Abstract
Genetic estimation of population sizes has been critical for monitoring cryptic and rare species; however, population estimates do not inherently reveal the permanence or stability of the population under study. Thus, it is important to monitor not only the number of individuals in a population, but also how they are associated in groups and how those groups are distributed across the landscape. Adding to the challenge of obtaining such information with high precision for endangered and elusive species is the need for long-term collection of such data. In this study we compare sampling approaches and genotype non-invasive genetic samples to estimate the number and distribution of wild western lowland gorillas occupying a ~ 100 km2 area in Loango National Park, Gabon, for the periods 2005–2007 and 2014–2017. Based on the number of genotyped individuals we inferred a minimum of 83 gorillas during the first and 81 gorillas during the second study period. We also obtained similar capture–recapture population size estimates for the two periods despite variance in social dynamics like group formations, group dissolutions and individual dispersal. We furthermore found area fidelity for two groups that were sampled for 10–12 years, despite variation in group membership. Our results revealed how individual movements link groups in a ‘network’ and show that western lowland gorilla populations can show a high degree of temporal and geographic stability concurrent with substantial social dynamics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The world`s biodiversity is declining and over the past half century the global population size of vertebrate species has decreased by more than half (WWF 2018). Non-human primates are no exception, with currently ~ 60% of primate species threatened with extinction while ~ 75% have declining populations (reviewed in Estrada et al. 2017). To assess the status of wildlife, identify future trends and enable effective population management, accurate and precise population size estimates of species are crucial (Keith et al. 2015; Pacifici et al. 2015). However, many endangered species are elusive or live in inaccessible habitats, which makes direct counting impossible. Indirect counting methods have been developed and improved over the years based on indirect signs (e.g. red foxes: Cortázar-Chinarro et al. 2018), non-invasive genetics (e.g. gray wolves: Stansbury et al. 2014), acoustics (e.g. vaquitas: Jaramillo-Legorreta et al. 2017; e.g. primates: Kalan et al. 2015) and camera traps (e.g. great apes: Després-Einspenner et al. 2017; Kühl et al. 2008; Head et al. 2013). Using these and other methods, population size estimates for the same area at different points in time may be used to infer population trends like growth and decline. However, obtaining sufficiently precise estimates to confidently infer growth or decline of a population, particularly for species with slow intrinsic rates of growth, is challenging and demands intensive effort (goose: Fox et al. 2010; eastern chimpanzees: Granjon et al. 2017; brown bears: Solberg et al. 2006; flying squirrels: Sulkava et al. 2008, mountain gorillas: Roy et al. 2014).
Furthermore, while a declining population is readily interpreted as sign for concern, even apparent stability in population size may mask various processes with differing long-term implications. The population structure for instance can fluctuate independent of the population size and changes might affect the population dynamics or lead to population crashes if occurring in combination with other factors (soay sheeps: Coulson et al. 2001; lizards: Le Galliard et al. 2005). Furthermore variation in habitat quality might also lead to source-sink dynamics resulting in habitat-specific birth and death rates. While ‘sinks’ are of poorer quality and have higher death as compared to birth rates, ‘sources’ are of higher quality and the birth rate exceeds the death rate. ‘Sink’ populations are therefore only viable if immigration from source populations counterbalances the local mortality (pikas: Kreuzer and Huntly 2003; root voles: Gundersen et al. 2001). These processes however can only be uncovered if information beyond the population size is available.
Genetic analysis of non-invasive samples is a particularly useful approach for the assessment of wild animal populations. Genotyping and identity analyses of samples found in a given area can provide both the minimum number and the sex of individuals using the area and, if the collection scheme is properly designed, can also supply the necessary detection frequencies for mark-recapture population estimation (mountain gorillas: Roy et al. 2014; amur tigers: Sugimoto et al. 2012; grizzly bears: Mowat and Strobeck 2000; european otters: Prigioni et al. 2006). Information on sampling location and co-association of samples can be used to reconstruct minimal group composition for group-living animals (gray wolf packs: Stenglein et al. 2011) and repeated detection of individuals and groups allows for inference of minimal home range or insights into temporal or seasonal use of space (central chimpanzees: Arandjelovic et al. 2011; otters: Hung et al. 2004; moose: Bao et al. 2017; eagles: Bulut et al. 2016; lynx: Davoli et al. 2013). Group dynamic processes such as group formations, dissolutions, dispersal patterns and changes in group composition can be monitored through genetic tracking of individuals and groups over time (western lowland gorillas: Arandjelovic et al. 2010; Hagemann et al. 2018; gray wolves: Caniglia et al. 2014).
Both species of gorillas (western gorillas, Gorilla gorilla; eastern gorillas, Gorilla beringei) are critically endangered (IUCN 2019), live in dense forests and are elusive, which makes them important candidates for population assessment using non-invasive genetic analyses. Western gorilla groups are characterized by the presence of a single adult ‘silverback’ male, which sires all offspring with the adult females in the group (Bradley et al. 2004; Parnell 2002; Robbins et al. 2004). Both males and females disperse from their natal group upon maturity (Gatti et al. 2004; Stokes et al. 2003; Harcourt and Stewart 2007). After dispersal, males typically become solitary or members of non-reproductive groups and some proportion of them eventually become silverbacks of their own breeding groups (Breuer et al. 2009; Levréro et al. 2006; Parnell 2002; Robbins et al. 2004; Robbins and Robbins 2018). In contrast, females are not found alone but disperse upon reaching adulthood (natal transfer) and commonly change group membership repeatedly (secondary transfer). Such transfers may occur after the death of the silverback and dissolution of the group or during intergroup encounters (Stokes et al. 2003).
