Abstract
Many approaches for modelling the biodiversity and ecosystem function (BEF) relationship have been developed over recent decades. Diversity-Interactions modelling, a regression-based approach, models the BEF relationship by expressing ecosystem functions as a linear combination of species-specific effects, species’ proportions, and species’ interactions. The species interactions in a Diversity-Interactions model can take different forms (e.g., a unique interaction term for each pair of species, or a single interaction term for any pair of species) and may include a non-linear parameter (\(\theta\)) as an exponent to the species interactions to capture non-linear relationships, giving rise to Generalized Diversity-Interactions (GDI) modelling. The structure of the interaction terms describes the underlying biological processes in the ecosystem, while the value of \(\theta\) can determine the shape of the BEF relationship. When fitting GDI models, it is unclear whether one should choose the interaction structure first and then estimate θ, or vice versa. It is also unknown whether the estimate of \(\theta\) is robust to changes in the structure of the linear interaction terms of the model. Using a simulation study, we test the robustness of \(\theta\) and compare multiple model selection approaches to identify an optimal and computationally efficient model selection procedure for GDI models. Results show that the estimate of \(\theta\) is robust and remains unbiased regardless of changes in the underlying structure of interaction terms, and that the most efficient model selection procedure is to first estimate \(\theta\) for one interaction structure and then reuse this estimate for the other interaction structures.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
An ecosystem function is a response measured on an ecosystem that may directly or indirectly capture the goods and services provided by the ecosystem (de Groot et al. 2002). Interest in quantifying the relationship between biodiversity and ecosystem functions (BEF) has driven a wealth of experiments and associated statistical modelling approaches over recent decades (Hector et al. 1999; Loreau and Hector 2001; Schmid et al. 2002; Bell et al. 2009; Kirwan et al. 2007; Cardinale et al. 2009; Isbell et al. 2011). Studies have shown that increasing the biodiversity of an ecosystem can improve the performance and stability of ecosystem functions across a range of ecosystem types (Tracy et al. 2004; Bell et al. 2005; Hooper et al. 2005; Balvanera et al. 2006; Worm et al. 2006; Cardinale et al. 2007; Finn et al. 2013; Gamfeldt et al. 2013). Species richness is often assumed to be the main driver of the BEF relationship (Spehn et al. 2005; Tillman et al. 1997); however, community evenness, species’ relative abundances, and their functional groupings may also be strongly influential (Wilsey and Polley 2004; Reich et al. 2004; Ebeling et al. 2014; Lembrechts et al. 2018). The Diversity-Interactions (DI; Kirwan et al. 2009; Brophy et al. 2011, 2017; Dooley et al. 2015) and Generalized Diversity-Interactions approaches (GDI; Connolly et al. 2013) model the BEF relationship using an alternative definition of species diversity by capturing species-specific effects, species’ abundances, and species’ interactions, in addition to richness patterns. The DImodels package (Moral et al. 2022) available for R software (R Core Team 2021) can be used to fit and compare DI and GDI models. The species interactions in Diversity-Interactions models can range in complexity from a single interaction term (assuming all pairs of species interact in the same way) to many interaction terms (e.g. assuming a separate interaction for all pairs of species) (Kirwan et al 2009) and may also include a non-linear parameter (Connolly et al. 2013). The structure of these interaction terms provides insight into the underlying biological processes and thus the proper estimation of the interaction structure is important. In this paper, we (1) explore and test (via simulation) the robustness of the non-linear parameter, and (2) compare different model fitting approaches for selecting the best interaction structure for Generalized Diversity-Interactions models in a computationally efficient way.
DI and GDI modelling are regression-based approaches that use species proportions and their interactions (defined as being proportional to the products of pairs of species proportions) as predictors to capture species diversity effects. Additional block or treatment effects may also be included. The interaction terms can take several different forms (Kirwan et al 2009) and may include a non-linear parameter if species interactions are not directly proportional to the product of their proportions, giving rise to Generalized Diversity-Interactions models (Connolly et al 2013). Figure 1 shows the effect of the non-linear parameter (\(\theta\)) on species interactions and model interpretation in GDI models using a hypothetical two-species example; the parameter \(\theta\) affects the realisation of the species interaction effects and can change the shape of the BEF relationship across the species proportions gradient. Model selection is an important part of modelling as it aids in a better understanding of the response-predictor relationship as well as the identification of significant and non-significant predictors (Mitchell and Beauchamp 1988). In the absence of the non-linear parameter (\(\theta\) set equal to 1, not estimated), model selection for the interaction terms in DI models can be carried out through a series of hierarchical comparisons (Kirwan et al 2009). Further, there is a plethora of techniques available to perform model selection for linear regression; these include F-tests, AIC (Akaike 1973), BIC (Schwarz 1978), stepwise regression (Breaux 1967), etc. The inclusion of the non-linear term in GDI models complicates the model selection process: should the user first identify the most appropriate interaction structure and then estimate \(\theta\) for only the selected interaction structure, or should they first estimate \(\theta\) for a particular interaction structure and then reuse this estimate for the remaining interaction structures, or should \(\theta\) be estimated for each interaction structure before an appropriate interaction structure can be selected. Estimating \(\theta\) for each interaction structure would be desirable, but is also computationally expensive as \(\theta\) may have to be re-estimated for any change to the interaction terms. Hence for increased user-friendliness we explore the viability of the following three possible approaches to selecting the best model: (a) Select the appropriate interaction structure first by ignoring \(\theta\) (i.e., assuming \(\theta =1\)) and then estimate and test for the inclusion of \(\theta\), (b) Estimate \(\theta\) and test its inclusion for the simplest interaction structure (average pairwise; Table 1) first and then reuse that estimate to fit the remaining interaction structures and select the best model, and (c) Estimate \(\theta\) and test its inclusion for each interaction structure and then perform model selection. Approach (c) is the most exhaustive method for model selection, but is computationally expensive (see Table A1 in Online Appendix A for comparison of model selection times for these three approaches for data from an experiment with 72 species), while approaches (a) and (b) aim for efficiency but rely on \(\theta\) being invariant across varying specifications of species interactions, making it important to test whether the estimate of \(\theta\) is robust to changes in the structure of the interaction terms of the model. In this paper, we address the following two questions using a simulation study:
-
1)
Is the estimation of the non-linear parameter (\(\theta\)) of a GDI model affected by changing the structure of the interactions?
