Abstract
Background
The PAM50 assay is used routinely in clinical practice to determine breast cancer prognosis and management; however, research assessing how technical variation and intratumoral heterogeneity contribute to misclassification and reproducibility of these tests is limited.
Methods
We evaluated the impact of intratumoral heterogeneity on the reproducibility of results for the PAM50 assay by testing RNA extracted from formalin-fixed paraffin embedded breast cancer blocks sampled at distinct spatial locations. Samples were classified according to intrinsic subtype (Luminal A, Luminal B, HER2-enriched, Basal-like, or Normal-like) and risk of recurrence with proliferation score (ROR-P, high, medium, or low). Intratumoral heterogeneity and technical reproducibility (replicate assays on the same RNA) were assessed as percent categorical agreement between paired intratumoral and replicate samples. Euclidean distances between samples, calculated across the PAM50 genes and the ROR-P score, were compared for concordant vs. discordant samples.
Results
Technical replicates (N = 144) achieved 93% agreement for ROR-P group and 90% agreement on PAM50 subtype. For spatially distinct biological replicates (N = 40 intratumoral replicates), agreement was lower (81% for ROR-P and 76% for PAM50 subtype). The Euclidean distances between discordant technical replicates were bimodal, with discordant samples showing higher Euclidian distance and biologic heterogeneity.
Conclusion
The PAM50 assay achieved very high technical reproducibility for breast cancer subtyping and ROR-P, but intratumoral heterogeneity is revealed by the assay in a small proportion of cases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is a heterogenous disease that includes subtypes with distinct biology, clinical characteristics, and prognosis. Thus, multigene assays for breast cancer subtype classification are used in research and treatment. Generally, these assays are run on a single sample from a representative tissue block. When assessing the gene expression patterns of a sampled specimen for diagnostic and treatment purposes, the aim is to sample a part of the tumor that best reflects tumor biology and predicts prognosis and treatment response. However, studies have demonstrated that spatial heterogeneity of biomarker expression, as well as chromosomal and genomic alterations, are common features of breast tumors. Morphologically distinct tumor regions may have distinct genetic aberrations [1, 2].
Considering standard clinical markers, heterogeneity of ER expression has been shown to be relatively low [3, 4], while heterogeneity of expression of HER2, PR, and Ki67 is known to be high [3,4,5,6,7]. For example, up to 15% of breast tumors show discordance of HER2 status by FISH across randomly sampled sections, with heterogeneity being most frequent (up to 27%) in tumors with an equivocal (2 +) HER2 score [8]. Further, a recent study found Ki67 heterogeneity present in 18% of sampled breast tumors, which included a high frequency of two types of heterogeneity: as a gradient of increasing staining towards the tumor edge (likely representing a common technical artifact) and as hot spots [9]. Beyond the clinical uncertainty associated with biomarker discordance, it is believed that intratumoral heterogeneity may have prognostic relevance, with high heterogeneity breast tumors associated with worse survival [10, 11].
In light of this spatial heterogeneity, multigene assays have been viewed as a more stable solution to breast tumor subtyping. In particular, the PAM50 assay was designed to identify five intrinsic subtypes (Luminal A, Luminal B, HER2-enriched, Basal-like, and Normal-like), which have both etiologic and prognostic significance. This assay and others (e.g., Oncotype DX and Mammaprint) are recommended clinically to guide treatment decisions [12,13,14,15], highlighting the importance of understanding the potential for outcome misclassification in studies of breast cancer subtype. Outcome misclassification could arise from technical or biological heterogeneity. To estimate the upper bound on the frequency of technical and biological variability in PAM50 classification, we selected morphologically heterogenous and/or spatially separate breast tumor regions and performed technical replication studies across formalin-fixed paraffin-embedded samples of breast tissue.
Methods
Study population
The UNC Normal Breast Study (NBS), described previously [16], was a hospital-based study conducted at UNC Hospitals in Chapel Hill, North Carolina from 2009 to 2013. The study recruited women undergoing breast surgery and samples of grossly normal-appearing breast tissue and tumor tissue were extracted, then snap frozen or paraffin embedded. The present study is of tumor tissue only.
