Abstract
Molecular mimicry between foreign and self-antigens has been implicated as a cause of autoimmune hepatitis in experimental models and cross-reacting antibodies in patients. This review describes the experimental and clinical evidence for molecular mimicry as a cause of autoimmune hepatitis, indicates the limitations and uncertainties of this premise, and encourages investigations that assess diverse environmental antigens as sources of disease-relevant molecular mimics. Pertinent articles were identified in PubMed using multiple search phrases. Several pathogens have linear or conformational epitopes that mimic the self-antigens of autoimmune hepatitis. The occurrence of an acute immune-mediated hepatitis after vaccination for severe acute respiratory syndrome (SARS)-associated coronavirus 2 (SARS-CoV-2) has suggested that vaccine-induced peptides may mimic disease-relevant tissue antigens. The intestinal microbiome is an under-evaluated source of gut-derived antigens that could also engage in molecular mimicry. Chaperone molecules may enhance the pathogenicity of molecular mimics, and they warrant investigation. Molecular mimics of immune dominant epitopes within cytochrome P450 IID6, the autoantigen most closely associated with autoimmune hepatitis, should be sought in diverse environmental antigens and assessed for pathogenicity. Avoidance strategies, dietary adjustments, vaccine improvement, and targeted manipulation of the intestinal microbiota may emerge as therapeutic possibilities. In conclusion, molecular mimicry may be a missing causality of autoimmune hepatitis. Molecular mimics of key immune dominant epitopes of disease-specific antigens must be sought in diverse environmental antigens. The ubiquity of molecular mimicry compels rigorous assessments of peptide mimics for immunogenicity and pathogenicity in experimental models. Molecular mimicry may complement epigenetic modifications as causative mechanisms of autoimmune hepatitis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autoimmune hepatitis is a consequence of dysregulated immune responses that overcome tolerance for self-antigens [1,2,3]. Cytokine-producing inflammatory cells, antigen-activated promiscuous CD4+ T cells, and liver-infiltrating cytotoxic T cells (CTLs) create a microenvironment that promotes liver damage [4,5,6,7]. The common clinical outcomes of this complex immune response are progressive hepatic injury and fibrosis [8, 9]. Environmental factors could trigger autoimmune hepatitis by overcoming homeostatic mechanisms that modulate immune tolerance and immune ignorance [10,11,12].
The linear or conformational mimicry of foreign antigens with self-antigens could promote the loss of self-tolerance and immune ignorance by increasing the array of immune targets and the avidity of promiscuous T cell effectors [4, 13,14,15,16,17,18,19,20,21,22]. Diverse pathogens have been evaluated as the principal sources of environmentally-derived molecular mimics of self-antigens in autoimmune hepatitis [23,24,25,26], and vaccination [27,28,29,30,31,32,33,34,35,36,37,38,39,40,41] and the intestinal microbiome [25, 26, 42,43,44,45,46,47] have emerged as sources of antigens that could also promote autoimmunity by molecular mimicry. Molecular mimicry could be a mechanism that complements epigenetic transformations in shaping the autoreactive response to environmental stimuli [48,49,50,51,52,53,54,55].
The molecular mimicry of environmental antigens has been implicated in the development of primary biliary cholangitis (PBC). Structural mimicries and immune cross-reactions have been established between the immune dominant pyruvate dehydrogenase complex-E2 (PDC-E2) of PBC and certain bacteria (Escherichia coli [56,57,58] and Novoshingobium aromaticivorans) [59,60,61]). Furthermore, limited population studies have supported the bacterial associations with the occurrence of clinical disease [58, 61,62,63]. Xenobiotics, especially 2-octynoic acid [64], have also been shown to mimic and modify the lipoyl domain of PDC-E2 by chemical conjugation [65]. Transformation of the altered native PDC-E2 into a homologue of native PDC-E2 could promote the production of antimitochondrial antibodies [66] and the development of a PBC-like disease in an experimental model [67,68,69]. The challenge has been to establish these environmentally-induced molecular mimicries as pathological bases for PBC in humans [70].
Improvement in the management of autoimmune hepatitis is recognized as an unmet clinical need [71, 72], and satisfaction of this need requires clarification of the factors that induce and sustain the autoreactive process. Identification of critical environmental elements and the mechanisms by which they can promote autoimmune hepatitis could also suggest strategies that prevent or limit the disease [73].
The goals of this review are to describe the experimental and clinical evidence that supports molecular mimicry as a causative mechanism of autoimmune hepatitis, indicate limitations and uncertainties of this premise, and encourage investigations that assess diverse environmental antigens as immunogenic and pathogenic molecular mimics of disease-relevant self-antigens. The validation of molecular mimicry as a pathogenic mechanism may direct future management strategies.
Methods
English abstracts were identified in PubMed using the search phrases, “environmental factors in autoimmune hepatitis”, “molecular mimicry”, “vaccination and autoimmune hepatitis”, and “intestinal microbiome and autoimmune hepatitis”. Key aspects of pertinent abstracts were recorded, and full-length articles that expanded pivotal concepts constituted the primary bibliography. Secondary and tertiary bibliographies were developed from the references cited in the selected full-length articles of the preceding bibliography. Several hundred abstracts and 184 full length articles were examined.
Molecular Mimicry and Environmental Pathogens
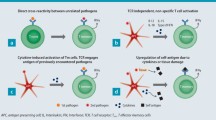
Molecular mimicry is a largely theoretical mechanism by which environmental pathogens could trigger autoimmune hepatitis [18, 21, 74]. The process requires the selection, processing, and presentation of an immunogenic peptide intrinsic to the pathogen [3, 75]. The peptide selected for presentation by antigen presenting cells (APCs) has to resemble, not duplicate, the structure of a self-antigen [75], and it has to activate an immune response by overcoming self-tolerance and immune ignorance [6, 21] (Fig. 1). Up-regulation of MHC molecules by the invading pathogen and an inflammatory milieu (cytokines, chemokines, and inflammatory cells) associated with the infection (bystander response) could promote the presentation of pathogen-derived immunogenic peptides [21, 76]. Activation and differentiation of CD4+ and CD8+ T cells sensitized to these peptides could then induce an immune-mediated, inflammatory liver disease depending on the number and avidity of the T cell effectors for the homologous tissue antigen [18, 21, 75].
Molecular mimicry and environmental antigens. Environmental antigens from exogenous and endogenous sources may have structural or conformational homology with a disease-related antigen. Peptides of at least 5 amino acids (aa) that are derived from these sources could constitute an immunogenic unit that is homologous to an immune dominant epitope of a self-antigen. Intestinal dysbiosis and gut-derived metabolic processes may affect levels of lipopolysaccharide (LSP) and short chain fatty acids (SCFA) which in turn may increase permeability of the intestinal mucosal barrier. Gut-derived immunogenic peptides could then translocate to systemic sites and encounter antigen presenting cells. In dendritic cells, the immunogenic homologues can undergo proteasomal processing and presentation by class I or class II molecules of the major histocompatibility complex (MHC). A molecular chaperone may facilitate the proteasomal processing and substrate presentation. Heat shock proteins (HSP) are theoretical candidates for this role (dotted dual-headed arrow). Antigen cross-presentation can allow a processed environmental antigen to be presented by MHC class I molecules to CD8+ T cells, and the activated CD8+ T cells may then differentiate to liver-infiltrating cytotoxic T cells (CTL). The processed homologue may also be presented by MHC class II molecules to CD4+ T cells. The activated CD4+ T cells may then induce a type 1 helper T cell (Th1) response with increased production of interleukin 2 (IL-2) and interferon-gamma (IFN-γ). Macrophages and natural killer (NK) cells may also be activated. The IFN-γ may in turn activate the class II transactivator (CIITA) gene which increases the expression of class II MHC molecules and presentation of the processed homologue. The activated CD4+ T cells may also produce a Th2 response characterized by certain interleukins (IL-4, IL-5, IL-13, and IL-10) and the activation of B cells and plasma cells which in turn may generate autoantibodies
Genetic Predisposition and Plasticity of the APCs
Antigenic peptides presented by the class II MHC molecules (HLA DR, DP, and DQ) are derived from extracellular proteins (foreign antigens) [77] that are processed mainly by proteases within endosomal compartments [78, 79] (Table 1). The antigen binding groove of class II MHC molecules consists of two chains of amino acids that are encoded by separate genes in the class II MHC region of chromosome 6 [80]. The antigen binding groove of the HLA-DR molecules associated with autoimmune hepatitis consists of a monomorphic α-chain and polymorphic β-chain [81,82,83,84]. The DRα-chain is encoded by the non-polymorphic DRA gene, and the DRβ-chain is encoded by the polymorphic DRB1 genes [85].
The selection, positioning, and presentation of the antigenic peptide to naïve CD4+ helper T cells in autoimmune hepatitis is influenced by the hypervariable regions encoded within the DRβ-chain [86] (Table 1). Sequence polymorphisms of the DRβ-chain determine the type of peptides that can bind in the pockets of the antigen binding groove [86], and this molecular alignment can affect susceptibility to autoimmune hepatitis [87, 88].
Structural alterations in the antigenic peptide and variations in the MHC alleles that encode the DRβ chain can change susceptibility by altering the conformation of the antigen-presenting complex [89,90,91] (Table 1). The conformational plasticity of the antigen binding groove of class II MHC molecules allows it to sample, select, and accommodate diverse antigenic peptides derived from the environment [91]. Furthermore, the antigen binding groove is open at both ends, and it can accommodate relatively large, environmentally derived peptides of 13–25 residues [91, 92].
The molecular mimicry of self-antigens by peptides of environmental origin requires presentation of the homologous peptides to CD4+ helper T cells, and the class II MHC molecules on dendritic cells, monocytes, macrophages, and B cells can met this requirement [80] (Table 1). Furthermore, the pro-inflammatory cytokine, interferon-gamma (IFN-γ), can induce the expression of class II MHC molecules on other cells by activating the class II transactivator (CIITA) gene [93,94,95] (Fig. 1). CIITA is the major regulator of MHC class II transcription [93], and it may expand the number and variety of APCs and the population of activated promiscuous CD4+ T cells.
Antigen cross-presentation by dendritic cells is another mechanism by which environmental peptides that mimic self-antigens can initiate an autoreactive response (Table 1). Antigen cross-presentation is a process by which antigens from the extracellular environment can be presented by MHC class I molecules and directly stimulate the development of antigen-specific CD8+ T cells [96,97,98,99] (Fig. 1). This process is distinguished from the canonical pathway in which extracellular (foreign) antigens are presented by MHC class II molecules and activate CD4+ helper T cells [100]. Dendritic cells can present processed antigens by MHC class I and MHC class II molecules [101,102,103], and they are the principal population engaged in antigen cross-presentation. Molecular chaperones that carry substrates generated by the proteasomal degradation of the foreign antigen can engage in the priming process [97]. Enhanced immune reactivity to the extrinsic antigen may be a consequence of this interaction depending on the antigen and the microenvironment [98, 104].
Causative Effect of Molecular Mimicry in Experimental Models
Molecular mimicry by a viral pathogen has been proposed as a basis for the immune-mediated liver damage in murine models of autoimmune hepatitis. Infection with an adenovirus expressing the human cytochrome P450 IID6 (hCYP2D6) [75, 105, 106] and immunization with hCYP2D6 and human formiminotransferase cyclodeaminase [107, 108] have induced laboratory, histological, and immunological changes of autoimmune hepatitis (Table 2). CYP2D6 [109,110,111] and formiminotransferase cyclodeaminase [112, 113] have been closely associated with type 2 autoimmune hepatitis, and molecular mimicry between the human antigens and mouse homologues in the experimental models has been proposed as the basis for the pathological [75, 105, 107, 108], serological [105, 107, 108], and fibrotic [114] changes resembling autoimmune hepatitis.