-
2)
What is the optimal and most computationally efficient model selection process for GDI models?
(Adapted from Grange et al. 2021). Illustration of the impact of the non-linear parameter (\(\theta\)) on species interactions and interpretation in Generalized Diversity-Interactions models: A hypothetical two-species mixture is considered, with the response being yield, giving the equation \( \widehat{\mathrm{y}}={\beta }_{1}{\mathrm{P}}_{1}+ {\beta }_{2}{\mathrm{P}}_{2}+\delta {\left({\mathrm{P}}_{1}{\mathrm{P}}_{2}\right)}^{\theta }\). The value of the response is expressed as a combination of the species identities and species interactions for all possible communities involving two species, ranging from a monoculture of species 1 (on left) to a monoculture of species 2 (on right) and all possible two-species mixtures in between. \({\beta }_{1}\)and \({\beta }_{2}\) are ‘identity effects’ for species 1 and species 2 respectively and are the expected performances of each species in monocultures. The expected performance of mixtures is the weighted average of the identity effects (\({\beta }_{1}{\mathrm{P}}_{1}+ {\beta }_{2}{\mathrm{P}}_{2}\)) plus the interaction effect (\(\delta *{\left({\mathrm{P}}_{1}{\mathrm{P}}_{2}\right)}^{\theta }\)). The non-linear parameter (\(\theta\)) feeds into the interaction effect as an exponent which scales the product of the species proportions (\({\left({\mathrm{P}}_{1}{\mathrm{P}}_{2}\right)}^{\theta }\)). Four different scenarios for the expected response are presented: (a) No interaction term (\(\delta =0\), \(\theta\) omitted), (b) No non-linear parameter (\(\theta =1\)), (c) \(\theta =0.3\), and (d) \(\theta =1.3\); in each panel the identity effects are kept constant, in panels (b)–(d) the value of the interaction parameter (\(\delta\)) is kept constant (and a positive value used). The expected ecosystem function for an example 0.75:0.25 mixture is shown across the four scenarios and is computed as \({\beta }_{1} \times 0.75 + {\beta }_{2} \times 0.25 +\delta \times {\left(0.75 \times 0.25\right)}^{\theta }\). These concepts scale up for systems with more than two species
2 Review of DI and GDI models
The DI modelling framework (Kirwan et al. 2009) models the BEF relationship by expressing an ecosystem function response as a linear function of the relative abundances of the species spread across the simplex space (Kirwan et al. 2007; Cornell 2011). BEF data suitable for applying the DI models framework would include a range of experimental units (species communities) where species diversity is manipulated across dimensions such as species composition (identity), richness, and/or evenness to assess the impact of these variables on the ecosystem function. It is also possible to apply the DI modelling approach to appropriate observational data. The general formulation of a DI model is
The response (y) is a community-level ecosystem function (e.g., biomass or weed resistance in a grassland ecosystem). The Identities and the Interactions components are the species-specific and the species interaction effects on the response, respectively, and are incorporated in the model using the initial proportions of the species and their products, respectively. Structures (experimental structures) are additional covariates or factors to capture experimentally manipulated treatments or blocks, or other measured descriptors of the experimental units. \(\varepsilon\) is a normally distributed error term.
Connolly et al. 2013 showed that modifying the formulation of the species interaction terms in DI models leads to Generalised Diversity-Interactions (GDI) models that provide a more flexible framework for modelling BEF relationships. GDI models incorporate all the benefits of DI models and provide deeper insight into how individual pairs of species interact and by extension, affect community-level responses whilst also enabling us to explore phenomena such as the effects of diversity loss, functional stability, saturation properties of the BEF relationship, and transgressive overyielding (Connolly et al. 2013). DI models characterise the contribution of two species \(i\) and \(j\) to an ecosystem function as being proportional to the product of their relative abundances (\({P}_{i}{P}_{j}\)), while GDI models assume a more general form for this contribution as \({\left({P}_{i}{P}_{j}\right)}^{\theta }\), where \(\theta\) is an additional parameter allowing for non-linearity in the relationship between the response and the interactions. A possible GDI model is:
where \({P}_{i}\) is the sown proportion of species \(i\), \(s\) is the number of species in the system, \({\beta }_{i}\) is the identity effect of species \(i\), the \({\delta }_{ij}\) parameters are the effects of the interactions between species \(i\) and \(j\), \(A\) is a vector (or matrix) of experiment structures, \(\alpha\) is a vector containing the effects of the experimental structures, and \(\epsilon\) is a normally distributed error term with mean 0 and variance \({\sigma }^{2}\), i.e. \(\varepsilon \sim N\left(0,{\sigma }^{2}\right)\). This variance is assumed to be constant, but it could be affected by the community structure; for example, it could differ for monoculture and mixture communities (Brophy et al. 2017; Cummins et al. 2021). \(\theta\) is a non-linear parameter that can affect the nature of the relationship between the species interactions and the ecosystem function (Fig. 1).
