Abstract
The negative impact of habitat fragmentation due to human activities may be different in different species that co-exist in the same area, with consequences on the development of environmental protection plans. Here we aim at understanding the effects produced by different natural and anthropic landscape features on gene flow patterns in two sympatric species with different specializations, one generalist and one specialist, sampled in the same locations. We collected and genotyped 194 wood mice (generalist species) and 199 bank voles (specialist species) from 15 woodlands in a fragmented landscape characterized by different potential barriers to dispersal. Genetic variation and structure were analyzed in the two species, respectively. Effective migration surfaces, isolation-by-resistance (IBR) analysis, and regression with randomization were used to investigate isolation-by-distance (IBD) and the relative importance of land cover elements on gene flow. We observed similar patterns of heterozygosity and IBD for both species, but the bank vole showed higher genetic differences among geographic areas. The IBR analysis suggests that (i) connectivity is reduced in both species by urban areas but more strongly in the specialist bank vole; (ii) cultivated areas act as dispersal corridors in both species; (iii) woodlands appear to be an important factor in increasing connectivity in the bank vole, and less so in the wood mouse. The difference in dispersal abilities between a generalist and specialist species was reflected in the difference in genetic structure, despite extensive habitat changes due to human activities. The negative effects of fragmentation due to the process of urbanization were, at least partially, mitigated by another human product, i.e., cultivated terrains subdivided by hedgerows, and this was true for both species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Habitat loss and fragmentation have negative impacts on populations, and are considered as one of the main causes of biodiversity loss and therefore a major issue in conservation biology (Fischer and Lindenmayer 2007; Wilson et al. 2016; Wu 2013). In particular, anthropogenic habitat fragmentation has modified the distribution and population sizes in many different organisms (Crooks et al. 2017; Haddad et al. 2015) with local and/or global reduction of genetic diversity and connectivity (DiBattista 2008; Leigh et al. 2019). Monitoring the genetic consequences of human activities that increase habitat fragmentation is therefore important to develop appropriate conservation and management strategies (Hoban et al. 2020).
The major consequence of habitat loss and fragmentation is to create discontinuities (i.e., patchiness) in the distribution of critical resources (e.g., food, habitat, water) or in environmental conditions (e.g., microclimate) (Segelbacher et al. 2010). Such discontinuities reduce connectivity among populations (Kindlmann and Burel 2008), threatening their long-term viability due to genetic (e.g., reduced evolutionary potential and inbreeding depression) and demographic factors (e.g., demographic stochasticity) (Balkenhol and Waits 2009). Habitat fragmentation may also have different short-term consequences in different species, for example, by reducing the suitable habitats or increasing the predation success, but these effects poorly predict long-term responses (Evans et al. 2017). Gene flow among subpopulations is necessary to alleviate the adverse genetic consequences of population fragmentation, reducing genetic drift and maintaining local genetic variation (Frankham et al. 2017). From a conservation perspective, inferring the functional connectivity of populations across landscapes becomes crucial (Segelbacher et al. 2010). Identifying the areas where gene flow is either facilitated or prevented, and the landscape factors responsible for that, is a high priority (Sork 2016; Trombulak and Baldwin 2010).
One interesting opportunity to investigate the causes and the genetic consequences of fragmentation is represented by sympatric species with partially overlapping ecological niches (Homola et al. 2019; Varudkar and Ramakrishnan 2015). Different species, in fact, may respond very differently to the same landscape matrix (Frantz et al. 2012; Gangadharan et al. 2016; Nevill et al. 2019; Robertson et al. 2018; Thatte et al. 2020). They may also react differently to the fragmentation of their previously continuous habitat, and these differences may be reflected in the geographic distribution of their genetic variation. In this work, we investigate the effects of habitat fragmentation present in agricultural landscapes in Central Italy on the genetic structure of two sympatric rodent species, the wood mouse (Apodemus sylvaticus) and the bank vole (Myodes glareolus).
The wood mouse is a generalist species known to inhabit a wide range of habitats including forests, hedgerows and agricultural fields (Marsh and Harris 2000; Montgomery and Dowie 1993; Mortelliti et al. 2010). In contrast, the bank vole is a “forest specialist”, i.e., it is more strictly associated with forest habitats, from mature stands to recently coppiced woodlands (Capizzi and Luiselli 1996; Ecke et al. 2002). In general, specialist species tend to be more affected by habitat fragmentation than generalist species. In fact, highly dispersed resources are harder to reach by the specialist species (Braschler and Baur 2005; Nupp and Swihart 2001; Youngentob et al. 2012), which also suffer from competitive exclusion by the generalist species (Sozio and Mortelliti 2016). Accordingly, the specialist bank vole seems to prefer sites with high connectivity (Mortelliti et al. 2009; Sozio and Mortelliti 2016), and the generalist wood mouse can also be found in highly fragmented habitats, being able to move across cultivated fields (Sozio and Mortelliti 2016; Sozio et al. 2013). We currently do not know whether these differences directly correspond to a stronger genetic structure in the bank vole compared to the wood mouse, and if (and how) different natural or anthropogenic habitat features have different relative impacts on gene flow. Our study addresses these questions following three steps: (i) neutral genetic markers will be used to estimate the genetic diversity and the population structure separately in each species; (ii) patterns of increased and reduced gene flow will be analyzed and compared between species; and (iii) species-specific landscape features with the largest influence on genetic variation patterns will be identified.
Materials and methods
Study area and sample method
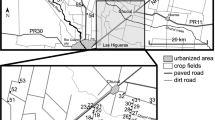
The study was conducted in a fragmented landscape (< 20% of residual woodland cover) located in central Italy (coordinates: 42°30′50″, 12°4′40″; elevation: 350 m; Fig. 1; Table S1).
