Abstract
Purpose
Up to 10% of all breast cancers (BC) are attributed to inherited pathogenic variants (PV) in BC susceptibility genes; however, most carriers of PVs remain unidentified. Here, we sought to determine the yield of hereditary cancer gene PVs among diverse women attending breast imaging centers, who could benefit from enhanced surveillance and/or risk reduction interventions.
Methods
This cross-sectional retrospective cohort study included consecutive women, unselected for personal or family cancer history, who were offered genetic testing for hereditary cancer genes at the time of breast imaging at three centers (November 2020–March 2022).
Results
Among 1943 patients (median age: 66 years), self-reported race/ethnicity was White (34.5%), Hispanic (27.7%), African American (17.9%), Asian (4.5%), Ashkenazi Jewish (0.6%), Other (3.5%), and missing (13.0%). Thirty-nine patients (2%) were identified as carriers of a PV in an autosomal dominant clinically actionable hereditary breast and ovarian cancer (HBOC)-related or Lynch syndrome gene, most frequently, BRCA2 (6/39; 15.4%), PALB2 (8/39; 20.5%), CHEK2 (10/39; 25.6%), and PMS2 (5/39; 12.8%). Of the 34 PVs with known race/ethnicity, 47% were detected among non-White patients. Overall, 354/1,943 (18.2%) of patients met NCCN guidelines for HBOC gene testing and only 15/39 (38.5%) patients with an autosomal dominant clinically actionable PV met guidelines.
Conclusion
This population health approach extended the reach of genetic cancer risk assessment in a diverse population and highlighted the limits of a guideline-based approach. This may help address inequity in access to risk-appropriate screening and cancer prevention.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A substantial proportion of cancers are associated with pathogenic variants (PV) in hereditary cancer genes [1, 2], including an estimated 10% of breast, 10% of colon, and 20% of ovarian cancers. Approximately, 1 in 300 to 500 people in the population will carry a PV in either BRCA1 or BRCA2 (BRCA; the genes most commonly associated with hereditary breast and ovarian cancer (HBOC)) [3,4,5,6], and 1 in 370 individuals will carry a pathogenic variant (PV) in one of the Lynch syndrome genes (the genes most commonly associated with hereditary colorectal cancer) [7]. Early identification of a PV in a cancer susceptibility gene provides individuals with the opportunity for enhanced surveillance and risk-reducing interventions which can significantly reduce the morbidity and mortality of these cancers [8,9,10,11,12,13]. Identification of carrier status also provides the opportunity for cascade testing in relatives [14,15,16].
Genetic testing for cancer susceptibility genes is currently offered predominantly to individuals who are considered high-risk for either BRCA or Lynch syndrome based on clinical and family history criteria. However, only around half of the carriers of PVs in BRCA and Lynch syndrome genes will meet clinical and family history criteria for genetic testing [5, 17, 18]. Additionally, current approaches to testing for hereditary cancer genes are associated with inequities in referral and access to testing [19, 20].
An alternative approach is population-based hereditary cancer genetic testing which may provide a more clinically effective strategy for early identification of high-risk individuals [5, 21]. To date, population-based testing has been primarily studied in the context of BRCA testing in the Ashkenazi Jewish population [22, 23], and data are limited in the general population, especially among under-represented racial and ethnic groups. The objective of this study was to determine the yield of hereditary cancer gene PVs among unselected diverse women attending breast imaging centers, as a potential strategy for more complete identification of high-risk individuals who could benefit from enhanced surveillance and/or risk reduction interventions.
Materials and methods
Study population
This retrospective cohort study included unselected female patients who were offered and underwent genetic testing at the time of breast imaging at three imaging centers (Memorial MRI and Diagnostics, Texas) from November 2020 through March 2022. All patients arriving at the imaging centers were given a written flier with an invitation to undergo genetic testing for a panel of hereditary cancer genes. Patients were also offered the option of an online genetic information session with a board-certified genetic counselor prior to testing. The lead clinician investigator (DM) served as the ordering clinician for the testing at all three centers. A limited number of providers using the imaging centers opted out of having their patients participate. Only patients (including those with a previous history of breast cancer) undergoing routine breast imaging, either by mammogram or ultrasound, were included. Patients undergoing imaging for newly diagnosed breast cancer were excluded.