Equation (2) can be adjusted in multiple different ways by modifying the specification of the interaction terms to describe different biological hypotheses. These adjustments serve the purpose of reducing the number of interaction terms when the species pool is large. Table 1 gives a list of the several different GDI models (Kirwan et al. 2007, 2009; Connolly et al. 2013) along with their equations and the biological aspects that they describe. These models are a subset of a range of different possible models. Traditional model selection methods using F-tests or information criteria can be used to select the best model which strikes a balance between parsimony and explaining the BEF relationship. The models in Table 1 can be further expanded by crossing the identities and interaction terms with the variables such as year or treatment, as appropriate. They could also be extended to have multivariate responses (Dooley et al. 2015) or to include random pairwise interaction effects for modelling numerous species interactions over a single (Brophy et al. 2017) or multi-year setting (McDonnell et al. 2023). Diversity-Interactions models have been widely used in understanding the BEF relationship in several experiments where the diversity was varied across the flora, fauna, or bacteria within the ecosystem (Kirwan et al. 2007; Connolly et al. 2009, 2011, 2018; Frankow-Lindberg et al. 2009; Nyfeler et al. 2009; O’Hea et al. 2010; Brophy et al. 2011; Grange et al. 2022). A key advantage that DI and GDI models have over the other approaches, such as richness-only or anova models, is that they can be used to make predictions for the entire simplex space (provided the initial communities were sufficiently spread across the simplex space).
The \(\theta\) parameter forms an integral part of GDI models. A value of \(\theta =1\) describes a linear interaction, proportional to the product of the species proportions, whilst a value of \(\theta <1\) corresponds to a stronger than expected contribution of species’ pairs to ecosystem functioning, particularly at low abundances of the species, resulting in a stronger interaction effect. This is akin to a scenario where there is a strong niche separation of resources between the species resulting in little or no interspecific competition for the resources and is highlighted in Figs. 1 and 2, which show the impact of varying the \(\theta\) parameter in two-species and three-species systems, respectively. Varying the value of \(\theta\) affects the shape of the BEF relationship; e.g. in the middle column of ternary diagrams in Fig. 2, for small \(\theta\) values the species interaction effect is flatter for a larger range of communities across the entire simplex, in contrast to high \(\theta\) values where the interaction effect is high for some communities in the centre and then declines as we move away from these central communities.
Ternary diagram illustrations of the effect of \(\theta\) on an ecosystem function response in GDI models (final column) for a range of \(\theta\) values and decomposed into the identity component (first column) and interactions component (middle column). A single dataset was simulated from a three-species design assuming the average pairwise model, with identity effect coefficient values \({\beta }_{1}=9, {\beta }_{2}= 5, {\beta }_{3}=3\), for species 1, 2 and 3 respectively (denoted S1, S2 and S3 in the ternary diagrams), a value of 9 for the average interaction effect (\({\delta }_{AV}\)), and \(\theta\) equal to 0.8. The random error term is added to the response from a normal distribution with \(\mu =0\) and \(\sigma\) = 0.8. Five versions of the average pairwise interaction model were then fit to this data, but where \(\theta\) was not estimated, instead it was fixed for a range of values: (a) 0.3, (b) 0.7, (c) 1, (d) 1.15, and (e) 1.3 giving us five different estimated models (estimates of the \({\beta }_{i}\)’s and \({\delta }_{AV}\) only and not the \(\theta\) parameter). The model predictions across the simplex space are shown for each of these models, as well as the decomposition into identities and interactions components. The model parameter estimates are also shown for each row. The identity effects aren’t strongly influenced by forcing the value of \(\theta\) to change; however, the interaction component (and hence the total response) changes considerably depending on the forced value of \(\theta\). For low values of \(\theta\), the interactions (and hence total response) are high and are quite flat over a wide range of communities across the simplex
3 Methods
A simulation study was performed under two different experimental designs, one with four species and one with nine species. Under both designs, the true underlying model was assumed to be the full pairwise model with equation
where \(s=4\) for the four-species design and \(s=9\) for the nine-species design, the \({P}_{i}\)’s were the proportions of the respective species, \({\beta }_{i}\)’s were the identity effects of the species, and \({\delta }_{ij}\) was the interaction effect between species \(i\) and \(j\), with \(\theta\) and \(\epsilon\) being the non-linear parameter and the random normal error term respectively.