Tumor samples
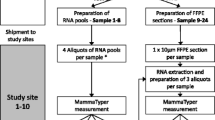
Tumors from 49 patients were assayed repeatedly (2–3 times) at different spatial locations (central versus peripheral tumor), specifically targeting regions that had distinct histological characteristics (e.g., tumor cellularity, tumor grade, admixture of DCIS or benign epithelium, and inflammatory response). From the formalin-fixed paraffin-embedded (FFPE) tumor block, a top slide was cut and stained with hematoxylin & eosin. Using this slide as a guide, up to three 1-mm cores were cut from histologically distinct regions of each tumor block. RNA was isolated from each core using the RNeasy FFPE kit (cat# 73504) from Qiagen and run on a Nanostring codeset containing 417 probes, including 11 housekeeping genes. There were 95 samples analyzed. We excluded 5 samples in technical quality control due to high missingness across genes, which left 90 tumor samples from 48 patients. Samples with no pair were excluded (n = 11), resulting in a final analytic sample of 85 tumor samples across 40 patients (35 patients with n = 2 samples and 5 patients with n = 3 samples). A second pathologist scored the tumor specimens for several histologic characteristics. Mitotic activity was assessed by recording the maximum number of mitoses in a single 20 × field, with heterogeneous cores defined as those that differ by two or more mitotic figures per field. Additionally, presence of immune infiltration (present/absent), distance from the biopsy site (adjacent/away) and location of the core on the tumor (peripheral/central) were assessed for each core. The biopsy site, which is the focus of inflammation and healing induced by a previous core needle biopsy, was present/identifiable for half of the tumors in the analytic sample. Tumors were categorized as concordant or discordant for each histologic characteristic.
To assess technical reproducibility, we generated 1–2 technical replicates of all 90 tumor samples, wherein each RNA sample was run on the assay at least twice. After sample cleaning, six samples were left without a pair and were subsequently excluded, leaving a total of 207 replicate tumor samples across 87 patients (54 patients with n = 2 samples and 33 patients with n = 3 samples). When estimating technical reproducibility, we also included technical replicates of tumor samples from a second study—Phase 3 of the Carolina Breast Cancer Study (CBCS3)—which is a population-based study that enrolled breast cancer cases occurring in North Carolina during the same study interval (2008–2013). Tumors from 57 patients in CBCS3 were assayed repeatedly at different spatial regions and 1–2 technical replicates were generated for 57 tumors. Throughout this manuscript when we refer to “technical reproducibility” or “technical replicates” we are referring to the repeated samples of the same RNA. When we refer to “intratumoral reproducibility” or “intratumoral samples” we are referring to the separate cores from a patient’s tumor block.
Gene expression was quantified using a research version of the PAM50 assay on the NanoString platform. The expression data were quality checked and cleaned using our previously described pipeline, then normalized as previously described using the remove unwanted variation (RUV) method [17,18,19]. Samples were classified for intrinsic subtype (Luminal A, Luminal B, HER2-enriched, Basal-like, or Normal-like) and for risk of recurrence (ROR-P) score (continuous) and group (low, medium, or high).
Statistical analysis
Reproducibility was assessed by calculating percent agreement of intrinsic subtype and ROR-P group across paired intratumoral and replicate samples. Additionally, we calculated the Euclidean distances of the expression levels of the 50 genes (stratified by discordance/concordance of intrinsic subtype) and of the ROR-P score (stratified by discordance/concordance of ROR-P group) within each set of intratumoral and replicate samples. Euclidean distance is a statistical metric commonly used to quantify the similarity between vectors. Wilcoxon rank sum tests were used to test whether the median of the distances differed between concordant and discordant samples.
Results
To assess technical reproducibility of the PAM50 assay, we performed Nanostring analysis in duplicate on the same RNA extract from 144 FFPE tumor samples from two different studies (87 NBS samples and 57 CBCS samples). Technical agreement was very high, but varied according to categorical predictor. For the three-class predictor—ROR-P (high, medium, low)—133 of 144 pairs were concordant for ROR-P group (93% agreement; Figs. 1, 2). All the discordant pairs changed between adjacent ordinal categories (n = 10 from low to medium and n = 1 from medium to high; Fig. 1). For PAM50 subtype, classification includes a larger number of categories and therefore we expected lower concordant rates. Of 144 pairs, 127 were concordant for PAM50 subtype (90% agreement; Figs. 1, 2). About half of the discordant cases (9 of 17) were Luminal A in one replicate and Luminal B in the second replicate (Fig. 1). Only one sample changed between a Luminal subtype and Basal-like. These technical reproducibility findings provide a comparison point for the intratumoral comparisons.