Clinical, demographic, and family cancer history information were ascertained through test requisition forms and family cancer history questionnaires completed by the patients. Patient questionnaires included clinical questions needed for Tyrer–Cuzick breast cancer risk assessment. Race/ethnicity (ancestry) was self-reported. National Comprehensive Cancer Network (NCCN) guidelines for Genetic/Familial High-Risk Assessment: Breast, Ovarian, and Pancreatic Cancer [24] and Lynch Syndrome (LS) were reviewed to determine if genetic testing would have been guideline indicated for a particular patient [25]. For the patients unaffected with breast cancer who completed the clinical portion of their questionnaire for breast cancer risk assessment, a Tyrer–Cuzick breast cancer risk 5 years and lifetime risk score was calculated [26]. This score was only reported back to patients if their genetic testing (described below) was negative/uninformative for PVs associated with increased breast cancer risk. It was not available if enough information about personal/family history was not provided to compute a Tyrer–Cuzick score or the patient had a personal history of breast cancer.
If the patients’ clinical and/or family history met NCCN guidelines for hereditary cancer testing, they were given the option to request insurance coverage for testing or self-pay. Patients who did not meet guidelines were offered the option to self-pay and financial assistance options were available to eligible patients, based on their income and family size. Genetic information sessions performed by board-certified genetic counselors (Natera, Inc.) were available to patients on a pre- and post- test basis. All patients with a PV in a breast cancer-associated predisposition gene were offered in person risk counseling by the lead clinician investigator (DM).
This study was granted a waiver of consent process under 45 CFR 46.116(d), a waiver of the requirement for documentation of informed consent according to 45 CFR 46.117(c)(2), and a waiver from the HIPAA Authorization Requirement according to 45 CFR 46.164.512(i) (Salus IRB, ID# 21204—01A).
Hereditary cancer testing
Next-generation sequencing (NGS)-based hereditary cancer risk assessment was carried out utilizing a multiplex gene panel (40 or 53 genes) testing (Empower™, Natera, Inc. in collaboration with Baylor Genetics). The targeted regions of the genes associated with hereditary cancer syndromes are enriched using a capture-based method and sequenced by next-generation sequencing (NGS) using the Illumina platform. The variants detected in exons and within 20 bp of the exon/intron boundary are reported, unless otherwise specified. Read depth analysis is used to detect copy number variation (CNV) for genes. Positive sequencing results from certain genes or regions with highly homologous sequences in the genome are confirmed by gene-specific long-range PCR and Sanger sequencing. Multiplex ligation-dependent probe amplification (MLPA), PCR-based methods, and/or array comparative genomic hybridization (aCGH) are used to confirm copy number changes involving the genes in the test.
All patients underwent testing for at least 25 clinically actionable genes (Table 1). Genes were considered clinically actionable based on the presence of established NCCN and/or peer-reviewed consensus management recommendations for enhanced surveillance, or risk-reducing interventions, and family cascade testing if a PV was detected. Clinically actionable genes were categorized as “high risk” or “moderate risk” based on their reported relative risk for cancer, relative risk of > 4, and relative risk of 2–4, respectively (Table 1). Testing included both HBOC and Lynch syndrome (LS) genes (NCCN HBOC, NCCN Colorectal Cancer (CRC)). Variants were classified consistent with guidelines from the American College of Medical Genetics and Genomics and the Association for Molecular Pathology as previously described [26,27,28]. Only likely pathogenic and pathogenic variants (PVs) were considered in the analysis: benign and likely benign variants, and variants of unknown significance were not considered.
For the purposes of the current analysis, variants were not considered clinically actionable (i.e., had no potential impact on patient care) and were not included if they:
-
1)
Were in genes only associated with autosomal recessive disease association (e.g., monoallelic MUTYH carriers which may conflate estimates of pathogenic variant prevalence) or
-
2)
Were low penetrance variants in clinically actionable genes, such as CHEK2 c.470 T > C [p.Ile157Thr].