For the four-species simulations, the species \(S{P}_{1}\), \(S{P}_{2}\), \(S{P}_{3}\), and \(S{P}_{4}\) were spread across the simplex space in a design that consisted of 37 different communities (shown in Fig. 3). The design comprised of 15 equi-proportional and 22 imbalanced mixtures. The equi-proportional communities included the four monocultures, six two-species mixtures (50%, 50%, 0%, 0%), four three-species mixtures (33.33%, 33.33%, 33.33%, 0%), and one centroid community (25%, 25%, 25%, 25%). The imbalanced communities included four mixtures with each species being dominant in turn at three different levels of dominance (90%, 3.33%, 3.33%, 3.33%), (70%, 10%, 10%, 10%) and (40%, 20%, 20%, 20%), six mixtures with two species being dominant in turn at (40%, 40%, 10%, 10%), and four mixtures with three species being dominant in turn at (30%, 30%, 30%, 10%). For each of these 37 communities, three values of the response were simulated, giving a design with 111 data points (per simulated dataset). The four species were assumed to be grouped into two functional groups (groupings based on the function they perform) with \(S{P}_{1}\) and \(S{P}_{2}\) being in the first functional group and \(S{P}_{3}\) and \(S{P}_{4}\) being in the second functional group.
Graphical representation of the 4 species simplex design: Each point in the tetrahedron represents a four-species community and its position is determined by the relative abundances of the species (P1, P2, P3, and P4). The points are coloured according to the richness level of the community. The black points represent communities where richness = 1, which are the monocultures and are positioned at the vertices of the tetrahedron. The red points represent the two-species mixtures and are positioned at points along the edges of the tetrahedron, determined by the relative abundances of the two species. The green points represent the three-species mixtures and are positioned along the faces of the tetrahedron. Finally, the blue points represent the four-species mixtures and are placed in the interior of the tetrahedron according to the relative proportions of the species
For the nine species simulations, the species were named \(S{P}_{1}\), \(S{P}_{2}\), \(S{P}_{3}\), …, \(S{P}_{9}\) and the design opted was the same as the one in the Jena dominance experiment (Roscher et al. 2005). There were nine monocultures, 36 two-species communities, 24 three-species communities, 18 four-species communities, 12 six-species communities, and one centroid community with nine species. This resulted in a simplex design with 100 unique equi-proportional communities. For each of these 100 communities three values of the response were simulated, giving a design with 300 data points (per simulated dataset). The functional grouping structure assumed was \(S{P}_{1}\), \(S{P}_{2}\), \(S{P}_{3}\), \(S{P}_{4}\), and \(S{P}_{5}\) being in functional group one, \(S{P}_{6}\) and \(S{P}_{7}\) being in functional group two and \(S{P}_{8}\) and \(S{P}_{9}\) being in functional group three. The full simplex design for both the four- and nine-species simulations are shown in Figs. A1 and A2 in Online Appendix A.
The species identities and interaction effects used to simulate the responses for the four- and nine-species models are shown in Online appendix Table A2a and Table A2b respectively. The identity effects for the four- and nine-species models were simulated from \(N\left(3, 9\right)\) and \(N\left(7, 4\right)\) distributions, respectively, and were rounded off to integers. To give a net non-zero interaction effect between the species, the interaction effects for the four-species model were simulated from a \(N\left(8, 16\right)\) distribution, while for the nine-species model they were simulated from a \(N\left(9, 36\right)\) distribution (the means and variances for these distributions were chosen to reflect species identity and interaction effects similar to what has been observed in real-world grassland biodiversity experiments measuring annual above ground biomass as the response e.g. Kirwan et al. 2007). The functional groupings of the species weren’t taken into consideration when simulating the interaction coefficients for these datasets as the true model was assumed to be the full pairwise model.
The response variable was simulated (using Eq. (3) and the values in Online Appendix Table A2) for ten different \(\theta\) values ranging from 0.05 up to 1.33. A random normal error term with mean 0 and constant standard deviation \(\sigma\) was then added to the response, \(\sigma\) was varied to have five different values: 0.8, 0.9, 1.0, 1.1 and 1.2. This gave us a total of 100 settings for the full simulation study (2 experimental designs × 10 \(\theta\) values × 5 \(\sigma\) values). A total of 200 datasets were simulated for each setting; datasets for the four species experimental design each consisted of 111 rows (37 communities * 3 replicates) and datasets for the nine species experimental design consisted of 300 rows (100 communities * 3 replicates). These datasets were simulated in R (version 4.0.3) and reproducible scripts are available in Online Appendix D and at https://github.com/rishvish/Theta-Simulation-Study. Increasing the number of simulated datasets up to 1000 was tested for a small number of settings, but the results stabilised at around 200 simulations and hence 200 was chosen as the number of datasets to simulate per setting (results for these preliminary simulations have not been provided).
To test the robustness of \(\theta\) estimation across different interaction specifications, the final four GDI models in Table 1 (the average pairwise, functional group, additive species and full pairwise models) were fit to each of the 200 simulated datasets across each of the 100 simulation settings using the DI function from the DImodels package (v1.2; Moral et al. 2022) in R (R Core Team 2021). The value of \(\theta\) was estimated by maximising the profile log-likelihood using the DImodels package and the distributional properties of the estimator were assessed graphically to determine whether the estimate of \(\theta\) differed across the different models and from its true underlying value. Profile log-likelihood confidence intervals (CI) were also calculated. This is the interval for \(\theta\) where the log-likelihood function \(l\left(\theta \right)\) is greater than \({l}_{max}\left(\theta \right)-0.5\times {\chi }_{1-\alpha }^{2}\left(1\right)\). Here, \({l}_{max}\left(\theta \right)\) is the maximum log-likelihood value and \({\chi }_{1-\alpha }^{2}\left(1\right)\) is the \(\left(1-\alpha \right)\times 100\%\) percentile of the chi-squared distribution with 1 d.f. (Morgan 1992, p.63). The corresponding coverage of the CI was assessed by taking the proportion of times that the true value of \(\theta\) fell within the computed CI.