Description of the population size and dynamics of multiple western gorilla groups across a landscape is rendered difficult by the challenges of habituation and continued observation (Arandjelovic et al. 2010; Williamson and Feistner 2003). Using a mixture of indirect (nest sites and attempts to follow movements) and direct inferences during a ten-year study, Tutin described extensive overlap between groups at Lope Reserve, Gabon (Tutin 1996). In contrast, direct monitoring of one group of western lowland gorillas over 5 years at Lossi Forest, Republic of Congo, revealed limited home range overlap (Bermejo 2004). A study of one group undergoing habituation at Bai Hokou, Central African Republic, suggested some degree of site fidelity over a three-year period while ranging patterns were influenced by the sudden reduction of the group by half after an ‘attack’, and seasonal changes in fruit availability (Cipolletta 2004). Studies at bais (swampy forest clearings) have reported that gorilla groups converge in the clearings, indicating home range overlap, the extent of which remains unknown (e.g. Maya Bai: Magliocca et al. 1999; Mbeli Bai: Stokes et al. 2003). It seems apparent that overlap between home ranges might foster intergroup encounters and promote female transfer. This in turn facilitates disease transmission between groups, especially after the death of a silverback which in this uni-male species leads to the dissolution of the group and females moving to other groups (Nunn et al. 2008). A study at Odzala-Kokoua National Park, Republic of Congo, based on the observation of three habituated groups over 5 years plus non-invasive genetics over 4 month postulated a highly dynamic social system with tolerant intergroup encounters and interactions like social play between members of different groups (Forcina et al. 2019). Accordingly, the outbreak of infectious diseases like Ebola have led to documented population declines of 90% from October 2002 to January 2003 in Lossi Sanctuary, Republic of Congo or 95% from December 2003 to July 2004 at Lokoué clearing, Odzala-Kokoua National Park, Republic of Congo (Bermejo et al. 2006; Caillaud et al. 2006). Thus, it is important to better understand the group membership dynamics and space use of multiple nearby groups over a longer period of time.
Here we use results from genetic analysis of non-invasive samples collected over a 12-year period to assess a population of western lowland gorillas (Gorilla gorilla gorilla) occupying a 100 km2 area at the edge of the western gorilla range. We compared genotypes from samples collected during multiple surveys at two periods in time in Loango National Park, Gabon, 2005–2007 (Arandjelovic et al. 2010) and 2014–2017 (Hagemann et al. 2018). We compared the efficacy of different sampling approaches and derived both minimum as well as mark-recapture estimates of the number of gorillas for the two periods. We next used information from co-association of samples to examine changes in group composition and distribution of individuals over time, as well as space use by the groups. Our results provide a detailed population assessment illuminating the links between population stability, space use and social dynamics.
Methods
Sample collection
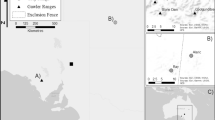
Between 2014 and 2017 we sampled gorilla feces within a ~ 100 km2 area in Loango National Park, Gabon. This area is bordered by a lagoon to the north-east and the Atlantic Ocean to the south-west and was also the setting of two previous studies on gorillas based on non-invasive genetic samples in 2005–2007 and 2009. In the first, a 101 km2 area was sampled over 2.5 years (Arandjelovic et al. 2010) and in the second 23 samples from two groups were added to the dataset (Arandjelovic et al. 2014). We utilize the 2005–2007 data for analyses regarding comparison between the periods on population size and refer to it as the ‘previous study’. For analyses spanning the entire period 2005–2017 on space use and population size estimates and analyses on population dynamics, we additionally utilized the data from 2009. For the present study, we divided the area into 25 2 km × 2 km grid cells. As previously described in detail (Hagemann et al. 2018), sampling was conducted by teams of two to three people and three different sampling approaches were applied: systematic sweeps, flexible design and opportunistic sampling, while always prioritizing fresh gorilla fecal samples (meaning assumed to be defecated within the last 2 days). The systematic sweeps targeted grid cells in order and were conducted between January 2014 and December 2015 with a slight modification after sweep four by allowing for increased search effort in grid cells with fresh signs of apes. Due to a low rate of sample acquisition, in January 2016 we switched to a flexible design, allowing for prioritization of grid cells with higher chances of success, for example due to fruit availability, during certain times of the year. During the entire study period samples were also collected opportunistically during other research tasks (Fig. 1). In the following we distinguish among (i) collected samples, which are all collected samples identified as gorilla feces in the field (Arandjelovic et al. 2010) and used for assessment of sampling success and (ii) genotyped samples, which are the portion of collected samples genotyped successfully and verified to originate from gorillas. These were used for identity analyses, parent–offspring analyses, group reconstruction, home range reconstruction and age estimation. Finally, we refer to (iii) captures, which are the subset of genotyped samples used for population size estimates. We derived monthly means of rainfall and maximum temperature of the respective year sweeps were conducted based on weather data collected in Yatouga camp (8–31 measurements per month for rainfall, average: 6.9 mm and 12–31 for max. temperature, average: 28.1 °C) (Loango gorilla project, long term data).
Genotyping and identity analyses
Of the 1317 samples collected, we prioritized 932 samples for analysis by freshness, occurrence at nest sites and proximity to other samples (Hagemann et al. 2018). As previously described in detail, a subset of the DNA extracts that passed a quality assessment was amplified at a total of 13 microsatellite loci and a sex-determining locus (Hagemann et al. 2018). After excluding 57 samples that amplified at fewer than 5 loci and 20 samples that STRUCTURE analyses revealed as originating from chimpanzees (Hagemann et al. 2018), we used the program CERVUS (version 3.0.7) (Kalinowski et al. 2007) to identify unique genotypes and compile replicates of the same genotype (pIDsib < 0.01) by comparing 681 gorilla samples that amplified at five to 13 microsatellite loci. To identify sampled gorillas who were also present in the population during the earlier study period, we compared our new data with the 85 consensus gorilla genotypes identified previously from samples collected in 2005–2007 and 2009 (Arandjelovic et al. 2010, 2014) using the same criteria but allowing for one mismatch to account for possible genotyping inconsistency. Two genotypes identified during 2005–2007/2009 matched genotypes identified during 2014–2017 but at fewer than five loci. Only one of these pairs was found within the same group across periods and was thus considered to belong to the same individual, the other was considered as two individuals.
Population size estimates
We used two different genetic capture–recapture approaches to estimate the population size. One approach requires discrete sampling periods in which individuals are considered as detected or not detected (White and Burnham 1999). The other approach allows for continuous sampling and considers the number of times individuals were (re)captured (Miller et al. 2005). Hence, the former is solely based on samples collected during the systematic sweeps with every completed sweep representing a discrete sampling period and the latter is based on samples collected during the entire collection period.
For the population size estimate approach relying on systematic sweeps (n = 6), we used the package RMark implemented in R (Laake 2013). We applied three different closed capture models, one which assumed capture probability was equal across systematic sweeps and individuals (M0), one which assumed equal capture probability across individuals but allowed for differences in capture probabilities across systematic sweeps (Mt) and one which assumed equal capture probability across systematic sweeps but allowed for two different capture probabilities across individuals (Mh). Model fit was evaluated using corrected Akaike Information Criterion (AICc) (Hurvich and Tsai 1989; Burnham et al. 1995).