Map of the study area in the Province of Viterbo, Central Italy. Landscape is reclassified according to the features utilized in the IBR analysis. RA represent the only railway intersecting the study area. Population codes are shown in Table 1
Woodland patches, consisting of mixed deciduous forest dominated by downy and turkey oaks (Quercus pubescens and Quercus cerris), were embedded in an agricultural matrix (mainly wheat fields) crossed by a network of hedgerows providing structural connectivity to habitat patches. The S2 highway and a railway bisect the study area, potentially acting as barriers to wildlife movements (Grilo et al. 2009). Urban areas are also present and represent approximately 5% of the total area. Twelve trapping sessions were conducted over a 2 year period, with bi-monthly trapping from April 2011 to February 2013. During each session, grids were trapped for three consecutive nights. A total of 199 bank voles and 194 wood mice samples were obtained. Sample sizes for each of the 15 different woodland patches is reported in Table 1. All the procedures of trapping and manipulation of animals took place in compliance with the European Council Directive 92/43EEC (Italian law D.Lgs 157/92 and LR 3/1994) and with the European Council Directive 86/609/EEC (Italian law D.Lgs 116/92). The capture and handling of species listed in the EU Habitat Directive was covered by permit number PNM 0024822 granted to A. M. by the Ministry of Environment, Rome, Italy.
Genotyping
Genomic DNA was extracted from the mouse ear lobe samples using the NucleoSpin® Tissue kit (Macherey–Nagel, Düren, Germany) according to the manufacturer’s protocol or using the Chelex-based DNA extraction method (Casquet et al. 2012). Eight microsatellite loci were used for the bank vole: Cg13B8, Cg6A1, Cg3F12, Cg13H9, Cg2E2, Cg3E10, Cg2A4 and Cg3A8 (Rikalainen et al. 2008). Seven microsatellite loci, described for members of the genus Apodemus, were used for the wood mouse: As-7, As11, As-12, As-20, As-34, GTTD9A and MsAf-8 (Gockel et al. 1997; Harr et al. 2000; Makova et al. 1998). A two-step PCR with the following conditions was carried out: initial denaturation at 95 °C for 15 min, followed by 30 cycles at 95 °C for 30 s, 56 °C for 45 s and 72 °C for 45 s, followed by eight cycles at 95 °C for 30 s, 53 °C for 45 s and 72 °C for 45 s, and a final elongation at 72 °C for 30 min. The forward primers were 5’ labelled with one of the following fluorescent labels: FAM, VIC, NED and PET. Fragments were analyzed on an ABI3130 capillary analyzer (Applied Biosystems, Life Technologies Corporation). Fragment data were analyzed using Peak Scanner Software (Applied Biosystems, Life Technologies Corporation).
Genetic diversity
Descriptive statistics of nuclear genetic diversity were estimated separately for each population (woodland patch) in each species. The mean number of alleles, and the observed and expected heterozygosities, were estimated using Genalex 6.5 (Peakall and Smouse 2006), and the same program was used to test for deviation from Hardy–Weinberg equilibrium. Allelic richness (AR) was calculated using the rarefaction procedure in the Fstat 2.9.4 software (Goudet 2001). Arlequin 3.5.2.2 (Excoffier and Lischer 2010) was used to test for linkage disequilibrium between each pair of loci for each sampling population following a likelihood-ratio statistic, whose null distribution was obtained by a permutation procedure. We applied sequential Bonferroni corrections to account for multiple comparisons (Rice 1989). Micro-Checker 2.2.3 (Van Oosterhout et al. 2004) was used to check for null alleles and scoring errors. FREENA (Chapuis and Estoup 2007) was used to compare uncorrected and corrected FST values to test for the impact of null alleles, when present. Genetic differentiation measured as FST values (Weir and Cockerham 1984) was estimated for each pair of sampling population with Arlequin. Statistical significance of the FST values was tested using 10000 permutations, and P values were multiplied by the total number of comparisons following the conservative Bonferroni approach for multiple testing.
Genetic structure
A Bayesian clustering method was used to identify the number of genetic groups (STRUCTURE v2.3.4; Pritchard et al. 2000). A burn-in length of 50,000 iterations and a run length of 100,000 iterations were used in an admixture model with correlated allele frequencies among populations testing each K value between 1 and 15. Each K value was run 10 times. The optimal K value was determined using the ∆K method (Evanno et al. 2005) implemented in the online tool STRUCTURE Harvester (Earl and VonHoldt 2012). CLUMPP (Jakobsson and Rosenberg 2007) was then applied to average the multiple runs given by STRUCTURE and to verify correct label switching. To display the results, the output from CLUMPP was visualized with DISTRUCT (Rosenberg 2004).
Visualizing deviation from Isolation by Distance
Genetic diversity between populations often exhibits patterns consistent with Isolation-by-Distance (IBD) (Wright 1943), where populations far apart in the geographic space receive less gene flow than neighbouring ones. Given the ubiquity of this phenomenon (Kuchta and Tan 2005; Sharbel et al. 2000) it is interesting to see locations where this does not hold true, as they might represent barriers or zones of high contact. Global deviation from IBD can be identified, for example, studying the decrease of similarity or autocorrelation with geographic distance. However, specific deviations in some areas, but not in others, cannot be easily investigated and visualized by standard methods. One recent solution to this problem comes from the use of Estimated Effective Migration Surfaces (EEMS; Petkova et al. 2015). EEMS employs individual based migration rates in order to visualize zones with higher or lower migration with respect to the overall rate. These areas represent locations in which the pattern of gene flow predicted by IBD is facilitated or hindered. The region under study was first divided in a grid of demes and the individuals were assigned to the deme closest to their sampling location. The matrix of effective migration rates was then computed by EEMS based on the stepping-stone model (Kimura and Weiss 1964) and on resistance distances (McRae 2006). We used the EEMS script for microsatellites analysis runeems_sats available from Github (https://github.com/dipetkov/eems) to construct EEMS surfaces for the bank vole and the wood mouse. Considering that the number of demes simulated during the grid construction phase can influence the scale of the deviation from the overall migration rate, we averaged three runs with 50, 100, 200, 300 and 400 demes to produce the final EEMS surface. Each single run consisted of 200,000 burn in steps followed by 1,000,000 MCMC iterations sampled every 10000 steps. We plotted the averaged EEMS and checked for MCMC convergence using the rEEMSplots package in R.