To explore the efficacy of different model selection procedures, using the same 200 simulated datasets for each of the 100 simulation settings, we checked the proportion of times that the true underlying interaction structure was selected as the best model using three different model selection procedures. Table 2 gives a detailed description of these model selection procedures.
Additional simulations for both the robustness of \(\theta\) estimation and the model selection efficacy were carried out under different conditions, including a high number of species (up to 72), the presence of experimental structures, different true underlying models to the full pairwise interaction model, different structure of functional groupings, and higher variance of error terms (results presented in Online Appendix C). Simulations were also performed to test the robustness of a re-parameterisation of \(\theta\) suggested by Connolly et al. 2018, where the \(\delta\) coefficients are scaled by a factor of \(\frac{2{s}^{2\theta } }{s\left(s-1\right)}\) to reduce the correlation between \(\theta\) and \(\delta\) coefficients (results presented in Online Appendix B).
4 Results
For each of the four models (average pairwise interaction, functional group effects, additive species contributions, and full pairwise interactions) fit to the datasets in the simulation study, the mean estimate of \(\theta\) was almost identical and was approximately equal to the true value of \(\theta\) (Fig. 4 and Table 3). Splitting the results up by the five \(\sigma\) values (0.8, 0.9, 1, 1.1, and 1.2), it was found that the results were invariant to a changing \(\sigma\), with the only effect of \(\sigma\) being an increase in the variation of the distribution of the estimates of \(\theta\) as the value of \(\sigma\) increased (select results shown in Online Figure A3 in Appendix A). The results obtained from the study were similar for both the four- and nine-species cases. The mean estimate of \(\theta\) was approximately equal to the true value of \(\theta\) and the average coverage of the 95% confidence interval for the estimated \(\theta\) was unusual for low values of \(\theta\) and approached 0.95 as the value of \(\theta\) increased, for both the four- and nine-species cases (select results shown in Table 3; full results shown in Online Table A4 in Online Appendix A). The unusual coverages for low \(\theta\) values were due to a combination of convergence problems near the boundary of \(\theta =0\) and the interval being too precise (see Online Appendix A for more details). The standard deviations of the estimates for \(\theta\) tend to increase as the true value of \(\theta\) increases. This is because we are simulating different datasets for each value of \(\theta\) and the range of the response variable is different for each value of \(\theta\), which causes the change in the standard deviations of the estimates. Scaling the standard deviations of predicted estimates by the interquartile range for each unique \(\theta\)-model combination results in the standard deviations being similar for each unique \(\theta\)-model combination (Figure A5 in Online Appendix A).
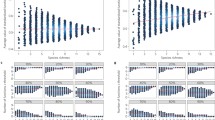
Mean estimated \(\theta\) vs true \(\theta\): (a) For the four-species model and (b) For the nine-species model. The black line in the centre is the x = y line. The \(\theta\) estimates of each of the four models (average pairwise, functional group, additive species, and full pairwise) for the different \(\theta\) values across the 1000 realizations (200 simulations × 5 \(\sigma\) values) are averaged and represented as points. The corresponding bands around each point give the 95% dispersion of the respective estimate of \(\theta\) (calculated using the 2.5% and 97.5% percentiles of the estimates of \(\theta\)). In each case, the true underlying model was the full pairwise interactions model
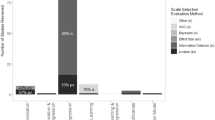
The simulations for testing the efficacy of different model selection methods showed that method ‘b’, where we first estimate \(\theta\) and then select the best interaction structure, was better than method ‘a’, where we select the appropriate interaction structures first and then estimate the value of \(\theta\). For model selection procedure ‘a’, we found that for lower values of \(\theta\) (\(\theta <0.5\)), irrespective of the number of species and the underlying true structure of interaction terms, the true underling interaction structure was hardly ever selected, instead the average pairwise interaction model was selected as the chosen model almost 100% of the time (Figure A6 in Online Appendix A). However, as the value of \(\theta\) increased, the proportion of times that the true underlying (full pairwise in our example) interaction structure was chosen increased (Table 4). Different selection metrics besides AIC, like F-tests and BIC, were also tested, but similar results were observed. A possible reason for this could be that for low values of \(\theta\) (\(\theta <0.5\)), the initial assumption of \(\theta\) being equal to 1 is incorrect, and thus all estimated models fit the data poorly and the selection criteria end up selecting the model with the simplest structure, resulting in the average pairwise interaction model being selected every time. As the value of \(\theta\) increases over 0.5, the initial assumption of \(\theta\) being 1 isn’t far off from the true value of \(\theta\) and thus the models fit the data better and the selection metrics have more power to select the best interaction structure.