For the continuous sampling approach we used the R package capwire (Pennell and Miller 2012) to estimate the number of gorillas in the study area following the idea of random sampling of mixing individuals (‘simple urn design’) and employing a maximum likelihood principle. We compared three models; the equal capture model (ECM), the two innate rates model (TIRM) and the partitioned two innate rates model (TIRMpart). The ECM assumes an equal detection rate and TIRM assumes two capture probability classes (high and low). TIRMpart works under the assumption that more than two capture classes exist, and thus partitions the data in three capture classes (Stansbury et al. 2014) and applies TIRM to a reduced dataset excluding individuals within the highest capture class as these may be an artifact of non-independence of sampling. We specified a maximum population size of 500 individuals and used a likelihood ratio test implemented in the package to assess the goodness-of-fit of the three different models. We derived 95% confidence intervals by parametric bootstrapping (n = 5000). This procedure was conducted based on samples found during the previous study (2005–2007) and the current study (2014–2017, systematic sweeps, flexible design and opportunistic) separately as well as over the entire time span 2005–2017, including samples from 2009, in order to compare the estimates derived from the two three-year periods with that based on the entire 12 year study. Such an extended sampling period might increase the probability of violating the assumption of population closure; however, it is noteworthy that a sampling period of 3 years yielded TIRM estimates consistent with shorter (3–12 months) sampling periods in long-lived species like western lowland gorillas (Arandjelovic et al. 2010) and eastern chimpanzees (Granjon et al. 2017).
Samples from the same individual found on the same day or estimated to be defecated on the same day within 20 m (Hagemann et al. 2018) of one another were treated as a single capture event and samples found more than 2 km outside the defined study area were excluded. Because the habituated research group Atananga was sampled frequently in 2014–2017 while it was subjected to near daily follows in the course of research, we excluded samples from this group for datasets including the period 2014–2017 for the population size assessment. For every estimate we added the known number of Atananga members born before 2016 (n = 14) to the minimum number of individuals, the point estimates and the lower and upper confidence interval. This procedure is statistically valid because we are adding a fixed number without uncertainty.
Group membership and transfers
Group membership was reconstructed as explained in detail in Arandjelovic et al. (2010, 2014) and Hagemann et al. (2018) for the period 2005–2007/2009 and 2014–2017. In brief: individuals whose samples were found on the same day within 20 m were assumed to be members of the same group. In addition we used patterns of co-association, meaning if individuals A, B and C are found together on an occasion and on another occasion D is found with A and B, then A, B, C and D are assumed to belong to the same group. Because western lowland gorillas live in social groups containing a single silverback, and females are rarely unassociated with a group, females whose samples were never found in proximity to other samples are deemed to have unknown group membership. However, a male who was found unassociated at least twice was considered to be a solitary silverback. Group composition was assumed to remain constant and transfers and dispersal events were inferred if an individual was found with one group and at a later point with another group or if a male was repeatedly found solitary. We further conducted parentage analysis implemented in the program CERVUS to infer parent–offspring relationships using all genotypes complete at eight or more loci and entered all individuals as potential offspring, all males as potential fathers and all females as potential mothers. We used this information to identify the silverback of the group by assuming that a male that fathered at least one offspring within a group is the group’s silverback. We only define a group as persisting across periods if the confirmed or unconfirmed silverback and at least one other member were found together during both time periods.
Gorilla group home range estimates
To estimate the minimum home range of groups we constructed minimum convex polygons (MCPs) encompassing all GPS-referenced samples from individuals belonging to the respective group. MCPs and their centroids (arithmetic mean position of all points in the MCP) were determined for all groups sampled at two or more occasions between 2005–2007/2009 and 2014–2017 separately. Analyses were conducted in R (R Core Team 2018) using the packages adehabitatHR (Calenge 2006) and rgeos (Bivand and Rundel 2017). We also calculated the percentage of MCP that did not overlap with the MCP of any other group to investigate minimum home range overlap. Furthermore we calculated the percent overlap of MCPs of the same groups found repeatedly over time during both time periods (n = 2) to assess site fidelity by dividing the overlapping area by the total area of the MCP derived for the respective group based on the first period. We chose this procedure since the number of data points varied for the groups between the study periods. The MCPs are based on pooled data from one up to 2.8 years.
Results
Sampling success rates and minimum number of gorillas
We collected 11–57 (average 33) samples during each of six complete systematic sweeps lasting between 60 and 91 days (Fig. 1). The change in protocol implementing increased effort in grid cells with fresh signs of apes, applied after sweep four, led to slight increase in the number of samples found. Although comments by the collectors suggested that sampling success was affected by temperature and rainfall, average monthly data do not reveal an obvious pattern (Fig. 1). The systematic sweep samples were found to represent 35 gorillas, each of which was ‘captured’ in a maximum of three of the six sweeps and between one and five times per sweep (Fig. 1).
The switch to a flexible collection plan in January 2016 dramatically improved sampling rates, yielding an additional 783 samples corresponding to 270 captures of 52 individuals over 14 months. In addition, 302 samples were collected opportunistically over 38 months between January 2014 and February 2017, resulting in 69 captures of 30 individuals. Overall, we genetically detected 98 gorillas (66 female, 32 male) between February 2014 and February 2017.
Population size estimates
Using the Mark M0 model, which requires discrete sampling sessions and assumes constant capture probabilities across individuals and time based on the capture histories of the 35 individuals found during the systematic sweeps, we obtain an estimate of 61 (95% CIM0 54–80) gorillas. Using the Mt model that allows for variation in capture probability across time we obtained a similar estimate of 59 (95% CIMt 53–76) gorillas. The Mh model allowing for capture heterogeneity across individuals likewise produced a similar estimate of 61 (95% CIMh 54–80) (Fig. 2). Model comparison revealed the most support for the Mt model with a relative AICc weight of > 0.99.
The estimated number of gorillas inferred using three detection models (M0, Mt and Mh) based on samples collected during systematic sweeps between 2014–2015 and three continuous sampling models (ECM, TIRM, TIRMpart) based on samples collected in the periods 2014–2017, 2005–2007 and 2005–2017. Points represent the point estimates while the black lines represent the 95% confidence intervals. Asterisks represent the number of unique genotypes and thus the minimum number of individuals in the study period
We next inferred the population size based on a continuous sampling period, which allowed us to use more data and incorporate the capture histories of 67 individuals which were genetically captured between one and 30 times (average 6.8) in 2014–2017. The three estimates obtained using Capwire (Miller et al. 2005; R package: Pennell et al. 2013) were highly similar (Fig. 2), and models allowing for capture heterogeneity between individuals received the most support (TIRMpart supported over TIRM and TRIM over ECM, p < 0.001), with the TIRMpart model returning an estimate of 89 gorillas (95% CITIRMpart 81–92).