Molecular mimicry has also been demonstrated in a study exposing wild-type and transgenic mice to the same human antigen associated with autoimmune hepatitis [75]. The infection of wild-type mice with adenovirus expressing hCYP2D6 has generated T cell activity against epitopes that have had intermediate homology between hCYP2D6 and mouse cytochrome (mCYP) homologues [75]. In contrast, the same adenovirus infection of transgenic mice expressing hCYP2D6 has failed to generate an hCYP2D6-specific T cell response [75]. Molecular mimicry rather than identity between the adenoviral hCYP2D6 and the mCYP homologues was the likely basis for the hCYP2D6-specific T cell response [75].
Molecular mimicry has not been established as a cause of autoimmune hepatitis in humans [21, 115], but the structural and conformational similarities between environmental pathogens and disease-associated antigens have compelled its consideration [109, 111, 116,117,118,119,120,121,122,123]. Undiscovered pathogens [124], isolated subclinical or antecedent infections [21, 74], and multiple previous infections by diverse viruses mimicking the same self-antigen may contribute to the risk of autoimmune disease [21, 74, 111].
Contributory Effects of Molecular Mimicry
Molecular mimicry may also be a mechanism that accelerates or sustains autoimmune hepatitis [44, 73, 125] (Table 2). The release of neo-antigens from damaged liver tissue [126, 127] or the translocation of bacterial products from a permeable intestine [44, 47, 128,129,130,131,132,133,134,135] could promote molecular mimicry and affect the severity and course of autoimmune hepatitis. Furthermore, molecular mimicry could contribute to the ongoing inflammatory process by expanding the number of targeted epitopes (epitope spread) [136,137,138,139] and the population of memory CD4+ and CD8+ T cells (bystander activation) [140, 141].
Mimicry Between CYP2D6 and Environmental Pathogens
CYP2D6, the principal autoantigen associated with one form of autoimmune hepatitis [109,110,111], has amino acid sequences within its structure that resemble sequences in the hepatitis C virus (HCV) [109, 119, 120], herpes simplex type 1 virus [109, 117], and human T lymphotrophic virus types 1 and 2 [117] (Table 2). A conformational epitope on CYP2D6 in patients with chronic hepatitis C and antibodies to liver kidney microsome type 1 (anti-LKM1) spans the major linear epitope associated with type 2 autoimmune hepatitis [122], and cross-reacting anti-LKM1 have been demonstrated in patients with chronic hepatitis C [122, 142, 143]. Furthermore, a homologous region between human CYP2D6 and protein from Salmonella typhimurium suggests that bacteria could be another environmental source of molecular mimicry [117].
None of the pathogens with amino acid sequences that resemble those of CYP2D6 has been recognized as a cause of type 2 autoimmune hepatitis in humans [21, 115, 144]. The immunogenicity of the shared sequences within the pathogen [145, 146] and the composition of the host-specific microenvironment [21, 76] may be the critical factors that influence the risk of disease.
Mimicry Between Other Immunogenic Antigens and Environmental Pathogens
Molecular mimicry has also been demonstrated between key antigenic targets in type 1 autoimmune hepatitis and environmental pathogens [116, 121, 123]. The deoxyribonucleic acid (DNA) polymerase of the hepatitis B virus (HBV) has structural similarities to smooth muscle and nuclear antigens [116], and homologies have also been described between other regions in these same antigens and amino acid sequences in the polyprotein of HCV [121] (Table 2). Neither virus has been recognized as a cause of type 1 autoimmune hepatitis [21, 144], and the major clinical consequence of the structural homologies may be to lower the diagnostic specificity of smooth muscle antibodies (SMA) and antinuclear antibodies (ANA) [147].
Structural mimicry has also been demonstrated between the immune dominant regions of the SepSecS protein (O-phosphoserine [Sep] transfer RNA:selenocysteine (Sec) transfer RNA synthase) and the PS 120 surface antigen of Rickettsia species [123] (Table 2). The SepSecS protein [148,149,150,151] generates antibodies to soluble liver antigen (anti-SLA) which are highly specific for autoimmune hepatitis [152,153,154]. The clinical impact of the structural homology between the SepSecS protein and the PS 120 surface antigen of Rickettsia species is uncertain [123, 151, 155].
Molecular Mimicry and Vaccination
Human vaccines have been developed against deleterious environmental pathogens by stimulating the recipient’s immune system to react against critical structural components of the invading organism [156,157,158,159,160,161]. The generation of neutralizing antibodies and the activation of an antigen-specific, adaptive immune response can eliminate the targeted organism [162,163,164,165]. Similarities between peptides generated by the vaccine and normal proteins within the vaccine recipient may generate immune cross-reactivity that is manifested as an autoimmune disease [166,167,168,169,170] (Table 2). Recent experiences in humans after vaccination against the severe acute respiratory syndrome (SARS)-associated coronavirus 2 (SARS-CoV-2) have suggested an etiological connection between the vaccines and autoimmune hepatitis which may relate to molecular mimicry [27,28,29,30,31,32,33,34,35,36,37,38,39,40,41].
Autoimmune Hepatitis-Like Disease After SARS-CoV-2 Vaccination
An acute onset hepatitis has been described in 87 patients within 3–65 days (median 15 days) after vaccination for SARS-CoV-2 [35]. The vaccines mainly contained messenger ribonucleic acid (mRNA) for the spike (S) protein of SARS-CoV-2, and 57% of patients with acute hepatitis had features of autoimmune hepatitis (Table 2). Glucocorticoids had been administered to most patients, and the liver disease had resolved in all treated and untreated patients with one exception (frequency of liver failure and transplantation, 1.1%). Relapse did not occur during an observation period that spanned 44–140 days after treatment withdrawal and 35–172 days after spontaneous resolution.
Similar findings have been reported in two systematic reviews of vaccine-associated liver disease [32, 34] (Table 2). In the largest review, an acute hepatitis with autoimmune features had developed within 7–21 days (median, 14 days) after vaccination in 138 patients; 98% had received mRNA-based vaccines against SARS-CoV-2; most patients had been treated with immunosuppressive drugs; 89% had achieved full recovery; and 2.2% had died [32]. In the smaller review involving 32 patients who had developed an autoimmune hepatitis-like disease after vaccination, 7 improved spontaneously (22%); 24 of the 25 treated patients improved or resolved (96%); and one patient died (3%) [34].
The acute onset of liver injury; its spontaneous resolution in some patients; the near-universal resolution after immunosuppressive therapy; and the absence of relapse after drug withdrawal are atypical features of autoimmune hepatitis [171] (Table 2). The atypical features, however, do not discount the actual or potential occurrence of autoimmune hepatitis after vaccination [34, 172] or the possible role of vaccine-induced molecular mimicry [27,28,29,30,31,32,33, 35, 37,38,39,40,41]. The overall safety and efficacy of vaccination against SARS-CoV-2 have far outweighed the rarity of vaccine-related liver dysfunction, and the recommendation for vaccination in the general population has remained strong [32, 34, 35, 173,174,175].
The apparent rarity of de novo autoimmune hepatitis after coronovirus disease 2019 (COVID-19) [176,177,178] suggests that the vaccine and its delivery system of lipid nanoparticles [158, 161] generate a more intense immunogenic signal than community-acquired infection.
Vaccine-Based Molecular Mimicry and Autoimmunity
Sequence analyses of 20,365 human proteins have identified 3781 proteins that share peptides of at least 6 amino acids with structural proteins of SARS-CoV-2 [179] (Table 2). Pentapeptides are recognized as the minimal immunogenic epitopes [145, 146], and the extensive sharing of hexapeptides between SARS-CoV-2 and human proteins suggests that molecular mimicry after vaccination is common and rarely pathogenic [21, 168, 170]. The immunogenicity and pathogenicity of the molecular mimic may depend on its relevance to a pivotal disease-pertinent self-antigen and the circumstances that affect its processing and presentation by the MHC molecules (Fig. 1).
Antibodies to the spike protein of SARS-CoV-2 have had reactivity to human tissue transglutaminase, extractable nuclear antigen, myosin basic protein, mitochondria, and nuclear antigens. These homologies have been expressed in serum as nuclear, actin, and mitochondrial antibodies [180] (Table 2). The serological findings indicate that molecular mimicry after vaccination can be a basis for immune reactivity associated with autoimmune hepatitis. The investigational challenges are to identify the key molecular mimic of an immune dominant epitope that can trigger autoimmune hepatitis and the circumstances that promote its immunogenicity. [21, 22, 170]
Molecular Mimicry and Gut-Derived Microbial Products
The intestinal microbiome is a reservoir of bacterial products and immune cells that have the potential to translocate to extra-intestinal sites and generate an immune response [131, 181]. CD4+ and CD8+ T cells activated by gut-derived peptides may target homologous tissue antigens and contribute to a loss of self-tolerance [44, 182, 183] (Fig. 1). Molecular mimicry based on changes within the intestinal microbiome constitutes a potentially powerful mechanism by which environmental factors could shape the intestinal microbiome.
Diet [25, 134], supplements [24], antibiotics [23], alcohol [23], pollutants [23], and toxins [26] have been associated with the risk of autoimmune hepatitis, and this risk may reflect changes in the intestinal microbiome induced by these diverse environmental factors [25, 26, 42,43,44,45,46,47] (Table 2). Findings in experimental models of autoimmune hepatitis [129, 184] and other autoimmune diseases [185,186,187,188] have supported this premise, albeit there are many other factors outside of molecular mimicry that may be contributory [45,46,47, 135, 189].
Dysbiosis and Disease Severity
Immune tolerances to commensal bacteria within the intestine may be lost by environmentally-induced dysbiosis [190], and toll-like receptors in the intestine may generate pathogenic T and B lymphocytes that are capable of systemic translocation and reactivity against homologous tissue antigens [188, 191] (Table 2). Intestinal dysbiosis has been demonstrated in an experimental model [128] and in patients with autoimmune hepatitis [132,133,134,135]. The key distinguishing features from healthy individuals have been reduced biodiversity of the intestinal microbiome [128, 132,133,134, 192], increased aerobic and facultative anaerobic bacteria [134], increased Veillonella species [132, 134], and decreased Bifidobacterium species [129, 134] (Fig. 1). The composition of the intestinal microbiome has been specific for autoimmune hepatitis when compared to that of healthy individuals and patients with primary biliary cholangitis (PBC) or ulcerative colitis [134].
Changes in the size of certain bacterial populations within the intestine have also been associated with the severity of autoimmune hepatitis [135]. An increased Veillonella population in patients with autoimmune hepatitis has correlated with the serum level of aspartate aminotransferase [132], and a decreased population of Bifidobacterium has been associated with failure to achieve remission during treatment [134] (Table 2). Furthermore, the abundance of Bifidobacterium in the intestinal microbiome has correlated directly with the average protein intake of patients with autoimmune hepatitis [134]. This finding underscores the possibility that dietary effects on the intestinal microbiota might impact on disease severity.
Increased Intestinal Permeability
Alterations in the structural proteins (zona occludens 1 and occludin) that bind intestinal epithelial cells have been described in patients with autoimmune hepatitis [129], and increased intestinal permeability has been demonstrated in experimental models of the disease [193,194,195] (Table 2). Multiple microbial products that are manufactured within the intestinal microbiome may be enhanced by the dysbiosis and account for the barrier disruption [135, 196].
Lipopolysaccharides (LSP) from commensal gram-negative bacteria may increase, and they may enhance intestinal permeability by disrupting signaling pathways that maintain barrier integrity [135, 197] (Fig. 1). The breakdown of indigestible carbohydrates to short chain fatty acids (acetic, proprionic, and butyric acids) may also be impaired by changes in the populations of commensal anaerobic bacteria [198,199,200,201]. Clostridia [201] and Bifidobacterium [202] are key carbohydrate-fermenting, bacterial populations, and reductions in their abundance could reduce concentrations of short chain fatty acids, especially butyrate, within the gut [200].
