Abstract
More Phytophthora species are detected from environmental samples through DNA metabarcoding than are isolated by baiting. We investigated whether bulking soil samples and baiting ~ 300 g samples in standard 1 L tubs resulted in the isolation of fewer Phytophthora and Phytophythium species compared with using 15 – 20 g samples of non-bulked soil in individual 200 ml tubs. At each of four sites with dying vegetation, 50 soil samples were collected and baited separately in small tubs, followed by plating of lesioned baits over 7 days. The number of Phytophthora species obtained was compared with those obtained from bulking the 50 samples and baiting subsamples in large tubs. Half of the asymptomatic baits were plated on day 7 and the remaining were assessed for the presence of Phytophthora using metabarcoding. Root samples with rhizosphere soil from the bulked soil in each site were also assessed using metabarcoding. A higher number of Phytophthora species was recovered from each site from baiting small volumes of non-bulked soil than large volumes of bulked soils. Metabarcoding of the asymptomatic baits revealed species not isolated from lesioned baits. More species were detected from the roots using metabarcoding than were isolated from baits. Metabarcoding did not reveal any species from the rhizosphere soil and roots that were not also detected from metabarcoding of the asymptomatic baits and/or plating. The numbers of Phytopythium species detected using the different methods followed the same trend as for Phytophthora. It was concluded that baiting small samples from across a site in separate small tubs results in the isolation of a higher number of Phytophthora species than the standard technique of baiting large samples of bulked soils, and that this, together with metabarcoding of asymptomatic baits, detects the highest total number of species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The numbers of Phytophthora species detected in environmental samples through DNA metabarcoding are higher than the numbers isolated when using traditional methods such as soil baiting ( Bose et al., 2018; Burgess et al., 2017; Khaliq et al., 2018; Riddell et al., 2019; Vannini et al., 2013; Vannini et al., 2013). Many Phytophthora species identified using metabarcoding have rarely or never been isolated into culture. Some non-cultured species, such as P. taxon ballota are widespread and associated with diseased trees and known pathogenic Phytophthora species, but whether or not P. taxon ballota is a pathogen cannot be determined (Català et al., 2017). Isolation into culture is essential not only for the study of pathogenicity but also for species biology and disease management. Detection by metabarcoding of environmental DNA (eDNA) only indicates the presence of the DNA of Phytophthora in a particular habitat; it does not indicate if viable organisms are present (Burgess et al., 2017; Català et al., 2015; Khaliq et al., 2018). The use of qPCR assays based on the extraction of eDNA has raised similar issues (Kunadiya et al., 2019, 2021). In both cases, a positive detection may not indicate a living organism's presence, and consequently, the reliability of eDNA based methods for regulatory purposes is compromised.
Several factors have been investigated to determine the basis of the disparity between traditional isolation and molecular detection. One study showed that the antimicrobials in the isolation medium are not responsible for the difficulty of isolating Phytophthora species (Sarker et al., 2020). However, species vary in their timing and abundance of sporangial and zoospore production after the commencement of baiting, and species slow to produce sporangia may be difficult to detect through baiting when in competition with species that sporulate quickly (Sarker et al., 2021). The bait species used may not be appropriate for some Phytophthora species, and inoculum levels and dormant propagules that do not germinate may also affect isolation success (Drenth, 2001; Jeffers & Martin, 1986).
Baiting of soils usually involves the collection of soil samples, including fine roots from the rhizosphere of plants to a depth of 10—20 cm, bulking the collections then baiting a subsample of the bulked soils (Bose et al., 2018; Jung, 2009; Khaliq et al., 2018). Various authors have used different volumes (100 ml – 500 ml) of soil samples in tubs 1L or larger, and various baits have included leaves, petals, seeds and fruits. The soil to water ratio is usually 1:3, and the baits are floated 1–4 cm above the soil (Burgess et al., 2021).
We hypothesized that bulking the soil samples from a site results in the detection of a lower diversity of Phytophthora species than when using a small volume of soil in each bait container. Using small containers with small soil volumes should reduce competition due to either the speed at which the different species sporulate or the ease with which they colonise the baits. To test the hypothesis, we compared the number of Phytophthora species detected from soils from four sites by (1) baiting and plating of lesioned baits from small samples of non-bulked soils in small tubs, (2) baiting of samples of bulked soils from each site in standard-sized tubs, and (3) metabarcoding of root samples including adhering rhizosphere soil from the bulked soils, and samples of asymptomatic baits from small and large tubs.