Analysis
The sociodemographic characteristics and personal and family history of cancer of the study population were explored. Patients’ characteristics were stratified based on the presence and type of PV. The prevalence of P/LP variants was calculated. For patients with a P/LP variant, the proportion who did and did not meet NCCN guidelines for genetic testing was evaluated. We also evaluated the proportion of patients with PV who would have qualified for additional screening based on the empiric risk model (i.e., Tyrer–Cuzick score > = 20%).
Results
Study population
A total of 1,943 women undergoing breast imaging elected to have hereditary cancer genetic testing during the study period (Table 2). Median age was 66 years (range 18–89 years). Self-reported race and ethnicity were Asian 5.0% (N = 85); Black 20% (N = 339); White 38% (N = 650); and Hispanic 32% (N = 534).
A personal history of cancer was documented for 7.5% (N = 146) (Fig. 1), a family history of cancer in 42.3% (N = 822), and no personal or family history of cancer in 50.2% (N = 975). A personal history of breast or ovarian-related cancers was recorded in 4% (N = 80), endometrial cancer in 0.8% (N = 15), and colorectal cancer in 0.5% (N = 10).
Overall, 18.2% (354/1943) of patients met current NCCN guidelines for hereditary breast and ovarian cancer (HBOC) gene testing, 3.7% (71/1943) met NCCN guidelines for Lynch syndrome genetic testing, and 1.0% (19/1943) met both HBOC and Lynch syndrome guidelines for testing (Table 3).
Prevalence of pathogenic variants
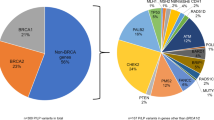
Among 1943 patients who received genetic testing, 39 (2%) were identified as carriers of a PV in an autosomal dominant clinically actionable HBOC-related or LS gene (Table 3). Of the 39 PVs identified, 84.6% (N = 33) were in HBOC-related genes which corresponds to a prevalence of 1.7% (33/1943) in the total cohort. The remaining 15.4% (N = 6) PVs were in LS genes which corresponds to a prevalence of 0.3% (6/1943) in the total cohort. The most common PVs were in CHEK2 (10/39; 25.6%); PALB2 (8/39; 20.5%); BRCA2 (6/39; 15.4%); and PMS2 (5/39; 12.8%) (Fig. 2). Of the 34 PVs where race/ethnicity were known, 47% were detected among non-White patients (Table 2). The PV prevalence (%) was distributed across the respective ancestral groups as follows: Black 3/339 (0.89%); White 16/650 (2.4%); Asian 3/85 (3.5%), and Hispanic 9/534 (1.7%).
Patients with a PV were over 50 years of age at the time of their testing in 82.1% (32/39) of cases. This is similar to the 1587/1943 (81.7%) of patients aged 51 or older who were tested in the total cohort and consistent with National screening guidelines for women at average risk [27] (Table 2).
Guideline eligibility for genetic testing or enhanced surveillance breast cancer
Only 38.5% (15/39) of PV carriers met either NCCN guidelines for HBOC testing or LS testing prior to genetic testing. Of these, 25.6% (10/39) met HBOC criteria for genetic testing with PVs in HBOC-related genes and 7.7% (3/39) had a PV in an LS gene, and 5.1% (2/39) met criteria for LS testing with PV in HBOC-related genes.
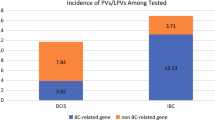
Notably, 5 out of 6 patients with a BRCA2 PV and 5 out of 8 patients with a PALB2 PV did not meet criteria for HBOC testing. The frequencies of clinically actionable PVs identified in high- or moderate-risk genes in patients who either met or did not meet NCCN guidelines for inclusion of the genes are shown in Fig. 2.
Data allowing calculation of the patient’s Tyrer–Cuzick score were available for 64.1% (25/39) of patients with a PV in an autosomal dominant clinically actionable HBOC (Table 3). Of these, only 8% (2/25) had a Tyrer–Cuzick score ≥ 20 which would have triggered health insurance coverage for increased surveillance for breast cancer, absent in the PV finding (Table 3).