For temporal comparison we re-estimated the number of gorillas using the area in 2005-2007 using the capture histories of 83 individuals who were detected between one and 12 times each (average 3.4) (Arandjelovic et al. 2010). Again, models permitting capture heterogeneity were highly supported (TIRMpart supported over TIRM and TRIM over ECM, p < 0.001) and we inferred an estimate of 104 (95% CITIRMpart 93–120) gorillas in the first period (2005–2007).
Finally, we combined the data, resulting in capture histories of 114 individuals detected between one and 30 times each (average 6.4), across the entire 12-year period (2005–2017) and, as before, the TIRMpart model was the most supported (p < 0.001). This yielded an estimate of 147 (95% CITIRMpart 134–153) gorillas.
Spatial distribution of captures
As might be expected given the heterogeneity of the landscape (Head et al. 2012), captures were not evenly distributed across the study area (Fig. 3). In both time periods (2005–2007 and 2014–2017) we found the majority of samples in proximity to swamps and in the coastal forest by the Atlantic Ocean, while few samples were found in the middle and the southeastern part of the study area. Accordingly, the MCP encompassing all captures during the systematic sweeps spans a 71 km2 area even though the search effort was constant across the 100 km2 area (Fig. 3a). Adding captures from opportunistic sampling and the flexible design expands the MCP to 103 km2, as compared to the 91 km2 surveyed in the previous study (2005–2007) (Fig. 3b, c). If we combine the captures for the current and previous study, adding samples found in 2009, the MCP enlarges slightly to 111 km2 (Fig. 3d).
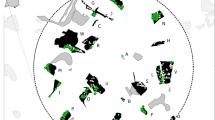
Capture locations and area utilized for the capture–recapture analyses according to time period. The outer polygons represent MCPs, circles represent captures with the area proportional to the number of captures. Capture locations for a only systematic samples collected during 2014–2015 in green, b all captures from 2005 to 2007 in blue, c all captures from 2014 to 2017 in pink d all captures from 2005 to 2007/2009 and 2014 to 2017 combined (with 2009 data in orange). The squares represent the grids implemented in 2014–2017 and thus the main sampling area of this study. In gray is forest area within the grid cells (darker grey when also within the respective MCP). Green lines represent edges of swamps and the lagoon which is inaccessible while yellow lines represent edges of savanna which is accessible but rarely used. (Color figure online)
Home ranges and site fidelity
In order to assess how the heterogeneous use of landscape is linked to social group ranges, we used previous inferences on the minimum composition of 11 mixed sex groups present in 2014–2017 obtained through sampling dates and occasions (Hagemann et al. 2018). Seven groups (Atananga, Group Green, Group Purple, Group Orange, Group Pink, Indegho and GroupC) had sufficient GPS data, between 29 and 186 GPS points per group (average 84.7), for estimation of MCPs. MCP areas varied between 15 and 32 km2 and over 70% of the respective areas overlapped with one another (Fig. 4a). We found that group MCPs overlapped with two (for example Group Green) to six (Group Purple) other groups but nevertheless groups can roughly be attributed to certain areas: four groups (Pink, Purple, Indegho and Atananga) range in the northern part, Groups Orange and Green in the south-eastern part and GroupC in the west (Fig. 4a).
a MCPs of the groups found in 2014–2017 with points indicating the respective centroids. The area of MCPs, indicated by filled polygons varies between 15 km2 for Group Green and 32 km2 for GroupC (MCPs in km2: Atananga = 23, Group Green = 15, Group C = 32, Indegho = 18, Group Orange = 18, Group Pink = 22, Group Purple = 28). b MCPs of the two groups that were sampled intensively in both study periods. (Color figure online)
Several of the 11 groups found in 2014–2017 were present during 2005–2007/2009 (Atananga, Indegho, LayonA and GroupC), allowing us to investigate long-term site fidelity for two of the groups that were sampled repeatedly in both periods, ranging from 9.9 years (Indegho) to 11.8 years (GroupC) (Hagemann et al. 2018). Both groups exhibited some consistency in home range use across the two time periods with a percentage of overlapping area in relation to the first MCP of 46% for GroupC and 48% for Indegho (Fig. 4b).
Between group transfer
After showing that at least some groups occupy consistent portions of the surveyed area over long time periods, we next turned to consideration of how individuals move across the landscape by dispersing. We previously inferred 33 dispersal events (28 female, five male) between mixed sex groups along with five group dissolutions and six group formations between 2005 and 2017 (Hagemann et al. 2018). All 12 groups that were sampled multiple times within the study area are connected to another group via individual transfers, thus forming a ‘network’ (Fig. 5), here defined as groups interconnected by individual movements. Nevertheless, we detected that certain groups exchange more individuals than others. One example is Group Purple, all three females that left this group transferred to Group Orange, a neighboring group with 4.3 km2 overlap in MCP (24% with regard to the smaller MCP). Similarly, eight females and one male of one of the two well-sampled groups that dissolved (Achilles) ended up in only four different groups, all in rough proximity with regard to the MCP centroid to the group of origin. In contrast four females and three males that left the other dissolved group (Madondo) were subsequently found in seven separate groups, mostly not in proximity to the group of origin.
An overview of 26 female transfers between 12 well-sampled mixed sex groups within the 2005–2017 period. Symbols indicate the centroids of the MCPs, triangles are the groups that were only observed in the previous study, circles the groups only observed during this study and squares the groups observed in both studies (based on combined samples across studies). The thickness of arrows is proportional to the number of detected transfers
Discussion
Sampling design
Our results reveal that by yielding only 232 samples over 24 months, strictly systematic sampling was inefficient when applied exclusively to a group-living species with heterogeneous use of space, in difficult habitat and with low abundance. Previous studies at the same locality suggest that the gorilla density is ~ one gorilla per km2 (Arandjelovic et al. 2010; Head et al. 2013) and that the gorillas do not use the area evenly but show varying, season dependent, preferences for particular habitat types (Head et al. 2012). As a consequence, some of the grid cells surveyed by systematic sampling teams may have represented areas that are rarely used by gorillas during certain times. In addition, in contrast to mountain gorilla sites where groups can regularly be tracked from one nest site to the next (Gray et al. 2013; Roy et al. 2014), the ground cover in Loango is predominantly open, which makes it difficult to follow gorilla tracks particularly in less swampy areas, leading to poor tracking success even if fresh signs were encountered. Fortunately, a change of protocol permitting the use of prior knowledge and experience at the site to prioritize areas with higher chances of gorilla presence during certain times led to a substantial increase in sampling success with the additional advantage that the impact of logistical, labor and weather related constraints was reduced due to increased flexibility. These changes enabled the collection of 783 samples during the 14 months of flexible sampling.
