Abstract
With the recent increased availability of ultra-high field (UHF) magnetic resonance imaging (MRI), substantial progress has been made in visualizing the human brain, which can now be done in extraordinary detail. This review provides an extensive overview of the use of UHF MRI in visualizing the human subcortex for both healthy and patient populations. The high inter-subject variability in size and location of subcortical structures limits the usability of atlases in the midbrain. Fortunately, the combined results of this review indicate that a large number of subcortical areas can be visualized in individual space using UHF MRI. Current limitations and potential solutions of UHF MRI for visualizing the subcortex are also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the last 25 years, the number of ultra-high field (UHF) (7.0 T and higher) magnetic resonance imaging (MRI) scanner sites has steadily increased globally (> 70 UHF MRI scanners worldwide at the time of writing). Previous reviews have highlighted the benefits of UHF MRI in the clinical domain (Beisteiner et al. 2011; van der Kolk et al. 2013; Kraff et al. 2014; Benjamin et al. 2015), in functional (f)MRI (Barth and Poser 2011; Francis and Panchuelo 2014), and in the visualization of specific subcortical structures such as the basal ganglia (BG) (Plantinga et al. 2014). For the subcortex as a whole, ultra-high field imaging is especially important, because of the possibility of identification and parcellatation of subcortical structures per individual. The use of atlases is well-spread for the larger cortical and subcortical regions, but atlases only exist for a relatively small number of the subcortical structures (Alkemade et al. 2013). In addition the size and location of subcortical regions vary substantially between individuals (Keuken et al. 2014; Tona et al. 2017), necessitating visualization of these areas in individual space. The subcortex is approximately five times smaller than the neocortex but consists of a large number of unique subcortical structures [approximately 455 structures (Dunbar 1992; Federative Committee on Anatomical Terminology 1998; Alkemade et al. 2013; Forstmann et al. 2017a)]. See Fig. 1 for a number of subcortical structures.
A visualization of a number of subcortical nuclei. Note that a number of nuclei, such as the STN, barely show any contrast on the T1-weighted scans but are clearly visible on the T2*-weighted scans. Image is adapted from (Forstmann et al. 2014)
As noted by Johansen-Berg, recent empirical studies on human cognition seem to neglect this part of the brain (Johansen-Berg 2013). To understand how cognitive functions are implemented in the brain, it is, however, vital to study the entire network of structures that might be functionally involved. The so called cortical-basal ganglia-thalamic loops exemplify how studying both cortical and subcortical areas is essential for fully understanding cognitive function (Alexander and Crutcher 1990; Alexander et al. 1990; Haber and Calzavara 2009; Ding and Gold 2013). These structural loops have a general topographic organization, whereby distinct cortical areas project to both the striatum (STR) and subthalamic nucleus (STN). The STR and STN are strongly connected to other BG nuclei, which via thalamic sub-nuclei project back to the cortex. It is thought that as a result of these distinct structural connections, the cortical-BG-thalamic loops are involved in motor, limbic, and cognitive functions (Alexander et al. 1990; Middleton and Strick 2000a; Haber and Calzavara 2009). For instance, within the thalamus the motor loop projects from the cortical motor areas to the ventral lateral nucleus pars oralis, whereas the cognitive loops, involving cortical areas such as the DLPFC, are thought to involve the directly adjacent ventral anterior nucleus pars parvocellularis (Middleton and Strick 2000b). To be able to study these functional domains it is therefore crucial to separate the distinct areas in the subcortex just as it is essential to identify the structural and functional distinct cortical areas (Turner 2013; Turner and Geyer 2014; Forstmann et al. 2017a).
With the increase of field strength, substantial progress has been made in visualizing the human brain in extraordinary detail (Robitaille and Berliner 2007; Duyn 2010; van der Zwaag et al. 2015; Cho 2016; Setsompop et al. 2016; Marrakchi-Kacem et al. 2016; Budinger et al. 2016; Turner and De Haan 2017; Dumoulin et al. 2017; Marques et al. 2017; Giuliano et al. 2017; Sclocco et al. 2017; Kemper et al. 2017; Gallichan 2017). Using UHF MRI, it has become possible to visualize intracortical anatomical structures, such as the bands of Baillarger, in vivo where before they could only be identified using post mortem myelin stains (Turner 2011; Fracasso et al. 2016).