Discussion
We found that 2% (39/1943) of women undergoing hereditary cancer gene testing as part of their clinical care at the time of breast imaging had PVs in hereditary cancer genes that had implications for cancer surveillance and clinical management. Importantly, the population undergoing testing was racially and ethnically diverse compared to previous studies [20, 28], predominantly over the age of 50 and without a personal history of cancer (92.5%). This was similar or more diverse compared to CDC population data on race and ethnicity for females aged 55 to 74 years who were resident in Texas in 2021 (4.8% Asian, 12.5% Black, 80.7% White, and 28.1% Hispanic) [29].
Among women who had a PV in either a HBOC or LS gene, only 38.5% met NCCN guidelines for testing of either of these conditions. Of those who had a PV in an HBOC gene and had data needed for a Tyrer–Cuzick score, only 8% had an estimated lifetime score of ≥ 20%. Therefore, while testing as part of clinical care at the time of breast imaging identified some women who were eligible for hereditary cancer gene testing based on NCCN guidelines, it predominantly identified women with PVs who neither met NCCN guidelines for testing nor the Tyrer–Cuzick threshold for increased surveillance for breast cancer [30]. Consequently, without this opportunity for genetic testing, women with PVs may not have accessed risk-appropriate enhanced surveillance.
Identification of individuals with PVs in HBOC and LS genes provides opportunities for cancer prevention and earlier detection [13, 31]. For people with PVs in HBOC genes, there are recommendations and options for earlier mammography, screening breast MRI, chemoprophylaxis, and risk-reducing surgeries, such as mastectomy and bilateral salpingo-oophorectomy [8, 10, 11, 13]. Knowledge of specific PVs can also impact decisions about surgery and adjuvant therapies [32,33,34]. The median age of the population in the current cohort was 66 years and previous research has suggested that HBOC gene testing in younger women would be the most cost-effective approach to testing [35]. Nonetheless, a study of the remaining lifetime risk for the subset of women who were older than 65 years in the population-based CARRIERS project [36] indicated that BRCA1, BRCA2, and PALB2 PVs were associated with enough breast cancer risk to warrant high-risk screening [37]. For people who have PVs in LS genes, there is also compelling evidence of the benefit of colonoscopy in reducing mortality from CRC [9, 38]. Additionally, in women with PVs in LS genes, the lifetime risk of endometrial cancer is similar to that of CRC, which can largely be prevented by hysterectomy [39]. For women with PVs who chose to share this information with family members, it can provide more accurate risk assessment and cascade testing [14,15,16]. The yield of PVs is typically higher (~ 10%) [13] when we use the guideline criteria to screen the eligibility for genetic testing. However, this study demonstrated a meaningful yield (~ 2%) of clinically actionable PVs among participants who did not meet any guideline. Amplifying this point, 5 out of 6 BRCA2 carriers identified by this unselected approach did not meet any guideline and there is ample evidence of reduction in cancer-specific and all-cause mortality by standard of care gene-specific clinical management [8, 40, 41]. Thus, the universal testing approach and increasingly cost-effective genetic testing are likely to have a high impact despite a modest yield.
Perhaps one of the most striking findings in this clinical cohort was the diversity of the population that accessed the testing. Later stages of cancer at the time of diagnosis and lower cancer survival rates are clearly documented in Black people in the USA [42]. There are data indicating that people of color or Hispanic ancestry are less likely to be referred for genetic counseling or testing and are less likely to take up genetic testing in the absence of a personal history of cancer [20, 42, 43]. As demonstrated in our study, increasing access and convenience of testing may help overcome barriers to more equitable testing [43, 44].