The sampling situation is reflected in the population assessment. While the use of all data from 2014 to 2017 yielded a minimum count of 81 gorilla genotypes and a relatively precise population size estimate of 81–92 gorillas (derived with Capwire), the exclusive use of samples collected during the six completed systematic sweeps yielded a minimum count of 49 and a less precise estimate of 53–76 gorillas (derived with MARK), thus underestimating the population size. This might be attributed to the difference in sample size (MARK: 56 captures of 35 individuals; Capwire: 454 captures of 67 individuals) amplified by the tendency of MARK to underestimate and Capwire to overestimate abundance (Woodruff et al. 2018). The prolongation of the time period to comprise all data available from 2005 to 2017 naturally led to an increase in sample size (733 captures of 114 individuals) but resulted in an overestimation of the population size (134–153 gorillas), in addition the TIRM estimate indicated poor model fit with the point estimate outside the 95% confidence interval. This shows that while a three-year-period does not seem to strongly violate the closed-population assumption (no birth, no death, no immigration and no emigration) in long-lived species with slow reproductive and low mortality rates like great apes (Arandjelovic et al. 2010; Granjon et al. 2017), there is a limit to how long the sampling period may be extended and 12 years violates the assumption excessively. The risk of violating the population closure assumption has been widely debated since it is expected to produce a positive bias due to inflation of detected individuals (grizzly bears: Boulanger and McLellan 2001; Proctor et al. 2010; bobcats: Ruell et al. 2009). In sum, our findings highlight the importance of the sampling design since it not only influences the precision but also the accuracy of the estimate.
Gorilla numbers and group dynamics
The minimum numbers of genotyped gorillas for the periods 2005–2007 and 2014–2017 were highly similar (83 compared to 81) despite the increased number of captures during the later period (284 compared to 454). This suggests that the majority of available individuals were captured during each period and thus the increase in sample size did not lead to an increase of detected individuals. Comparing the derived population size estimates between the two periods 2005–2007 and 2014–2017 revealed higher precision for the later time period which is to be expected due to the increase in sample size (Arandjelovic and Vigilant 2018; Miller et al. 2005; Woodruff et al. 2018). The derived point estimates are slightly higher for the period 2005–2007 compared to 2014–2017 and the confidence intervals do not overlap for the model with the most support (TIRMpart), suggesting a possible slight decrease in the population size in the 2014–2017 period. Confidence intervals for the other models (ECM and TIRM) did however overlap over the two study periods. In sum, our mark-recapture analysis does not reveal evidence for an obvious change in population size and suggests population stability. The study site in Loango National Park is characterized by low levels of poaching and human disturbance which may allow for this observed stability over time.
Of the 83 individuals identified in 2005–2007, 38 were found again in 2014–2017 and were thus present in the study area for 6–12 years. Individuals that were found only in one of the sampling periods may have died or been born, immigrated or emigrated or not sampled in either sampling period. One of the features of our study site is that it is bordered by a lagoon on one side, the Atlantic Ocean on the other and neighbored by large savanna patches on a third, so exchange or expansion is not possible in all directions. In a previous study we reconstructed group dynamic events (group formations, dissolutions and individuals dispersal) and revealed that the time period 2005–2007/2009 was more dynamic compared to 2014–2017. Thus interestingly, we observed overall stability in population size over time despite this varying social dynamics and change in individuals. The observed variance in social dynamics across time periods might be a product of stochasticity due to the observation of relatively rare events within relatively short time periods stressing the value of long-term studies.
Space use over time and group overlap
Consistent with a lack of notable climate or human-driven effects on the ecology and landscape of the study region, we did not observe differences in the distribution of samples across the sampling area between 2005–2007 and 2014–2017. The majority of samples were found on the edge of swamps or in the coastal forest. We found some long-term area fidelity across 9.9–11.8 years (Fig. 4b), despite varying group membership (Hagemann et al. 2018). For two groups we found evidence of seasonal variation in habitat use in 2014–2017 (data not shown), consistent with an earlier study of multiple groups based on camera traps at the same locality suggesting that gorillas show varying landscape preferences depending on rainfall and fruit abundance (Head et al. 2012). Indeed, it has been shown in several species that spatial knowledge can have adaptive benefits. These can occur via reduced predation risk, as in ungulates (Forrester et al. 2015), and foraging advantages, as seen in fur seals (Arthur et al. 2015). Fruits are an energy-rich but challenging food source due to their seasonal and patchy distribution (Milton 2003; Van Schaik et al. 1993; Masi et al. 2009; Remis 1997), and knowledge of the area may be advantageous for relocating fruiting trees at the right time (reviewed for primates: Zuberbühler and Janmaat 2010).
All groups that were sampled several times within this study showed substantial overlap in MCP with up to six other groups. The fraction of MCP that did not overlap with any other group was typically less than 20%. This is consistent with other studies on western lowland gorillas revealing variable extent of home range overlap between groups and frequent non-aggressive intergroup encounters (Seiler et al. in prep., Bermejo 2004; Doran-Sheehy et al. 2004; Doran and McNeilage 1998; Tutin 1996; Forcina et al. 2019). In addition to the inherent problems of the MCP method (Burgman and Fox 2003; Börger et al. 2006) a caveat of our study is that the MCPs encompass areas that are accessible for gorillas but also areas that are not accessible for gorillas (like water surfaces) and are constructed over a time frame of up to 2.8 years thus they do not necessarily represent home ranges. It is possible that groups use the same area, but not at the same time. However, all groups within the study area observed between 2005 and 2017 are connected with each other by individual dispersal, contributing to the picture of network in which some groups exchange more individuals than other groups. This could be due to spatial proximity and a higher chance of intergroup encounters facilitating female transfers or immediate involuntary female transfers following the sudden dissolution of a group due to the death of the silverback (Stokes et al. 2003). This impression of connectedness fits the observation that infectious disease outbreaks, like Ebola, have severe consequences on western lowland populations due to their changing group membership and incorporation of new individuals (Bermejo et al. 2006; Genton et al. 2015; Le Gouar et al. 2009; Forcina et al. 2019).