Concordance and percent agreement of PAM50 subtype and ROR-P group between technical replicates (A) and between intratumoral samples (B). Grey cells indicate unavailable samples. CBCS = Carolina Breast Cancer Study, NBS = Normal Breast Study, PAM50 = Prediction Analysis of Microarray 50, ROR-P = relapse score based on subtype and proliferation
To assess intratumoral (i.e., biological) reproducibility of the PAM50 assay, we performed a ‘challenge’ study that specifically oversampled histologically distinct regions within each tumor. Of the 40 tumors analyzed, 33 were concordant for ROR-P group (81% agreement) and 29 were concordant for PAM50 subtype (76% agreement; Figs. 1, 2). The pathologist-selected histologically distinct regions were independently scored for a variety of parameters by a second pathologist and 83% showed discordance in at least one histological characteristic. Compared to subtype-concordant samples, paired samples that were discordant for PAM50 subtype had more heterogeneity with respect to the distance of each core from the biopsy site and presence of stromal TILs (Table 1). Heterogeneity of mitotic activity (as recorded by the pathologist) was observed for 57% of samples discordant and 39% of samples concordant for ROR-P group (n = 4/7 and n = 12/31, respectively; Table 1).
Multigene classifiers can vary in their strength of association with the single sample predictor vector, suggesting that not all samples have high concordance with the standard for that class. Confidence scores and other diagnostics that reflect the strength of the classifier correlation can be a useful indicator of sample quality and may also impact both technical and biological reproducibility. To assess whether the strength of the classification was a factor influencing reproducibility, we evaluated technical and biological replicates for confidence scores using the method of Parker et al. [20] Among technical replicates, samples discordant for PAM50 subtype were less likely to have high confidence subtype calls compared to concordant samples (76% compared to 93%; Table 1). Among intratumoral samples, however, concordant and discordant tumors had similar rates of high confidence calls (93% and 82%, respectively; Table 1). For both the technical replicates and intratumoral samples, the majority of discordant subtype calls were Luminal A with Luminal B, or Luminal A with Normal-like (Figs. 1, 2).
To identify possible predictors of technical or biological discordance, we assessed whether tumor characteristics were associated with discordance. For both the technical replicates and intratumoral samples, the distributions of tumor grade and size were not associated with discordance of PAM50 subtype call. However, we observed non-statistically significant differences in tumor size between ROR-P-concordant and discordant tumors, with discordant samples generally arising from larger tumors (Table 1). Among technical replicates, 82% of discordant samples came from tumors larger than 2 cm compared to 55% of concordant samples. Among intratumoral samples, 86% of discordant samples came from tumors larger than 2 cm compared to 52% of concordant samples.
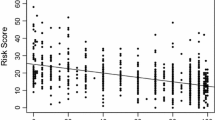
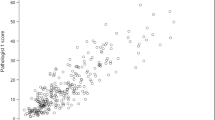
While our emphasis was to assess reproducibility of categorical classification, we also evaluated the differences in multigene continuous scores. Specifically, we calculated Euclidean distance between pairs of two or three replicates from each tumor. Figure 3 illustrates the magnitude of the difference in the continuous ROR-P score and in the gene expression levels of the genes used by the PAM50 classifier between paired samples. Among the technical replicates (Fig. 3A), the distances between ROR-P scores and between the gene expression levels were higher for discordant compared to concordant cores, but the distributions were similar and largely overlapping. In contrast, among intratumoral samples (Fig. 3B), the distances between cores were higher for discordant samples—particularly for samples discordant for ROR-P group (p = 0.002). The distributions among samples discordant for ROR-P group or PAM50 subtype appear bimodal, with the first peak largely overlapping with the distributions for concordant samples and the second peak corresponding to samples with greater distances among ROR-P scores or among gene expression levels, suggesting that discordant biological replicates had differences beyond expectation for technical variation.