Limitations of this study include that women under 40 years of age without documented increased risk are unlikely to have routine mammography and therefore, may not be represented in this cohort. Thus, BRCA carriers may be under-represented. Nonetheless, several BRCA2 carriers, the majority of which did not meet any testing guidelines, were identified representing a critical opportunity for screening and prevention. Another limitation of the current cohort is that the number of patients who declined hereditary cancer testing or who may have had genetic testing prior to this study was unknown. Given the uncertainty about the total number of women who received the invitation to have genetic testing, the potential benefit of our strategy with regard to the yield of actionable PVs in the imaging center population could be over- or underestimated. Finally, though the racial and ethnic composition of the study participants were exceptionally diverse, we do not know if there were significant differences in the uptake among the respective groups [45]. Nonetheless, (as cited above) the population who underwent testing is diverse and generally representative of the ethnic makeup of the state of Texas. Further, we do not have qualitative or quantitative data on how the patients made their decision to participate and receive testing as it was beyond the scope of our study. However, we believe that this will be important for future research in the context of population health implementation. Finally, the presence of a PV or elevated empiric risk (e.g., > 20%) does not guarantee insurance coverage, and access has been problematic across different healthcare systems and among the underinsured. Nonetheless, we believe that this report provides additional evidence supporting access to risk-appropriate care. Without granular insurance data, we note that all of the individuals in the study at the least had access to the imaging centers.
Additionally, while genetic information sessions both before and after testing were available to all patients who underwent testing, formal pre-test genetic counseling was not required. Requiring pre-test genetic counseling may itself present a barrier to testing [46], so there may be a trade-off between the benchmark of full-genetic counseling and the use of abbreviated genetic information sessions to improve access to potentially life-saving information, especially in population health settings. Additional research is needed to establish what constitutes optimal pre- and post-test counseling and informed consent for patients receiving genetic testing in non-genetic/population health settings. However, in the meantime, the results of this and similar studies suggest that testing in a diverse imaging center population can extend the reach of genetic cancer risk assessment, has a clinically meaningful and actionable yield of cancer-associated PVs, and can help address inequity in access to testing and risk-appropriate screening and prevention.
Data availability
This is descriptive summary data on genetic testing outcome. The specific variant level data are contributed to ClinVar by Baylor Genetics, Houston, Texas.
References
Ceyhan-Birsoy O, Jayakumaran G, Kemel Y, Misyura M, Aypar U, Jairam S et al (2022) Diagnostic yield and clinical relevance of expanded genetic testing for cancer patients. Genome Med 14:92. https://doi.org/10.1186/s13073-022-01101-2
Pearlman R, Frankel WL, Swanson B, Zhao W, Yilmaz A, Miller K et al (2017) Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol 3:464–471. https://doi.org/10.1001/jamaoncol.2016.5194
Anglian Breast Cancer Study Group (2000) Prevalence and penetrance of BRCA1 and BRCA2 mutations in a population-based series of breast cancer cases. Anglian breast cancer study group. Br J Cancer 83:1301–1308. https://doi.org/10.1054/bjoc.2000.1407
Antoniou AC, Pharoah PD, McMullan G, Day NE, Stratton MR, Peto J et al (2002) A comprehensive model for familial breast cancer incorporating BRCA1, BRCA2 and other genes. Br J Cancer 86:76–83. https://doi.org/10.1038/sj.bjc.6600008
Manchanda R, Patel S, Gordeev VS, Antoniou AC, Smith S, Lee A et al (2018) (2018) Cost-effectiveness of population-based BRCA1, BRCA2, RAD51C, RAD51D, BRIP1, PALB2 mutation testing in unselected general population women. J Natl Cancer Inst 110:714–725. https://doi.org/10.1093/jnci/djx265
Peto J, Collins N, Barfoot R, Seal S, Warren W, Rahman N et al (1999) Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J Natl Cancer Inst 91:943–949. https://doi.org/10.1093/jnci/91.11.943