Conclusion
Our study highlights the power of non-invasive genetics to provide estimates of population sizes as well as illuminate population dynamics. Our results showed that systematic sample collection was rendered difficult by seasonality, landscape and social group effects which led to non-homogenous distribution of individuals across the study area. A flexible yet targeted sampling design is thus worth considering for a group living species in difficult habitat with low abundance and heterogeneous use of space. Only the combination of information on the number of gorillas and the use of space combined enabled us to conclude long-term population stability at our study site. This is despite variation in social dynamics, thus confirming that fluctuation in social dynamics is not necessarily coupled with population instability.
Data availablity
The consensus microsatellite genotypes analysed during the current study are available in the KNB repository, https://doi.org/10.5063/F1M61HK9.
References
Arandjelovic M, Vigilant L (2018) Non-invasive genetic censusing and monitoring of primate populations. Am J Primatol. https://doi.org/10.1002/ajp.22743
Arandjelovic M, Kühl H, Boesch C et al (2010) Effective non-invasive genetic monitoring of multiple wild western gorilla groups. Biol Conserv 143:1780–1791. https://doi.org/10.1016/j.biocon.2010.04.030
Arandjelovic M, Head J, Rabanal LI et al (2011) Non-invasive genetic monitoring of wild central chimpanzees. PLoS ONE 6:1–11. https://doi.org/10.1371/journal.pone.0014761
Arandjelovic M, Head J, Boesch C et al (2014) Genetic inference of group dynamics and female kin structure in a western lowland gorilla population (Gorilla gorilla gorilla). Primate Biol 1:29–38. https://doi.org/10.5194/pb-1-29-2014
Arthur B, Hindell M, Bester M et al (2015) Return customers: foraging site fidelity and the effect of environmental variability in wide-ranging antarctic fur seals. PLoS ONE 10:1–19. https://doi.org/10.1371/journal.pone.0120888
Bao H, Fryxell JM, Hui L et al (2017) Effects of interspecific interaction-linked habitat factors on moose resource selection and environmental stress. Sci Rep 7:1–10. https://doi.org/10.1038/srep41514
Bermejo M (2004) Home-range use and intergroup encounters in Western Gorillas (Gorilla g. gorilla) at Lossi Forest, North Congo. Am J Primatol 232:223–232. https://doi.org/10.1002/ajp.20073
Bermejo M, Domingo Rodríguez-Teijeiro J, Illera G et al (2006) Ebola outbreak killed 5000 gorillas. Science 314:1564. https://doi.org/10.1126/science.1133105
Bivand R, Rundel C. (2017) rgeos: Interface to geometry engine open source (‘GEOS’). R package
Börger L, Franconi N, De Michele G et al (2006) Effects of sampling regime on the mean and variance of home range size estimates. J Anim Ecol 75:1393–1405. https://doi.org/10.1111/j.1365-2656.2006.01164.x
Boulanger J, McLellan B (2001) Closure violation in DNA-based mark-recapture estimation of grizzly bear populations. Can J Zool 79:642–651. https://doi.org/10.1139/cjz-79-4-642
Bradley BJ, Doran-Sheehy DM, Lukas D et al (2004) Dispersed male networks in Western Gorillas. Curr Biol 14:510–513
Breuer T, Hockemba MB, Olejniczak C et al (2009) Physical maturation, life-history classes and age estimates of free-ranging Western Gorillas—insights from Mbeli Bai, Republic of Congo. Am J Primatol 71:106–119. https://doi.org/10.1002/ajp.20628
Bulut Z, Bragin EA, DeWoody JA et al (2016) Use of noninvasive genetics to assess nest and space use by white-tailed eagles. J Raptor Res 50:351–362
Burgman MA, Fox JC (2003) Bias in species range estimates from minimum convex polygons: implications for conservation and options for improved planning. Anim Conserv 6:19–28. https://doi.org/10.1017/S1367943003003044
Burnham KP, White GC, Anderson DR (1995) Model selection strategy in the analysis of capture-recapture data. Biometrics 51:888–898
Caillaud D, Levrero F, Cristescu R et al (2006) Gorilla susceptibility to Ebola virus: the cost of sociality. Curr Biol 16:489–491
Calenge C (2006) The package “adehabitat” for the R software: a tool for the analysis of space and habitat use by animals. Ecol Modell 197:516–519. https://doi.org/10.1016/j.ecolmodel.2006.03.017
Caniglia R, Fabbri E, Galaverni M et al (2014) Noninvasive sampling and genetic variability, pack structure, and dynamics in an expanding wolf population. J Mammal 95:41–59
Cipolletta C (2004) Effects of group dynamics and diet on the ranging patterns of a western gorilla group (Gorilla gorilla gorilla) at Bai Hokou, Central African Republic. Am J Primatol 64:193–205. https://doi.org/10.1002/ajp.20072
Cortázar-Chinarro M, Halvarsson P, Virgós E (2018) Sign surveys for red fox (Vulpes vulpes) censuses: evaluating different sources of variation in scat detectability. Mam Res 64:183. https://doi.org/10.1007/s13364-018-0404-y
Coulson T, Catchpole EA, Albon SD et al (2001) Age, sex, density, winter weather, and population crashes in soay sheep. Science 292:1528–1532. https://doi.org/10.1126/science.292.5521.1528
Davoli F, Schmidt K, Kowalczyk R et al (2013) Hair snaring and molecular genetic identification for reconstructing the spatial structure of Eurasian lynx populations. Mamm Biol 78:118–126. https://doi.org/10.1016/j.mambio.2012.06.003
Després-Einspenner ML, Howe EJ, Drapeau P, Kühl HS (2017) An empirical evaluation of camera trapping and spatially explicit capture-recapture models for estimating chimpanzee density. Am J Primatol 79:1–12. https://doi.org/10.1002/ajp.22647
Doran DM, Mcneilage A (1998) Gorilla ecology and behavior. Evol Anthropol 6:120–131
Doran-Sheehy DM, Greer D, Mongo P, Schwindt D (2004) Impact of ecological and social factors on ranging in western gorillas. Am J Primatol 64:207–222. https://doi.org/10.1002/ajp.20075
Estrada A, Garber PA, Rylands AB et al (2017) Impending extinction crisis of the world’s primates: why primates matter. Sci Adv 3:1–16. https://doi.org/10.1126/sciadv.1600946