Materials and methods
Soil sampling
Soil samples were collected from four sites within 35 km of Perth, Australia. In each site, some trees were declining. Sites A and B (collected in August and November 2019) were in an urban forest with mixed vegetation dominated by Eucalyptus rudis; site C (collected in February 2020) was an irrigated orchard of avocado, apple, pear and nectarine trees; site D (collected in June 2020) was an urban forest with mixed Eucalyptus species, Banksia attenuata, B. menziesii midstory, Melaleuca teretifolia and Astartea aff. fascicularis understory. From each site, 50 soil samples were collected within a 5 m radius of declining trees located at least 25 m apart. At each sampling site, approximately 5 cm of the topsoil was removed, and then approximately 100 g of soil and fine roots were collected to a depth of 15–20 cm. Each sample was placed in a separate zip-lock bag and transported to the laboratory in an insulated container.
Baiting and processing of samples for metabarcoding
On the day after soil collection, a baiting trial was set up using small and large volumes of soil in different-sized tubs. Approximately 15–20 g of soil (~ 10 cm3) along with fine roots from each of the 50 samples were placed in 50 separate 200 ml (9 cm high and 6.5 cm diameter) clear plastic tubs for baiting (Fig. 1). The remaining soils from all the collections from a site were bulked and mixed thoroughly. Small segments of randomly selected fine roots from throughout each bulked soil sample were collected and then chopped into 1–2 mm segments, and approximately 1000 mg was placed into each of three Eppendorf tubes and stored frozen at -20 ◦C until used for DNA extraction. From each of the bulked soils, approximately 300 g (~ 150 cm3) was placed in each of the three large plastic tubs (take-away containers 16.5 × 11 × 7 cm) routinely used for baiting (Fig. 1). The weight of the soil used in each container varied between the trials which had different soil types and different soil moisture contents. Water was added to a depth of 4–5 cm in large tubs and 5–6 cm in small tubs. Floating particles were removed with paper towels. Young leaves of Scholtzia involucrata, Pimelea ferruginea, Chamelaucium uncinatum, and Quercus suber were used as baits, and the number of leaves was proportional to the total surface area of water. Baiting tubs were incubated at 21 °C (± 1 °C). The bait leaves were observed daily for seven days for the appearance of lesions. Necrotic baits and those with water-soaked lesions were removed, blotted dry in between paper towels, and plated onto NARH medium: a cornmeal agar medium containing nystatin 22.72 ppm, ampicillin 100 ppm, rifampicin 10 ppm, hymexazol 50 ppm but without pentachloronitrobenzene (PCNB) (Sarker et al., 2020). After seven days, all the lesioned leaves and approximately half of the non-lesioned leaves (randomly selected) were plated for both sized containers. The remaining non-lesioned leaves from all the small tubs were bulked, blotted dry, chopped into fine pieces, placed into three Eppendorf tubes, and stored at -20 °C until DNA extraction. Non-lesioned baits from the three large containers were also bulked and stored following the same procedure.
Plates were incubated at 22 °C (± 1 °C) in the dark and observed daily for typical growth of Phytophthora. When mycelial growth of Phytophthora was observed, plates were examined under 10x magnification; hyphal tips were excised and plated onto NARH twice to eliminate any contaminants, then hyphal tips were finally transferred onto half-strength potato dextrose agar (PDA) and vegetable juice agar (V8A) plates [100 ml/L filtered vegetable juice (Campbells V8 vegetable juice; Campbell Grocery products Ltd., Norfolk, UK), 900 ml/L distilled water, 0.1 g/L CaCO3, pH adjusted to 7 and 17 g Grade A Agar (Becton, Dickenson and Company, Sparks, MD, USA] for identification of the species using morphology and DNA analysis.