Model selection procedures ‘b’ and ‘c’ offered an improvement on this as instead of assuming \(\theta\) to be 1, we first estimate it for a specific interaction structure and then reuse that estimate of \(\theta\) to fit the remaining interaction structures in method ‘b’ or estimate \(\theta\) separately for each interaction structure in method ‘c’. Thus, these model selection procedures outperformed method ‘a’ and selected the true underlying interaction structure most of the time across the different values of \(\theta\) (Table 4). The proportion of times the true interaction structure gets selected did decrease sometimes as the value of \(\sigma\) increased (see Figures A6, A7, and A8 in Online Appendix A) and this can be expected as increasing \(\sigma\) had the effect of increasing the noise in the data. Similar results were observed when testing with different selection criteria besides AIC. Comparing the efficacy that model selection procedures ‘b’ and ‘c’ showed that method ‘b’ had comparable performance to method ‘c’ whilst giving a four- to seven-fold (depending on the number of species in the experiment) reduction in computation time (Figure A9 and Table A1 in Online Appendix A). Thus, our recommendation is to use approach ‘b’ for model selection within the Diversity-Interactions modelling framework.
Simulations testing additional scenarios like a higher number of species (up to 72 species), a different true underlying interaction structure, a different structure of functional groups, the presence of additional experimental structures, and the reparameterisation of \(\theta\), all yielded similar results in that the mean estimate of \(\theta\) was approximately equal to the true value of \(\theta\) and didn’t differ much across the four (average pairwise interactions, functional group effects, additive species contributions, and full pairwise interactions) estimated models. Results for model selection too were similar to the results observed for the four- and nine-species cases (Tables and figures presented in Online Appendix B for \(\theta\) reparameterization and Online Appendix C for all other factors).
5 Discussion
Our simulation study showed that the estimator of \(\theta\) is unbiased, with the estimate of \(\theta\) being robust across the different interaction structures in GDI models. This robustness of \(\theta\) is consistent across a wide range of \(\theta\) values (0.05 up to 1.3) and a variety of different scenarios, including changes to the number of species (up to 72), true underlying interaction structure, functional grouping of species, presence of experimental structures, and crossing of interaction terms with experimental structures.
The results of our study give conclusive evidence for the robustness of \(\theta\) and thus help us in deducing a model fitting procedure for GDI models which is parsimonious and informative. We recommend first estimating \(\theta\) using profile likelihood and testing its inclusion for the simplest interaction structure (average pairwise) and then using that estimate of \(\theta\) to fit the remaining interaction structures and perform model selection. This approach is desirable due to the speed (compared to the approach of estimating \(\theta\) for each interaction structure (method ‘c’)) with which we can converge to an appropriate GDI model and is used by the autoDI function in version 1.2 of the DImodels R package to perform model selection (note that the autoDI function in earlier versions 1.0 and 1.1 of the DImodels R package used method ‘a’ for model selection but we have switched to this recommended method of model selection from version 1.2). There is precedent for fixing a model parameter to a specific value when performing model selection in wider statistics. For example, in negative binomial models, the dispersion parameter may be fixed when testing for model effects; in generalized linear mixed models, including observational-level random effects (e.g. Poisson-normal, see Demétrio et al. 2014), the variance of the random effects may also be fixed when testing fixed effects. We note that joint estimation of all parameters in a GDI model using a non-linear estimation framework would also be possible, however, our profile likelihood solution scales up well when the data structure becomes more complex (e.g., multiple responses, multiple time points and/or multiple study locations).
The robustness in \(\theta\) estimation does rely on a few prerequisites being satisfied. Firstly, the species proportions should be reasonably spread around the simplex space (rather than restricted to a small subspace). Further, if there is lack-of-fit in the models or if all, or a majority, of the interaction terms aren’t significant then this would have trickle-down effects on \(\theta\) and its estimation would be affected, resulting in estimates which are considerably different across the different GDI models while at the same time being far off from the true underlying value of \(\theta\). Thus, it is important to check for any data issues and model fit before finalising model selection. Examples with such issues are highlighted in Online Appendix C. In this study, we have examined over 300,000 datasets, which gives assurance in the reliability of our results. Our study assumed a single \(\theta\) parameter across all interaction terms; there is scope for allowing different \({\theta }\)’s for each interaction term in GDI models, perhaps in a multivariate or repeated measures setting with complex variance structures.
6 Conclusion
The aim of our research was to discern if the non-linear parameter (\(\theta\)) in GDI models was robust to changes in the structure of the underlying interaction terms and compare different multiple model selection approaches to identify an optimal and computationally efficient model selection procedure for GDI models. The results of our simulation study show that for our experimental designs, \(\theta\) estimation is invariant to different interaction structures and that the most efficient model selection procedure for GDI models is to first estimate \(\theta\) for the simplest interaction structure (average pairwise) and test for its significance, and then use that estimate to fit the different interaction structures and finally select the most appropriate structure using selection criteria.
References
Akaike H (1973) Information theory and an extension of the maximum likelihood principle. In: Petrow BN, Czaki F (Eds.), Proceedings of the 2nd international symposium on information. Akademiai Kiado, Budapest
Balvanera P, Pfisterer AB, Buchmann N, He JS, Nakashizuka T, Raffaelli D, Schmid B (2006) Quantifying the evidence for biodiversity effects on ecosystem functioning and services. Ecol Lett 9(10):1146–1156
Bell T, Newman JA, Silverman BW, Turner SL, Lilley AK (2005) The contribution of species richness and composition to bacterial services. Nature 436:1157–1160
Bell T, Lilley AK, Hector A, Schmid B, King L, Newman JA (2009) A linear model method for biodiversity-ecosystem functioning experiments. Am Nat 174:836–849
Breaux HJ (1967) On stepwise multiple linear regression. Army Ballistic Research Lab Aberdeen Proving Ground MD.