Butyrate has strengthened tight junction assembly and improved barrier function in experimental models [203,204,205,206], and its deficiency in autoimmune hepatitis could facilitate translocation of bacterial products and enhance the opportunity for molecular mimicry. A butyrate deficiency could also affect other signaling pathways that modulate immune and inflammatory responses pivotal for the occurrence and maintenance of autoimmune hepatitis [207,208,209,210]. Dietary fiber and butyrate supplementations have increased mucosal junction proteins, elevated the ratio of regulatory T cells (Tregs) to T helper 17 (Th17) cells, and decreased translocation of microbial protein in experimental autoimmune hepatitis [211].
The critical impact of dysbiosis on the integrity of the intestinal mucosal barrier has been demonstrated further by manipulating the gut microbiota. Probiotics are preparations of living enteric commensal micro-organisms that are intended to induce a salutary microbiome [212,213,214]. The administration of probiotics containing species of Bifidobacterium with [195] or without Lactobaccillus [193] has had a broad spectrum of beneficial effects in experimental models of autoimmune hepatitis (Table 2). It has suppressed hepatic inflammation [195], decreased serum aminotransferase levels [195], diminished proliferation of Th17 cells [193, 195], reduced serum endotoxin levels [193], enhanced the abundance of short chain fatty acids [193], inhibited transcription of cytokines [195], and maintained or strengthened the intestinal mucosal barrier [193, 195].
Interactive Modulatory Metabolic Effects
Metabolic products from different bacterial populations within the intestine can become shared energy sources that sustain the composition of the microbiome and the integrity of the intestinal mucosal barrier [42, 199]. The degree of communal interdependence is unclear, but cross-feeding among commensal populations may enhance their resiliency. The acetate produced by Bifidobacterium can be utilized by Fecalibacterium prausnitzii, a dominant intestinal inhabitant, to produce butyrate [215]. The cross-feeding of energy sources may be a mechanism by which Bifidobacterium can up-regulate tight junctions, improve barrier function, reduce endotoxemia, impair proliferation of Th17 cells, and mitigate experimental autoimmune hepatitis [193, 215].
The intestinal microbiota may also produce metabolites that modulate the inflammatory and immune responses. Glycolipids derived from the intestinal bacteria can be presented by the MHC class I-like molecule, CD1d, and activate natural killer T (NKT) cells [216]. NKT cells activated by this mechanism have promoted liver injury in an experimental model of hepatitis [217]. Secondary bile acids (lithocholic acid and deoxycholic acid) derived from the actions of enteric bacteria on the primary bile acids (cholic acid and chenodeoxycholic acid) may counter a pro-inflammatory response by activating the G-protein-coupled bile acid receptor 1 (GPBAR1). GPBAR1 can direct the differentiation of NKT cells to an anti-inflammatory, interleukin 10-producing, subset that ameliorates experimental immune-mediated hepatitis [189].
Molecular Mimicry of Gut-Derived Antigens in Autoimmune Hepatitis
Atypical perinuclear anti-neutrophil cytoplasmic antibodies (pANCA) occur in 49–96% of patients with type 1 autoimmune hepatitis [218,219,220,221,222,223]. They are also found in patients with primary sclerosing cholangitis (PSC) [219, 224, 225] and inflammatory bowel disease [226,227,228]. Atypical pANCA recognize β-tubulin isotype 5 in conjunction with its evolutionary bacterial precursor protein, filamenting temperature-sensitive mutant Z protein (FtsZ) [229] (Table 2). The dual reactivity is dependent on an intact intestinal microbiome in a mouse model of inflammatory bowel disease [229]. The absence of this dual reactivity in germ-free animals supports the premise that gut-derived antigens can escape the intestine and trigger an antibody response that is disease-related [229,230,231]. The high prevalence and high titers of atypical pANCA in some studies of type 1 autoimmune hepatitis [218, 223] compel this consideration, albeit the antibody production may not translate into a pathogenic immune response.
Prospect of Molecular Mimicry as a Cause of Autoimmune Hepatitis
The prospect that molecular mimicry of environmental antigens will emerge as a valid cause of autoimmune hepatitis depends on the demonstration of its immunogenicity and pathogenicity. Examinations of 30 viral proteomes [232] and 40 bacterial proteomes [233] for similarities to the human proteome have disclosed massive sharing of amino acid motifs at the pentapeptide level and higher. The ubiquity of molecular mimicry between pathogenic viral and bacterial species and normal tissue proteins indicates that molecular mimicry by itself is insufficient to cause autoimmune disease [179, 232,233,234].
Future investigations must evaluate diverse environmental antigens in addition to pathogens as candidates for molecular mimicry [131, 188, 235,236,237,238]. They must demonstrate the immunogenicity [21] and pathogenicity [188] of the implicated homologues, and they must determine the factors that influence their potency as triggering peptides. The intestinal microbiome must also be rigorously assessed as a source of molecular mimicries that can initiate, enhance, and sustain a deleterious autoreactive immune response [44, 73, 131, 188, 238, 239]. Demonstration of the pathogenic nature of the molecular mimicries will require a humanized animal model of autoimmune hepatitis that expresses human leukocyte antigen (HLA) risk alleles.
Discovering Diverse Environmental Homologues and Demonstrating Immunogenicity
Pentapeptides are the shortest functional units recognized by the immune system [145, 146]. The search for immunogenic and pathogenic molecular mimics in autoimmune hepatitis must focus on identifying peptides in environmental antigens that not only mimic the immune dominant epitopes of CYP2D6 but also have sufficient size to be immunogenic [109, 111, 120] (Table 3). CYP2D6 is the pivotal antigen for the discovery of disease-relevant molecular mimics in the environment because of its high specificity for type 2 autoimmune hepatitis [109, 111, 120]. Type 2 autoimmune hepatitis, however, is less common in North American adult patients than type 1 autoimmune hepatitis [110, 147, 153, 240]. The key antigen associated with type 1 autoimmune hepatitis remains unclear, and this deficiency limits discovery of its disease-relevant molecular mimics within the environment.
The epitope of CYP2D6 which spans the amino acid (aa) region, aa193-212, is a prime immune dominant epitope associated with antibodies to liver kidney microsome type 1 (anti-LKM1) and type 2 autoimmune hepatitis [111]. The CYP2D6 epitope spanning the amino acid sequence, aa316 and 327, is expressed on the molecular surface, and its surface location may enhance its candidacy as another immune dominant epitope of CYP2D6 [120]. A 33 amino acid segment of the CYP2D6 molecule contains a shorter segment of 8 amino acids which resembles the herpes simplex type 1 virus [109], and it is another epitope of CYP2D6 that could be used to probe for homologies in diverse environmental antigens. Homologues of each immune dominant epitope of CYP2D6 can be sought using established databases (Genbank, Protein Information Resource [PIR], SWISS-PROT, or the International Nucleotide Sequence Database Collaboration [INSCD]) [109, 111, 241].
The immunogenicity of the implicated homologue can be demonstrated by detecting its antibodies in sera from patients with type 2 autoimmune hepatitis. Pre-absorption of the sera with LKM1 enhances its specificity for the candidate homologue [109, 242]. Immunogenic peptide mimics of the pivotal CYP2D6 epitopes could also be assessed by using peptide libraries of environmental antigens expressed on a microarray and probed with the pre-absorbed serum [242,243,244,245]. The identification of disease-pertinent peptide mimics in diverse environmental antigens (foodstuffs, vaccines, toxins, and micro-organisms [235,236,237, 246]) would support the causal role of molecular mimicry in type 2 autoimmune hepatitis and encourage discovery of the immune dominant epitopes in type 1 disease.
Defining Conditions That Enhance Immunogenicity
The ubiquity of molecular mimicry between foreign and self-proteins and the relative rarity of autoimmune diseases [233] have driven the search for factors that enhance the causative impact of molecular mimicry [21]. The factors that have been associated with the loss of self-tolerance (bystander activation of pre-primed T cells, memory cells and APCs [73, 140, 141]; epitope spread to tissue antigens of progressively lower homology [136,137,138,139, 247]; the release of neo-antigens from damaged tissue [126, 127]; and the production of superantigens that directly activate T lymphocytes [248, 249]) intensify the immune response [6], but they are not the primary causes of it. Adjuvant mechanisms that can particularly enhance the immunogenicity of molecular mimics would strengthen their candidacy as primary interfaces between the environment and autoimmune hepatitis (Table 3).
Molecular Chaperones and Peptide Immunogenicity
Molecular chaperones are highly conserved proteins that are present in all cellular compartments [250]. They stabilize the processing, maturation, and activity of intracellular proteins by aiding in their folding, assembly, and trafficking [250,251,252]. Heat shock proteins (HSPs) are molecular chaperones that reside constitutively within the cytoplasm, and they can be induced by oxidative stress, nitrosative stress, inflammation, or infection [253, 254]. HSPs can also affect the immunogenicity of intra- and extracellular peptides [255,256,257].
HSPs can bind to intracellular antigens containing both MHC class I and MHC class II epitopes, and as molecular chaperones, they can facilitate cross-presentation of acquired antigens by dendritic cells [257] (Fig. 1). HSPs may be adjuvants that could strengthen the immunogenicity of an environmental peptide that mimics a self-antigen (Table 3). The investigation of HSPs as modulators of the immune response to foreign antigens may strengthen the candidacy of molecular mimicry as a causative factor in autoimmune hepatitis. It may also suggest HSP-directed therapeutic strategies for future study [254, 258,259,260].
Other molecular chaperones that could affect the immunogenicity of environmental antigens are tapasin [261, 262] and the transporter associated with antigen processing (TAP)-binding protein (TAPBPR) [263,264,265,266]. Both chaperone molecules facilitate the selection, processing, and editing of high affinity peptides for MHC class I molecules. The lysosome-associated membrane protein-2 isoform (LAMP-2a) promotes the presentation of peptides by MHC class II molecules [267, 268], and the human leukocyte antigen (HLA)-B-associated transcript 3 (Bat3) protects autoreactive T cell responses by preventing T cell exhaustion [269, 270]. Chaperone molecules that contribute to the selection, editing, and presentation of peptides and the preservation of the autoreactive T cell response are factors that can affect the immunogenicity of environmental antigens. Their roles are unassessed in autoimmune hepatitis.
Demonstrating Pathogenicity of the Implicated Homologue
The implicated homologue must generate T and B cell reactivity to the principal antigen associated with autoimmune hepatitis (CYP2D6) [271], and the cross-reactivity must produce the disease in a genetically predisposed experimental model [188, 272]. The homologue must be non-identical to CYP2D6, and the experimental model must be susceptible to the disease in order to mitigate immune tolerance [75, 188]. The pathogenicity of a homologue derived from the intestinal microbiome has been elegantly demonstrated in patients with antiphospholipid syndrome, [188] and a similar methodology can be applied in patients with autoimmune hepatitis.
Immunogenic epitopes of CYP2D6 can be used to search for mimics within the intestinal microbiome by examining available databases, including resources developed by the Human Microbiome Project Consortium [273, 274] (Table 3). The enteric commensal bacterial species expressing the critical homologue can be identified, sought in the fecal specimens of patients with type 2 autoimmune hepatitis, and assessed for pathogenicity by evaluating T and B cell cross-reactivity between the epitopes of CYP2D6 and the molecular mimics expressed in the bacterial species. The methodology can be expanded to identify and assess homologues derived from other environmental sources.