Identification of isolates
All isolates from both large and small tubs were divided into morphotypes based on their colony morphology and hyphal characteristics observed at 10x magnification. Two isolates from each morphotype were selected for sequence-based identification using the internal transcribed spacer (ITS) gene region of the ribosomal DNA. DNA was extracted using ZR Fungal/Bacterial DNA Miniprep™ (Zymo Research, Irvine, California). The region spanning the ITS was amplified using the primers DC6 (Cooke et al., 2000) and ITS-4 (White et al., 1990). Templates were sequenced in both directions with primers used to amplify ITS gene regions. Clean-up of PCR products and sequencing were performed. ITS sequence data were obtained for all isolates, and their identity was confirmed initially by BLAST search in GenBank (www.ncbi.nlm.nih.gov/genbank/). The identity of the Phytophthora species was verified by phylogenetic analysis in which they were compared to the type isolates of related species (Abad et al., 2019). Several of the Phytopythium species appear to be undescribed, and a phylogenetic tree of all Phytopythium species is shown in Fig. S1, where the undescribed species were designated as Phytopythium A-E.
DNA extraction from fine roots and asymptomatic baits and metabarcoding
DNA was extracted from stored samples of fine roots with adhering rhizosphere soil and asymptomatic baits using DNeasy Plant Pro Kit (Qiagen) following the manufacturer’s protocol. For DNA extraction three replicate samples of 50 mg were taken from each bulked soil and from the bulked samples of non-lesioned leaves from the small and large baiting containers. Samples were ground using the TissueLyser LT (Qiagen, Hilden, Germany). Appropriate precautions were taken at every step of the extraction process to avoid possible contamination, and controls were also included. After extraction, all DNA was stored at -20 °C before amplicon generation. PCR and Illumina sequencing was conducted as described by Khaliq et al. (2021). Briefly, in the first PCR round, ITS 18 s rRNA gene sequences (~ 250 bp) were amplified using Phytophthora-specific primers 18h2f and 5.8RBis by nested PCR, and nested primers ITS6 and 5.8S-IR (Català et al., 2015; Scibetta et al., 2012) in the second round with Illumina MiSeq adapter sequences attached to the 5’ end. The PCRs were performed in 25 µl tubes containing 12.5 µl of PCR buffer KAPA HiFi HotStart ReadyMix (KAPA Biosystems) to minimize errors, 8 µl of PCR grade water, 1 µM of each primer and 2.5 µl of genomic DNA (first round) or 1 µl of the PCR product (second round). Negative PCR controls were included each time a PCR reaction was set up. PCR was run in several cycles with variable conditions of temperature and time. The products were discarded if any band was visualised in the negative PCR controls. First-round PCR was conducted in duplicate, and second-round PCR products were combined based on the intensity of bands on 2% agarose gels. PCR products were cleaned with Agencourt AMPure XP magnetic beads (Beckman Coulter) as described in the Illumina protocol. All the samples were then uniquely barcoded and prepared for sequencing according to the recommended protocol and sequenced on an Illumina Miseq using 500-cycle V2 chemistry (250 bp paired-end reads), following the manufacturer’s recommendation.
Bioinformatics
Paired-end reads were merged using USEARCH v10 with a minimum overlap length of 50 bp with no gaps allowed in the merged alignments. Sequence deconvolution, such as quality control and clustering, was also carried out using USEARCH v10 (Edgar, 2010). Specifically, sequences less than 200 bp and with low mean quality (< 20) were removed. Sequences that passed quality control were clustered into amplicon sequence variants (ASVs). From this point, the sequences were processed within Geneious Prime ® version 2019. 2.3 (www.geneious.com) in order to use phylogeny to assign species names. ASVs were aligned using MAFFT alignment within Geneious with default parameters. The primers are highly specific to Phytophthora but can amplify other oomycetes in the Peronosporales and Phytopythium species. For Phytophthora species, identities were first assigned to ASVs by conducting an internal blast search against a customised reference database. The reference database consisted of ITS1 sequence of 300 Phytophthora species and undescribed (but designated) taxa. A read number of 2 was accepted as evidence of the presence of a species. After that, all ASVs were separated into phylogenetic clades, and phylogenetic analyses were conducted using Geneious tree builder using verified sequences of all known Phytophthora species. An ASV was considered a putative new species if it did not match any known species in the phylogenetic analysis. These ASVs were named with a number representing the phylogenetic clade and a letter to distinguish between putative new phylotypes in the same clade. To identify ASVs of Phytopythium sp., a blast search was conducted in NCBI, and related reference sequences (including sequence of type isolates where possible) were imported into Geneious. Thereafter, the process of assigning an identity to the ASVs was the same as for Phytophthora.