Forcina G, Vallet D, Le Gouar PJ et al (2019) From groups to communities in western lowland gorillas. Proc R Soc B 286:1–9. https://doi.org/10.1098/rspb.2018.2019
Forrester TD, Casady DS, Wittmer HU (2015) Home sweet home: fitness consequences of site familiarity in female black-tailed deer. Behav Ecol Sociobiol 69:603–612. https://doi.org/10.1007/s00265-014-1871-z
Fox AD, Ebbinge BS, Mitchell C et al (2010) Current estimates of goose population sizes in western Europe, a gap analysis and an assessment of trends. Ornis Svecica 20:115–127. https://doi.org/10.34080/os.v20.19922
Gatti S, Levréro F, Ménard N, Gautierhion A (2004) Population and group structure of western Lowland Gorillas (Gorilla gorilla gorilla) at Lokoué, Republic of Congo. Am J Primatol 63:111–123. https://doi.org/10.1002/ajp.20045
Genton C, Pierre A, Cristescu R et al (2015) How Ebola impacts social dynamics in gorillas: a multistate modelling approach. J Anim Ecol 84:166–176. https://doi.org/10.1111/1365-2656.12268
Granjon AC, Rowney C, Vigilant L, Langergraber KE (2017) Evaluating genetic capture-recapture using a chimpanzee population of known size. J Wildl Manag 81:279–288. https://doi.org/10.1002/jwmg.21190
Gray M, Roy J, Vigilant L et al (2013) Genetic census reveals increased but uneven growth of a critically endangered mountain gorilla population. Biol Conserv 158:230–238. https://doi.org/10.1016/j.biocon.2012.09.018
Gundersen G, Johannesen E, Andreassen HP, Ims RA (2001) Source—sink dynamics: how sinks affect demography of sources. Ecol Lett 4:14–21
Hagemann L, Boesch C, Robbins MM et al (2018) Long-term group membership and dynamics in a wild western lowland gorilla population (Gorilla gorilla gorilla) inferred using non-invasive genetics. Am J Primatol. https://doi.org/10.1002/ajp.22898
Harcourt AH, Stewart KJ (2007) Gorilla society: conflict, compromise, and cooperation between the sexes. University of Chicago Press, Chicago
Head JS, Robbins MM, Mundry R et al (2012) Remote video-camera traps measure habitat use and competitive exclusion among sympatric chimpanzee, gorilla and elephant in Loango National Park, Gabon. J Trop Ecol 28:571–583. https://doi.org/10.1017/S0266467412000612
Head JS, Boesch C, Robbins MM et al (2013) Effective sociodemographic population assessment of elusive species in ecology and conservation management. Ecol Evol 3:2903–2916. https://doi.org/10.1002/ece3.670
Hung C, Li S, Lee L (2004) Faecal DNA typing to determine the abundance and spatial organisation of otters (Lutra lutra) along two stream systems in Kinmen. Anim Conserv 7:301–311
Hurvich CM, Tsai C-L (1989) Regression and time series model selection in small samples. Biometrika 76:297–307. https://doi.org/10.2307/2336663
IUCN (2019) The IUCN red list of threatened species. Version 2019-2. http://www.iucnredlist.org
Jaramillo-Legorreta A, Cardenas-Hinojosa G, Nieto-Garcia E et al (2017) Passive acoustic monitoring of the decline of Mexico’s critically endangered vaquita. Conserv Biol 31:183–191. https://doi.org/10.1111/cobi.12789
Kalan AK, Mundry R, Wagner OJJ et al (2015) Towards the automated detection and occupancy estimation of primates using passive acoustic monitoring. Ecol Indic 54:217–226. https://doi.org/10.1016/j.ecolind.2015.02.023
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106. https://doi.org/10.1111/j.1365-294X.2007.03089.x
Keith D, Akçakaya HR, Butchart SHM et al (2015) Temporal correlations in population trends: conservation implications from time-series analysis of diverse animal taxa. Biol Conserv 192:247–257. https://doi.org/10.1016/j.biocon.2015.09.021
Kreuzer MP, Huntly NJ (2003) Habitat-specific demography: evidence for source-sink population structure in a mammal, the pika. Oecologia 134:343–349. https://doi.org/10.1007/s00442-002-1145-8
Kühl H, Maisels F, Ancrenaz M, Williamson EA (2008) Best practice guidelines for surveys and monitoring of great ape populations. IUCN
Laake JL (2013) RMark: An R interface for analysis of capture-recapture data with MARK. AFSC Process Rep 2013-01
Le Galliard J-F, Fitze PS, Ferriere R, Clobert J (2005) Sex ratio bias, male aggression, and population collapse in lizards. PNAS 102:18231–18236
Le Gouar PJ, Vallet D, David L et al (2009) How Ebola impacts genetics of western lowland gorilla populations. PLoS ONE. https://doi.org/10.1371/journal.pone.0008375
Levréro F, Gatti S, Ménard N et al (2006) Living in nonbreeding groups: an alternative strategy for maturing gorillas. Am J Primatol 68:275–291. https://doi.org/10.1002/ajp.20223
Magliocca F, Querouil S, Gautier-Hion A (1999) Population structure and group composition of western lowland gorillas in North-Western Republic of Congos. Am J Primatol 48:1–14
Masi S, Cipolletta C, Robbins MM (2009) Western Lowland Gorillas (Gorilla gorilla gorilla) change their activity patterns in response to frugivory. Am J Primatol 71:91–100. https://doi.org/10.1002/ajp.20629
Miller CR, Joyce P, Waits LP (2005) A new method for estimating the size of small populations from genetic mark-recapture data. Mol Ecol 14:1991–2005. https://doi.org/10.1111/j.1365-294X.2005.02577.x
Milton K (2003) Animal source foods to improve micronutrient nutrition and human function in developing countries the critical role played by animal source foods in human (Homo) evolution. Am Soc Nutr Sci 133:3886–3892
Mowat G, Strobeck C (2000) Estimating population size of grizzly bears using hair capture, DNA profiling, and mark-recapture analysis. J Wildl Manag 64:183–193
Nunn CL, Thrall PH, Stewart K, Harcourt AH (2008) Emerging infectious diseases and animal social systems. Evol Ecol 22:519–543. https://doi.org/10.1007/s10682-007-9180-x
Pacifici M, Foden WB, Visconti P et al (2015) Assessing species vulnerability to climate change. Nat Clim Change 5:215–225. https://doi.org/10.1038/nclimate2448