Abu-Ghazaleh N, Kaushik V, Gorelik A, Jenkins M, Macrae F (2022) Worldwide prevalence of Lynch syndrome in patients with colorectal cancer: systematic review and meta-analysis. Genet Med 24:971–985. https://doi.org/10.1016/j.gim.2022.01.014
Domchek SM, Friebel TM, Singer CF, Evans DG, Lynch HT, Isaacs C et al (2010) Association of risk-reducing surgery in BRCA1 or BRCA2 mutation carriers with cancer risk and mortality. JAMA 304:967–975. https://doi.org/10.1001/jama.2010.1237
Jarvinen HJ, Renkonen-Sinisalo L, Aktan-Collan K, Peltomaki P, Aaltonen LA, Mecklin JP (2009) 10 years after mutation testing for Lynch syndrome: cancer incidence and outcome in mutation-positive and mutation-negative family members. J Clin Oncol 27:4793–4797. https://doi.org/10.1200/JCO.2009.23.7784
Kauff ND, Domchek SM, Friebel TM, Robson ME, Lee J, Garber JE et al (2008) Risk-reducing salpingo-oophorectomy for the prevention of BRCA1- and BRCA2-associated breast and gynecologic cancer: a multicenter, prospective study. J Clin Oncol 26:1331–1337. https://doi.org/10.1200/JCO.2007.13.9626
Kotsopoulos J, Huzarski T, Gronwald J, Singer CF, Moller P, Lynch HT et al (2017) Bilateral oophorectomy and breast cancer risk in BRCA1 and BRCA2 mutation carriers. J Natl Cancer Inst. https://doi.org/10.1093/jnci/djw177
Warner E, Hill K, Causer P, Plewes D, Jong R, Yaffe M et al (2011) Prospective study of breast cancer incidence in women with a BRCA1 or BRCA2 mutation under surveillance with and without magnetic resonance imaging. J Clin Oncol 29:1664–1669. https://doi.org/10.1200/JCO.2009.27.0835
Weitzel JN, Blazer KR, MacDonald DJ, Culver JO, Offit K (2011) Genetics, genomics, and cancer risk assessment: state of the art and future directions in the era of personalized medicine. CA Cancer J Clin 61:327–359. https://doi.org/10.3322/caac.20128
Bednar EM, Sun CC, McCurdy S, Vernon SW (2020) Assessing relatives’ readiness for hereditary cancer cascade genetic testing. Genet Med 22:719–726. https://doi.org/10.1038/s41436-019-0735-3
Caswell-Jin JL, Zimmer AD, Stedden W, Kingham KE, Zhou AY, Kurian AW (2019) Cascade genetic testing of relatives for hereditary cancer risk: results of an online initiative. J Natl Cancer Inst 111:95–98. https://doi.org/10.1093/jnci/djy147
Lee J, Ham JY, Park HY, Jung JH, Kim WW, Kang B et al (2022) Feasibility of targeted cascade genetic testing in the family members of BRCA1/2 gene pathogenic variant/likely pathogenic variant carriers. Sci Rep 12:1842. https://doi.org/10.1038/s41598-022-05931-3
LaDuca H, Polley EC, Yussuf A, Hoang L, Gutierrez S, Hart SN et al (2020) A clinical guide to hereditary cancer panel testing: evaluation of gene-specific cancer associations and sensitivity of genetic testing criteria in a cohort of 165,000 high-risk patients. Genet Med 22:407–415. https://doi.org/10.1038/s41436-019-0633-8
Wood ME, Kadlubek P, Pham TH, Wollins DS, Lu KH, Weitzel JN et al (2014) Quality of cancer family history and referral for genetic counseling and testing among oncology practices: a pilot test of quality measures as part of the American society of clinical oncology quality oncology practice initiative. J Clin Oncol 32:824–829. https://doi.org/10.1200/JCO.2013.51.4661
Chapman-Davis E, Zhou ZN, Fields JC, Frey MK, Jordan B, Sapra KJ et al (2021) Racial and ethnic disparities in genetic testing at a hereditary breast and ovarian cancer center. J Gen Intern Med 36:35–42. https://doi.org/10.1007/s11606-020-06064-x
Khan A, Rogers CR, Kennedy CD, Lopez A, Jeter J (2022) Genetic evaluation for hereditary cancer syndromes among African Americans: a critical review. Oncologist 27:285–291. https://doi.org/10.1093/oncolo/oyab082
Foulkes WD, Knoppers BM, Turnbull C (2016) Population genetic testing for cancer susceptibility: founder mutations to genomes. Nat Rev Clin Oncol 13:41–54. https://doi.org/10.1038/nrclinonc.2015.173
Manchanda R, Loggenberg K, Sanderson S, Burnell M, Wardle J, Gessler S et al (2015) Population testing for cancer predisposing BRCA1/BRCA2 mutations in the Ashkenazi-Jewish community: a randomized controlled trial. J Natl Cancer Inst 107:379. https://doi.org/10.1093/jnci/dju379
Metcalfe KA, Poll A, Royer R, Llacuachaqui M, Tulman A, Sun P et al (2010) Screening for founder mutations in BRCA1 and BRCA2 in unselected Jewish women. J Clin Oncol 28:387–391. https://doi.org/10.1200/JCO.2009.25.0712
Network NCC (2023) Genetic/familial high-risk assessment: breast, ovarian, and pancreatic. https://www.nccn.org/professionals/physician_gls/pdf/genetics_bop.pdf
NCCN.org. (2023). https://www.nccn.org/professionals/physician_gls/pdf/genetics_colon.pdf.