Isolation by resistance
Understanding the effect of environmental components on the genetic makeup of natural populations is the goal of landscape genetics, which integrates population genetics, landscape ecology and spatial statistics (Holderegger and Wagner 2008; Manel et al. 2003; Storfer et al. 2007). One of the techniques more commonly used in landscape genetics to identify discontinuities in gene flow and determine the relative resistance to movement imposed by different landscape elements is Isolation-by-Resistance (IBR; McRae 2006). IBR offers a conceptual model in which landscape resistance is the analogue of electrical resistance, and the movements of individuals and flow of genes are analogues of electrical current (Amos et al. 2012). It greatly extends the ability to model multiple complementary paths of connectivity, while being sufficiently computationally efficient to allow its use over large landscapes at relatively fine resolution (McRae and Beier 2007; McRae et al. 2008). In order to analyze the effect of specific landscape components on gene flow, we tested for the presence of IBR. We first constructed a raster grid encompassing all our study area reclassifying the land cover based on features that were a priori most likely to affect gene flow in both the bank vole and the wood mouse: woodland, urban areas, cultivated terrain and hedgerows (Fig. 1). We obtained a raster file encompassing the entire studied area with a resolution of 10 m that was previously classified based on major land cover types (CORINE Land Cover; Büttner et al. 2004). Aerial photographs were used to add the locations of hedgerows with the same resolution (Sozio and Mortelliti 2016). We also included in our raster grid the major roads intersecting our study area from OpenStreetMap (http://www.openstreetmap.org) and the railways tracks from the DIVA-GIS database (http://www.diva-gis.org/gdata).
In order to determine the relative importance of land cover elements in hindering or facilitating gene flow, we modified the created raster under two different scenarios. The first set (resistance set) was aimed at determining the resistance caused by a specific land cover feature with respect to the others. We assigned a varying maximum resistance (REmax) to a target component, keeping the other landscape features to a uniform minimum resistance (REmin = 1). The second set of grids (permeability set) was built to establish the possible role of a specific landscape feature in facilitating the connection between different populations. Due to the challenges in determining the effective resistance posed by each environmental feature to gene flow, we employed a optimization procedure encompassing a wide range of resistance values (Ortego et al. 2015; Roffler et al. 2016). We assigned a minimum resistance value to a target landscape component and a varying REmax to all remaining features. For both set of grids we employed eight maximum resistance values (REmax = 5, 10, 50, 100, 500, 1000, 5000 and 10000) obtaining a total of 96 different surfaces. We computed pairwise resistance distances between populations for both the bank vole and the wood mouse using the different sets of grids. Distances were obtained considering the eight-neighbour cell connection scheme in CIRCUITSCAPE v4.0 (McRae, Shah, and Mohapatra, 2013) with the sampled woodland patches as focal regions. We also computed an IBD scenario considering a homogeneous resistance surface (all RE = 1) (Castillo et al. 2014; Ortego et al. 2015). We then compared the resistance and the FST matrices using multiple matrix regression with randomization (MMRR) (Wang 2013). We considered a consistent trend of increase of R2, at least for 4 levels of resistance values, with P values always smaller than 0.05 in single tests, as evidence of the impact of the corresponding land cover feature. For each landscape variable, the most supported model was identified as the one corresponding to the highest supported R2 value. In case of plateau, we preferred the model corresponding to the onset of the plateau (Castillo et al. 2014). Statistical significance of the coefficients was determined using 9999 permutations with the MMRR function (Wang 2013). All statistical analyses were conducted in R.
Results
Genetic diversity
All loci were polymorphic in both species. The average expected heterozygosity was very similar in the two different sets of markers typed in the two species (0.74 in the bank vole and 0.72 in the wood mouse), and the number of alleles varied between 2 and 16 in the wood mouse and between 3 and 11 in the bank vole, respectively. All the genetic variation statistics are reported in Table 1.
No systematic deviation from linkage equilibrium was observed between loci for any population in both species, and none of the pairwise tests was significant after Bonferroni correction. Some loci showed evidence of the presence of null alleles, but only in some populations. We analyzed the effect of these alleles by comparing matrices of pairwise FST values computed from the complete data set with values corrected for null alleles as estimated by FreeNA. Multilocus global FST values had identical values when calculated with and without correcting for null alleles in both species (wood mouse: FST = 0.03, with or without correction; bank vole: FST = 0.08, with or without correction), with identical or very similar confidence intervals in the two analyses (0.01–0.05 in wood mouse, with and without correction, 0.07–0.09 and 0.06–0.08 in bank vole, with and without correction, respectively). Multilocus pairwise FST values with and without correction were also highly correlated (wood mouse: r = 0.99; p = 0.001; bank vole: r = 0.99; p = 0.001; Mantel test). We decided therefore to use the complete data set for all downstream analyses. Pairwise FST values in the wood mouse were significant after sequential Bonferroni correction only in 7 out of 105 comparisons, all involving the PRV population (with FST values never larger than 0.08) (Fig. 2; Table S2). On the contrary, the bank vole showed a finer geographic structure. Approximately half of the FST values were significant, with the highest divergence values observed in comparisons including PRV, and, as reported above, the average FST was much higher than that estimated in the wood mouse (Fig. 2; Table S2).
Genetic structure
The most likely partition implied three genetic groups (K = 3) in both species. Here we present individual assignment plots for K equal to 2, 3, and 4 (Fig. 3a, b) to better visualize different aspects of the genetic structure, and we also report the geographic distribution of the most supported clades in both species (Fig. 3c). In the wood mouse (Fig. 3a), the isolation of PRV already suggested by the pairwise FST matrix was supported at different values of K. With the most supported K = 3, or with K = 4, a large fraction of individuals and populations (with the exception of PRV) showed a mixed ancestry. In the bank vole (Fig. 3b), populations appeared more internally homogeneous, with three distinct genetic groups prevailing in the northern areas (ALB, BRN, FDT, FRR and GST), in the western areas (API, IUG, MCD, PRV and YAH), and in a single eastern population (CRC), respectively, and the other populations having a more mixed and less geographically localized genetic composition. The results are robust with increasing burn-in and replicates.
Population assignment test performed with STRUCTURE. Bar plots represent the genetic composition of single individuals (thin vertical columns) from K = 2 to K = 4. A wood mouse; b bank vole. Maps of the study area with the genetic composition of each population for K = 3 in the wood mouse (left) and the bank vole (right)
Visualizing deviation from IBD
The spatial visualization of the geographic areas with higher or lower gene flow compared to IBD expectations was similar in the two species (Fig. 4). The main pattern consisted of a central area of reduced gene flow, centered around PRV, extended only in the bank vole towards the southern and the eastern borders of the region. These branches of reduced migration clearly produced the higher genetic structure observed in the bank vole when compared to the wood mouse, with the latter having a much higher connectivity in most of the areas we considered.