Euclidean distances between the ROR-P scores and expression levels of the 50 genes that comprise the PAM50 classifier for technical replicates (A) and intratumoral samples (B). p-values correspond to the test statistic from a Wilcoxon rank sum test comparing the median of the distances in each group. PAM50 = Prediction Analysis of Microarray 50, ROR-P = relapse score based on subtype and proliferation
Discussion
This study examined the extent to which clinically relevant multigene scores for breast cancer are subject to misclassification. Despite observing heterogeneity across replicate samples (from both biological and technical sources), there was moderate-to-strong concordance of PAM50 subtype call and ROR-P groups across technical replicates and spatially distant biological replicates. A slightly higher rate of discordance among samples selected for histologic heterogeneity suggests meaningful variation across tumors, particularly in tumors that are larger size, have higher mitotic variability, and more infiltration of TILs. This biological heterogeneity detected within blocks may be even more pronounced when comparing across separate blocks, suggesting that especially for Luminal A and B tumors, there is potential for outcome misclassification in a proportion (up to 20%) of samples.
We expected to observe some degree of discordance across a tumor, as prior studies have revealed intratumoral genomic heterogeneity of breast tumors. Raychaudhuri et al. assessed heterogeneity in expression levels of several pro-metastatic miRNAs across multiple regions from each tumor and observed considerable intratumoral heterogeneity (coefficient of variation = 40%) [21]. Similarly, substantial heterogeneity of putative driver genes has been observed across spatially separated breast tumor samples [22]. Interestingly, a study of the impact of intratumoral heterogeneity on the performance of microarray-based assays in breast cancer found the amount of intratumoral variance directly correlated with expression level, with lowly expressed genes having higher heterogeneity [23]. A previous paper by Lopez-Knowles et al. found that poor reproducibility could be linked with low confidence [24]. We observed this pattern for technical replicates; however, we could not explain intratumoral reproducibility by confidence scores alone.
Despite previous reports suggesting high genomic intratumoral instability, heterogeneity of multigene scores has been found to be relatively minimal. The robustness of the PAM50 classifier and other multigene classifiers has been attributed to improved filtration of background noise and aggregation across multiple signals, presumably resulting in more accurate classification. For example, the intrinsic gene subset that gave rise to the PAM50 classifier was devised to maximize heterogeneity of expression between tumors and minimize heterogeneity of expression within tumors [25]. High agreement of PAM50 intrinsic subtype classification has been reported, with 11/12 and 4/6 tumors with concordant samples (based on RNA-sequencing and microarray-based gene expression profiling, respectively) [22]. Several studies have also found minimal intratumoral heterogeneity of the recurrence score from the Oncotype DX assay, with the continuous score having high concordance between core biopsies and resections (n = 49 patients, Pearson’s r = 0.86) [26] and between multiple tumor blocks (n = 19 patients, r = 0.94) [27]. When categorizing the Oncotype DX recurrence score into low, intermediate, and high categories, three of eight patients had one core in a higher risk category than the other three cores [28]. Low intratumoral heterogeneity has also been reported for the EndoPredict [29] and MammaPrint [30, 31] risk of recurrence scores. Results from these studies of intratumoral heterogeneity of multigene scores are comparable to the present study, where there was 81% and 76% intratumoral agreement of ROR-P group and PAM50 subtype, respectively.
While rare, some discordance in ROR-P group between intratumoral samples was observed (19% disagreement), with large tumor size (> 2 cm) and heterogeneity of mitotic activity between cores observed as potential predictors of ROR-P group discordance. This raises the question of whether tissue from larger tumors should be reviewed for histologic features prior to taking samples for RNA extraction and testing. There is no clear indication in the literature that there is more variation in larger tumors, but it is a reasonable expectation that might be true in some cases. For example, different blocks might yield more heterogeneity of ER expression [32]. This also may depend on patterns for ordering the assays. One paper suggests that multigene assays are ordered more frequently on large tumors [33], but according to Medicare data they seem to be ordered for small tumors if they appear to be high risk [34]. The current evidence may not justify targeted selection of sampling regions because the clinical significance of the potential heterogeneity is unknown. Nevertheless, the use of this technology seems to be growing and future work might consider whether large tumors that produce multiple blocks require multiple assays.
