which number was variable) were taken and labelled according to their relative position from the axis, grading them from the tip (L1) to the basal leaf (LX) (Fig. 1B). The samples taken from every dormant shoot were registered. All samples were weighed, photographed, fast frozen in liquid nitrogen, and stored at -80 °C until further use.
Viral presence was determined by real time RT-PCR. For that, 100 mg of plant material was ground to a fine powder. Total RNA isolation was performed using Spectrum Plant Total RNA Kit (Sigma-Aldrich, Oakville, ON, Canada) following manufacturer instructions with slight modifications:—2% Polyvinylpyrrolidone (PVPP) and 5 µl β-Mercaptoethanol were added to the lysis buffer to avoid polyphenols and proteins, and—the elution step was repeated twice to increase RNA yield. 500 ng of total RNA was reverse-transcribed using the PrimeScript RT Reagent Kit (Takara Bio Inc., Shiga, Japan) following manufacturer instructions. Real-time amplification was carried out in an ABI StepOne Plus thermocycler (Applied Biosystems, Foster City, CA, USA). PCR mixture included 10 ng of cDNA, 1 × TB Green Premix Ex Taq II and 1 × ROX reference dye from a kit (Takara Bio Inc., Shiga, Japan), 0.4 μM forward and reverse primers (Table 1) (Thermo Fisher, Waltham, MA USA) in a final volume of 10 µl. Amplification conditions for GFLV and GFkV were: initial denaturation at 95ºC for 5 s, followed by 40 consecutive cycles of 95ºC for 15 s (denaturation), and at 60ºC for 1 min (annealing and extension). Actin, GLRaV-1, and GLRaV-3 were run according to the published conditions (Table 1). According to amplification results, virus presence/absence could be determined. To determine the viral load, real time RT-PCR data were normalized according to the method proposed by Pfaffl (2001). Relative expression (R) was calculated with the expression of actin as the reference gene and the amplification efficiency of the specific genes. As a control, for every sample, the gene expression of the primary dormant shoot was used. The viral load was expressed as mean values ± Standard Error (SE). Statistical differences between tissues and phenological stages were tested with a one-way analysis of variance (ANOVA) (p < 0.05) and significantly different stages were determined with Tukey’s HSD test. Instead, leaves were tested according to their position and the correlation between leaf position and viral load was tested with Kendall’s tau-b nonparametric correlation test. Statistic tests were done using SPSS (Corp, 2021) and R4.2.2 (R Core Team, 2022) with RStudio (RStudio Team, 2020) and the packages “ggpubr” (Alboukadel Kassambara, 2023) and “Kendall” (McLeod, 2022).
Results and discussion
Short-distance transport
In the first part of the study, virus short-distance transport was evaluated through the analysis of viral presence in buds at different development stages. The presence of viral infections did not affect the development of the buds, as shown by the weight of the samples in all the phenological stages (data not shown). GFLV, GFkV, and GLRaV-1 were detected in all buds sampled. Contrastingly, only one-third of the buds coming from GLRaV-3-infected cuttings tested positive at BBCH03, BBCH05, and BBCH07 stages (Table 2). To our knowledge, only Fiore et al. (2009) have analysed previously the presence of GFLV, GFkV, GLRaV-1, and GLRaV-3 in developing buds. These authors reported buds to be positive by RT-PCR for all the viruses, except for some of those sampled from cuttings infected by GLRaV-1. Although the results obtained by Fiore et al. (2009) and those reported here are not in total agreement, both agents which were not detected in a fraction of the samples belong to the Closterovirus family, sharing many common characteristics and mechanisms.