Individual-based EEMS analysis of effective migration rates (m) for the wood mouse (left) and the bank vole (right). The effective migration rate is represented on a log10 scale. Areas showing negative values (orange) represent possible barriers to gene-flow while zones with positive values (blue) correspond to places of increased gene-flow, both with respect to the Isolation-by-Distance background (white). Migration surfaces are averages of 3 runs each with 50, 100, 200, 300, and 400 demes. The scale is different in the two panels, indicating that the increase (blue colors) or decrease (red colors) of the rates relative to the average rate are more pronounced in the wood mouse compared to the bank vole
Isolation by resistance (IBR)
Both the wood mouse and the bank vole populations showed in the IBR a significant pattern of IBD (Tables S3–S6). However, we also found consistently higher association between pairwise FST and resistance distance in some models including land cover features (Fig. 5; Tables S3–S6). Highest R2 values in the best models were always smaller in the wood mouse, reflecting probably the fact that gene flow is abundant in this species and factors limiting or favouring gene flow in this species are more difficult to detect.
Goodness of fit for models of landscape resistance. Panels show the coefficient of determination (R2) for models analyzing genetic differentiation (panel a–c: wood mouse; panel b–d: bank vole) in relation to resistance (a, b) and permeability (c, d) distance matrices plotted against resistance values for different landscape features. Circles with black outline showed significant P-values
In both species, the first set of distances (resistance, RE) reached the highest value of R2 when urban areas presented intermediate resistance values (RE = 500) with respect to the surrounding environmental feature (see Fig. 5a, b; Tables S3–S4). Compared to the scenario including only IBD, urban areas produced an increase of R2 by approximately 37 and 69% in the wood mouse and the bank vole, respectively. Only hedges in the wood mouse seemed also to produce a small additional fit to the model. The second set of resistance values (permeability), designed to highlight factors facilitating connectivity between different populations, supported the role of cultivated areas in both species, with a 50% increase of R2 compared to IBD at RE = 1/100 (see Fig. 5c,d; Tables S5–S6). In addition, woodlands seem also to play a role especially in the bank vole, and areas comprising and surrounding major roads increased the fit of the model only in the wood mouse.
A summary of the major results obtained in the two species is in Table 2.
Discussion
In this study we investigated the relationships between human-related changes in habitat amount and configuration (i.e., habitat structure), habitat use and genetic structure. We applied an identical sampling scheme within the same fragmented area to two sympatric rodent species, the wood mouse and the bank vole. Our main result is that the same factors appear to limit (urban areas) and favour (woodland and cultivated areas) gene flow in both species despite their difference in ecological niche. The genetic structure in the same region before human intervention is not known, but it is interesting to note that the joint and compensatory effects of urban and cultivated areas did not modify the relative levels of genetic structure expected in the two species: the generalist wood mouse has a population structure much more genetically connected than the forest-specialized bank vole. More specifically, gene flow favored by cultivated areas likely increases the genetic exchanges in the wood mouse even above the high level expected in natural conditions, now limited only by urban areas. In the bank vole, on the other hand, cultivated areas possibly act compensating the genetic fragmentation due to the loss of the preferred niche (woodland) and the increase of urban areas. Any further replacement of woodlands with urban and not cultivated areas will clearly reduce this compensatory effect. Overall, we conclude that the difference between wood mouse and bank vole is still reflected in the difference between their current genetic structure, but if woodlands will be further replaced by urban settlements and not cultivated areas this difference will likely increases.
Genetic diversity
Habitat fragmentation did not produce a detectable loss of genetic variation in the two species. Levels of diversity in different populations are comparable to those reported for other rodent species (Czarnomska et al. 2018; Dominguez et al. 2021; Gerlach and Musolf 2000; Martin Cerezo et al. 2020). When the global genetic divergence between populations was analyzed, the wood mouse showed much weaker population structure than the bank vole. This pattern is expected considering that, at a short geographic scale (distances < 30 km), genetic structure is commonly found only in rodents with a specialized ecological niche (Bani et al. 2017; Fasanella, et al. 2013; Kozakiewicz et al. 2009; Łopucki et al. 2022).
With the exclusion of the population sampled in PRV (see below), the wood mouse appears rather homogenous at this geographic scale, indicating that gene flow was not prevented by the human-induced fragmentation of their natural habitat. This result reflects the enormous capacity of adaptation and mobility in this species, which can be found in all types of forests and even in cultivated fields during certain periods of the year (Harris and Woollard 1990; Szacki et al. 1993; Tew 1994). Conversely, populations of the bank vole sampled in the same patches showed the presence of a significant genetic differentiation with a lower degree of genetic admixture and higher FST values. Similar studies on bank vole confirmed that there is a significant reduction of gene flow already at geographical distance of about 8 km (Stacy et al. 1997), and that environmental features, such as seasonal temperature variations, can contribute in a decisive way in increasing the genetic structure of this species (Gȩbczyński and Ratkiewicz 1998).
Fragmentation of the natural environment as a consequence of human activity (e.g., urbanization, agriculture, pastures) can reduce the genetic diversity of populations by decreasing population size and gene-flow, while increasing genetic drift. Generalist species can be considered as “urban adapters” (Blair 2001), since they can occupy novel environments/ecosystems given their wider niche breath. In white-footed mice (Peromyscus leucopus), urbanization increased population differentiation while suffering minimal loss in genetic diversity (Munshi-South and Kharchenko 2010) and keeping IBR between patches of urban vegetation (Munshi-South 2012). Our results also show a similar pattern in which both species retained their genetic diversity and showed population structure. While we cannot assess the presence of historical population differentiation prior to environment fragmentation, we show that different habitat disturbance similarly affected sympatric species with different ecological specializations.
Spatial patterns of gene flow
Isolation-by-distance was significant for both species indicating that geographic distance is an important factor for isolating the various populations. An additional shared feature appears to be the isolation of PRV in all analyses, supporting the hypothesis that individuals of both species have difficulty reaching this area. This result may be related to the fact that urban areas are highly diffused around PRV, and the IBR analysis suggested that they act as a barrier for both species.