Amir E, Evans DG, Shenton A, Lalloo F, Moran A, Boggis C et al (2003) Evaluation of breast cancer risk assessment packages in the family history evaluation and screening programme. J Med Genet 40:807–814. https://doi.org/10.1136/jmg.40.11.807
Siu AL, Force USPST (2016) Screening for Breast Cancer: U.S. preventive services task force recommendation statement. Ann Intern Med 164:279–296. https://doi.org/10.7326/M15-2886
Kurian AW, Ward KC, Howlader N et al (2019) Genetic testing and results in a population-based cohort of breast cancer patients and ovarian cancer patients. J Clin Oncol 37(15):1305–1315. https://doi.org/10.1200/JCO.18.01854
Prevention CfDCa: CDC WONDER. https://wonder.cdc.gov/
Tyrer J, Duffy SW, Cuzick J (2004) A breast cancer prediction model incorporating familial and personal risk factors. Stat Med 23:1111–1130. https://doi.org/10.1002/sim.1668
Evans O, Gaba F, Manchanda R (2020) Population-based genetic testing for women’s cancer prevention. Best Pract Res Clin Obstet Gynaecol 65:139–153. https://doi.org/10.1016/j.bpobgyn.2020.02.007
Shubeck S, Sevilimedu V, Berger E, Robson M, Heerdt AS, Pilewskie ML (2022) Comparison of outcomes between BRCA pathogenic variant carriers undergoing breast-conserving surgery versus mastectomy. Ann Surg Oncol. 29:4706–4713. https://doi.org/10.1245/s10434-022-11756-1
Tung NM, Boughey JC, Pierce LJ, Robson ME, Bedrosian I, Dietz JR et al (2020) Management of hereditary breast cancer: American society of clinical oncology, American society for radiation oncology, and society of surgical oncology guideline. J Clin Oncol 38:2080–2106. https://doi.org/10.1200/JCO.20.00299
Tutt ANJ, Garber JE, Kaufman B, Viale G, Fumagalli D, Rastogi P et al (2021) Adjuvant olaparib for patients with BRCA1- or BRCA2-mutated breast cancer. N Engl J Med 384:2394–2405. https://doi.org/10.1056/NEJMoa2105215
Evans DG, van Veen EM, Byers HJ, Evans SJ, Burghel GJ, Woodward ER et al (2022) High likelihood of actionable pathogenic variant detection in breast cancer genes in women with very early onset breast cancer. J Med Genet 59:115–121. https://doi.org/10.1136/jmedgenet-2020-107347
Hu C, Hart SN, Gnanaolivu R, Huang H, Lee KY, Na J et al (2021) A Population-based study of genes previously implicated in breast cancer. N Engl J Med 384:440–451. https://doi.org/10.1056/NEJMoa2005936
Boddicker NJ, Hu C, Weitzel JN, Kraft P, Nathanson KL, Goldgar DE et al (2021) Risk of late-onset breast cancer in genetically predisposed women. J Clin Oncol 39:3430–3440. https://doi.org/10.1200/JCO.21.00531
Kastrinos F, Ingram MA, Silver ER, Oh A, Laszkowska M, Rustgi AK et al (2021) Gene-specific variation in colorectal cancer surveillance strategies for Lynch syndrome. Gastroenterology 161(453–462):e15. https://doi.org/10.1053/j.gastro.2021.04.010
Schmeler KM, Lynch HT, Chen LM, Munsell MF, Soliman PT, Clark MB et al (2006) Prophylactic surgery to reduce the risk of gynecologic cancers in the Lynch syndrome. N Engl J Med 354:261–269. https://doi.org/10.1056/NEJMoa052627
Giannakeas V, Narod SA (2018) The expected benefit of preventive mastectomy on breast cancer incidence and mortality in BRCA mutation carriers, by age at mastectomy. Breast Cancer Res Treat 167:263–267. https://doi.org/10.1007/s10549-017-4476-1