When quantifying viral load, no significant differences were obtained between bud developmental stages. However, significant differences were obtained between buds and the dormant shoots from which they sprouted. The viral load of GFLV in the buds was as high as in the dormant shoots (Fig. 2A), while the load of the remaining viruses (GFkV, GLRaV-1, and GLRaV-3) was significantly lower in the buds than in the dormant shoots (Fig. 2 B-D). This result suggests a more efficient spread of Nepovirus (GFLV) compared to that of the phloem-limited viruses, such as Maculavirus or Ampelovirus (GFkV, GLRaV-1, and GLRaV-3). Winter and developing buds (BBCH01-BBCH03) are not hydraulically connected to the dormant shoot, but the vascular tissues are created through bud burst, starting between stages BBCH03-BBCH05 (Xie et al., 2018). Thus, viral spreading at early developing bud tissues is made through plasmodesmata. Plasmodesmata have a narrow diameter and, specifically in grapevine, their molecular exclusion size is around 2 nm at bud burst (Signorelli et al., 2020). This pore size is not sufficient for the transport of viral agents such as GFLV and GFkV, whose diameter is around 30 nm, or of GLRaV-1 and GLRaV-3, which are 12 nm in diameter. Rather, plasmodesmata must be modified to enlarge the pore size exclusion limit and enable viral transit: Nepovirus use the formation of tubules as channels to promote viral transfer (Amari et al., 2010, 2011; Kalasjan et al., 1979; Kasteel et al., 1993); in the Closteroviridae family, transport is mediated by the association of HSP70h with plasmodesmata (Alzhanova et al., 2001; Peremyslov et al., 1999); the cell-to-cell transport mechanism of GFkV is not yet known. Our results suggest that tubule formation of GFLV is a successful strategy. These results are in concordance with the difficulty in producing GFLV virus-free tissues, which shows its particular ability to infect and disseminate callus tissues (Gambino et al., 2010).
Relative virus content (R) of the primary dormant shoots and the bud stages (BBCH01, BBCH03, BBCH05, and BBCH07) for the different viruses under study A) GFLV, B) GFkV, C) GLRaV-1 and D) GLRaV-3. Bars represent standard errors (SE). Different letters indicate statistically significant differences (p < 0.05)
Long-distance transport
In the second part of the study, developing shoots with five to eight leaves were used to evaluate long-distance transport. Viral infections did not affect leaf area or shoot development (data not shown). Regarding the viral presence, all the single leaves sampled from GFLV, GFkV, GLRaV-1, or GLRaV-3 infected dormant cuttings were positive (Table 3).
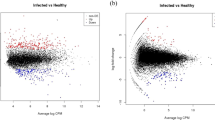
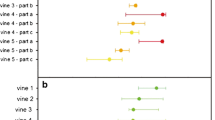
Variations in viral load were examined based on leaf position (i.e.: age). According to Kendall’s tau-b correlation calculations (tau), viral load and leaf position were significantly correlated for GFLV (p-value = 0.036) (Fig. 3A) and GFkV (0.003) (Fig. 3B), while viral load distribution did not have a pattern for GLRaV-1 (Fig. 3C) and GLRaV-3 (Fig. 3D). For GFLV and GFkV, Pearson’s correlation coefficient (tau load/leaf position) was negative as showed a higher load in leaves closer to the tip/younger. A similar correlation trend has been previously reported for GFLV, for which an increasing load towards apical leaves was observed in spring in field plants (Bouyahia et al., 2003). Here we further demonstrate that the same distribution trend is followed by GFkV in infecting young shoots.
Scatter plot showing the relative virus content (R) of the single leaves (L1, …, L8) and its regression line for the viruses A) GFLV, B) GFkV, C) GLRaV-1 and D) GLRaV-3. Kendall’s Tau correlation coefficients (tau, p-value) are included in every chart. A negative statistically significant correlation between viral load and leave position was found for the virus GFLV (rload, leaf position = -0.391, p = 0.036) and GFkV (-0.567, 0.003)
The success of GFLV, GFkV, GLRaV-1, and GLRaV-3 colonizing young leaves becomes evident with these results. At that developmental stage, when young shoots are growing, the four viruses are spread through the phloem which confirms the effectiveness of the long-distance transport for viral dispersion and the establishment of a systemic infection (Gilbertson & Lucas, 1996). Long-distance transport of viruses and photoassimilates takes place together so, if mechanisms for viral flow restriction are triggered, they may endanger plant development as they would restrict not only viral flow but also that of assimilates (Hipper et al., 2013). On the other hand, our results point out that, at least for some viruses, the distribution of viral load in young shoots contrasts with that of mature tissues and storage organs. For the last, the general assumption is that the viral load increases when tissues become older and those arguments are considered in sampling proceedings for virus testing (Monis & Bestwick, 1996; Osman et al., 2018). Even symptomatology is often observed earlier in older leaves (Baozhong et al., 2017). Here, we report that not all tissues follow the same viral distribution, and age might be a factor that determines it. This changeable distribution trend might be related to the source-to-sink relations: at developing tissues, the sink strength for photoassimilates of the apical meristem may attract a higher viral flow too, while that strength lessens through tissue maturity (Bendix & Lewis, 2018; Gutiérrez et al., 2012; Scott and Leisner., 1993).