For the bank vole, we found additional areas of enhanced or reduced gene flow, in comparison with the IBD pattern in the background. Specifically, three main areas showed higher gene flow than expected, corresponding to western, eastern and northern patches. Barriers separating them are composed of a mix of different environmental features but the IBR modelling suggests that urban areas play a major role, as observed for other rodent species (Mapelli et al. 2020; Munshi-South 2012).
Interestingly, our results show that the predicted difference in genetic structure (less structured generalist vs. more structured specialist) was maintained despite habitat alterations. We hypothesize a sort of compensatory effect, similarly acting in both species, between a decrease of gene flow in urban areas, while cultivated terrains increased population connectivity. Compensatory dynamics in disturbed habitats have been observed before in minks (Neovison vison) in Scotland and in the western slimy salamander (Plethodon albagula) in the USA (Oliver et al. 2016; Peterman et al. 2014). For salamanders, a compensatory dispersal behavior was observed through suboptimal habitats (i.e., high resistance surfaces) (Peterman et al. 2014). For our species, we observed higher gene-flow than predicted in some patches, which may be due to a compensatory behavior (increased gene-flow) through the landscape so previous patterns of population connectivity are maintained. This pattern, combined with compensatory landscape modifications (reduced connectivity in urban areas and increased in cultivated terrains) may have contributed to the maintenance of previous relative differences in the two investigated species. Further genetic monitoring of these populations can elucidate if this pattern will be maintained in time.
Our analysis highlighted the importance of woodland patches acting as a corridor between different sampled locations in both studied species, with particular emphasis for the bank vole. Urban areas were shown to be the major barriers experienced by gene-flow in the bank vole and wood mouse, albeit to a different degree. Moreover, railways and roads (never wider than 10 m in this area) cannot be considered as barriers to the dispersal of these species, consistently with previous studies (Gerlach and Musolf 2000; Redeker et al. 2006). Indeed, roads appear as a factor that favors gene flow in the wood mouse. This may be because, for this species, the size of the roads present in the study area should not be considered as a barrier and/or that roads, in the environmental matrix, were included in (or surrounded by) a suitable ground. Similarly, cultivated fields do not limit dispersal, but may even play a role as corridors (Johnson and Munshi-South 2017; Miles et al. 2019; Tattersall et al. 2001). The only anthropogenic factor that seems to negatively affect the dispersal pattern is the presence of urban areas. Clearly, if woodlands will be further reduced by urbanization, genetic drift due to fragmentation could increase in our two studied species.
Spatial scale effects may be of particular importance in landscape genetic studies, especially on species having dispersal abilities limited. Different spatial scale could have revealed other environmental factors, but our sampling was spatially designed to test only a few factors related to anthropic activities.
Conclusions and implications for conservation
Overall, our results show that despite extensive habitat changes due to human activities, levels of genetic variation are quite high in both species, and their difference in the dispersal abilities is still reflected in the difference of genetic structure. The wood mouse, a generalist species with high dispersal ability, shows in fact higher genetic connectivity than the bank vole, which is a less mobile species closely linked to woodland areas. Nevertheless, we found also that cultivated fields and urban areas modifies the natural dispersion patterns in both species, probably in a way that will, in the future, increase the difference between their genetic structure. Our study supports the view that patterns of gene flow can be similarly affected, even in sympatric species with different ecological characteristics, by the same changes of land use. Locally, this implies that future monitoring efforts should prioritize the bank vole, the species with the highest genetic structure where genetic fragmentation is more likely to increase due to urbanization.
Data availability
Microsatellite datasets of the wood mouse and the bank vole are available for download from Zenodo (https://doi.org/10.5281/zenodo.6527316).
References
Amos JN, Bennett AF, Mac Nally R, Newell G, Pavlova A, Radford JQ, Sunnucks P (2012) Predicting landscape-genetic consequences of habitat loss, fragmentation and mobility for multiple species of woodland birds. PLoS ONE 7:e30888. https://doi.org/10.1371/journal.pone.0030888
Balkenhol N, Waits LP (2009) Molecular road ecology: exploring the potential of genetics for investigating transportation impacts on wildlife. Mol Ecol 18:4151–4164. https://doi.org/10.1111/j.1365-294X.2009.04322.x
Bani L, Orioli V, Pisa G, Fagiani S, Dondina O, Fabbri E, Mortelliti A (2017) Population genetic structure and sex-biased dispersal of the hazel dormouse (Muscardinus avellanarius) in a continuous and in a fragmented landscape in central Italy. Conserv Genet 18:261–274. https://doi.org/10.1007/s10592-016-0898-2
Blair RB (2001) Birds and butterflies along urban gradients in two ecoregions of the United States: is urbanization creating a homogeneous fauna? In: Lockwood JL, McKinney ML (eds) Biotic Homogenization. Academic Press, Netherlands. https://doi.org/10.1007/978-1-4615-1261-5_3
Braschler B, Baur B (2005) Experimental small-scale grassland fragmentation alters competitive interactions among ant species. Oecologia 143:291–300. https://doi.org/10.1007/s00442-004-1778-x
Büttner G, Feranec J, Jaffrain G, Mari L, Maucha G, Soukup T (2004) The CORINE land cover 2000 project. EARSeL EProceedings 3:331–346