Results
In total, 26 Phytophthora ASVs from eight clades were detected, of which 10 species were isolated by plating of lesioned leaves, and 14 were detected only from metabarcoding (excluding P. cryptogea and P. erythroseptica as these two species were the parent of the hybrid species isolated). The metabarcoding analysis of root samples detected 13 species of Phytophthora from sites A, B and D, (no amplification was found from site C). Of these 13 species, 4 were also isolated from symptomatic baits, while the remainder were detected only using metabarcoding of roots or asymptomatic baits (Table 1, Fig. 2).
A comparison of the number of species cultured from lesions of bait leaves in small tubs containing material from individual small volumes of soil and those from large tubs with subsamples of the bulked soil from each site showed that five species P. multivora, P. inundata, P. thermophila, P. cinnamomi, and P. pseudocryptogea were isolated by baiting both from large and small soil volumes, while an additional four were isolated only using small volumes; P. pseudorosacearum, P. personensis, P. condilina from clade 6 and P. pseudocryptogea x cryptogea from Clade 8. P. cryptogea x erythroseptica was isolated only from large tubs. Only a single species was isolated from any individual small tub, while from large tubs, different baits could show the presence of more than one species.
In addition to the species isolated through plating, an additional 14 species were found to be present on asymptomatic bait leaves analysed using DNA metabarcoding, and the number was similar using asymptomatic baits from either small or large tubs. No Phytophthora species were evident from plating non-lesioned leaves from small or large containers. Metabarcoding of subsamples of the bulked roots of each site showed 13 species which was less than the total (26) revealed by a combination of plating of lesions and metabarcoding analysis of asymptomatic baits. No species was detected solely from metabarcoding of roots (Fig. 2). However, some species detected from most sites using metabarcoding (Phytophthora capensis, Phytophthora citricola, and Phytophthora versiformis) were not isolated from lesioned baits from either the large or small tubs.
Soils from sites A, B and D, areas which had dead or dying vegetation, had a similar number of Phytophthora species present, and this was higher than the number recorded from site C, an irrigated orchard in which Phytophthora is managed through phosphite application and the addition of organic mulches. From the orchard, only one species (P. cinnamomi) was identified from baiting, though a total of six species were detected from metabarcoding of asymptomatic baits (Table 2).
A total of 11 Phytopythium species were detected, 7 from the plating of lesioned leaves, including 2 only isolated from lesions and not from metabarcoding (Table 3). As with the Phytophthora species, small volumes of non-bulked soil samples in small tubs resulted in a slightly higher number of species from lesioned leaves than from larger volumes in the large tubs, while the numbers detected using metabarcoding were higher and similar from small and large tubs. The highest total number of Phytopythium species was detected at site D and the lowest at site C (Table 4). No Phytopythium species were cultured from site B. Five putative new Phytopythium species were detected in this study (Figure S1), and four of these were cultured.
Discussion
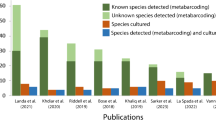
Collecting and baiting soil samples separately without bulking and mixing soils from a site can increase the number of Phytophthora species isolated from a site. A higher number of Phytophthora species (9) was recovered from the sites when individual small volumes of soil samples were baited in small tubs compared with the number recovered (6) when soil samples from each site were bulked and baited using larger volumes in large tubs. Previous work has shown significant differences between Phytophthora species in the speed of sporangial production and zoospore release (Sarker et al., 2021). When species with different timing of sporulation are bulked together, slow sporulating species are hard to isolate through baiting when in competition with fast sporulating species (Sarker et al., 2021). When small soil samples are baited separately in different tubs, there are fewer Phytophthora species present, and thus competition between those fast or slow to sporulate is reduced, resulting in a greater number of total species detected. Numerous studies have shown vastly more species to be detected by metabarcoding compared with traditional isolation methods (Khdiar et al., 2020; Landa et al., 2021; Mora-Sala et al., 2019; Riddell et al., 2019), which raises questions about the ecological roles of the species as well as why there should be such a difference in the number of species detected. For example, Khdiar et al. (2020) detected 44 Phytophthora phylotypes from remnant bushland in an urban environment, but only four species were isolated when baiting from bulked soil in large tubs. Illumina sequencing detected 23 Phytophthora species from 14 public parks and gardens in Britain, while only six species were isolated from baits (Riddell et al., 2019). However, a recent study of Amazonian forest soils reported no difference in the number of species detected using baiting or metabarcoding of baits and root samples. In their study, Legeay et al. (2020) found only six species in total, with two P. heveae-like species dominant in all sites suggesting that there were few species present at these sites and thus low interspecies competition during baiting.