Focusing on the Intestinal Microbiome
Gut-derived homologous peptides must translocate from the intestine to extra-intestinal sites to generate pathogenic, cross-reactive, T and B cell responses [188]. The genetic and biological factors that contribute to these responses must be clarified in autoimmune hepatitis [231] (Table 3). The enteric milieu of short chain fatty acids [198,199,200,201], LSP [197], glycolipids [217], and bile acids [189, 209] must be characterized to define the conditions for increased intestinal permeability. Furthermore, the principal factors that alter or restore the function of the intestinal epithelial cells must be determined [129, 275]. The investigational emphasis must shift from measuring the size of the enteric populations in autoimmune hepatitis to assessing their function and interactivity [47]. The pathogenic sequence from environmental factors to intestinal dysbiosis to a disease-producing immune response by molecular mimicry awaits experimental validation [214, 275, 276].
Prospective Management Strategies
The feasible management strategies for pathogenic molecular mimicry include life-style changes [277], dietary adjustments or supplements [204, 211, 231, 277,278,279,280], improved vaccine production [161], and targeted manipulation of the intestinal microbiome [44, 47, 135, 214, 231, 281] (Table 3). Diets emphasizing fiber [211] and favoring the intestinal production of short chain fatty acids [280] and butyrate [203,204,205, 211] may preserve or strengthen the intestinal mucosal barrier. Probiotics [195, 213, 278], pharmacological mucosal protectors [275, 282, 283], and fecal microbiota transplantation (FMT) [284, 285] are other interventions that could mitigate the impact of gut-derived molecular mimicry [193, 195]. Studies that clarify the principal environmental sources for pathogenic molecular mimicry will direct the evaluation of appropriate management strategies.
Molecular Mimicry and Epigenetic Changes
Environmental factors may influence the causality of autoimmune hepatitis outside of molecular mimicry [52, 53, 55, 286]. Epigenetic changes induced by the environment can alter the transcriptional activity of immune regulatory genes without changing the nucleotide sequence of DNA [54, 287, 288]. Induced alterations in the methylation, acetylation, and phosphorylation of histones within the nucleosomes can reversibly up- or downregulate gene expression and influence the immune response [48, 53, 55].
Unlike molecular mimicry which is antigen-dependent, epigenetic changes can be induced by non-antigenic environmental factors that affect the activity of enzymes that modify histone structure [48]. Nutritional deficiencies [51], alcohol [289], tobacco [290], pollutants [291, 292], and psychological stress [50, 293] are environmental factors that may affect disease susceptibility by altering pivotal gene transcription rather than mimicking a self-antigen. Non-antigenic environmental stimuli may also change the intestinal microbiome and indirectly promote intestinal dysbiosis and gut-derived, antigen-induced, molecular mimicry [23, 26, 42, 47].
Molecular mimicry and epigenetic changes may contribute independently to the causality of autoimmune hepatitis depending on the nature of the environmental stimulus. Future investigations must establish the pathogenicity and relevance of molecular mimicry compared to epigenetic mechanisms. Therapeutic interventions could emerge that are mechanism-dependent.
Conclusions
Molecular mimicry may translate diverse environmental antigens into autoimmune hepatitis by increasing the avidity of immune cells for multiple homologous tissue antigens. Vaccinations and the intestinal microbiome may be prime environmental sources for peptides that mimic immune dominant epitopes of key self-antigens associated with autoimmune hepatitis. Disease-pertinent peptide mimics can be sought among diverse environmental antigens by searching for homologues of the known immune dominant epitopes of CYP2D6. Immune dominant epitopes of key antigens associated with type 1 autoimmune hepatitis await clarification. The conditions required for immunogenicity of the mimicking epitope must be clarified and include chaperone molecules, such as HSPs. Translocated gut-derived bacterial antigens should be evaluated as molecular mimics, and the conditions for increased intestinal permeability clarified. The implicated environmental homologue must be shown to be pathogenic by generating T and B cell reactivity to CYP2D6 and producing experimental autoimmune hepatitis. The relative contributions of molecular mimicry and epigenetic alterations to the causality of autoimmune hepatitis should be determined, and management strategies evaluated that are appropriate for the principal pathogenic mechanism.
References
Longhi MS, Ma Y, Mieli-Vergani G, Vergani D. Aetiopathogenesis of autoimmune hepatitis. J Autoimmun. 2010;34:7–14.
Liberal R, Longhi MS, Mieli-Vergani G, Vergani D. Pathogenesis of autoimmune hepatitis. Best Prac Res Clin Gastroenterol. 2011;25:653–664.
Czaja AJ. Transitioning from idiopathic to explainable autoimmune hepatitis. Dig Dis Sci. 2015;60:2881–2900.
Doherty DG, Penzotti JE, Koelle DM et al. Structural basis of specificity and degeneracy of T cell recognition: pluriallelic restriction of T cell responses to a peptide antigen involves both specific and promiscuous interactions between the T cell receptor, peptide, and HLA-DR. J Immunol. 1998;161:3527–3535.
Doherty DG. Immunity, tolerance and autoimmunity in the liver: A comprehensive review. J Autoimmun. 2016;66:60–75.
Annunziato F, Cosmi L, Liotta F, Maggi E, Romagnani S. Human T helper type 1 dichotomy: origin, phenotype and biologic activities. Immunology. 2014;144:343–351.
Radziszewska A, Moulder Z, Jury EC, Ciurtin C. CD8(+) T cell phenotype and function in childhood and adult-onset connective tissue disease. Int J Mol Sci. 2022;23:11431.
Roberts SK, Therneau TM, Czaja AJ. Prognosis of histological cirrhosis in type 1 autoimmune hepatitis. Gastroenterology. 1996;110:848–857.
Czaja AJ, Carpenter HA. Progressive fibrosis during corticosteroid therapy of autoimmune hepatitis. Hepatology. 2004;39:1631–1638.
Ferber I, Schonrich G, Schenkel J et al. Levels of peripheral T cell tolerance induced by different doses of tolerogen. Science. 1994;263:674–676.
Kurts C, Sutherland RM, Davey G et al. CD8 T cell ignorance or tolerance to islet antigens depends on antigen dose. Proc Natl Acad Sci USA. 1999;96:12703–12707.
Voehringer D, Blaser C, Grawitz AB et al. Break of T cell ignorance to a viral antigen in the liver induces hepatitis. J Immunol. 2000;165:2415–2422.
Oldstone MB. Molecular mimicry as a mechanism for the cause and a probe uncovering etiologic agent(s) of autoimmune disease. Curr Top Microbiol Immunol. 1989;145:127–135.
Wucherpfennig KW, Strominger JL. Molecular mimicry in T cell-mediated autoimmunity: viral peptides activate human T cell clones specific for myelin basic protein. Cell. 1995;80:695–705.
Bogdanos DP, Choudhuri K, Vergani D. Molecular mimicry and autoimmune liver disease: virtuous intentions, malign consequences. Liver. 2001;21:225–232.
Oldstone MB. Molecular mimicry, microbial infection, and autoimmune disease: evolution of the concept. Curr Top Microbiol Immunol. 2005;296:1–17.
Bowen DG. Of mice and molecular mimicry: modeling autoimmune hepatitis. Hepatology. 2008;48:1013–1015.
Christen U, Hintermann E, Holdener M, von Herrath MG. Viral triggers for autoimmunity: is the “glass of molecular mimicry” half full or half empty? J Autoimmun. 2010;34:38–44.
Cusick MF, Libbey JE, Fujinami RS. Molecular mimicry as a mechanism of autoimmune disease. Clin Rev Allergy Immunol. 2012;42:102–111.
Oldstone MB. Molecular mimicry: its evolution from concept to mechanism as a cause of autoimmune diseases. Monoclon Antib Immunodiagn Immunother. 2014;33:158–165.
Christen U, Hintermann E. Pathogens and autoimmune hepatitis. Clin Exp Immunol. 2019;195:35–51.
Vahabi M, Ghazanfari T, Sepehrnia S. Molecular mimicry, hyperactive immune system, and SARS-COV-2 are three prerequisites of the autoimmune disease triangle following COVID-19 infection. Int Immunopharmacol. 2022;112:109183.
Ngu JH, Gearry RB, Frampton CM, Stedman CA. Autoimmune hepatitis: the role of environmental risk factors: a population-based study. Hepatol Int. 2013;7:869–875.
Lammert C, Chalasani SN, Green K et al. Patients with autoimmune hepatitis report lower lifetime coffee consumption. Dig Dis Sci. 2022;67:2594–2599.
Vieira SM, Pagovich OE, Kriegel MA. Diet, microbiota and autoimmune diseases. Lupus. 2014;23:518–526.
Wang H, Banerjee N, Liang Y et al. Gut microbiome-host interactions in driving environmental pollutant trichloroethene-mediated autoimmunity. Toxicol Appl Pharmacol. 2021;424:115597.
Avci E, Abasiyanik F. Autoimmune hepatitis after SARS-CoV-2 vaccine: New-onset or flare-up? J Autoimmun. 2021;125:102745.
Bril F, Al Diffalha S, Dean M, Fettig DM. Autoimmune hepatitis developing after coronavirus disease 2019 (COVID-19) vaccine: causality or casualty? J Hepatol. 2021;75:222–224.
Garrido I, Lopes S, Simoes MS et al. Autoimmune hepatitis after COVID-19 vaccine - more than a coincidence. J Autoimmun. 2021;125:102741.
McShane C, Kiat C, Rigby J, Crosbie O. The mRNA COVID-19 vaccine - A rare trigger of autoimmune hepatitis? J Hepatol. 2021;75:1252–1254.
Tan CK, Wong YJ, Wang LM, Ang TL, Kumar R. Autoimmune hepatitis following COVID-19 vaccination: true causality or mere association? J Hepatol. 2021;75:1250–1252.
Alhumaid S, Al Mutair A, Rabaan AA et al. New-onset and relapsed liver diseases following COVID-19 vaccination: a systematic review. BMC Gastroenterol. 2022;22:433.
Boettler T, Csernalabics B, Salie H et al. SARS-CoV-2 vaccination can elicit a CD8 T-cell dominant hepatitis. J Hepatol. 2022;77:653–659.
Chow KW, Pham NV, Ibrahim BM, Hong K, Saab S. Autoimmune hepatitis-like syndrome following COVID-19 vaccination: a systematic review of the literature. Dig Dis Sci. 2022;67:4574–4580.
Efe C, Kulkarni AV, Terziroli Beretta-Piccoli B et al. Liver injury after SARS-CoV-2 vaccination: features of immune-mediated hepatitis, role of corticosteroid therapy and outcome. Hepatology. 2022;76:1576–1586.
Eslami O, Lashkarizadeh M. Autoimmune hepatitis after coronavirus disease vaccination. J Gastrointestin Liver Dis. 2022;31:355–356.
Ferronato M, Lenzi M, Muratori L. Liver injury with autoimmune features after vaccination against SARS-CoV-2: the verdict is still open. Eur J Intern Med. 2023;108:108–110.
Floreani A, De Martin S. COVID-19 and autoimmune liver diseases. J Clin Med. 2022;11:2681.
Izagirre A, Arzallus T, Garmendia M et al. Autoimmune hepatitis following COVID-19 vaccination. J Autoimmun. 2022;132:102874.
Palla P, Vergadis C, Sakellariou S, Androutsakos T. Letter to the editor: autoimmune hepatitis after COVID-19 vaccination: A rare adverse effect? Hepatology. 2022;75:489–490.
Zafar M, Gordon K, Macken L et al. COVID-19 vaccination-induced cholangiopathy and autoimmune hepatitis: a series of two cases. Cureus. 2022;14:e30304.
Scott KP, Gratz SW, Sheridan PO, Flint HJ, Duncan SH. The influence of diet on the gut microbiota. Pharmacol Res. 2013;69:52–60.
Miyake Y, Yamamoto K. Role of gut microbiota in liver diseases. Hepatol Res. 2013;43:139–146.
Czaja AJ. Factoring the intestinal microbiome into the pathogenesis of autoimmune hepatitis. World J Gastroenterol. 2016;22:9257–9278.