Generally however, imaging the human subcortex with MRI has been particularly challenging for a number of reasons (Forstmann et al. 2017a). The subcortex consists of a large number of small, directly adjunct structures of which a large number have anatomical properties that makes them very hard to distinguish with standard anatomical T1-weighted MRI and require tailored MRI contrasts (Tourdias et al. 2014; Visser et al. 2016a; Priovoulos et al. 2017). Other general MRI factors that hinder the visualization of the subcortex include the substantially lower absolute SNR in the middle of the brain than in the cortex due to the increased distance from the elements of the modern head coils (Wiggins et al. 2009; de Hollander et al. 2017). In addition, g-factor penalties associated with parallel imaging, are larger in the middle of the brain (Larkman 2007; Vaughan and Griffiths 2012; Pohmann et al. 2015).
The visualization of small subcortical structures benefits from UHF for a number of reasons. The first is the linear increase of signal-to-noise ratio (SNR) with field strength (McRobbie et al. 2006; Robitaille and Berliner 2007; Duyn 2012; van der Zwaag et al. 2015; Pohmann et al. 2015). This increased SNR can be used to improve the spatial resolution and visualize fine grained details due to reduced partial volume effects (PVE) (Lüsebrink et al. 2013; Federau and Gallichan 2016). Further, UHF MRI can provide increased T1-contrast between grey and white matter (van der Zwaag et al. 2015). Similarly, T2* differences tend to be larger at 7T than at lower fields, leading to larger contrasts which has been used for the identification of anatomical borders between the substantia nigra (SN) and STN which were previously challenging to visualize (Dula et al. 2010; Abosch et al. 2010; Cho et al. 2011b). Finally, the g-factor penalties in the middle of the brain are lower on 7T than on 3T, which means that higher acceleration factors can be achieved on 7T with a smaller SNR loss than on 3T (Wen et al. 2015). These advantages of UHF MRI make it a powerful tool for visualizing small nuclei in vivo.
Using UHF MRI several of the thalamic subnuclei can now be visualized in individual space without the need to refer to standardized atlases (Tourdias et al. 2014; Saranathan et al. 2014; Kanowski et al. 2014). However, a large and growing number of subcortical structures can be visualized using UHF MRI, many of which have been demonstrated in a single publication. This paper provides and overview of the 169 subcortical structures which have so far been visualized in the human brain using UHF MRI and the methods used to achieve this. The review will focus on the type of MRI sequence, participant demographics and methods used to parcellate the structure of interest.
Materials and Methods
Search Strategy
A comprehensive literature search was conducted using the Entrez search tools implemented in the Biopython’s Bio.Entrez module (Cock et al. 2009). This is a python application programming interface (API) tool that queries the PubMed database (http://www.pubmed.org). The query date was the 1st of December 2017 and used the following inclusion criteria: publication date was before the 1st of December 2017, focused on humans, used an MRI scanner with a static B0 field strength ≥ 7.0 T, and report the visualization of a subcortical (either in the cerebrum, cerebellum or brainstem) nucleus or region. The search terms that were used were for example “ultra-high field magnetic resonance imaging”, “7 T structural MRI”, “7T neuroimaging”, and “7.0 T magnetic resonance imaging”. All search terms were used with the different common B0 field strengths for UHF MRI (7.0, 8.0, 9.4, 10.5, and 11.7).
Inclusion Procedure
All 5818 resulting abstracts were read by two raters (MCK & BRI) and based on the inclusion criteria detailed above, a decision was made to read the full-text paper or not. The abstracts that both raters did not agree on were checked again. The potential 388 full-text papers were read by a single rater (MCK) and were separated into reviews and empirical papers. The 299 empirical papers were checked for all inclusion criteria and if there was a match, the paper was included in the final list. The 58 review papers were cross referenced, which entailed that the 5252 abstracts of all cited papers were read and checked for additional potential full-text papers.
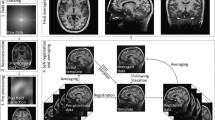
Finally, to test whether the employed search strategy resulted in a comprehensive set of papers, the included papers were compared to the publications of the authors of this review. The included papers were compared to the list of publications which were a priori known to fit the inclusion criteria. This comparison indicated that two out of the 27 papers by our own group were not found via the PubMed search, implying that approximately 7% of the empirical papers that would fit the inclusion criteria were not identified. The literature search resulted in the inclusions of 169 papers (see Fig. 2 for an overview of the article selection procedure).