This study has several strengths. First, we conducted a detailed histopathology review of samples to try to maximize the opportunity to detect heterogeneity. Additionally, the estimate of baseline technical variation provides a metric with which to compare the observed discordance to the assay-based discordance. It is also important that although the probes used may differ slightly from those in the commercial version of the assay, we have used the same genes and algorithm as the Prosigna assay. With proper data normalization, this has been shown to be robust across datasets [20]. Any discrepancies relative to actual clinical subtype is expected to be random.
This study also has several limitations. This analysis was designed to challenge intra-block variation (reproducibility within a block) and does not inform on the level of variation across lesions from different tumor blocks. We did not assess whether the level of reproducibility observed in the present study was significantly different than some established threshold. Rather, our goal was to describe the reproducibility in a diverse cohort where we had multiple measures. Additionally, the modest sample size does not allow for stratification by other tumor characteristics (e.g., grade or subtype). Lastly, the goal of this study was to identify the maximum amount of misclassification that may occur, yet a cohort of breast cancer cases with more aggressive tumors may observe higher rates of intratumoral heterogeneity. However, the distribution of tumor characteristics in the present study is similar to those in other population-based studies (e.g., CBCS).
Our findings indicate that intratumoral heterogeneity is not likely to be a major impediment to the interpretation of multigene scores for breast tumors. The PAM50 classifier is relatively robust to repeated sampling and, in most applications, the ROR-P score seems highly reproducible. Achieving the level of discordance observed in this study required us to selectively sample heterogenous appearing tumor regions, therefore representing an upper bound on the level of discordance expected from a single tumor block. While intratumoral heterogeneity is not expected to be a major cause of misclassification when applying multigene signatures, heterogeneity does exist within breast tumors and sampling may cause some misclassification of tumors, with relevance for breast cancer treatment decisions.
Data availability
The datasets generated during and/or analyzed during the current study are not publicly available due to some human subjects restrictions, but may be available from the corresponding author on reasonable request.
References
Geyer FC, Weigelt B, Natrajan R et al (2010) Molecular analysis reveals a genetic basis for the phenotypic diversity of metaplastic breast carcinomas. J Pathol 220(5):562–573. https://doi.org/10.1002/path.2675
Patani N, Barbashina V, Lambros MB et al (2011) Direct evidence for concurrent morphological and genetic heterogeneity in an invasive ductal carcinoma of triple-negative phenotype. J Clin Pathol 64(9):822–828. https://doi.org/10.1136/jclinpath-2011-200135
Allott EH, Geradts J, Sun X et al (2016) Intratumoral heterogeneity as a source of discordance in breast cancer biomarker classification. Breast Cancer Res 18(1):68. https://doi.org/10.1186/s13058-016-0725-1
Nadji M, Gomez-Fernandez C, Ganjei-Azar P, Morales AR (2005) Immunohistochemistry of estrogen and progesterone receptors reconsidered: experience with 5,993 breast cancers. Am J Clin Pathol 123(1):21–27. https://doi.org/10.1309/4wv79n2ghj3x1841
Nassar A, Radhakrishnan A, Cabrero IA, Cotsonis GA, Cohen C (2010) Intratumoral heterogeneity of immunohistochemical marker expression in breast carcinoma: a tissue microarray-based study. Appl Immunohistochem Mol Morphol 18(5):433–441. https://doi.org/10.1097/PAI.0b013e3181dddb20
Pertschuk LP, Axiotis CA, Feldman JG, Kim YD, Karavattayhayyil SJ, Braithwaite L (1999) Marked intratumoral heterogeneity of the proto-oncogene Her-2/neu determined by three different detection systems. Breast J 5(6):369–374. https://doi.org/10.1046/j.1524-4741.1999.97088.x
Seol H, Lee HJ, Choi Y et al (2012) Intratumoral heterogeneity of HER2 gene amplification in breast cancer: its clinicopathological significance. Modern Pathol 25(7):938–948. https://doi.org/10.1038/modpathol.2012.36