Manzoor R, Ahmed W, Afify N et al. Trust your gut: the association of gut microbiota and liver disease. Microorganisms. 2022;10:1045.
Xu M, Luo K, Li J et al. Role of intestinal microbes in chronic liver diseases. Int J Mol Sci. 2022;23:12661.
Liwinski T, Heinemann M, Schramm C. The intestinal and biliary microbiome in autoimmune liver disease-current evidence and concepts. Semin Immunopathol. 2022;44:485–507.
Czaja AJ. Missing causality and heritability of autoimmune hepatitis. Dig Dis Sci. 2023;68:1585–1604.
Jirtle RL, Skinner MK. Environmental epigenomics and disease susceptibility. Nat Rev Genet. 2007;8:253–262.
Mathers JC, Strathdee G, Relton CL. Induction of epigenetic alterations by dietary and other environmental factors. Adv Genet. 2010;71:3–39.
Donohoe DR, Bultman SJ. Metaboloepigenetics: interrelationships between energy metabolism and epigenetic control of gene expression. J Cell Physiol. 2012;227:3169–3177.
Boyce WT, Kobor MS. Development and the epigenome: the “synapse” of gene-environment interplay. Dev Sci. 2015;18:1–23.
Czaja AJ. Epigenetic changes and their implications in autoimmune hepatitis. Eur J Clin Invest. 2018;48:e12899.
Aristizabal MJ, Anreiter I, Halldorsdottir T et al. Biological embedding of experience: a primer on epigenetics. Proc Natl Acad Sci USA. 2020;117:23261–23269.
Czaja AJ. Epigenetic aspects and prospects in autoimmune hepatitis. Front Immunol. 2022;13:921765.
Bogdanos DP, Baum H, Grasso A et al. Microbial mimics are major targets of crossreactivity with human pyruvate dehydrogenase in primary biliary cirrhosis. J Hepatol. 2004;40:31–39.
Wang JJ, Yang GX, Zhang WC et al. Escherichia coli infection induces autoimmune cholangitis and anti-mitochondrial antibodies in non-obese diabetic (NOD).B6 (Idd10/Idd18) mice. Clin Exp Immunol. 2014;175:192–201.
Smyk DS, Bogdanos DP, Kriese S et al. Urinary tract infection as a risk factor for autoimmune liver disease: from bench to bedside. Clin Res Hepatol Gastroenterol. 2012;36:110–121.
Padgett KA, Selmi C, Kenny TP et al. Phylogenetic and immunological definition of four lipoylated proteins from Novosphingobium aromaticivorans, implications for primary biliary cirrhosis. J Autoimmun. 2005;24:209–219.
Kaplan MM. Novosphingobium aromaticivorans: a potential initiator of primary biliary cirrhosis. Am J Gastroenterol. 2004;99:2147–2149.
Olafsson S, Gudjonsson H, Selmi C et al. Antimitochondrial antibodies and reactivity to N. aromaticivorans proteins in Icelandic patients with primary biliary cirrhosis and their relatives. Am J Gastroenterol. 2004;99:2143–2146.
Burroughs AK, Rosenstein IJ, Epstein O et al. Bacteriuria and primary biliary cirrhosis. Gut. 1984;25:133–137.
Bogdanos DP, Baum H, Butler P et al. Association between the primary biliary cirrhosis specific anti-sp100 antibodies and recurrent urinary tract infection. Dig Liver Dis. 2003;35:801–805.
Amano K, Leung PS, Rieger R et al. Chemical xenobiotics and mitochondrial autoantigens in primary biliary cirrhosis: identification of antibodies against a common environmental, cosmetic, and food additive, 2-octynoic acid. J Immunol. 2005;174:5874–5883.
Rieger R, Leung PS, Jeddeloh MR et al. Identification of 2-nonynoic acid, a cosmetic component, as a potential trigger of primary biliary cirrhosis. J Autoimmun. 2006;27:7–16.
Leung PS, Quan C, Park O et al. Immunization with a xenobiotic 6-bromohexanoate bovine serum albumin conjugate induces antimitochondrial antibodies. J Immunol. 2003;170:5326–5332.
Leung PS, Park O, Tsuneyama K et al. Induction of primary biliary cirrhosis in guinea pigs following chemical xenobiotic immunization. J Immunol. 2007;179:2651–2657.
Wakabayashi K, Lian ZX, Leung PS et al. Loss of tolerance in C57BL/6 mice to the autoantigen E2 subunit of pyruvate dehydrogenase by a xenobiotic with ensuing biliary ductular disease. Hepatology. 2008;48:531–540.
Wakabayashi K, Yoshida K, Leung PS et al. Induction of autoimmune cholangitis in non-obese diabetic (NOD).1101 mice following a chemical xenobiotic immunization. Clin Exp Immunol. 2009;155:577–586.
Gershwin ME, Mackay IR. The causes of primary biliary cirrhosis: convenient and inconvenient truths. Hepatology. 2008;47:737–745.
Jones D, Manns MP, Terracciano L, Torbenson M, Vierling JM. Unmet needs and new models for future trials in autoimmune hepatitis. Lancet Gastroenterol Hepatol. 2018;3:363–370.
Vierling JM, Kerkar N, Czaja AJ et al. Immunosuppressive treatment regimens in autoimmune hepatitis: systematic reviews and meta-analyses supporting American Association for the Study of Liver Diseases guidelines. Hepatology. 2020;72:753–769.
Czaja AJ. Examining pathogenic concepts of autoimmune hepatitis for cues to future investigations and interventions. World J Gastroenterol. 2019;25:6579–6606.
Christen U, Hintermann E. Pathogen infection as a possible cause for autoimmune hepatitis. Int Rev Immunol. 2014;33:296–313.
Ehser J, Holdener M, Christen S et al. Molecular mimicry rather than identity breaks T-cell tolerance in the CYP2D6 mouse model for human autoimmune hepatitis. J Autoimmun. 2013;42:39–49.
von Herrath MG, Fujinami RS, Whitton JL. Microorganisms and autoimmunity: making the barren field fertile? Nat Rev Microbiol. 2003;1:151–157.
Wang P, Sidney J, Kim Y et al. Peptide binding predictions for HLA DR, DP and DQ molecules. BMC Bioinform. 2010;11:568.
Fineschi B, Miller J. Endosomal proteases and antigen processing. Trends Biochem Sci. 1997;22:377–382.
Rock KL, Reits E, Neefjes J. Present yourself! by MHC class I and MHC class II molecules. Trends Immunol. 2016;37:724–737.
Ting JP, Trowsdale J. Genetic control of MHC class II expression. Cell. 2002;109:S21-33.
Brown JH, Jardetzky T, Saper MA et al. A hypothetical model of the foreign antigen binding site of class II histocompatibility molecules. Nature. 1988;332:845–850.
Stern LJ, Brown JH, Jardetzky TS et al. Crystal structure of the human class II MHC protein HLA-DR1 complexed with an influenza virus peptide. Nature. 1994;368:215–221.
Dessen A, Lawrence CM, Cupo S, Zaller DM, Wiley DC. X-ray crystal structure of HLA-DR4 (DRA*0101, DRB1*0401) complexed with a peptide from human collagen II. Immunity. 1997;7:473–481.
Tollefsen S, Hotta K, Chen X et al. Structural and functional studies of trans-encoded HLA-DQ2.3 (DQA1*03:01/DQB1*02:01) protein molecule. J Biol Chem. 2012;287:13611–13619.
Doherty DG, Donaldson PT, Underhill JA et al. Allelic sequence variation in the HLA class II genes and proteins in patients with autoimmune hepatitis. Hepatology. 1994;19:609–615.
Jones EY, Fugger L, Strominger JL, Siebold C. MHC class II proteins and disease: a structural perspective. Nat Rev Immunol. 2006;6:271–282.
Strettell MD, Donaldson PT, Thomson LJ et al. Allelic basis for HLA-encoded susceptibility to type 1 autoimmune hepatitis. Gastroenterology. 1997;112:2028–2035.
Czaja AJ, Donaldson PT. Genetic susceptibilities for immune expression and liver cell injury in autoimmune hepatitis. Immunol Rev. 2000;174:250–259.
Ferrante A, Templeton M, Hoffman M, Castellini MJ. The thermodynamic mechanism of peptide-MHC class II complex formation is a determinant of susceptibility to HLA-DM. J Immunol. 2015;195:1251–1261.
Wieczorek M, Sticht J, Stolzenberg S et al. MHC class II complexes sample intermediate states along the peptide exchange pathway. Nat Commun. 2016;7:13224.
Wieczorek M, Abualrous ET, Sticht J et al. Major histocompatibility complex (MHC) class I and MHC class II proteins: conformational plasticity in antigen presentation. Front Immunol. 2017;8:292.
Chicz RM, Urban RG, Lane WS et al. Predominant naturally processed peptides bound to HLA-DR1 are derived from MHC-related molecules and are heterogeneous in size. Nature. 1992;358:764–768.
Steimle V, Siegrist CA, Mottet A, Lisowska-Grospierre B, Mach B. Regulation of MHC class II expression by interferon-gamma mediated by the transactivator gene CIITA. Science. 1994;265:106–109.
Muhlethaler-Mottet A, Otten LA, Steimle V, Mach B. Expression of MHC class II molecules in different cellular and functional compartments is controlled by differential usage of multiple promoters of the transactivator CIITA. EMBO J. 1997;16:2851–2860.
Morris AC, Beresford GW, Mooney MR, Boss JM. Kinetics of a gamma interferon response: expression and assembly of CIITA promoter IV and inhibition by methylation. Mol Cell Biol. 2002;22:4781–4791.
Norbury CC, Malide D, Gibbs JS, Bennink JR, Yewdell JW. Visualizing priming of virus-specific CD8+ T cells by infected dendritic cells in vivo. Nat Immunol. 2002;3:265–271.
Norbury CC, Basta S, Donohue KB et al. CD8+ T cell cross-priming via transfer of proteasome substrates. Science. 2004;304:1318–1321.
Rock KL, Shen L. Cross-presentation: underlying mechanisms and role in immune surveillance. Immunol Rev. 2005;207:166–183.
Embgenbroich M, Burgdorf S. Current concepts of antigen cross-presentation. Front Immunol. 2018;9:1643.
Cresswell P. Assembly, transport, and function of MHC class II molecules. Annu Rev Immunol. 1994;12:259–293.
Villadangos JA, Cardoso M, Steptoe RJ et al. MHC class II expression is regulated in dendritic cells independently of invariant chain degradation. Immunity. 2001;14:739–749.
Dudziak D, Kamphorst AO, Heidkamp GF et al. Differential antigen processing by dendritic cell subsets in vivo. Science. 2007;315:107–111.
Villadangos JA, Schnorrer P. Intrinsic and cooperative antigen-presenting functions of dendritic-cell subsets in vivo. Nat Rev Immunol. 2007;7:543–555.
Kurts C, Robinson BW, Knolle PA. Cross-priming in health and disease. Nat Rev Immunol. 2010;10:403–414.
Holdener M, Hintermann E, Bayer M et al. Breaking tolerance to the natural human liver autoantigen cytochrome P450 2D6 by virus infection. J Exp Med. 2008;205:1409–1422.
Hintermann E, Ehser J, Christen U. The CYP2D6 animal model: how to induce autoimmune hepatitis in mice. J Vis Exp. 2012;60:e3644.
Lapierre P, Djilali-Saiah I, Vitozzi S, Alvarez F. A murine model of type 2 autoimmune hepatitis: xenoimmunization with human antigens. Hepatology. 2004;39:1066–1074.
Lapierre P, Beland K, Djilali-Saiah I, Alvarez F. Type 2 autoimmune hepatitis murine model: the influence of genetic background in disease development. J Autoimmun. 2006;26:82–89.