In our study, the species most commonly isolated from baits (and also detected through metabarcoding) were P. multivora, P. inundata, P. pseudocryptogea and P. thermophila. Recovery of these species was achieved regardless of whether or not the soil was bulked. These species were also frequently detected by soil baiting and metabarcoding in other studies (Bose et al., 2018; Khaliq et al., 2018; Khdiar et al., 2020; Riddell et al., 2019). We have shown that P. multivora, P. pseudocryptogea and P. thermophila were among the fastest to sporulate and release zoospores from infected tissue placed in baiting water (Sarker et al., 2021). A comparison of the growth rates of mycelium on cornmeal agar showed that P. multivora, P. inundata, and P. pseudocryptogea were amongst the fastest-growing species, while P. thermophila was in the medium range (Sarker et al., 2020). The recovery of fast-growing or fast-sporulating species during baiting is not affected by whether samples are bulked or sample volume size, but manipulating these factors increases the chance of isolating the slower-growing species.
DNA metabarcoding of roots and asymptomatic baits revealed more species than could be cultured from lesioned baits. Analysis of asymptomatic baits using metabarcoding revealed several additional species, but plating samples of asymptomatic baits did not reveal any Phytophthora or Phytophythium species. Metabarcoding detection has two problems. Firstly, it cannot detect hybrid Phytophthora species, but only the parent species individually, which results in problems for species that frequently hybridise (Burgess et al., 2022). In the present study, two hybrid species P. cryptogea x erythroseptica and P. pseudocryptogea x cryptogea, were isolated by baiting. ASVs matching P. cryptogea and P. erypthroseptia could have been from either the parent species, or from a hybrid. Secondly, some pairs of species cannot be separated on the ITS gene region during metabarcoding of ITS1 gene region. Thus, in this study, ASVs corresponding to P. inundata could include P. condilina. The number of Phytophthora species detected using metabarcoding of DNA extracted directly from roots was less than the total recorded from isolations from baits and metabarcoding of asymptomatic baits. Phytophthora pseudorosacearum and P. personensis were detected through baiting of small tubs, but not detected using metabarcoding. This suggests some species are missed during sampling for metabarcoding, possibly because the extraction uses such a tiny sample size (50 mg). There are other studies in which metabarcoding has not revealed all the species isolated from baiting (Khaliq et al., 2021), but in most studies, all species from baiting have been detected using metabarcoding.
ASVs matching P. versiformis were frequently detected at all sites from roots and asymptomatic baits, but it was not isolated. When a bait is infected with multiple Phytophthora species, those with slow mycelial growth may be overgrown by fast-growing species. For example, P. versiformis was not isolated despite the baiting of small samples of non-bulked soil, and plating of both lesioned and asymptomatic baits from small tubs. P. versiformis is a very slow-growing species. Its growth rate was found 2.3 mm/day on nutrient-rich V8 agar (Paap et al., 2017) and even slower 1.27 mm/day on nutrient-poor cornmeal agar (Sarker et al., 2020). It was not observed to sporulate during 48 h from the commencement of baiting (Sarker et al., 2021). It is possible that the bait species and/or the selective agar are not optimal for this and other slow-growing species. This study also revealed 11 Phytopythium species, and the use of small tubs also resulted in the isolation of a slightly higher number of Phytopythium species than those from large tubs.