Search strategy. Using the Entrez search tools implemented in the Biopython’s Bio.Entrez module the PubMed database was queried for a number of search terms. This resulted in a number of abstracts that were read and double-checked by two independent readers. The resulting full texts were then downloaded and separated in empirical studies and reviews. The empirical papers were read to check if they matched the inclusion criteria, resulting in the inclusion of 131 papers. The reviews were cross referenced and resulting abstracts were read by one rater. The resulting full text empirical papers were read, and an additional 9 papers were added. Finally, the 140 papers from the PubMed search were compared to the publications by the authors of this review. This resulted in 2 papers that were not identified by our search strategy
The information extracted from the papers was as follows: which subcortical structures were visualized or parcellated, whether the measurements were from in vivo or post mortem samples, whether the population consisted of healthy or clinical subjects, which MRI contrast was used to visualize the subcortical structures, and the accompanying MRI parameters.
Identification Versus Parcellation
The subcortical structure(s) in each paper was classified as being either ‘identified’ or ‘parcellated’. Identification was defined as the placement of abbreviations, arrows or other visual markers that corresponded to an anatomical label in an image of a structural MRI scan. Parcellation was defined as the manual, automatic, or semi-automatic delineation of the entire or partial structure. Manual parcellation is defined as the process where an expert delineates and labels the borders of a region of interest (ROI) manually [e.g., (Lenglet et al. 2012; Kwon et al. 2012)]. Automatic parcellation is defined as the process where the ROI is parcellated using a software package without any manual editing [e.g., (Zhang et al. 2001; Visser et al. 2016a)]. Semi-automatic parcellation is defined as automatic parcellation whereby the resulting parcellation is manually edited if needed [e.g. (Mestres-Missé et al. 2014)].
The parcellation method had to employ the actual contrast of the nuclei and the surrounding tissue. Single atlas label propagations, where an individual anatomical MRI scan is registered to a pre-labeled standard structural template, were excluded. The reason for this exclusion is that label propagation is a registration problem between the template and the entire individual anatomical MRI volume and is unable to capture large anatomical variation (Doan et al. 2010; Cabezas et al. 2011).
MRI Sequence Classes
The MRI contrasts which were used to visualize the structures of interest were grouped according to the main classes of contrasts: T1, T2, T2*, functional (regardless of underlying mechanism—T2* BOLD, T2 BOLD, T1 VASO, fQSM, etc), diffusion weighted imaging (DWI), susceptibility weighted imaging (SWI), including phase imaging and quantitative susceptibility mapping (QSM), magnetization transfer (MT), proton density (PD), multiple, and other. The multiple MRI sequence category entails those studies that visualized the structure of interest in a number of MRI sequences. Inclusion in the ‘other’ category was either a single MRI sequence that was not specific to a given contrast mechanism (e.g., both PD and T2 weighted) or did not fit the above classification scheme (e.g., magnetic resonance spectroscopy).
It is beyond the scope of this review to go into a detailed description of the separate contrast mechanisms and we refer to the following literature (McRobbie et al. 2006; Robitaille and Berliner 2007). Very briefly, a T1 contrast is based on the recovery time of the longitudinal component of the magnetization following the application of a radio frequency excitation pulse, while T2 refers to the decay of the transverse magnetization component as a result of proton interactions (McRobbie et al. 2006). The T2* contrast is based on the decay of the transverse magnetization component as a result of proton interactions and the magnetic field inhomogeneity (McRobbie et al. 2006; Chavhan et al. 2009). The DWI contrast is based on the dephasing of the protons due to the diffusion of water molecules (Jones et al. 2013; Chilla et al. 2015). SWI and QSM contrasts are based on a combination of T2*-weighted magnitude and filtered phase images (Haacke et al. 2008; Liu et al. 2014). The MT contrast is based on the effect of off-resonance RF pulses on bound and free moving protons (Grossman et al. 1994; McRobbie et al. 2006). Finally the PD contrast reflects the density of the protons (McRobbie et al. 2006). To be able to summarize across the large number of sequence categories no distinction was made between quantitative or qualitative MRI scans (e.g., T1 maps versus T1 weighted scans or QSM versus SWI).