Conclusion
As far as we are aware, this study is the first to examine the presence of the grapevine GFLV, GFkV, GLRaV-1, and GLRaV -3 viruses through bud and early leaf development in grapevine. Regarding short-distance viral transport, the results indicate that all viruses tested can invade tissues from the start of development (stages BBCH01 in buds). The capacity of GFLV to disseminate by symplastic means was outstanding: the viral load detected in young buds (BCCH01 R=0.58±0.09 and BCCH03 R=1.60±1.23) was as high as in the primary dormant shoot (R=1). This may be explained by its well-known plasmodesmata modification strategy, which enlarges plasmodesmata through tubules. Furthermore, short-distance viral transport showed infected leaves as small as 50mm2 which confirms their early ability to cause systemic infection. Additionally, it was shown that the GFLV and GFkV viral load in young shoots did not follow the general assumption that older tissues accumulate a higher viral load. In contrast, at early shoot development stages, the viral load appears to be driven by the apical meristem sink strength of the shoot tip (for GLFV rload, leaf position = -0.391 and for GFkV rload, leaf position =-0.567).
Abbreviations
- HSP70=:
-
Heat shock protein
References
Alzhanova, D. V., Napuli, A. J., Creamer, R., & Dolja, V. V. (2001). Cell-to-cell movement and assembly of a plant closterovirus: Roles for the capsid proteins and Hsp70 homolog. EMBO Journal, 20, 6997–7007. https://doi.org/10.1093/emboj/20.24.6997
Amari K., Boutant E., Hofmann C., Schmitt-Keichinger C., Fernandez-Calvino L., Didier, P., Lerich, A., Mutterer, J., Thomas, C.L., & Heinlein, M. (2010). A family of plasmodesmal proteins with receptor-like properties for plant viral movement proteins. PLoS Pathogens 6(9): e1001119. https://doi.org/10.1371/journal.ppat.1001119
Amari K., Lerich A., Schmitt-Keichinger C., Dolja V. V., & Ritzenthaler C. (2011). Tubule-guided cell-to-cell movement of a plant virus requires class XI myosin motors. PLoS Pathogens, 7(10), e1002327. https://doi.org/10.1371/journal.ppat.1002327
Andret-Link, P., Laporte, C., Valat, L., Ritzenthaler, C., Demangeat, G., Vigne, E., Laval, V., Pfeiffer, P., Stussi-Garaud, C., Fuchs, M., 2004. Grapevine fanleaf virus: Still a major threat to the grapevine industry. J. Plant Pathol. 183–195. http://www.jstor.org/stable/41992424
Baozhong M, Giovanni P. Martelli, Deborah A. Golino, M.F. (Ed.), 2017. Grapevine Viruses: Molecular Biology, Diagnostics and Management, 1st ed. Springer Cham. https://doi.org/10.1007/978-3-319-57706-7
Bendix, C., & Lewis, J. D. (2018). The enemy within: Phloem-limited pathogens. Molecular Plant Pathology, 19, 238–254. https://doi.org/10.1111/mpp.12526
Bouyahia H., Potere, O., & Boscia, D. (2003). Sampling methodology for the detection of Grapevine fanleaf virus by ELISA. Extended Abstracts of the 14th Meeting of ICVG, Locorotondo 2003, 204.
Carrington, J. C., Kasschau, K. D., Mahajan, S. K., & Schaad, M. C. (1996). Cell-to-cell and long-distance transport of viruses in plants. The Plant Cell, 8, 1669. https://doi.org/10.1105/tpc.8.10.1669
Čepin, U., Gutiérrez-Aguirre, I., Balažic, L., Pompe-Novak, M., Gruden, K., & Ravnikar, M. (2010). A one-step reverse transcription real-time PCR assay for the detection and quantitation of Grapevine fanleaf virus. Journal of Virological Methods, 170, 47–56. https://doi.org/10.1016/j.jviromet.2010.08.018
Chooi, K. M., Cohen, D., & Pearson, M. N. (2016). Differential distribution and titre of selected grapevine leafroll-associated virus 3 genetic variants within grapevine rootstocks. Archives of Virology, 161, 1371–1375. https://doi.org/10.1007/s00705-016-2791-1
Corp, I., 2021. IBM SPSS Statistics for Windows, Version 28.0.
Directive, H. A. S. A. T. (1968). Council Directive 68/193/EEC of 9 April 1968 on the marketing of material for the vegetative propagation of the vine. Off. J. L, 93, 15–23.