Capizzi D, Luiselli L (1996) Ecological relationships between small mammals and age of coppice in an oak-mixed forest in central Italy. Revue D’ecologie (la Terre Et La Vie) 51:277–291
Casquet J, Thebaud C, Gillespie RG (2012) Chelex without boiling, a rapid and easy technique to obtain stable amplifiable DNA from small amounts of ethanol-stored spiders. Mol Ecol Resour 12:136–141. https://doi.org/10.1111/j.1755-0998.2011.03073.x
Castillo JA, Epps CW, Davis AR, Cushman SA (2014) Landscape effects on gene flow for a climate-sensitive montane species, the American pika. Mol Ecol 23:843–856. https://doi.org/10.1111/mec.12650
Chapuis MP, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24:621–631. https://doi.org/10.1093/molbev/msl191
Crooks KR, Burdett CL, Theobald DM, King SR, Di Marco M, Rondinini C, Boitani L (2017) Quantification of habitat fragmentation reveals extinction risk in terrestrial mammals. Proc Natl Acad Sci 114:7635–7640. https://doi.org/10.1073/pnas.1705769114
Czarnomska SD, Niedziałkowska M, Borowik T, Jędrzejewska B (2018) Regional and local patterns of genetic variation and structure in yellow-necked mice-the roles of geographic distance, population abundance, and winter severity. Ecol Evol 8:8171–8186. https://doi.org/10.1002/ece3.4291
DiBattista JD (2008) Patterns of genetic variation in anthropogenically impacted populations. Conserv Genet 9:141–156. https://doi.org/10.1007/s10592-007-9317-z
Dominguez JC, Calero-Riestra M, Olea PP, Malo JE, Burridge CP, Proft K, García JT (2021) Lack of detectable genetic isolation in the cyclic rodent Microtus arvalis despite large landscape fragmentation owing to transportation infrastructures. Sci Rep 11:1–14. https://doi.org/10.1038/s41598-021-91824-w
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Ecke F, Löfgren O, Sörlin D (2002) Population dynamics of small mammals in relation to forest age and structural habitat factors in Northern Sweden. J Appl Ecol 39:781–792. https://doi.org/10.1046/j.1365-2664.2002.00759.x
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Evans MJ, Banks SC, Driscoll DA, Hicks AJ, Melbourne BA, Davies KF (2017) Short-and long-term effects of habitat fragmentation differ but are predicted by response to the matrix. Ecology 98:807–819. https://doi.org/10.1002/ecy.1704
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under linux and windows. Mol Ecol Resour 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Fasanella M, Bruno C, Cardoso Y, Lizarralde M (2013) Historical demography and spatial genetic structure of the subterranean rodent Ctenomys magellanicus in Tierra del Fuego (Argentina). Zool J Linn Soc 169:697–710. https://doi.org/10.1111/zoj.12067
Fischer J, Lindenmayer DB (2007) Landscape modification and habitat fragmentation: a synthesis. Glob Ecol Biogeogr 16:265–280. https://doi.org/10.1111/j.1466-8238.2007.00287.x
Frankham R, Ballou JD, Ralls K, Eldridge M, Dubash MR, Fenster CB et al (2017) Genetic management of fragmented animal and plant populations. Oxford University Press, Oxford. https://doi.org/10.1093/oso/9780198783398.001.0001
Frantz AC, Bertouille S, Eloy MC, Licoppe A, Chaumont F, Flamand MC (2012) Comparative landscape genetic analyses show a Belgian motorway to be a gene flow barrier for red deer (cervus elaphus), but not wild boars (sus scrofa). Mol Ecol 21:3445–3457. https://doi.org/10.1111/j.1365-294X.2012.05623.x
Gangadharan A, Vaidyanathan S, St. Clair, C. C. (2016) Categorizing species by niche characteristics can clarify conservation planning in rapidly-developing landscapes. Anim Conserv 19:451–461. https://doi.org/10.1111/acv.12262
Gȩbczyński M, Ratkiewicz M (1998) Does biotope diversity promote an increase of genetic variation in the bank vole population? Acta Theriol 43:163–173. https://doi.org/10.4098/AT.arch.98-12
Gerlach G, Musolf K (2000) Fragmentation of landscape as a cause for genetic subdivision in bank voles. Conserv Biol 14:1066–1074. https://doi.org/10.1046/j.1523-1739.2000.98519.x
Gockel J, Harr B, Schlötterer C, Arnold W, Gerlach G, Tautz D (1997) Isolation and characterization of microsatellite loci from Apodemus flavicollis (Rodentia, Muridae) and Clethrionomys glareolus (Rodentia, Cricetidae). Mol Ecol 6:597–599. https://doi.org/10.1046/j.1365-294X.1997.00222.x
Goudet, J. (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Retrieved from http://www2.unil.ch/popgen/softwares/fstat.htm
Grilo C, Bissonette JA, Santos-Reis M (2009) Spatial-temporal patterns in Mediterranean carnivore road casualties: consequences for mitigation. Biol Cons 142:301–313. https://doi.org/10.1016/j.biocon.2008.10.026
Haddad NM, Brudvig LA, Clobert J, Davies KF, Gonzalez A, Holt RD, Townshend JR (2015) Habitat fragmentation and its lasting impact on Earth’s ecosystems. Sci Adv 1:e1500052. https://doi.org/10.1126/sciadv.1500052
Harr B, Musolf K, Gerlach G (2000) Characterization and isolation of DNA microsatellite primers in wood mice (Apodemus sylvaticus, Rodentia). Mol Ecol 9:1664–1665. https://doi.org/10.1046/j.1365-294X.2000.01043-3.x
Harris S, Woollard T (1990) The dispersal of mammals in agricultural habitats in Britain. In: Bunce RGH, Howard DC (ed) Species dispersal in agricultural habitats. Belhaven Press, London, pp 159–188
Hoban S, Bruford M, Jackson JDU, Lopes-Fernandes M, Heuertz M, Hohenlohe PA, Laikre L (2020) Genetic diversity targets and indicators in the CBD post-2020 Global biodiversity framework must be improved. Biol Cons 248:108654. https://doi.org/10.1016/j.biocon.2020.108654
Holderegger R, Wagner HH (2008) Landscape genetics. Bioscience 58:199–207. https://doi.org/10.1641/B580306
Homola JJ, Loftin CS, Kinnison MT (2019) Landscape genetics reveals unique and shared effects of urbanization for two sympatric pool-breeding amphibians. Ecol Evol 9:11799–11823. https://doi.org/10.1002/ece3.5685
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. https://doi.org/10.1093/bioinformatics/btm233
Johnson MTJ, Munshi-South J (2017) Evolution of life in urban environments. Science 358:8327. https://doi.org/10.1126/science.aam8327