(Near) Isotropic Voxel Size
Isotropic voxels are essential when visualizing small structures, as they have equal biases in all directions when determining the borders. Using anisotropic voxels has the advantage of high in-plane resolution, but determining the border in the z-direction becomes problematic as PVE are increased and can result in measurement biases of subcortical structures (Wonderlick et al. 2009). We determined whether a study acquired isotropic or near isotropic voxels by first calculating the reported voxel volume. For a given volume, the corresponding isotropic voxel dimension was calculated, and compared to the actual acquired voxel size. If the acquired voxel dimensions were within a 10% margin of the isotropic dimensions, the acquired voxel was deemed (near) isotropic, all other voxels were classified as anisotropic.
Open Access and Interactive Use
All data and code used to analyze and generate the summary figures can be found online (osf.io/fwc2p/, https://doi.org/10.17605/OSF.IO/FWC2P). In addition, a condensed R script is provided which can be used to generate the list of subcortical structures identified with UHF as well to create a summary figure (such as Fig. 6,7 and 8) for a given structure of interest. The R code contains a description of the software requirements as well as instructions for use.
Results
A total of 169 papers were published between 1993 and 2018 that together report the visualization of 163 subcortical structures using 7 T or higher, including both in vivo and post mortem studies. The most frequently employed field strength was 7.0 T (7.0 T: 147 studies; 8.0 T: 7 studies; 9.4 T: 11 studies; 11.7 T: 2 study; 21.1 T: 2 studies; see Fig. 3a). This was expected as the number of 7.0T MRI scanner sites is much larger than that of the higher field strengths (Plantinga et al. 2014). The most frequently employed MRI contrast across the different field strengths and structures were T2* based scans, followed by T1, SWI, and T2 contrasts (see Fig. 3b for the frequency of using a given MRI contrast).
Overview results. a The frequency that a MRI scanner with a given B0 field strength was used in the 142 studies. b The frequency of using a certain MRI sequence type to visualize a subcortical area. c Of the 658 cases of identifying a subcortical area, most were done using in vivo samples. d The thirteen most frequently reported structures. Funct functional MRI sequences that employed functional localizer stimuli, DWI diffusion weighted imaging; SWI susceptibility weighted imaging, MT magnetization transfer; PD proton density, N.s. not stated
Demographics
The overall sample size ranged between 1 and 152 participants, with a mean sample size of 18.99 (SD 21.81) and a median of 11 participants across the 169 papers. The in vivo sample size was on average 19.09 (SD 17.93) with a median of 13 participants. The post mortem sample size was on average 15.67 (SD 31.90) with a median of 3.5 specimens. 108 studies included only healthy controls, 13 studies included only patients, 43 studies included both patient and healthy participants, and for 5 studies the participants’ status was not disclosed. The most frequently measured patient groups with UHF MRI were people suffering from Parkinson’s Disease (PD) and Multiple Sclerosis followed by studies that focused on fetal development and or fetal abnormalities. Two out of the six studies that included fetal samples used a wide-bore UHF MRI scanner (see Table 1).
Subcortical Structures
The frequency with which a structure was reported ranged between 1 and 51, with a mean reported frequency of 4.62 (SD 8.88) and a median of 1. There are 55 UHF MRI studies that only reported a single structure, whereas for 83 structures there was only a single UHF MRI study that visualized that specific structure [e.g., for the locus coeruleus (Keren et al. 2015); the field of Forel (Massey et al. 2012); and a number of thalamic sub-nuclei such as the magno- and parvocellular part of the lateral geniculate nucleus (Denison et al. 2014)]. The SN was reported most frequently (51 reports), closely followed by the red nucleus (50 reports) and putamen (48 reports; see Fig. 3d for the seventeen most frequently reported structures).
Identification Versus Parcellation
Of the 753 reports across the 169 papers, there were 344 reports where the authors (partially) parcellated a subcortical structure. This was either done by manual parcellation (208 reports), placing a ROI in a visually identified area (51 reports), semi-automatic procedures (22 reports), fully automatic procedures (26 reports), using a functional localizer (5 reports), or otherwise parcellated in a way that was unclear from the manuscript (32 reports). Overall, regardless of method, the most frequently parcellated structure was the putamen (31 reports) whereas the STN was the most frequently manually parcellated structure (21 reports).
Of the 344 parcellated reports there were 75 structures parcellated in vivo, and 36 structures parcellated using post mortem samples. There is an overlap of 17 structures that are parcellated in both in vivo and post mortem data (see Fig. 4 for a comparison between the image quality achievable with in vivo versus post mortem scanning).