Ohlschlegel C, Zahel K, Kradolfer D, Hell M, Jochum W (2011) HER2 genetic heterogeneity in breast carcinoma. J Clin Pathol 64(12):1112–1116. https://doi.org/10.1136/jclinpath-2011-200265
Boros M, Moncea D, Moldovan C, Podoleanu C, Georgescu R, Stolnicu S (2017) Intratumoral heterogeneity for Ki-67 Index in invasive breast carcinoma: a Study on 131 consecutive cases. Appl Immunohistochem Mol Morphol 25(5):338–340. https://doi.org/10.1097/pai.0000000000000315
McDonald KA, Kawaguchi T, Qi Q et al (2019) Tumor heterogeneity correlates with less immune response and worse survival in breast cancer patients. Ann Surg Oncol 26(7):2191–2199. https://doi.org/10.1245/s10434-019-07338-3
Lindström LS, Yau C, Czene K et al (2018) Intratumor heterogeneity of the estrogen receptor and the long-term risk of fatal breast cancer. J Natl Cancer Inst 110(7):726–733. https://doi.org/10.1093/jnci/djx270
Harris LN, Ismaila N, McShane LM et al (2016) Use of biomarkers to guide decisions on adjuvant systemic therapy for women with early-stage invasive breast cancer: American Society of clinical oncology clinical practice guideline. J Clin Oncol 34(10):1134–1150. https://doi.org/10.1200/jco.2015.65.2289
Krop I, Ismaila N, Andre F et al (2017) Use of biomarkers to guide decisions on adjuvant systemic therapy for women with early-stage invasive breast cancer: American Society of clinical oncology clinical practice guideline focused update. J Clin Oncol 35(24):2838–2847. https://doi.org/10.1200/jco.2017.74.0472
National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology. Breast Cancer Version 5.2020. 2020; Available from: www.nccn.org. Accessed 8 July 2022
Thomssen C, Balic M, Harbeck N, Gnant M (2021) St. Gallen, Vienna 2021: a brief summary of the consensus discussion on customizing therapies for women with early breast cancer. Breast Care (Basel) 16(2):135–143. https://doi.org/10.1159/000516114
Sandhu R, Chollet-Hinton L, Kirk EL, Midkiff B, Troester MA (2016) Digital histologic analysis reveals morphometric patterns of age-related involution in breast epithelium and stroma. Hum Pathol 48:60–68. https://doi.org/10.1016/j.humpath.2015.09.031
Risso D, Ngai J, Speed TP, Dudoit S (2014) Normalization of RNA-seq data using factor analysis of control genes or samples. Nat Biotechnol 32(9):896–902. https://doi.org/10.1038/nbt.2931
Bhattacharya A, Hamilton AM, Furberg H et al (2021) An approach for normalization and quality control for NanoString RNA expression data. Brief Bioinform. https://doi.org/10.1093/bib/bbaa163
Hamilton AM, Hurson AN, Olsson LT et al (2022) The landscape of immune microenvironments in racially-diverse breast cancer patients. Cancer Epidemiol Biomarkers Prev. https://doi.org/10.1158/1055-9965.Epi-21-1312
Parker JS, Mullins M, Cheang MC et al (2009) Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 27(8):1160–1167. https://doi.org/10.1200/jco.2008.18.1370
Raychaudhuri M, Schuster T, Buchner T et al (2012) Intratumoral heterogeneity of microRNA expression in breast cancer. J Mol Diagnos 14(4):376–384. https://doi.org/10.1016/j.jmoldx.2012.01.016
Karthik GM, Rantalainen M, Stålhammar G et al (2017) Intra-tumor heterogeneity in breast cancer has limited impact on transcriptomic-based molecular profiling. BMC Cancer 17(1):802. https://doi.org/10.1186/s12885-017-3815-2
Barry WT, Kernagis DN, Dressman HK et al (2010) Intratumor heterogeneity and precision of microarray-based predictors of breast cancer biology and clinical outcome. J Clin Oncol 28(13):2198–2206. https://doi.org/10.1200/jco.2009.26.7245
López-Knowles E, Gao Q, Cheang MC et al (2016) Heterogeneity in global gene expression profiles between biopsy specimens taken peri-surgically from primary ER-positive breast carcinomas. Breast Cancer Res 18(1):39. https://doi.org/10.1186/s13058-016-0696-2
Perou CM, Sørlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752. https://doi.org/10.1038/35021093
Habel LA, Shak S, Jacobs MK et al (2006) A population-based study of tumor gene expression and risk of breast cancer death among lymph node-negative patients. Breast Cancer Res 8(3):R25. https://doi.org/10.1186/bcr1412