Esau, K. (1967). Anatomy of plant virus infections. Annual Review of Phytopathology, 5, 45–74. https://doi.org/10.1146/annurev.py.05.090167.000401
Espinoza, C., Vega, A., Medina, C., Schlauch, K., Cramer, G., & Arce-Johnson, P. (2007). Gene expression associated with compatible viral diseases in grapevine cultivars. Functional & Integrative Genomics, 7, 95–110. https://doi.org/10.1007/s10142-006-0031-6
Fiore, N., Prodan, S., Pino, A.M., 2009. Monitoring grapevine viruses by ELISA and RT-PCR throughout the year. J. Plant Pathol. 91, 489–493. https://www.jstor.org/stable/41998649
Fuchs, M. (2020). Grapevine viruses: A multitude of diverse species with simple but overall poorly adopted management solutions in the vineyard. Journal of. Plant Pathology, 102, 643–653. https://doi.org/10.1007/s42161-020-00579-2
Gambino, G., Vallania, R., & Gribaudo, I. (2010). In situ localization of Grapevine fanleaf virus and phloem-restricted viruses in embryogenic callus of Vitis vinifera. European Journal of Plant Pathology, 127, 557–570. https://doi.org/10.1007/s10658-010-9619-8
Gasparro, M., Milella, R. A., Alba, V., Giannandrea, M. A., & Caputo, A. R. (2019). Seasonal dynamics and spatial distribution of main Grapevine viruses in field-grown grapevine cultivars. European Journal of Plant Pathology, 155, 193–205. https://doi.org/10.1007/s10658-019-01761-8
Gautheret, R. J. (1983). Plant tissue culture: A history. Bot. Mag. Shokubutsu-Gaku-Zasshi, 96, 393–410. https://doi.org/10.1007/BF02488184
Gilbertson, R. L., & Lucas, W. J. (1996). How do viruses traffic on the ‘vascular highway’? Trends in Plant Science, 1, 250–251. https://doi.org/10.1016/1360-1385(96)10029-7
Griesser, M., Martinez, S. C., Eitle, M. W., Warth, B., Andre, C. M., Schuhmacher, R., & Forneck, A. (2018). The ripening disorder berry shrivel affects anthocyanin biosynthesis and sugar metabolism in Zweigelt grape berries. Planta, 247(2), 471–481. https://doi.org/10.1007/s00425-017-2795-4
Gutiérrez, S., Yvon, M., Pirolles, E., Garzo, E., Fereres, A., Michalakis, Y., & Blanc, S. (2012). Circulating virus load determines the size of bottlenecks in viral populations progressing within a host. PLoS Pathog., 8, e1003009. https://doi.org/10.1371/journal.ppat.1003009
Hipper, C., Brault, V., Ziegler-Graff, V., & Revers, F. (2013). Viral and cellular factors involved in phloem transport of plant viruses. Frontiers in Plant Science, 4, 154. https://doi.org/10.3389/fpls.2013.00154
Kalasjan, J. A., Litvak, L. A., & Marinescu, V. G. (1979). Tubular structures in grapevine tissue after infection with grapevine fanleaf virus. Arch Phytopathol Pflanzenschutz, 15, 373–376.
Alboukadel Kassambara, 2023. ggpubr: “ggplot2” Based Publication Ready Plots.
Kasteel, D., Wellink, J., Verver, J., Van Lent, J., Goldbach, R., & Van Kammen, A. (1993). The involvement of cowpea mosaic virus M RNA-encoded proteins in tubule formation. Journal of General Virology, 74, 1721–1724. https://doi.org/10.1099/0022-1317-74-8-1721
Komínek, P., Glasa, M., & Komínková, M. (2009). Analysis of multiple virus-infected grapevine plant reveals persistence but uneven virus distribution. Acta Virologica, 53, 281. https://doi.org/10.4149/av_2009_04_281
Krebelj, A. J., Čepin, U., Ravnikar, M., & Novak, M. P. (2015). Spatio-temporal distribution of Grapevine fanleaf virus (GFLV) in grapevine. European Journal of Plant Pathology, 142, 159–171. https://doi.org/10.1007/s10658-015-0600-4
Limasset, P., & Cornuet, P. (1949). Recherche du virus de la mosaique du tabac (Marmor-tabaci Holmes) dans les meristemes des plantes infectees. Comptes Rendus Hebdomadaires Des Séances De L’académie Des Sciences, 228, 1971–1972.