In vivo versus post mortem comparison. The left panel shows the MNI152 template with a highlighted subcortical region. The middle panel highlights this subcortical region using a 7T in vivo 0.5 mm isotropic resolution T2*-weighted structural scan where the globus pallidus externa (GPe), globus pallidus interna (GPi), STN and SN can be visualized. The right panel illustrates a similar region in a post mortem sample scanned with a 0.1 mm isotropic resolution T2*-weighted scan where a number of subcortical areas can be identified which are not clearly visible in the in vivo scans such as the fields of Forel (H1, H2), zona incerta and the comb system. Image is adapted from (Forstmann et al. 2017a)
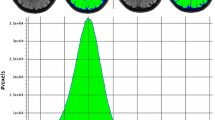
Structures which were only parcellated using post mortem data include a number of small structures in the lower brainstem such as the abducens nucleus, primary olivary nucleus, cuneate nucleus, a number of sub-nuclei of the hypothalamus, and the claustrum. That the claustrum has never been parcellated in vivo was somewhat surprising as it is a relatively large structure, medial to the striatum. A potential explanation why such small structures in the brainstem are only parcellated using post mortem data is the employed voxel volume (see Fig. 5 for an overview of voxel volumes used per MRI sequence and sample type). One of the benefits of post mortem scanning is the possibility to employ longer scan times in the absence of motion, which allows for the acquisition of smaller voxels, and/or the possibility of scanning a smaller sample at higher fields than available in vivo [e.g., 0.05 mm isotropic voxels with an acquisition of 4.3 h using 21.1T (Foroutan et al. 2013) or 0.09 mm isotropic voxels with an acquisition of 10.5 h using 7.0T (Makris et al. 2013b)].
Voxel volume for the different MRI sequences. Each dot represents the voxel volume used to visualize a subcortical structure across the 169 studies. The in vivo samples are displayed in red, whereas the post mortem samples are shown in blue. The color intensity corresponds to the number of studies using the same voxel volume. Funct functional MRI sequences that employed functional localizer stimuli, DWI diffusion weighted imaging, SWI susceptibility weighted imaging, MT magnetization transfer, PD proton density, N.s. not stated, PD (patient type) Parkinson’s disease, MS multiple sclerosis
Voxel Volume and Isotropic Voxels
The voxel volume across the different structural MRI contrasts including the DWI scans for the in vivo scans ranged between 0.0144 and 42.875 mm3, with a mean volume of 1.09 mm3 (SD 3.71 mm3) and a median of 0.245 mm3. The voxel volume for the functional MRI contrasts for the in vivo scans ranged between 0.422 and 39.051 mm3, with a mean volume of 4.50 mm3 (SD 7.72 mm3) and a median of 1.33 mm3. For the post mortem scans the volume varied between 0.000125 and 1.47 mm3 with a mean voxel volume of 0.075 mm3 (SD 0.23 mm3) and a median of 0.01 mm3. See Fig. 5 for an overview of voxel volumes used per MRI sequence.
Of all the structures that were identified using a T1 based contrast, 128 reports of structures were achieved using isotropic or near isotropic voxels, and 83 reports were based on anisotropic voxels. For the T2 based contrasts sequences, 26 reports were based on isotropic voxels, and 90 reports were based on anisotropic voxels. Using a T2* sequence, 114 reports were based on isotropic voxels, whereas 138 reports were not. For the functional sequences, all 25 reports were based on isotropic voxels. The DWI sequence resulted in 60 reports using isotropic voxels and 27 reports using anisotropic voxels. SWI sequences that were used to identify structures were isotropic in 82 cases and in 21 cases anisotropic. All three reports that identified a structure using an MT based sequence were based on anisotropic voxels. The PD sequences that were used to identify structures were isotropic for 6 reports and 18 reports were based on anisotropic voxels.
Volumetric Reports
With a total of 51 reports, the SN is the most frequently visualized structure, of which only 9 papers provide an explicit volume estimate (see Table 2). For the STN, directly adjacent to the SN, there are 42 reports, of which there are 12 reports that provide a volume estimate. There is substantial variability in volume estimates for both structures. For the SN, volumes range between 224.75 and 1300 mm3. For the STN the volumes range between 37.32 and 223 mm3. The volumes are based on a range of different MRI contrasts and parcellation methods, such as automatic segmentations or the conjunction of two manual raters. This variability in methods makes it problematic to provide a summary of volume estimates and whether there is a systematic difference due to acquisition technique.