Kimura M, Weiss GH (1964) The stepping stone model of population structure and the decrease of genetic correlation with distance. Genetics 49:561–576
Kindlmann P, Burel F (2008) Connectivity measures: a review. Landscape Ecol 23:879–890. https://doi.org/10.1007/s10980-008-9245-4
Kozakiewicz M, Gortat T, Panagiotopoulou H, Gryczyńska-Siemia̧tkowska A, Rutkowski R, Kozakiewicz A, Abramowicz K (2009) The spatial genetic structure of bank vole (Myodes glareolus) and yellow-necked mouse (Apodemus flavicollis) populations: The effect of distance and habitat barriers. Anim Biol 59:169–187. https://doi.org/10.1163/157075609X437691
Kuchta SR, Tan ANM (2005) Isolation by distance and post-glacial range expansion in the rough-skinned newt, Taricha granulosa. Mol Ecol 14:225–244. https://doi.org/10.1111/j.1365-294X.2004.02388.x
Leigh DM, Hendry AP, Vázquez-Domínguez E, Friesen VL (2019) Estimated six per cent loss of genetic variation in wild populations since the industrial revolution. Evol Appl 12:1505–1512. https://doi.org/10.1111/eva.12810
Łopucki R, Mróz I, Nowak-Życzyńska Z, Perlińska-Teresiak M, Owadowska-Cornil E, Klich D (2022) Genetic structure of the root vole Microtus oeconomus: resistance of the habitat specialist to the natural fragmentation of preferred moist habitats. Genes 13:434. https://doi.org/10.3390/genes13030434
Makova KD, Patton JC, Krysanov EY, Chesser RK, Baker RJ (1998) Microsatellite markers in wood mouse and striped field mouse (genus Apodemus). Mol Ecol 7:247–249. https://doi.org/10.1111/j.1365-294X.1998.00315.x
Manel S, Schwartz MK, Luikart G, Taberlet P (2003) Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol 18:189–197. https://doi.org/10.1016/S0169-5347(03)00008-9
Mapelli FJ, Boston ES, Fameli A, Gómez Fernández MJ, Kittlein MJ, Mirol PM (2020) Fragmenting fragments: landscape genetics of a subterranean rodent (Mammalia, Ctenomyidae) living in a human-impacted wetland. Landscape Ecol 35:1089–1106. https://doi.org/10.1007/s10980-020-01001-z
Marsh ACW, Harris S (2000) Partitioning of woodland habitat resources by two sympatric species of apodemus: lessons for the conservation of the yellow-necked mouse (A. flavicollis) in Britain. Biol Cons 92:275–283. https://doi.org/10.1016/S0006-3207(99)00071-3
Martin Cerezo ML, Kucka M, Zub K, Chan YF, Bryk J (2020) Population structure of Apodemus flavicollis and comparison to Apodemus sylvaticus in northern Poland based on RAD-seq. BMC Genomics 21:1–14. https://doi.org/10.1186/s12864-020-6603-3
McRae BH, Shah VB, Mohapatra T (2013) Circuitscape 4 User Guide. The Nature Conservancy
McRae BH (2006) Isolation by Resistance. Evolution 60:1551–1561. https://doi.org/10.1554/05-321.1
McRae BH, Beier P (2007) Circuit theory predicts gene flow in plant and animal populations. Proc Natl Acad Sci USA 104:19885–19890. https://doi.org/10.1073/pnas.0706568104
McRae BH, Dickson BG, Keitt TH, Shah VB (2008) Using circuit theory to model connectivity in ecology, evolution, and conservation. Ecology 89:2712–2724. https://doi.org/10.1890/07-1861.1
Miles LS, Rivkin LR, Johnson MTJ, Munshi-South J, Verrelli BC (2019) Gene flow and genetic drift in urban environments. Mol Ecol 28:4138–4151. https://doi.org/10.1111/mec.15221
Montgomery WI, Dowie M (1993) The distribution and population regulation of the wood mouse Apodemus sylvaticus on field boundaries of pastoral farmland. J Appl Ecol 30:783–783. https://doi.org/10.2307/2404256
Mortelliti A, Amori G, Annesi F, Boitani L (2009) Testing for the relative contribution of patch neighborhood, patch internal structure, and presence of predators and competitor species in determining distribution patterns of rodents in a fragmented landscape. Can J Zool 87:662–670. https://doi.org/10.1139/Z09-054
Mortelliti A, Amori G, Capizzi D, Rondinini C, Boitani L (2010) Experimental design and taxonomic scope of fragmentation studies on European mammals: current status and future priorities. Mammal Rev 40:125–154. https://doi.org/10.1111/j.1365-2907.2009.00157.x
Munshi-South J (2012) Urban landscape genetics: canopy cover predicts gene flow between white-footed mouse (Peromyscus leucopus) populations in New York City. Mol Ecol 21:1360–1378. https://doi.org/10.1111/j.1365-294X.2012.05476.x
Munshi-South J, Kharchenko K (2010) Rapid, pervasive genetic differentiation of urban white-footed mouse (Peromyscus leucopus) populations in New York City. Mol Ecol 19:4242–4254. https://doi.org/10.1111/j.1365-294X.2010.04816.x
Nevill PG, Robinson TP, Di Virgilio G, Wardell-Johnson G (2019) Beyond isolation by distance: what best explains functional connectivity among populations of three sympatric plant species in an ancient terrestrial island system? Divers Distrib 25:1551–1563. https://doi.org/10.1111/ddi.12959
Nupp TE, Swihart RK (2001) Assessing competition between forest rodents in a fragmented landscape of midwestern USA. Mamm Biol 66:345–356
Oliver MK, Piertney SB, Zalewski A, Melero Y, Lambin X (2016) The compensatory potential of increased immigration following intensive American mink population control is diluted by male-biased dispersal. Biol Invasions 18:3047–3061. https://doi.org/10.1007/s10530-016-1199-x
Ortego J, Aguirre MP, Noguerales V, Cordero PJ (2015) Consequences of extensive habitat fragmentation in landscape-level patterns of genetic diversity and structure in the Mediterranean esparto grasshopper. Evol Appl 8:621–632. https://doi.org/10.1111/eva.12273
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Peterman WE, Connette GM, Semlitsch RD, Eggert LS (2014) Ecological resistance surfaces predict fine-scale genetic differentiation in a terrestrial woodland salamander. Mol Ecol 23:2402–2413. https://doi.org/10.1111/mec.12747
Petkova D, Novembre J, Stephens M (2015) Visualizing spatial population structure with estimated effective migration surfaces. Nat Genet 48:94–100. https://doi.org/10.1038/ng.3464
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1111/j.1471-8286.2007.01758.x