Kim C, Paik S (2010) Gene-expression-based prognostic assays for breast cancer. Nat Rev Clin Oncol 7(6):340–347. https://doi.org/10.1038/nrclinonc.2010.61
Drury S, Salter J, Baehner FL, Shak S, Dowsett M (2010) Feasibility of using tissue microarray cores of paraffin-embedded breast cancer tissue for measurement of gene expression: a proof-of-concept study. J Clin Pathol 63(6):513–517. https://doi.org/10.1136/jcp.2010.075754
Müller BM, Brase JC, Haufe F et al (2012) Comparison of the RNA-based EndoPredict multigene test between core biopsies and corresponding surgical breast cancer sections. J Clin Pathol 65(7):660–662. https://doi.org/10.1136/jclinpath-2012-200716
Delahaye LJ, Wehkamp D, Floore AN, Bernards R, Van’t Veer LJ, Glas AM (2013) Performance characteristics of the MammaPrint(®) breast cancer diagnostic gene signature. Pers Med 10(8):801–811. https://doi.org/10.2217/pme.13.88
Crozier JA, Barone J, Whitworth P et al (2022) High concordance of 70-gene recurrence risk signature and 80-gene molecular subtyping signature between core needle biopsy and surgical resection specimens in early-stage breast cancer. J Surg Oncol 125(4):596–602. https://doi.org/10.1002/jso.26780
Chung GG, Zerkowski MP, Ghosh S, Camp RL, Rimm DL (2007) Quantitative analysis of estrogen receptor heterogeneity in breast cancer. Lab Invest 87(7):662–669. https://doi.org/10.1038/labinvest.3700543
Schwedhelm TM, Rees JR, Onega T et al (2020) Patient and physician factors associated with Oncotype DX and adjuvant chemotherapy utilization for breast cancer patients in New Hampshire, 2010–2016. BMC Cancer 20(1):847. https://doi.org/10.1186/s12885-020-07355-6
Dinan MA, Mi X, Reed SD, Hirsch BR, Lyman GH, Curtis LH (2015) Initial trends in the use of the 21-gene recurrence score assay for patients with breast cancer in the medicare population, 2005–2009. JAMA Oncol 1(2):158–166. https://doi.org/10.1001/jamaoncol.2015.43
Funding
This research was supported by a grant from UNC Lineberger Comprehensive Cancer Center, which is funded by the University Cancer Research Fund of North Carolina, the Susan G Komen Foundation, the National Cancer Institute of the National Institutes of Health (Grant No. P01CA151135), and the National Cancer Institute Specialized Program of Research Excellence (SPORE) in Breast Cancer (NIH/NCI P50-CA058223). This research recruited participants &/or obtained data with the assistance of Rapid Case Ascertainment, a collaboration between the North Carolina Central Cancer Registry and UNC Lineberger. RCA is supported by a grant from the National Cancer Institute of the National Institutes of Health (Grant No. P30CA016086). The authors would like to acknowledge the University of North Carolina BioSpecimen Processing Facility for sample processing, storage, and sample disbursements (http://bsp.web.unc.edu/). We are grateful to CBCS participants and study staff.
Author information
Authors and Affiliations
Contributions
JG and MAT conceptualized the study and its design. ANH led the analysis and drafted the manuscript. ELK led sample handling and processing. AMH, LTO, and ELK contributed to data processing. BCC and JG provided visual scoring of tumor slides. ANH, MES, JG, and MAT interpreted the study results. All authors critically revised and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The University of North Carolina, Chapel Hill has a license of intellectual property interest in GeneCentric Diagnostics and BioClassifier, LLC, which may be used in this study. The University of North Carolina, Chapel Hill may benefit from this interest that is/are related to this research. The terms of this arrangement have been reviewed and approved by the University of North Carolina, Chapel Hill Conflict of Interest Program in accordance with its conflict of interest policies.
Ethics approval
All study procedures were approved by the University of North Carolina School of Medicine Institutional Review Board.
Informed consent
All participants provided written informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hurson, A.N., Hamilton, A.M., Olsson, L.T. et al. Reproducibility and intratumoral heterogeneity of the PAM50 breast cancer assay. Breast Cancer Res Treat 199, 147–154 (2023). https://doi.org/10.1007/s10549-023-06888-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-023-06888-1