MRI Contrasts for Visualizing the SN, STN, and Thalamus
It is interesting to note the variability in MRI contrasts used to visualize a number of subcortical structures. For the SN by far the most commonly used contrast is a T2* based sequence followed by SWI contrasts (Fig. 6d). Given that the SN contains relatively large amounts of iron, which increases the magnetic susceptibility, it is not surprising that T2* and SWI seem to be the contrasts of choice (Hallgren and Sourander 1958; Chavhan et al. 2009). In terms of demographics, the SN is regularly visualized in PD patients, which is expected due to the underlying pathology occurring in the SN in PD (Fig. 6c).
Overview of the use of UHF MRI for visualizing the substantia nigra. a Of the 51 studies that identified the SN, most were done using in vivo samples. b Most studies only used healthy controls, whereas a substantial number also included patients. c The studies that included a clinical group mainly focused on Parkinson’s Disease patients or abnormal fetal developments. d The frequency of using a certain MRI sequence type to visualize the SN. The most frequently used contrast was a T2* type of sequence. Funct functional MRI sequences that employed functional localizer stimuli, DWI diffusion weighted imaging, SWI susceptibility weighted imaging, MT magnetization transfer, PD proton density, N.s. not stated, PD (patient type) Parkinson’s Disease, MS multiple sclerosis
Another structure which is implicated in the pathophysiology of PD is the STN, a structure also high in iron content and located directly adjacent to the SN. As with the SN, the most frequently used contrast mechanism to visualize the STN is T2* (Fig. 7d). The ratio for identification versus parcellation of the STN is larger than for the SN. Additionally, the STN is more commonly visualized in the healthy population, compared to the SN which included relatively more clinical groups (Fig. 6c versus Fig. 7c).
Overview of the use of UHF MRI for visualizing the subthalamic nucleus. a Of the 42 studies that identified the STN, most were done using in vivo samples. b Most studies only used healthy controls. Compared to the SN there were substantially fewer studies that also included patients. c The studies that included a clinical group mainly focused on Parkinson’s Disease patients or abnormal fetal developments. d The frequency of using a certain MRI sequence type to visualize the STN. The most frequently used contrast was a T2* type of sequence. Funct functional MRI sequences that employed functional localizer stimuli, DWI diffusion weighted imaging, SWI susceptibility weighted imaging, MT magnetization transfer, PD proton density, N.s. not stated, PD (patient type) Parkinson’s Disease, MS multiple sclerosis
The thalamus (Th), a structure that contains roughly four times less iron than the SN (Hallgren and Sourander 1958) is visualized with a much wider range of MRI sequences (Fig. 8d). A T2* based contrast is used most frequently which is surprising given the lower iron concentrations in the Th, but is closely followed by T1 based sequences.
Overview of the use of UHF MRI for visualizing the thalamus. a Of the 36 studies that identified the Th, most were done using in vivo samples. b Most studies only used healthy controls. Compared to the SN, there were substantially fewer studies that also included patients. c The studies that included a clinical group mainly focused on abnormal fetal developments. d The frequency of using a certain MRI sequence type to visualize the Th. The most frequently used contrast was a T2* type of sequence, followed closely by T1 type of sequences. Funct functional MRI sequences that employed functional localizer stimuli, DWI diffusion weighted imaging, SWI susceptibility weighted imaging, MT magnetization transfer, PD proton density, N.s. not stated, PD (patient type) Parkinson’s disease, MS multiple sclerosis
Optimal MRI Contrast
There are a number of studies that explicitly state that one MRI contrast is superior to other sequences for the identification or parcellation of the SN, STN, or Th. There were 7 papers for the SN (Abduljalil et al. 2003; Abosch et al. 2010; Deistung et al. 2013a, b; Eapen et al. 2011; Schäfer et al. 2012; Shmueli et al. 2009; Khabipova et al. 2015; Kerl et al. 2012), 6 papers for the STN (Abosch et al. 2010; Schäfer et al. 2012; Kerl et al. 2012; Deistung et al. 2013b; Zeineh et al. 2014; Alkemade et al. 2017), and 6 papers that compared sequences for the Th (Abduljalil et al. 2003; Hammond et al. 2008a; Abosch et al. 2010; Deistung et al. 2013b; Tourdias et al. 2014; Kanowski et al. 2014). For the SN, the consensus for visualization seems to be that either a T2* or SWI based sequence is optimal, which are highly similar contrasts. For the STN, this is not as clear as there are roughly an equal number of studies that prefer T2*, SWI or T2 based images. The Th was preferentially visualized using a T2* contrast (see Table 3).