Manns MP, Griffin KJ, Sullivan KF, Johnson EF. LKM-1 autoantibodies recognize a short linear sequence in P450IID6, a cytochrome P-450 monooxygenase. J Clin Invest. 1991;88:1370–1378.
Czaja AJ, Manns MP. The validity and importance of subtypes in autoimmune hepatitis: a point of view. Am J Gastroenterol. 1995;90:1206–1211.
Kerkar N, Choudhuri K, Ma Y et al. Cytochrome P4502D6(193–212): a new immunodominant epitope and target of virus/self cross-reactivity in liver kidney microsomal autoantibody type 1-positive liver disease. J Immunol. 2003;170:1481–1489.
Lapierre P, Hajoui O, Homberg JC, Alvarez F. Formiminotransferase cyclodeaminase is an organ-specific autoantigen recognized by sera of patients with autoimmune hepatitis. Gastroenterology. 1999;116:643–649.
Muratori L, Sztul E, Muratori P et al. Distinct epitopes on formiminotransferase cyclodeaminase induce autoimmune liver cytosol antibody type 1. Hepatology. 2001;34:494–501.
Hintermann E, Ehser J, Bayer M, Pfeilschifter JM, Christen U. Mechanism of autoimmune hepatic fibrogenesis induced by an adenovirus encoding the human liver autoantigen cytochrome P450 2D6. J Autoimmun. 2013;44:49–60.
Vento S, Cainelli F. Is there a role for viruses in triggering autoimmune hepatitis? Autoimmun Rev. 2004;3:61–69.
Gregorio GV, Choudhuri K, Ma Y et al. Mimicry between the hepatitis B virus DNA polymerase and the antigenic targets of nuclear and smooth muscle antibodies in chronic hepatitis B virus infection. J Immunol. 1999;162:1802–1810.
Gueguen M, Boniface O, Bernard O et al. Identification of the main epitope on human cytochrome P450 IID6 recognized by anti-liver kidney microsome antibody. J Autoimmun. 1991;4:607–615.
Yamamoto AM, Cresteil D, Boniface O, Clerc FF, Alvarez F. Identification and analysis of cytochrome P450IID6 antigenic sites recognized by anti-liver-kidney microsome type-1 antibodies (LKM1). Eur J Immunol. 1993;23:1105–1111.
Kitazawa E, Igarashi T, Kawaguchi N et al. Differences in anti-LKM-1 autoantibody immunoreactivity to CYP2D6 antigenic sites between hepatitis C virus-negative and -positive patients. J Autoimmun. 2001;17:243–249.
Ma Y, Thomas MG, Okamoto M et al. Key residues of a major cytochrome P4502D6 epitope are located on the surface of the molecule. J Immunol. 2002;169:277–285.
Gregorio GV, Choudhuri K, Ma Y et al. Mimicry between the hepatitis C virus polyprotein and antigenic targets of nuclear and smooth muscle antibodies in chronic hepatitis C virus infection. Clin Exp Immunol. 2003;133:404–413.
Marceau G, Lapierre P, Beland K, Soudeyns H, Alvarez F. LKM1 autoantibodies in chronic hepatitis C infection: a case of molecular mimicry? Hepatology. 2005;42:675–682.
Paiardini A, Pascarella S. Structural mimicry between SLA/LP and Rickettsia surface antigens as a driver of autoimmune hepatitis: insights from an in silico study. Theor Biol Med Model. 2013;10:25.
Czaja AJ, Abdulkarim AS, Carpenter HA et al. GB virus-C infection in type 1 autoimmune hepatitis. Mayo Clin Proc. 1998;73:412–418.
Christen U, von Herrath MG. Induction, acceleration or prevention of autoimmunity by molecular mimicry. Mol Immunol. 2004;40:1113–1120.
McRae BL, Vanderlugt CL, Dal Canto MC, Miller SD. Functional evidence for epitope spreading in the relapsing pathology of experimental autoimmune encephalomyelitis. J Exp Med. 1995;182:75–85.
Rosen A, Casciola-Rosen L. Autoantigens in systemic autoimmunity: critical partner in pathogenesis. J Intern Med. 2009;265:625–631.
Yuksel M, Wang Y, Tai N et al. A novel “humanized mouse” model for autoimmune hepatitis and the association of gut microbiota with liver inflammation. Hepatology. 2015;62:1536–1550.
Lin R, Zhou L, Zhang J, Wang B. Abnormal intestinal permeability and microbiota in patients with autoimmune hepatitis. Int J Clin Exp Pathol. 2015;8:5153–5160.
Rojas M, Restrepo-Jimenez P, Monsalve DM et al. Molecular mimicry and autoimmunity. J Autoimmun. 2018;95:100–123.
Manfredo Vieira S, Hiltensperger M, Kumar V et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science. 2018;359:1156–1161.
Wei Y, Li Y, Yan L et al. Alterations of gut microbiome in autoimmune hepatitis. Gut. 2020;69:569–577.
Lou J, Jiang Y, Rao B et al. Fecal microbiomes distinguish patients with autoimmune hepatitis from healthy individuals. Front Cell Infect Microbiol. 2020;10:342.
Liwinski T, Casar C, Ruehlemann MC et al. A disease-specific decline of the relative abundance of Bifidobacterium in patients with autoimmune hepatitis. Aliment Pharmacol Ther. 2020;51:1417–1428.
Cheng Z, Yang L, Chu H. The gut microbiota: a novel player in autoimmune hepatitis. Front Cell Infect Microbiol. 2022;12:947382.
Lehmann PV, Forsthuber T, Miller A, Sercarz EE. Spreading of T-cell autoimmunity to cryptic determinants of an autoantigen. Nature. 1992;358:155–157.
Vanderlugt CL, Miller SD. Epitope spreading in immune-mediated diseases: implications for immunotherapy. Nat Rev Immunol. 2002;2:85–95.
Hintermann E, Holdener M, Bayer M et al. Epitope spreading of the anti-CYP2D6 antibody response in patients with autoimmune hepatitis and in the CYP2D6 mouse model. J Autoimmun. 2011;37:242–253.
Cornaby C, Gibbons L, Mayhew V et al. B cell epitope spreading: mechanisms and contribution to autoimmune diseases. Immunol Lett. 2015;163:56–68.
Eberl G, Brawand P, MacDonald HR. Selective bystander proliferation of memory CD4+ and CD8+ T cells upon NK T or T cell activation. J Immunol. 2000;165:4305–4311.
Boyman O. Bystander activation of CD4+ T cells. Eur J Immunol. 2010;40:936–939.
Lenzi M, Ballardini G, Fusconi M et al. Type 2 autoimmune hepatitis and hepatitis C virus infection. Lancet. 1990;335:258–259.
Lenzi M, Bellentani S, Saccoccio G et al. Prevalence of non-organ-specific autoantibodies and chronic liver disease in the general population: a nested case-control study of the Dionysos cohort. Gut. 1999;45:435–441.
Czaja AJ, Carpenter HA, Santrach PJ et al. Evidence against hepatitis viruses as important causes of severe autoimmune hepatitis in the United States. J Hepatol. 1993;18:342–352.
Lucchese G, Stufano A, Trost B, Kusalik A, Kanduc D. Peptidology: short amino acid modules in cell biology and immunology. Amino Acids. 2007;33:703–707.
Kanduc D. Pentapeptides as minimal functional units in cell biology and immunology. Curr Protein Pept Sci. 2013;14:111–120.
Czaja AJ. Performance parameters of the conventional serological markers for autoimmune hepatitis. Dig Dis Sci. 2011;56:545–554.
Gelpi C, Sontheimer EJ, Rodriguez-Sanchez JL. Autoantibodies against a serine tRNA-protein complex implicated in cotranslational selenocysteine insertion. Proc Natl Acad Sci USA. 1992;89:9739–9743.
Costa M, Rodriguez-Sanchez JL, Czaja AJ, Gelpi C. Isolation and characterization of cDNA encoding the antigenic protein of the human tRNP(Ser)Sec complex recognized by autoantibodies from patients withtype-1 autoimmune hepatitis. Clin Exp Immunol. 2000;121:364–374.
Wies I, Brunner S, Henninger J et al. Identification of target antigen for SLA/LP autoantibodies in autoimmune hepatitis. Lancet. 2000;355:1510–1515.
Herkel J, Heidrich B, Nieraad N et al. Fine specificity of autoantibodies to soluble liver antigen and liver/pancreas. Hepatology. 2002;35:403–408.
Baeres M, Herkel J, Czaja AJ et al. Establishment of standardised SLA/LP immunoassays: specificity for autoimmune hepatitis, worldwide occurrence, and clinical characteristics. Gut. 2002;51:259–264.
Czaja AJ, Carpenter HA, Manns MP. Antibodies to soluble liver antigen, P450IID6, and mitochondrial complexes in chronic hepatitis. Gastroenterology. 1993;105:1522–1528.
Czaja AJ, Donaldson PT, Lohse AW. Antibodies to soluble liver antigen/liver pancreas and HLA risk factors for type 1 autoimmune hepatitis. Am J Gastroenterol. 2002;97:413–419.
Frykholm BO. On the question of infectious aetiologies for multiple sclerosis, schizophrenia and the chronic fatigue syndrome and their treatment with antibiotics. Med Hypotheses. 2009;72:736–739.
Iavarone C, O’Hagan DT, Yu D, Delahaye NF, Ulmer JB. Mechanism of action of mRNA-based vaccines. Expert Rev Vaccines. 2017;16:871–881.
Walsh EE, Frenck RW Jr, Falsey AR et al. Safety and immunogenicity of two RNA-based Covid-19 vaccine candidates. N Engl J Med. 2020;383:2439–2450.
Lamb YN. BNT162b2 mRNA COVID-19 vaccine: first approval. Drugs. 2021;81:495–501.
Khoshnood S, Ghanavati R, Shirani M et al. Viral vector and nucleic acid vaccines against COVID-19: a narrative review. Front Microbiol. 2022;13:984536.
Heidary M, Kaviar VH, Shirani M et al. A comprehensive review of the protein subunit vaccines against COVID-19. Front Microbiol. 2022;13:927306.
Chen J, Chen J, Xu Q. Current developments and challenges of mRNA vaccines. Annu Rev Biomed Eng. 2022;24:85–109.
Corbett KS, Edwards DK, Leist SR et al. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness. Nature. 2020;586:567–571.
Sahin U, Muik A, Derhovanessian E et al. COVID-19 vaccine BNT162b1 elicits human antibody and TH1 T cell responses. Nature. 2020;586:594–599.
Sahin U, Muik A, Vogler I et al. BNT162b2 vaccine induces neutralizing antibodies and poly-specific T cells in humans. Nature. 2021;595:572–577.
Costa C, Scozzari G, Migliore E et al. Cellular immune response to BNT162b2 mRNA COVID-19 vaccine in a large cohort of healthcare workers in a tertiary care university hospital. Vaccines (Basel). 2022;10:1031.
Kanduc D. Peptide cross-reactivity: the original sin of vaccines. Front Biosci (Schol Ed). 2012;4:1393–1401.
Segal Y, Shoenfeld Y. Vaccine-induced autoimmunity: the role of molecular mimicry and immune crossreaction. Cell Mol Immunol. 2018;15:586–594.
Kanduc D, Shoenfeld Y. Molecular mimicry between SARS-CoV-2 spike glycoprotein and mammalian proteomes: implications for the vaccine. Immunol Res. 2020;68:310–313.
Kanduc D. From anti-SARS-CoV-2 immune responses to COVID-19 via molecular mimicry. Antibodies (Basel). 2020;9:33.
Rodriguez Y, Novelli L, Rojas M et al. Autoinflammatory and autoimmune conditions at the crossroad of COVID-19. J Autoimmun. 2020;114:102506.
Mack CL, Adams D, Assis DN et al. Diagnosis and management of autoimmune hepatitis in adults and children: 2019 practice guidance and guidelines from the American Association for the Study of Liver Diseases. Hepatology. 2020;72:671–722.