Mannini, F., Digiaro, M., 2017. The effects of viruses and viral diseases on grapes and wine, in: Grapevine Viruses: Molecular Biology, Diagnostics and Management. Springer, 453–482. https://doi.org/10.1007/978-3-319-57706-7_23
Martínez, L., Miranda, C., Royo, J. B., Urrestarazu, J., Martínez de Toda, F., Balda, P., & Santesteban, L. G. (2016). Direct and indirect effects of three virus infections on yield and berry composition in grapevine (Vitis vinifera L.) cv. ‘Tempranillo.’ Science Horticulturae, 212, 20–28. https://doi.org/10.1016/j.scienta.2016.09.023
McLeod, A.I., 2022. Kendall Rank Correlation and Mann-Kendall Trend Test.
Meier, U. (1997). Growth stages of mono-and dicotyledonous plants. Blackwell Wissenschafts-Verlag.
Monis, J., & Bestwick, R. K. (1996). Detection and localization of grapevine leafroll associated closteroviruses in greenhouse and tissue culture grown plants. American Journal of Enology and Viticulture, 47, 199–205. https://doi.org/10.5344/ajev.1996.47.2.199
Mori, K., & Hosokawa, D. (1977). Localization of viruses in apical meristem and production of virus-free plants by means of meristem and tissue culture. Acta Horticulturae, 78, 389–396. https://doi.org/10.17660/ActaHortic.1977.78.49
Moutinho-Pereira, J., Correia, C. M., Gonçalves, B., Bacelar, E. A., Coutinho, J. F., Ferreira, H. F., Lousada, J. L., & Cortez, M. I. (2012). Impacts of leafroll-associated viruses (GLRaV-1 and -3) on the physiology of the Portuguese grapevine cultivar “Touriga Nacional” growing under field conditions. The Annals of Applied Biology, 160, 237–249. https://doi.org/10.1111/j.1744-7348.2012.00536.x
Naidu, R., Rowhani, A., Fuchs, M., Golino, D., & Martelli, G. P. (2014). Grapevine leafroll: A complex viral disease affecting a high-value fruit crop. Plant Disease, 98, 1172–1185. https://doi.org/10.1094/PDIS-08-13-0880-FE
OEPP/EPPO. (2008). Certification scheme. No. PM 4/8 (2): Pathogen-tested material of grapevine varieties and rootstocks. EPPO Bull., 38, 422–429.
Osman, F., Golino, D., Hodzic, E., & Rowhani, A. (2018). Virus distribution and seasonal changes of Grapevine leafroll-associated viruses. American Journal of Enology and Viticulture, 69, 70–76. https://doi.org/10.5344/ajev.2017.17032
Osman, F., Leutenegger, C., Golino, D., Rowhani, A. (2007). Real-time RT-PCR (TaqMan®) assays for the detection of Grapevine Leafroll-associated viruses 1–5 and 9. Journal of Virological Methods, 141(1), 22–29. https://doi.org/10.1016/j.jviromet.2006.11.035
Pacifico, D., Caciagli, P., Palmano, S., Mannini, F., & Marzachì, C. (2011). Quantitation of Grapevine leafroll associated virus-1 and-3, Grapevine virus A, Grapevine fanleaf virus and Grapevine fleck virus in field-collected Vitis vinifera L. ‘Nebbiolo’by real-time reverse transcription-PCR. Journal of Virological Methods, 172, 1–7. https://doi.org/10.1016/j.jviromet.2010.12.002
Padmanabhan, M. S., Goregaoker, S. P., Golem, S., Shiferaw, H., & Culver, J. N. (2005). Interaction of the tobacco mosaic virus replicase protein with the Aux/IAA protein PAP1/IAA26 is associated with disease development. Journal of Virology, 79, 2549–2558. https://doi.org/10.1128/JVI.79.4.2549-2558.2005
Peremyslov, V. V., Hagiwara, Y., & Dolja, V. V. (1999). HSP70 homolog functions in cell-to-cell movement of a plant virus. Proceeding. National Academy of Science, 96, 14771–14776. https://doi.org/10.1073/pnas.96.26.14771
Pfaffl, M. W. (2001). A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Research, 29, 45. https://doi.org/10.1093/nar/29.9.e45
R Core Team, 2022. R: A Language and Environment for Statistical Computing.