Our study showed that metabarcoding of asymptomatic baits could be a useful tool for the detection of the diversity of Phytophthora species. The asymptomatic baits have access to a much larger sample of soil and roots than that used for metabarcoding of roots, and species found on asymptomatic baits must be alive and reproducing, while identifications limited to roots may include dead or dormant species. The sampling of asymptomatic bait leaves is similar to the analysis of bait water (Khaliq et al., 2018) in that both reveal more species than detected in mycelial outgrowths from baits. While analysis of bait water detects species that produce zoospores on the day of collection and species that are not attracted to the available baits, analysis of asymptomatic baits will detect encysted zoospores from species that have sporulated over the whole period of baiting and are attracted to the baits.
Conclusions
These experiments have shown that changes in baiting and isolation protocols may result in the isolation of a greater number of Phytophthora and Phytophythium species from a site. When the objective of a survey is to determine the diversity of species in an area, individual soil collections from the site should be retained and not bulked. Baiting individual small volumes of soil in small tubs allows the isolation of a greater number of Phytophthora species than the use of larger volumes of bulked soil in large tubs. We suggest this is due to the reduced number of species in small volumes of soil enabling the slow-growing or slow-sporulating species to infect baits. Metabarcoding of asymptomatic baits revealed that additional species were present as viable propagules in the baited samples. The presence of these additional viable species not isolated from the baits indicates that further improvements in baiting conditions and isolation media are required.
Data availability
Data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Abad, G., Burgess, T., Bienapfl, J., Redford, A., Coffey, M. & Knight, L. (2019). IDphy: molecular and morphological identification of Phytophthora based on the types. http://idtools.org/id/phytophthora/factsheet_index.php (Accessed on 26 May 2021).
Bose, T., Wingfield, M. J., Roux, J., Vivas, M., & Burgess, T. I. (2018). Community composition and distribution of Phytophthora species across adjacent native and non-native forests of South Africa. Fungal Ecology, 36, 17–25.
Burgess, T. I., White, D., McDougall, K. M., Garnas, J., Dunstan, W. A., Catala, S., et al. (2017). Distribution and diversity of Phytophthora across Australia. Pacific Conservation Biology, 23, 150–162.
Burgess, T. I., López-Villamor, A., Paap, T., Williams, B., Belhaj, R., Crone, M., et al. (2021). Towards a best practice methodology for the detection of Phytophthora species in soils. Plant Pathology, 70, 604–614.
Burgess, T. I., White, D., & Sapsford, S. J. (2022). Comparison of primers for the detection of Phytophthora (and other Oomycetes) from environmental samples. Journal of Fungi, 8, 980.
Català, S., Pérez-Sierra, A., & Abad-Campos, P. (2015). The use of genus-specific amplicon pyrosequencing to assess Phytophthora species diversity using eDNA from soil and water in northern Spain. PLoS ONE, 10, e0119311.
Català, S., Berbegal, M., Pérez-Sierra, A., & Abad-Campos, P. (2017). Metabarcoding and development of new real-time specific assays reveal Phytophthora species diversity in holm oak forests in eastern Spain. Plant Pathology, 66, 115–123.
Cooke, D. E. L., Drenth, A., Duncan, J. M., Wagels, G., & Brasier, C. M. (2000). A molecular phylogeny of Phytophthora and related Oomycetes. Fungal Genetics and Biology, 30, 17–32.
Drenth, A. (2001). Practical guide to detection and identification of Phytophthora. Australian Centre for International Agricultural Research.
Edgar, R. C. (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics, 26, 2460–2461.
Jeffers, S. N., & Martin, S. B. (1986). Comparison of two media selective for Phytophthora and Pythium species. Plant Disease, 70, 1038–1043.
Jung, T. (2009). Beech decline in Central Europe driven by the interaction between Phytophthora infections and climatic extremes. Forest Pathology, 39, 73–94.
Khaliq, I., Hardy, G. E. S. J., White, D., & Burgess, T. I. (2018). eDNA from roots: a robust tool for determining Phytophthora communities in natural ecosystems. FEMS Microbiology Ecology, 94, fiy048.
Khaliq, I., Burgess, T. I., Hardy, G. E. S. J., White, D., & McDougall, K. L. (2021). Phytophthora and vascular plant species distributions along a steep elevation gradient. Biological Invasions, 23, 1443–1459.