Parnell RJ (2002) Group size and structure in Western Lowland Gorillas (Gorilla gorilla gorilla) at Mbeli Bai, Republic of Congo. Am J Primatol 56:193–206. https://doi.org/10.1002/ajp.1074
Pennell MW, Miller CR (2012) Capwire: Estimates population size from non-invasive sampling. R package version 1.1.4. https://CRAN.R-project.org/package=capwire
Pennell MW, Stansbury CR, Waits LP, Miller CR (2013) Capwire: a R package for estimating population census size from non-invasive genetic sampling. Mol Ecol Resour 13:154–157. https://doi.org/10.1111/1755-0998.12019
Prigioni C, Remonti L, Balestrieri A et al (2006) Estimation of European Otter (Lutra Lutra) population size by fecal DNA typing in Southern Italy. J Mammal 87:855–858. https://doi.org/10.1017/S175173111800023X
Proctor M, Mclellan B, Boulanger J et al (2010) Ecological investigations of grizzly bears in Canada using DNA from hair, 1995–2005: a review of methods and progress. Ursus 21:169–188
R Core Team (2018) R: A language and environment for statistical computing. R foundation for statistical computing, Vienna, Austria. https://www.R-project.org/
Remis MJ (1997) Western lowland gorillas (Gorilla gorilla gorilla) as seasonal frugivores: use of variable resources. Am J Primatol 43:87–109
Robbins MM, Robbins AM (2018) Variation in the social organization of gorillas: life history and socioecological perspectives. Evol Anthropol 27:1–16
Robbins MM, Bermejo M, Cipolletta C et al (2004) Social structure and life-history patterns in western gorillas (Gorilla gorilla gorilla). Am J Primatol 64:145–159. https://doi.org/10.1002/ajp.20069
Roy J, Vigilant L, Gray M et al (2014) Challenges in the use of genetic mark-recapture to estimate the population size of Bwindi mountain gorillas (Gorilla beringei beringei). Biol Conserv 180:249–261. https://doi.org/10.1016/j.biocon.2014.10.011
Ruell EW, Riley SP, Douglas MR et al (2009) Estimating bobcat population sizes and densities in a fragmented urban landscape using noninvasive capture—recapture sampling. J Mammal 90:129–135
Solberg KH, Bellemain E, Drageset OM et al (2006) An evaluation of field and non-invasive genetic methods to estimate brown bear (Ursus arctos) population size. Biol Conserv 128:158–168. https://doi.org/10.1016/j.biocon.2005.09.025
Stansbury CR, Ausband DE, Zager P et al (2014) A long-term population monitoring approach for a wide-ranging carnivore: noninvasive genetic sampling of gray wolf rendezvous sites in Idaho, USA. J Wildl Manage 78:1040–1049. https://doi.org/10.1002/jwmg.736
Stenglein JL, Waits LP, Ausband DE et al (2011) Estimating gray wolf pack size and family relationships using noninvasive genetic sampling at rendezvous sites. J Mammal 92:784–795. https://doi.org/10.1644/10-MAMM-A-200.1
Stokes EJ, Parnell RJ, Olejniczak C (2003) Female dispersal and reproductive success in wild western lowland gorillas (Gorilla gorilla gorilla). Behav Ecol Sociobiol 54:329–339. https://doi.org/10.1007/s00265-003-0630-3
Sugimoto T, Nagata J, Aramilev VV, Mccullough DR (2012) Population size estimation of Amur tigers in Russian Far East using noninvasive genetic samples. J Mammal 93:93–101. https://doi.org/10.1644/10-MAMM-A-368.1
Sulkava R, Mäkelä A, Kotiaho JS, Mönkkönen M (2008) Difficulty of getting accurate and precise estimates of population size: the case of the Siberian flying squirrel in Finland. Ann Zool Fenn 45:521–526. https://doi.org/10.5735/086.045.0607
Tutin CEG (1996) Ranging and social structure of lowland gorillas in the Lopé Reserve, Gabon. In: McGrew WC, Archant LF, Nishida T (eds) Great Ape Societies. Cambridge University Press, Cambridge, pp 58–70
Van Schaik CP, Terborgh JW, Wright SJ (1993) The phenology of tropical forests: adaptive significance and consequences for primary consumers. Annu Rev Ecol Syst 24:353–377
White GC, Burnham KP (1999) Program mark: survival estimation from populations of marked animals. Bird Study 46:120–139. https://doi.org/10.1080/00063659909477239
Williamson EA, Feistner ATC (2003) Habituating primates: processes, techniques, variables and ethics. In: Curtis DJ, Setchell JM (eds) Field and laboratory methods in primatology: a practical guide. Cambridge University Press, Cambridge, pp 25–39
Woodruff SP, Lukacs PM, Waits LP (2018) Comparing performance of multiple non-invasive genetic capture—recapture methods for abundance estimation: a case study with the Sonoran pronghorn Antilocapra americana sonoriensis. Oryx. https://doi.org/10.1017/S003060531800011X
WWF (2018) Living planet report 2018: aiming higher. Gland, Switzerland
Zuberbühler K, Janmaat K (2010) Foraging cognition in nonhuman primates. In: Platt ML, Ghazanfar AA (eds) Primate neuroethology. Oxford University Press, Oxford, pp 64–83
Acknowledgements
Open access funding provided by Max Planck Society. We thank the Agence Nationale des Parcs Nationaux (ANPN) and the Centre National de la Recherche Scientifique et Technique (CENAREST) of Gabon for permission to conduct our research in Loango National Park. Funding was provided by the United States Fish and Wildlife Service Great Ape Fund, Tusk Trust, Berggorilla und Regenwald Direkthilfe e. V. and the Max Planck Society. We are very grateful to F. Baguette, T. Moubec, S. Luccesi, J. Richardson, K. Judson, K. Sweeny, A. Granjon, K. Mannion, M. Basile, E. Theleste, K. Dierks, C. Nebel and all involved trackers from Waka National Park for their help with sample collection and importation. We thank A. Nicklisch and K. Wuttig for laboratory support and V. Städele, H. Jang, M. Manguette, E. Wright, N. Seiler, J. Lester and A. Granjon for helpful discussion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Hagemann, L., Arandjelovic, M., Robbins, M.M. et al. Long-term inference of population size and habitat use in a socially dynamic population of wild western lowland gorillas. Conserv Genet 20, 1303–1314 (2019). https://doi.org/10.1007/s10592-019-01209-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-019-01209-w