Rowhani, A., & Walker, M. S. R. (1992). Sampling strategies for the detection of grapevine fanleaf virus and the grapevine strain of tomato ringspot virus. Vitis - Journal. Grapevine Research, 31, 35–44. https://doi.org/10.5073/vitis.1992.31.35-44
RStudio Team, 2020. RStudio: Integrated Development for R. RStudio.
Schwach, F., Vaistij, F. E., Jones, L., & Baulcombe, D. C. (2005). An RNA-dependent RNA polymerase prevents meristem invasion by potato virus X and is required for the activity but not the production of a systemic silencing signal. Plant Physiology, 138, 1842–1852. https://doi.org/10.1104/pp.105.063537
Scott, M., & Leisner, R. T. (1993). Movement of virus and photoassimilate in the phloem: A comparative analysis. BioEssays, 15, 741–748. https://doi.org/10.1002/bies.950151107
Shabanian, M., Xiao, H., & Meng, B. (2020). Seasonal dynamics and tissue distribution of two major viruses associated with grapevine Leafroll under cool climate condition. European Journal of Plant Pathology, 158, 1017–1031. https://doi.org/10.1007/s10658-020-02137-z
Signorelli, S., Shaw, J., Hermawaty, D., Wang, Z., Verboven, P., Considine, J. A., & Considine, M. J. (2020). The initiation of bud burst in grapevine features dynamic regulation of the apoplastic pore size. Journal of Experimental Botany, 71, 719–729. https://doi.org/10.1093/jxb/erz200
Vega, A., Gutiérrez, R. A., Pena-Neira, A., Cramer, G. R., & Arce-Johnson, P. (2011). Compatible GLRaV-3 viral infections affect berry ripening decreasing sugar accumulation and anthocyanin biosynthesis in Vitis vinifera. Plant Molecular Biology, 77, 261–274. https://doi.org/10.1007/s11103-011-9807-8
Vigne E., Komar V., Tannières M., Demangeat G., Duchêne E., Steyer, D., Lemarquis, G., Ritzenthaler, C., & Lemaire, O. (2015). Comparative pathogenic effects of distinct Grapevine fanleaf virus strains on Vitis vinifera cvs Gewurztraminer and Chardonnay. 18th Congress of the International Council for the Study of Virus and Virus-like Diseases of the Grapevine (ICVG), Ankara, Turkey.
Waigmann, E. & Heinlein, M. (2007). Viral transport in plants. Springer Science & Business Media. https://doi.org/10.1007/978-3-540-69967-5
Walter, B., & Etienne, L. (1987). Detection of the Grapevine Fanleaf Viruses Away from the Period of Vegetation. Journal of Phytopathology, 120, 355–364. https://doi.org/10.1111/j.1439-0434.1987.tb00499.x
Wu Q., Habili N., Constable F., Al Rwahnih M., Goszczynski D. E., Wang Y., Pagay V. (2020). Virus pathogens in Australian vineyards with an emphasis on Shiraz disease. Viruses, 12, 818. https://doi.org/10.3390/v12080818
Xie, Z., Forney, C. F., & Bondada, B. (2018). Renewal of vascular connections between grapevine buds and canes during bud break. Sci. Hortic, 233, 331–338. https://doi.org/10.1016/j.scienta.2018.02.019
Youssef, S. A., Al-Dhaher, M. M. A., & Shalaby, A. A. (2009). Elimination of Grapevine fanleaf virus (GFLV) and Grapevine leaf roll-associated virus-1 (GLRaV-1) from infected grapevine plants using meristem tip culture. International Journal of Virology, 5, 89–99. https://doi.org/10.3923/ijv.2009.89.99
Acknowledgements
This work was performed in the scope of the BEST-FEET project (Ref.: 0011-1365-2021-000068), with the financial support of co-funded with FEDER funds and regional funds of the Department of Economic Development of the Government of Navarra. We thank to the ETSIAB technicians (Manuel Chocarro Martín, Josepe Irigarai Gil and Juan José Urdiáin Asensio) for their help in the phytotron experiment establishment. We would also like to show our gratitude to Oihane Oneka and María Ancín for their assistance in the lab work.
Funding
Open Access funding provided by Universidad Pública de Navarra.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Crespo-Martínez, S., Ramírez-Lacunza, A., Miranda, C. et al. Dynamics of GFLV, GFkV, GLRaV-1, and GLRaV -3 grapevine viruses transport toward developing tissues. Eur J Plant Pathol 167, 197–205 (2023). https://doi.org/10.1007/s10658-023-02703-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-023-02703-1