Khdiar, M. Y., Barber, P. A., Hardy, G. E. S., Shaw, C., Steel, E., McMains, C., et al. (2020). Association of Phytophthora with declining vegetation in an urban forest environment. Microorganisms, 8, 973.
Kunadiya, M. B., Dunstan, W. D., White, D., Hardy, G. E. S. J., Grigg, A. H., & Burgess, T. I. (2019). A qPCR assay for the detection of Phytophthora cinnamomi including an mRNA protocol designed to establish propagule viability in environmental samples. Plant Disease, 103, 2443–2450.
Kunadiya, M. B., Burgess, T. I., Dunstan, W., White, D., & Hardy, G. E. S. (2021). Persistence and degradation of Phytophthora cinnamomi DNA and RNA in different soil types. Environmental DNA, 3, 92–104.
Landa, B. B., Arias-Giraldo, L. F., Henricot, B., Montes-Borrego, M., Shuttleworth, L. A., & Pérez-Sierra, A. (2021). Diversity of Phytophthora species detected in disturbed and undisturbed British soils using High-Throughput Sequencing targeting ITS rRNA and COI mtDNA regions. Forests, 12, 229.
Legeay, J., Husson, C., Boudier, B., Louissana, E., Baraloto, C., Schimann, C., et al. (2020). Surprising low diversity of the plant pathogen Phytophthora in Amazonian forests. Environmental Microbiology, 22, 5019–5032.
Mora-Sala, B., Gramaje, D., Abad-Campos, P., & Berbegal, M. (2019). Diversity of Phytophthora species associated with Quercus ilex L. in three Spanish regions evaluated by NGS. Forests, 10, 979. https://doi.org/10.3390/f10110979
Paap, T., Croeser, L., White, D., Aghighi, S., Barber, P., Hardy, G. E. S., et al. (2017). Phytophthora versiformis sp. nov., a new species from Australia related to P. quercina. Australasian Plant Pathology, 46, 369–378.
Riddell, C. E., Frederickson-Matika, D., Armstrong, A. C., Eliot, M., Forster, J., Hedley, P. E., et al. (2019). Metabarcoding reveals a high diversity of woody host-associated Phytophthora spp. in soils at public gardens and amenity woodlands in Britain. PeerJ, 7, e6931.
Sarker, S. R., McComb, J., Burgess, T. I., & Hardy, G. E. S. J. (2020). Antimicrobials in Phytophthora isolation media and the growth of Phytophthora species. Plant Pathology, 69, 1426–1436.
Sarker, S. R., McComb, J., Burgess, T. I., & Hardy, G. E. S. J. (2021). Timing and abundance of sporangia production and zoospore release influences the recovery of different Phytophthora species by baiting. Fungal Biology. https://doi.org/10.1016/j.funbio.2021.01.009
Scibetta, S., Schena, L., Chimento, A., Cacciola, S. O., & Cooke, D. E. (2012). A molecular method to assess Phytophthora diversity in environmental samples. Journal of Microbiological Methods, 88, 356–368.
Vannini, A., Bruni, N., Tomassini, A., Franceschini, S., & Vettraino, A. M. (2013). Pyrosequencing of environmental soil samples reveals biodiversity of the Phytophthora resident community in chestnut forests. FEMS Microbiology Ecology, 85, 433–442.
White, T. J., Bruns, T., Lee, S., & Taylor, J. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, & T. J. White (Eds.), A guide to methods and applications. Academic Press.
Acknowledgements
We thank Diane White (CPSM, Murdoch University, WA) for her technical help in molecular works and Sarah Sapsford (School of Biological Sciences, University of Canterbury, Christchurch, New Zealand) for providing technical assistance on the bioinformatics analysis.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions This research was supported by a Research Training Program Scholarship, from Murdoch University, Australia.
Author information
Authors and Affiliations
Contributions
Suchana R. Sarker conducted the experimental work, all authors contributed to the design of the experiments, interpretation of results and production of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
We confirm that we have no conflicts of interest to disclose.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sarker, S.R., McComb, J., Hardy, G.E.S.J. et al. Sample volume affects the number of Phytophthora and Phytopythium species detected by soil baiting. Eur J Plant Pathol 166, 303–313 (2023). https://doi.org/10.1007/s10658-023-02661-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-023-02661-8