Czaja AJ. Acute and acute severe (fulminant) autoimmune hepatitis. Dig Dis Sci. 2013;58:897–914.
Lleo A, Cazzagon N, Rigamonti C et al. Clinical update on risks and efficacy of anti-SARS-CoV-2 vaccines in patients with autoimmune hepatitis and summary of reports on post-vaccination liver injury. Dig Liver Dis. 2022;54:722–726.
Rodriguez Y, Rojas M, Beltran S et al. Autoimmune and autoinflammatory conditions after COVID-19 vaccination. New case reports and updated literature review. J Autoimmun. 2022;132:102898.
Ruther DF, Weltzsch JP, Schramm C, Sebode M, Lohse AW. Autoimmune hepatitis and COVID-19: no increased risk for AIH after vaccination but reduced care. J Hepatol. 2022;77:250–251.
Kabacam G, Wahlin S, Efe C. Autoimmune hepatitis triggered by COVID-19: a report of two cases. Liver Int. 2021;41:2527–2528.
Marabotto E, Ziola S, Sheijani AD, Giannini EG. COVID-19 and liver disease: not all evil comes to harm. Liver Int. 2021;41:237–238.
Volchkova EA, Legkova KS. Topchy TB [COVID-19 as a trigger of autoimmune hepatitis. case report]. Ter Arkh 2022;94:259–264.
Marino Gammazza A, Legare S, Lo Bosco G et al. Human molecular chaperones share with SARS-CoV-2 antigenic epitopes potentially capable of eliciting autoimmunity against endothelial cells: possible role of molecular mimicry in COVID-19. Cell Stress Chaperones. 2020;25:737–741.
Vojdani A, Kharrazian D. Potential antigenic cross-reactivity between SARS-CoV-2 and human tissue with a possible link to an increase in autoimmune diseases. Clin Immunol. 2020;217:108480.
Wiest R, Garcia-Tsao G. Bacterial translocation (BT) in cirrhosis. Hepatology. 2005;41:422–433.
Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R. Diversity, stability and resilience of the human gut microbiota. Nature. 2012;489:220–230.
Hollister EB, Gao C, Versalovic J. Compositional and functional features of the gastrointestinal microbiome and their effects on human health. Gastroenterology. 2014;146:1449–1458.
Yuksel M, Laukens D, Heindryckx F et al. Hepatitis mouse models: from acute-to-chronic autoimmune hepatitis. Int J Exp Pathol. 2014;95:309–320.
Sokol H, Seksik P, Rigottier-Gois L et al. Specificities of the fecal microbiota in inflammatory bowel disease. Inflamm Bowel Dis. 2006;12:106–111.
Wen L, Ley RE, Volchkov PY et al. Innate immunity and intestinal microbiota in the development of Type 1 diabetes. Nature. 2008;455:1109–1113.
Wu HJ, Ivanov II, Darce J et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity. 2010;32:815–827.
Ruff WE, Dehner C, Kim WJ et al. Pathogenic autoreactive T and B cells cross-react with mimotopes expressed by a common human gut commensal to trigger autoimmunity. Cell Host Microbe. 2019;26:100–113.
Biagioli M, Carino A, Fiorucci C et al. GPBAR1 functions as gatekeeper for liver NKT cells and provides counterregulatory signals in mouse models of immune-mediated hepatitis. Cell Mol Gastroenterol Hepatol. 2019;8:447–473.
Frank DN, Zhu W, Sartor RB, Li E. Investigating the biological and clinical significance of human dysbioses. Trends Microbiol. 2011;19:427–434.
Rogier R, Koenders MI, Abdollahi-Roodsaz S. Toll-like receptor mediated modulation of T cell response by commensal intestinal microbiota as a trigger for autoimmune arthritis. J Immunol Res. 2015;2015:527696.
Elsherbiny NM, Rammadan M, Hassan EA et al. Autoimmune hepatitis: shifts in gut microbiota and metabolic pathways among Egyptian patients. Microorganisms. 2020;8:1011.
Zhang H, Liu M, Liu X et al. Bifidobacterium animalis ssp. Lactis 420 mitigates autoimmune hepatitis through regulating intestinal barrier and liver immune cells. Front Immunol. 2020;11:569104.
Zhang H, Liu M, Zhong W et al. Leaky gut driven by dysbiosis augments activation and accumulation of liver macrophages via RIP3 signaling pathway in autoimmune hepatitis. Front Immunol. 2021;12:624360.
Liu Q, Tian H, Kang Y et al. Probiotics alleviate autoimmune hepatitis in mice through modulation of gut microbiota and intestinal permeability. J Nutr Biochem. 2021;98:108863.
Macia L, Thorburn AN, Binge LC et al. Microbial influences on epithelial integrity and immune function as a basis for inflammatory diseases. Immunol Rev. 2012;245:164–176.
Guo S, Nighot M, Al-Sadi R et al. Lipopolysaccharide regulation of intestinal tight junction permeability Is mediated by TLR4 signal transduction pathway activation of FAK and MyD88. J Immunol. 2015;195:4999–5010.
Cummings JH, Pomare EW, Branch WJ, Naylor CP, Macfarlane GT. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut. 1987;28:1221–1227.
Flint HJ, Scott KP, Duncan SH, Louis P, Forano E. Microbial degradation of complex carbohydrates in the gut. Gut Microbes. 2012;3:289–306.
Morrison DJ, Preston T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes. 2016;7:189–200.
Mizuno M, Noto D, Kaga N, Chiba A, Miyake S. The dual role of short fatty acid chains in the pathogenesis of autoimmune disease models. PLoS ONE. 2017;12:e0173032.
Milani C, Turroni F, Duranti S et al. Genomics of the genus Bifidobacterium reveals species-specific adaptation to the glycan-rich gut environment. Appl Environ Microbiol. 2016;82:980–991.
Wang HB, Wang PY, Wang X, Wan YL, Liu YC. Butyrate enhances intestinal epithelial barrier function via up-regulation of tight junction protein Claudin-1 transcription. Dig Dis Sci. 2012;57:3126–3135.
Peng L, Li ZR, Green RS, Holzman IR, Lin J. Butyrate enhances the intestinal barrier by facilitating tight junction assembly via activation of AMP-activated protein kinase in Caco-2 cell monolayers. J Nutr. 2009;139:1619–1625.
Peng L, He Z, Chen W, Holzman IR, Lin J. Effects of butyrate on intestinal barrier function in a Caco-2 cell monolayer model of intestinal barrier. Pediatr Res. 2007;61:37–41.
Kelly CJ, Zheng L, Campbell EL et al. Crosstalk between microbiota-derived short-chain fatty acids and intestinal epithelial HIF augments tissue barrier function. Cell Host Microbe. 2015;17:662–671.
Thangaraju M, Cresci GA, Liu K et al. GPR109A is a G-protein-coupled receptor for the bacterial fermentation product butyrate and functions as a tumor suppressor in colon. Cancer Res. 2009;69:2826–2832.
Furusawa Y, Obata Y, Fukuda S et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature. 2013;504:446–450.
Sheng L, Jena PK, Hu Y et al. Hepatic inflammation caused by dysregulated bile acid synthesis is reversible by butyrate supplementation. J Pathol. 2017;243:431–441.
Siddiqui MT, Cresci GAM. The immunomodulatory functions of butyrate. J Inflamm Res. 2021;14:6025–6041.
Hu ED, Chen DZ, Wu JL et al. High fiber dietary and sodium butyrate attenuate experimental autoimmune hepatitis through regulation of immune regulatory cells and intestinal barrier. Cell Immunol. 2018;328:24–32.
Hill C, Guarner F, Reid G et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol. 2014;11:506–514.
Bron PA, Kleerebezem M, Brummer RJ et al. Can probiotics modulate human disease by impacting intestinal barrier function? Br J Nutr. 2017;117:93–107.
Beyaz Coskun A, Sagdicoglu Celep AG. Therapeutic modulation methods of gut microbiota and gut-liver axis. Crit Rev Food Sci Nutr. 2022;62:6505–6515.
Rios-Covian D, Gueimonde M, Duncan SH, Flint HJ, de los Reyes-Gavilan CG. Enhanced butyrate formation by cross-feeding between Faecalibacterium prausnitzii and Bifidobacterium adolescentis. FEMS Microbiol Lett. 2015;362:176.
Montano-Loza AJ, Czaja AJ. Cell mediators of autoimmune hepatitis and their therapeutic implications. Dig Dis Sci. 2014;60:1528–1542.
Wei Y, Zeng B, Chen J et al. Enterogenous bacterial glycolipids are required for the generation of natural killer T cells mediated liver injury. Sci Rep. 2016;6:36365.
Targan SR, Landers C, Vidrich A, Czaja AJ. High-titer antineutrophil cytoplasmic antibodies in type-1 autoimmune hepatitis. Gastroenterology. 1995;108:1159–1166.
Bansi D, Chapman R, Fleming K. Antineutrophil cytoplasmic antibodies in chronic liver diseases: prevalence, titre, specificity and IgG subclass. J Hepatol. 1996;24:581–586.
Zauli D, Ghetti S, Grassi A et al. Anti-neutrophil cytoplasmic antibodies in type 1 and 2 autoimmune hepatitis. Hepatology. 1997;25:1105–1107.
Roozendaal C, Kallenberg CG. Anti-neutrophil cytoplasm autoantibodies (ANCA) in autoimmune liver diseases. Hepatogastroenterology. 1999;46:3034–3040.
Lindgren S, Nilsson S, Nassberger L, Verbaan H, Wieslander J. Anti-neutrophil cytoplasmic antibodies in patients with chronic liver diseases: prevalence, antigen specificity and predictive value for diagnosis of autoimmune liver disease. Swedish Internal Medicine Liver Club (SILK). J Gastroenterol Hepatol. 2000;15:437–442.
Terjung B, Bogsch F, Klein R et al. Diagnostic accuracy of atypical p-ANCA in autoimmune hepatitis using ROC- and multivariate regression analysis. Eur J Med Res. 2004;9:439–448.
Duerr RH, Targan SR, Landers CJ et al. Neutrophil cytoplasmic antibodies: a link between primary sclerosing cholangitis and ulcerative colitis. Gastroenterology. 1991;100:1385–1391.
Halbwachs-Mecarelli L, Nusbaum P, Noel LH et al. Antineutrophil cytoplasmic antibodies (ANCA) directed against cathepsin G in ulcerative colitis, Crohn’s disease and primary sclerosing cholangitis. Clin Exp Immunol. 1992;90:79–84.
Saxon A, Shanahan F, Landers C, Ganz T, Targan S. A distinct subset of antineutrophil cytoplasmic antibodies is associated with inflammatory bowel disease. J Allergy Clin Immunol. 1990;86:202–210.
Duerr RH, Targan SR, Landers CJ, Sutherland LR, Shanahan F. Anti-neutrophil cytoplasmic antibodies in ulcerative colitis. Comparison with other colitides/diarrheal illnesses. Gastroenterology. 1991;100:1590–1596.
Cambridge G, Rampton DS, Stevens TR et al. Anti-neutrophil antibodies in inflammatory bowel disease: prevalence and diagnostic role. Gut. 1992;33:668–674.
Terjung B, Sohne J, Lechtenberg B et al. p-ANCAs in autoimmune liver disorders recognise human beta-tubulin isotype 5 and cross-react with microbial protein FtsZ. Gut. 2010;59:808–816.
Terjung B, Spengler U. Atypical p-ANCA in PSC and AIH: a hint toward a “leaky gut”? Clin Rev Allergy Immunol. 2009;36:40–51.
Liu Q, He W, Tang R, Ma X. Intestinal homeostasis in autoimmune liver diseases. Chin Med J (Engl). 2022;135:1642–1652.