Redeker S, Andersen LW, Pertoldi C, Madsen AB, Jensen TS, Jørgensen JM (2006) Genetic structure, habitat fragmentation and bottlenecks in Danish bank voles (Clethrionomys glareolus). Mamm Biol 71:144–158. https://doi.org/10.1016/j.mambio.2005.12.003
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223. https://doi.org/10.2307/2409177
Rikalainen K, Grapputo A, Knott E, Koskela E, Mappes T (2008) A large panel of novel microsatellite markers for the bank vole (Myodes glareolus). Mol Ecol Resour 8:1164–1168. https://doi.org/10.1111/j.1755-0998.2008.02228.x
Robertson JM, Murphy MA, Pearl CA, Adams MJ, Páez-Vacas MI, Haig SM, Funk WC (2018) Regional variation in drivers of connectivity for two frog species (Rana pretiosa and R. luteiventris) from the US Pacific Northwest. Mol Ecol 27:3242–3256. https://doi.org/10.1111/mec.14798
Roffler GH, Schwartz MK, Pilgrim KL, Talbot SL, Sage GK, Adams LG, Luikart G (2016) Identification of landscape features influencing gene flow: how useful are habitat selection models? Evol Appl 9:805–817. https://doi.org/10.1111/eva.12389
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138. https://doi.org/10.1046/j.1471-8286.2003.00566.x
Segelbacher G, Cushman SA, Epperson BK, Fortin MJ, Francois O, Hardy OJ, Manel S (2010) Applications of landscape genetics in conservation biology: concepts and challenges. Conserv Genet 11:375–385. https://doi.org/10.1007/s10592-009-0044-5
Sharbel TF, Haubold B, Mitchell-Olds T (2000) Genetic isolation by distance in Arabidopsis thaliana: biogeography and postglacial colonization of Europe. Mol Ecol 9:2109–2118. https://doi.org/10.1046/j.1365-294X.2000.01122.x
Sork VL (2016) Gene flow and natural selection shape spatial patterns of genes in tree populations: implications for evolutionary processes and applications. Evol Appl 9:291–310. https://doi.org/10.1111/eva.12316
Sozio G, Mortelliti A (2016) Empirical evaluation of the strength of interspecific competition in shaping small mammal communities in fragmented landscapes. Landscape Ecol 31:775–789. https://doi.org/10.1007/s10980-015-0286-1
Sozio G, Mortelliti A, Boitani L (2013) Mice on the move: wheat rows as a means to increase permeability in agricultural landscapes. Biol Cons 165:198–202. https://doi.org/10.1016/j.biocon.2013.05.022
Stacy JE, Jorde PE, Steen H, Ims RA, Purvis A, Jakobsen KS (1997) Lack of concordance between mtDNA gene flow and population density fluctuations in the bank vole. Mol Ecol 6:751–759. https://doi.org/10.1046/j.1365-294X.1997.d01-470.x
Storfer A, Murphy MA, Evans JS, Goldberg CS, Robinson S, Spear SF, Waits LP (2007) Putting the “landscape” in landscape genetics. Heredity 98:128–142. https://doi.org/10.1038/sj.hdy.6800917
Szacki J, Babinska-Werka J, Liro A (1993) The influence of landscape spatial structure on small mammal movements. Acta Theriol 38:113–123. https://doi.org/10.4098/AT.arch.93-10
Tattersall FH, Macdonald DW, Hart BJ, Manley WJ, Feber RE (2001) Habitat use by wood mice (Apodemus sylvaticus) in a changeable arable landscape. J Zool 255:487–494. https://doi.org/10.1017/S095283690100156X
Tew TE (1994) Farmland hedgerows: habitat, corridors or irrelevant? A small mammal’s perspective. Hedgerow Manag Nat Conserv 80:94
Thatte P, Chandramouli A, Tyagi A, Patel K, Baro P, Chhattani H, Ramakrishnan U (2020) Human footprint differentially impacts genetic connectivity of four wide-ranging mammals in a fragmented landscape. Divers Distrib 26:299–314. https://doi.org/10.1111/ddi.13022
Trombulak SC, Baldwin RF (2010) Landscape-scale conservation planning. Landsc-Scale Conserv Plan. https://doi.org/10.1007/978-90-481-9575-6
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Varudkar A, Ramakrishnan U (2015) Commensalism facilitates gene flow in mountains: a comparison between two Rattus species. Heredity 115:253–261. https://doi.org/10.1038/hdy.2015.34
Wang IJ (2013) Examining the full effects of landscape heterogeneity on spatial genetic variation: a multiple matrix regression approach for quantifying geographic and ecological isolation. Evolution 67:3403–3411. https://doi.org/10.1111/evo.12134
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370. https://doi.org/10.1111/j.1558-5646.1984.tb05657.x
Wilson MC, Chen XY, Corlett RT, Didham RK, Ding P, Holt RD, Yu M (2016) Habitat fragmentation and biodiversity conservation: key findings and future challenges. Landscape Ecol 31:219–227. https://doi.org/10.1007/s10980-015-0312-3
Wright S (1943) Isolation by distance. Genetics 28:114
Wu J (2013) Key concepts and research topics in landscape ecology revisited: 30 years after the allerton park workshop. Landscape Ecol 28:1–11. https://doi.org/10.1007/s10980-012-9836-y
Youngentob KN, Yoon HJ, Coggan N, Lindenmayer DB (2012) Edge effects influence competition dynamics: a case study of four sympatric arboreal marsupials. Biol Cons 155:68–76. https://doi.org/10.1016/j.biocon.2012.05.015
Acknowledgements
RB’s stay at the University of Potsdam was supported by the Erasmus Placement grant (UU-ER/2010/010744, M.S.) from the European Commission Life Long Learning programme. RB thanks Ralph Tiedemann for his hospitality at the Unit of Evolutionary Biology/Systematic Zoology (Institute of Biochemistry and Biology, University of Potsdam) during the laboratory analysis. We thank all students that helped us with fieldwork.
Funding
Open access funding provided by Università degli Studi di Ferrara within the CRUI-CARE Agreement. Funded was provided by Erasmus Placement (Grant no. UU-ER/2010/010744)
Author information
Authors and Affiliations
Contributions
RB, AM, VK and GB: conceived and designed the study. AM and GS: obtained the samples. RB and KH: carried out the laboratory work. RB and AB: analyzed the data. RB, AB and GB: wrote the manuscript, with contributions from all the authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Biello, R., Brunelli, A., Sozio, G. et al. The genetic structure and connectivity in two sympatric rodent species with different life histories are similarly affected by land use disturbances. Conserv Genet 24, 59–72 (2023). https://doi.org/10.1007/s10592-022-01485-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-022-01485-z