Brophy C, Connolly J, Fagerli IL, Duodu S, Svenning MM (2011) A baseline category logit model for assessing competing strains of rhizobium bacteria. J Agric Biol Environ Stat 16:409–421
Brophy C, Dooley Á, Kirwan L, Finn JA, McDonnell J, Bell T, Cadotte MW, Connolly J (2017) Biodiversity and ecosystem function: making sense of numerous species interactions in multi-species communities. Ecology 98(7):1771–1778
Cardinale BJ, Wright JP, Cadotte MW, Carroll IT, Hector A, Srivastava DS, Loreau M, Weis JJ (2007) Impacts of plant diversity on biomass production increase through time because of species complementarity. Proc Natl Acad Sci 104(46):18123–18128
Cardinale BJ, Srivastava DS, Duffy JE, Wright JP, Downing AL, Sankaran M, Jouseau C, Cadotte MW, Carroll IT, Weis JJ, Hector A (2009) Effects of biodiversity on the functioning of ecosystems: a summary of 164 experimental manipulations of species richness: ecological archives E090–060. Ecology 90(3):854–854
Connolly J, Finn JA, Black AD, Kirwan L, Brophy C, Luscher A (2009) Effects of multi-species swards on dry matter production and the incidence of unsown species at three irish sites Irish. J Agric Food Res 48:243–260
Connolly J, Cadotte MW, Brophy C, Dooley A, Finn JA, Kirwan L, Roscher C, Weigelt A (2011) Phylogenetically diverse grasslands are associated with pairwise interspecific processes that increase biomass. Ecology 92:1385–1392
Connolly J, Bell T, Bolger T, Brophy C, Carnus T, Finn J, Kirwan L, Isbell F, Levine J, Lüscher A, Picasso V, Roscher C, Sebastia M, Suter M, Weigelt A (2013) An improved model to predict the effects of changing biodiversity levels on ecosystem function. J Ecol 101(2):344–355
Connolly J, Sebastià M, Kirwan L, Finn J, Llurba R, Suter M, Collins R, Porqueddu C, Helgadóttir Á, Baadshaug O, Bélanger G, Black A, Brophy C, Čop J, Dalmannsdóttir S, Delgado I, Elgersma A, Fothergill M, Frankow-Lindberg B, Ghesquiere A, Golinski P, Grieu P, Gustavsson A, Höglind M, Huguenin-Elie O, Jørgensen M, Kadziuliene Z, Lunnan T, Nykanen-Kurki P, Ribas A, Taube F, Thumm U, De Vliegher A, Lüscher A (2018) Weed suppression greatly increased by plant diversity in intensively managed grasslands: a continental-scale experiment. J Appl Ecol 55(2):852–862
Cornell JA (2011) The original mixture problem designs and models for exploring the entire simplex factor space. In: Shewhart WA, Wilks SS, Cornell JA (eds) A primer on experiments with mixtures. Wiley, New York, pp 22–95. https://doi.org/10.1002/9780470907443.ch2
Cummins S, Finn JA, Richards KG, Lanigan GJ, Grange G, Brophy C, Cardenas LM, Misselbrook TH, Reynolds CK, Krol DJ (2021) Beneficial effects of multi-species mixtures on N2O emissions from intensively managed grassland swards. Sci Total Environ 792:148163
De Groot RS, Wilson MA, Boumans RM (2002) A typology for the classification, description and valuation of ecosystem functions, goods and services. Ecol Econ 41(3):393–408
Demétrio CG, Hinde J, Moral RA (2014) Models for overdispersed data in entomology. In: Ferreira CP, Godoy WAC (eds) Ecological modelling applied to entomology. Springer, Cham, pp 219–259
Dooley Á, Isbell F, Kirwan L, Connolly J, Finn JA, Brophy C (2015) Testing the effects of diversity on ecosystem multifunctionality using a multivariate model. Ecol Lett 18(11):1242–1251
Ebeling A, Pompe S, Baade J, Eisenhauer N, Hillebrand H, Proulx R, Roscher C, Schmid B, Wirth C, Weisser WW (2014) A trait-based experimental approach to under-stand the mechanisms underlying biodiversity–ecosystemfunctioning relationships. Basic Appl Ecol 15:229–240
Finn JA et al (2013) Ecosystem function enhanced by combining four functional types of plant species in intensively managed grassland mixtures: a 3-year continental-scale field experiment. J Appl Ecol 50:365–375
Frankow-Lindberg BE, Brophy C, Collins RP, Connolly J (2009) Biodiversity effects on yield and unsown species invasion in a temperate forage ecosystem. Ann Bot 103:913–921
Gamfeldt L, Snäll T, Bagchi R, Jonsson M, Gustafsson L, Kjellander P, Ruiz-Jaen MC, Fröberg M, Stendahl J, Philipson CD, Mikusiński G (2013) Higher levels of multiple ecosystem services are found in forests with more tree species. Nat Commun 4(1):1–8
Grange G, Finn JA, Brophy C (2021) Plant diversity enhanced yield and mitigated drought impacts in intensively managed grassland communities. J Appl Ecol 58(9):1864–1875
Grange, G., Brophy, C. and Finn, J.A., 2022. Drought and plant diversity effects on the agronomic multifunctionality of intensively managed grassland. Grassland at the heart of circular and sustainable food systems, pp.403–405.