Kanduc D, Stufano A, Lucchese G, Kusalik A. Massive peptide sharing between viral and human proteomes. Peptides. 2008;29:1755–1766.
Trost B, Lucchese G, Stufano A et al. No human protein is exempt from bacterial motifs, not even one. Self Nonself. 2010;1:328–334.
Srinivasappa J, Saegusa J, Prabhakar BS et al. Molecular mimicry: frequency of reactivity of monoclonal antiviral antibodies with normal tissues. J Virol. 1986;57:397–401.
Vojdani A. A potential link between environmental triggers and autoimmunity. Autoimmune Dis. 2014;2014:437231.
Vojdani A, Pollard KM, Campbell AW. Environmental triggers and autoimmunity. Autoimmune Dis. 2014;2014:798029.
Vojdani A. Molecular mimicry as a mechanism for food immune reactivities and autoimmunity. Altern Ther Health Med. 2015;21:34–45.
Ruff WE, Kriegel MA. Autoimmune host-microbiota interactions at barrier sites and beyond. Trends Mol Med. 2015;21:233–244.
Ruff WE, Vieira SM, Kriegel MA. The role of the gut microbiota in the pathogenesis of antiphospholipid syndrome. Curr Rheumatol Rep. 2015;17:472.
Czaja AJ, Manns MP, Homburger HA. Frequency and significance of antibodies to liver/kidney microsome type 1 in adults with chronic active hepatitis. Gastroenterology. 1992;103:1290–1295.
Bairoch A, Apweiler R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 2000;28:45–48.
Whittemore K, Johnston SA, Sykes K, Shen L. A general method to discover epitopes from sera. PLoS ONE. 2016;11:e0157462.
Geysen HM, Rodda SJ, Mason TJ. A priori delineation of a peptide which mimics a discontinuous antigenic determinant. Mol Immunol. 1986;23:709–715.
Meloen RH, Puijk WC, Slootstra JW. Mimotopes: realization of an unlikely concept. J Mol Recognit. 2000;13:352–359.
Huang J, Ru B, Zhu P et al. MimoDB 2.0: a mimotope database and beyond. Nucleic Acids Res. 2012;40:D271-277.
Vojdani A, Vojdani E, Melgar AL, Redd J. Reaction of SARS-CoV-2 antibodies with other pathogens, vaccines, and food antigens. Front Immunol. 2022;13:1003094.
Vanderlugt CL, Begolka WS, Neville KL et al. The functional significance of epitope spreading and its regulation by co-stimulatory molecules. Immunol Rev. 1998;164:63–72.
Schiffenbauer J. Superantigens and their role in autoimmune disorders. Arch Immunol Ther Exp (Warsz). 1999;47:17–24.
Llewelyn M, Cohen J. Superantigens: microbial agents that corrupt immunity. Lancet Infect Dis. 2002;2:156–162.
Paladino L, Vitale AM, Caruso Bavisotto C et al. The role of molecular chaperones in virus infection and implications for understanding and treating COVID-19. J Clin Med. 2020;9:3518.
Frydman J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu Rev Biochem. 2001;70:603–647.
Young JC, Agashe VR, Siegers K, Hartl FU. Pathways of chaperone-mediated protein folding in the cytosol. Nat Rev Mol Cell Biol. 2004;5:781–791.
Czaja AJ. Nature and implications of oxidative and nitrosative stresses in autoimmune hepatitis. Dig Dis Sci. 2016;61:2784–2803.
Tukaj S, Kaminski M. Heat shock proteins in the therapy of autoimmune diseases: too simple to be true? Cell Stress Chaperones. 2019;24:475–479.
Androvitsanea A, Stylianou K, Drosataki E, Petrakis I. The pathophysiological role of heat shock response in autoimmunity: a literature review. Cells. 2021;10:2626.
Rajaiah R, Moudgil KD. Heat-shock proteins can promote as well as regulate autoimmunity. Autoimmun Rev. 2009;8:388–393.
Murshid A, Gong J, Calderwood SK. The role of heat shock proteins in antigen cross presentation. Front Immunol. 2012;3:63.
van Eden W. Immune tolerance therapies for autoimmune diseases based on heat shock protein T-cell epitopes. Philos Trans R Soc Lond B Biol Sci. 2018;373:20160531.
Tukaj S. Heat shock protein 70 as a double agent acting inside and outside the cell: insights into autoimmunity. Int J Mol Sci. 2020;21:5298.
Altinok S, Sanchez-Hodge R, Stewart M, Smith K, Schisler JC. With or without you: co-chaperones mediate health and disease by modifying chaperone function and protein triage. Cells. 2021;10:3121.
Sadasivan B, Lehner PJ, Ortmann B, Spies T, Cresswell P. Roles for calreticulin and a novel glycoprotein, tapasin, in the interaction of MHC class I molecules with TAP. Immunity. 1996;5:103–114.
Ortmann B, Copeman J, Lehner PJ et al. A critical role for tapasin in the assembly and function of multimeric MHC class I-TAP complexes. Science. 1997;277:1306–1309.
Boyle LH, Hermann C, Boname JM et al. Tapasin-related protein TAPBPR is an additional component of the MHC class I presentation pathway. Proc Natl Acad Sci USA. 2013;110:3465–3470.
Hermann C, Trowsdale J, Boyle LH. TAPBPR: a new player in the MHC class I presentation pathway. Tissue Antigens. 2015;85:155–166.
Hermann C, van Hateren A, Trautwein N et al. TAPBPR alters MHC class I peptide presentation by functioning as a peptide exchange catalyst. Elife. 2015;4:e09617.
Morozov GI, Zhao H, Mage MG et al. Interaction of TAPBPR, a tapasin homolog, with MHC-I molecules promotes peptide editing. Proc Natl Acad Sci USA. 2016;113:E1006-1015.
Cuervo AM, Dice JF. Unique properties of lamp2a compared to other lamp2 isoforms. J Cell Sci. 2000;113(Pt 24):4441–4450.
Zhou D, Li P, Lin Y et al. Lamp-2a facilitates MHC class II presentation of cytoplasmic antigens. Immunity. 2005;22:571–581.
Rangachari M, Zhu C, Sakuishi K et al. Bat3 promotes T cell responses and autoimmunity by repressing Tim-3-mediated cell death and exhaustion. Nat Med. 2012;18:1394–1400.
Zhu C, Dixon KO, Newcomer K et al. Tim-3 adaptor protein Bat3 is a molecular checkpoint of T cell terminal differentiation and exhaustion. Sci Adv. 2021;7:eabd2710.
Manns MP, Johnson EF, Griffin KJ, Tan EM, Sullivan KF. Major antigen of liver kidney microsomal autoantibodies in idiopathic autoimmune hepatitis is cytochrome P450db1. J Clin Invest. 1989;83:1066–1072.
Christen U, Holdener M, Hintermann E. Cytochrome P450 2D6 as a model antigen. Dig Dis. 2010;28:80–85.
Human Microbiome Project C. Structure. function and diversity of the healthy human microbiome. Nature. 2012;486:207–214.
Human Microbiome Project C. A framework for human microbiome research. Nature. 2012;486:215–221.
Lopetuso LR, Scaldaferri F, Bruno G et al. The therapeutic management of gut barrier leaking: the emerging role for mucosal barrier protectors. Eur Rev Med Pharmacol Sci. 2015;19:1068–1076.
Milosevic I, Vujovic A, Barac A et al. Gut-liver axis, gut microbiota, and its modulation in the management of liver diseases: a review of the literature. Int J Mol Sci. 2019;20:395.
Junior DST. Environmental and individual factors associated with protection and predisposition to autoimmune diseases. Int J Health Sci (Qassim). 2020;14:13–23.
Konieczna P, Akdis CA, Quigley EM, Shanahan F, O’Mahony L. Portrait of an immunoregulatory Bifidobacterium. Gut Microbes. 2012;3:261–266.
Bach Knudsen KE, Laerke HN, Hedemann MS et al. Impact of diet-modulated butyrate production on intestinal barrier function and inflammation. Nutrients. 2018;10:1499.
Pohl K, Moodley P, Dhanda A. The effect of increasing intestinal short-chain fatty acid concentration on gut permeability and liver injury in the context of liver disease: a systematic review. J Gastroenterol Hepatol. 2022;37:1498–1506.
Wang CH, Yen HR, Lu WL et al. Adjuvant probiotics of Lactobacillus salivarius subsp. salicinius AP-32, L. johnsonii MH-68, and Bifidobacterium animalis subsp. lactis CP-9 attenuate glycemic levels and inflammatory cytokines in patients with type 1 diabetes mellitus. Front Endocrinol (Lausanne). 2022;13:754401.
Frasca G, Cardile V, Puglia C, Bonina C, Bonina F. Gelatin tannate reduces the proinflammatory effects of lipopolysaccharide in human intestinal epithelial cells. Clin Exp Gastroenterol. 2012;5:61–67.
Scaldaferri F, Lopetuso LR, Petito V et al. Gelatin tannate ameliorates acute colitis in mice by reinforcing mucus layer and modulating gut microbiota composition: emerging role for “gut barrier protectors” in IBD? United Eur Gastroenterol J. 2014;2:113–122.
Wang WW, Zhang Y, Huang XB et al. Fecal microbiota transplantation prevents hepatic encephalopathy in rats with carbon tetrachloride-induced acute hepatic dysfunction. World J Gastroenterol. 2017;23:6983–6994.
Liang M, Liwen Z, Jianguo S et al. Fecal microbiota transplantation controls progression of experimental autoimmune hepatitis in mice by modulating the TFR/TFH immune imbalance and intestinal microbiota composition. Front Immunol. 2021;12:728723.
Zachou K, Arvaniti P, Lyberopoulou A, Dalekos GN. Impact of genetic and environmental factors on autoimmune hepatitis. J Transl Autoimmun. 2021;4:100125.
Bird A. Perceptions of epigenetics. Nature. 2007;447:396–398.
Czaja AJ. Examining micro-ribonucleic acids as diagnostic and therapeutic prospects in autoimmune hepatitis. Expert Rev Clin Immunol. 2022;18:591–607.
Seitz HK, Stickel F. Molecular mechanisms of alcohol-mediated carcinogenesis. Nat Rev Cancer. 2007;7:599–612.
Phillips JM, Goodman JI. Inhalation of cigarette smoke induces regions of altered DNA methylation (RAMs) in SENCAR mouse lung. Toxicology. 2009;260:7–15.
Salnikow K, Zhitkovich A. Genetic and epigenetic mechanisms in metal carcinogenesis and cocarcinogenesis: nickel, arsenic, and chromium. Chem Res Toxicol. 2008;21:28–44.
Baccarelli A, Wright RO, Bollati V et al. Rapid DNA methylation changes after exposure to traffic particles. Am J Respir Crit Care Med. 2009;179:572–578.
Yehuda R, Daskalakis NP, Bierer LM et al. Holocaust exposure induced intergenerational effects on FKBP5 methylation. Biol Psychiatry. 2016;80:372–380.
van Gerven NM, de Boer YS, Zwiers A et al. HLA-DRB1*03:01 and HLA-DRB1*04:01 modify the presentation and outcome in autoimmune hepatitis type-1. Genes Immun. 2015;16:247–252.
Funding
This review did not receive financial support from a funding agency or institution.
Author information
Authors and Affiliations
Contributions
AJC, MD researched, designed, and wrote this article. The three tables and one figure are original, constructed by the author, and developed solely for this article.
Corresponding author
Ethics declarations
Conflict of interest
Albert J. Czaja, MD has no conflict of interests to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Czaja, A.J. Incorporating the Molecular Mimicry of Environmental Antigens into the Causality of Autoimmune Hepatitis. Dig Dis Sci 68, 2824–2842 (2023). https://doi.org/10.1007/s10620-023-07967-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-023-07967-5