Evans DG, Harkness EF, Howell A, Wilson M, Hurley E, Holmen MM et al (2016) Intensive breast screening in BRCA2 mutation carriers is associated with reduced breast cancer specific and all cause mortality. Hered Cancer Clin Pract 14:8. https://doi.org/10.1186/s13053-016-0048-3
Reid S, Cadiz S, Pal T (2020) Disparities in genetic testing and care among black women with hereditary breast cancer. Curr Breast Cancer Rep 12:125–131. https://doi.org/10.1007/s12609-020-00364-1
McGuinness JE, Trivedi MS, Silverman T, Marte A, Mata J, Kukafka R et al (2019) Uptake of genetic testing for germline BRCA1/2 pathogenic variants in a predominantly Hispanic population. Cancer Genet 235–236:72–76. https://doi.org/10.1016/j.cancergen.2019.04.063
Rubinsak LA, Kleinman A, Quillin J, Gordon SW, Sullivan SA, Sutton AL et al (2019) Awareness and acceptability of population-based screening for pathogenic BRCA variants: do race and ethnicity matter? Gynecol Oncol 154:383–387. https://doi.org/10.1016/j.ygyno.2019.06.009
Coughlin SE, Heald B, Clark DF, Nielsen SM, Hatchell KE, Esplin ED et al (2022) Multigene panel testing yields high rates of clinically actionable variants among patients with colorectal cancer. JCO Precis Oncol 6:e2200517. https://doi.org/10.1200/PO.22.00517
Beitsch PD, Whitworth PW, Hughes K, Patel R, Rosen B, Compagnoni G et al (2019) Underdiagnosis of hereditary breast cancer: are genetic testing guidelines a tool or an obstacle? J Clin Oncol 37:453–460. https://doi.org/10.1200/JCO.18.01631
Acknowledgements
We would like to acknowledge Sofia Hurtado, BS and Mayra Rodas, BS from Natera, Inc. for data acquisition and Urmi Sengupta, PhD, from Natera, Inc., for assistance with the development of the manuscript.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
LW, VS, and JW have contributed to the study concept and design. Data acquisition, analysis, and interpretation were performed by LW, DM, MM, KH, MY, and NK. LW, DM, VS, AR, MM, KH, YS, MY, NK, and JW were involved in the administrative process. LW, VS, and JW supervised the study. Original drafting of the manuscript was performed by LW, VS, and MY. Critical revision and editing of the manuscript were performed by LW, DM, VS, AR, MM, KH, YS, MY, and JW. All authors read and approved the final version of the manuscript for submission.
Corresponding author
Ethics declarations
Competing interests
VS, MM, KH, YS, MY, RG, and WS are employees of Natera, Inc. with stocks or options to own stocks. LW, DM, and NK declare no conflict of interest.
Ethical approval
This study was granted a waiver of consent process under 45 CFR 46.116(d), a waiver of the requirement for documentation of informed consent according to 45 CFR 46.117(c)(2), and a waiver from the HIPAA Authorization Requirement according to 45 CFR 46.164.512(i) (Salus IRB, ID# 21204—01A).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Westbrook, L., Miltenburg, D., Souter, V. et al. Hereditary cancer testing in a diverse sample across three breast imaging centers. Breast Cancer Res Treat 203, 365–372 (2024). https://doi.org/10.1007/s10549-023-07137-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-023-07137-1