Hector A, Schmid B, Beierkuhnlein C, Caldeira MC, Diemer M, Dimitrakopoulos PG et al (1999) Plant diversity and productivity experiments in european grasslands. Science 286:1123–1127
Hooper DU, Chapin FS, Ewel JJ, Hector A, Inchausti P, Lavorel S et al (2005) Effects of biodiversity on ecosystem functioning: a consensus of current knowledge. Ecol Monogr 75:3–35
Isbell F, Calcagno V, Hector A, Connolly J, Harpole WS, Reich PB et al (2011) High plant diversity is needed to maintain ecosystem services. Nature 477:4
Kirwan L, Luscher A, Sebastia MT, Finn JA, Collins RP, Porqueddu C et al (2007) Evenness drives consistent diversity effects in intensive grassland systems across 28 European sites. J Ecol 95:530–539
Kirwan L, Connolly J, Finn JA, Brophy C, Luscher A, Nyfeler D, Sebastia MT (2009) Diversity-interaction modeling: estimating contributions of species identities and interactions to ecosystem function. Ecology 90:2032–2038
Lembrechts JJ, De Boeck HJ, Liao J, Milbau A, Nijs I (2018) Effects of species evenness can be derived from species richness ecosystem functioning relationships. Oikos 127:337–344. https://doi.org/10.1111/oik.04786
Loreau M, Hector A (2001) Partitioning selection and complementarity in biodiversity experiments. Nature 412(6842):72–76. https://doi.org/10.1038/35083573
McDonnell J, McKenna T, Yurkonis KA, Hennessy D, de Andrade Moral R, Brophy C (2023) A mixed model for assessing the effect of numerous plant species interactions on grassland biodiversity and ecosystem function relationships. J Agric Biol Environ Stat 28(1):1–19
Mitchell TJ, Beauchamp JJ (1988) Bayesian variable selection in linear regression. J Am Stat Assoc 83:1023–1036
Moral, R. A., Connolly, J., Vishwakarma, R., and Brophy, C. (2022). Dimodels: Diversity-Interactions (DI) Models. R package version 1.2. https://CRAN.R-project.org/package=DImodels.
Morgan BJT (1992) Maximum-likelihood fitting of simple models. Anal Quantal Resp Data. https://doi.org/10.1007/978-1-4899-4539-6_2
Nyfeler D, Huguenin-Elie O, Suter M, Frossard E, Connolly J, Luscher A (2009) Strong mixture effects among four species in fertilized agricultural grassland led to persistent and consistent transgressive overyielding. J Appl Ecol 46:683–691
O’Hea NM, Kirwan L, Finn JA (2010) Experimental mixtures of dung fauna affect dung decomposition through complex effects of species interactions. Oikos 119:1081–1088
R Core Team (2021) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Reich PB, Tilman D, Naeem S, Ellsworth DS, Knops J, Craine J, Wedin D, Trost J (2004) Species and functional group diversity independently influence biomass accumulation and its response to CO2 and N. Proc Natl Acad Sci USA 101:10101–10106
Roscher C, Temperton VM, Scherer-Lorenzen M, Schmitz M, Schumacher J, Schmid B, Buchmann N, Weisser WW, Schulze ED (2005) Overyielding in experimental grassland communities – irrespective of species pool or spatial scale. Ecol Lett 8:419–429
Schmid B, Hector A, Huston MA, Inchausti P, Nijs I, Leadley PW, Tilman D (2002) The design and analysis of biodiversity experiments. In: Loreau M, Naeem S, Inchausti P (eds) Biodiversity and Ecosystem Functioning: Systhesis and Perspectives. Oxford University Press, Oxford, pp 61–75
Schwarz G (1978) Estimating the dimension of a model. Ann Statist. https://doi.org/10.1214/aos/1176344136
Spehn EM et al (2005) Ecosystem effects of biodiversity manipulations in European grasslands. Ecol Monogr 75:37–63
Tilman D, Lehman CL, Thomas KT (1997) Plant diversity and ecosystem productivity: theoretical considerations. Proceedings of the National Academy of Sciences of the United States ofAmerica 94:1857–1861
Tracy BF, Renne IJ, Gerrish J, Sanderson MA (2004) Effects of plant diversity on invasion of weed species in experimental pasture communities. Basic Appl Ecol 5(6):543–550
Wilsey BJ, Polley HW (2004) Realistically low species evenness does not alter grassland species-richness-productivity relationships. Ecology 85:2693–2700
Worm B, Barbier EB, Beaumont N, Duffy JE, Folke C, Halpern BS, Jackson JB, Lotze HK, Micheli F, Palumbi SR, Sala E (2006) Impacts of biodiversity loss on ocean ecosystem services. Science 314(5800):787–790
Acknowledgements
All authors were supported by the Science Foundation Ireland Frontiers for the Future programme, grant number 19/FFP/6888 award to CB.
Funding
Open Access funding provided by the IReL Consortium.
Author information
Authors and Affiliations
Contributions
CB, JC, and RV: conceived the ideas. RV led the coding with contributions from LB and RAM. RV and CB: wrote the initial draft of the paper. LB, JC, and RAM: each contributed to the writing of the subsequent drafts of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Handling Editor: Luiz Duczmal.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vishwakarma, R., Byrne, L., Connolly, J. et al. Estimation of the non-linear parameter in Generalised Diversity-Interactions models is unaffected by change in structure of the interaction terms. Environ Ecol Stat 30, 555–574 (2023). https://doi.org/10.1007/s10651-023-00563-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10651-023-00563-w