Discussion
The subcortex can be parcellated into a large number of anatomically distinct structures (Federative Committee on Anatomical Terminology 1998). Only approximately 7% of these known structures are incorporated in standard anatomical MRI atlases (Alkemade et al. 2013). However, by reviewing the literature that utilized UHF MRI to visualize the subcortex, it became apparent that the number of observed subcortical structures is considerably larger. Specifically, at least 163 unique subcortical structures are identifiable in individual space using UHF MRI. We have provided R code to enable the reader to explore the use of UHF MRI for a given structure. A reader interested in structure ‘A’ can now obtain a list of the papers identifying this structure and the resolutions and methods used to do so.
The ability of UHF MRI to identify a large number of subcortical nuclei in individual space is of the utmost importance given the anatomical variability that exists across individuals (Mazziotta et al. 1995; Amunts et al. 1999; Uylings et al. 2005; Daniluk et al. 2009; Keuken et al. 2014). This anatomical variability is far from static as a number of factors including gene–environment interactions, healthy aging, and disease all influence individual anatomy over time (Thompson et al. 2001; Raz 2005; Lenroot and Giedd 2008; Daniluk et al. 2009; Keuken et al. 2013, 2017). These factors question the validity of using anatomical atlases which fail to incorporate anatomical variability or are not specific for an age group or clinical population (Devlin and Poldrack 2007; Alho et al. 2011).
The Clinical Use of UHF
There are numerous recent reviews highlighting the substantial benefits of UHF MRI in a clinical setting (Cho et al. 2010b; Beisteiner et al. 2011; Duchin et al. 2012; Plantinga et al. 2014; Kraff et al. 2014; Trattnig et al. 2015, 2016; Gizewski et al. 2015). A number of studies have directly compared clinically utilized 1.5 and 3.0T field strengths to UHF MRI, showing UHF MRI results in an improved visualization across a number of patient groups and structures (Peters et al. 2007; Cho et al. 2008a, 2010b, 2010a, 2011a; Hammond et al. 2008a; Kollia et al. 2009; Yao et al. 2009; Zwanenburg et al. 2009; Abosch et al. 2010; Blazejewska et al. 2013; Chalifoux et al. 2013; Derix et al. 2014; Saranathan et al. 2014; Cosottini et al. 2015). Based on our own review, it is clear that UHF MRI is already frequently used to visualize subcortical structures in a clinical setting for populations such as Parkinson’s Disease, Alzheimer’s Disease, and Multiple Sclerosis. The benefit of UHF MRI in a clinical setting can be illustrated by its use with regards to preoperative planning for Deep Brain Stimulation (DBS) procedures as a treatment for PD patients. DBS is a surgical procedure where an electrode is inserted into the STN with the goal of reducing the motor symptoms of the disease, while simultaneously minimizing the occurrence cognitive and limbic side-effects known to affect a number of patients (Limousin et al. 1995; Temel et al. 2005). The development of these side-effects can partially be attributed to the suboptimal placement of the electrode in the STN (Kleiner-Fisman et al. 2006; Cakmakli et al. 2009; Paek et al. 2011). Given that the location of the STN changes with both age and disease (Dunnen and Staal 2005; Kitajima et al. 2008; Keuken et al. 2013, 2017; Mavridis et al. 2014; Pereira et al. 2016) it is crucial to visualize such a structure as accurately as possible per individual, which is why the superior visualization of UHF MRI is so valuable to DBS. The same logic can be passed to alternative neurosurgical interventions such as tumor delineation and removal, proton beam, gamma knife and radiation therapies which all require precise anatomical visualization, best afforded by UHF MRI (Forstmann et al. 2017b).
Optimal MRI Sequence per Structure
Optimal MRI sequences providing sufficient Contrast-to-Noise Ratio (CNR) are essential for clinical research. It is crucial to visualize the structure of interest while maintaining a clinically feasible scanning time. Therefore, given that different tissues require different MR sequences and parameters, it is important to experimentally determine the optimal sequence for each structure of interest (Marques and Norris 2017).
To highlight the variability of preferred sequences, the studies that used multiple MRI sequences to visualize the SN, STN, and Th were compared. Based on the literature review, the preferred contrast to visualize any of these three structures, even the Th is a T2* sequence (Abduljalil et al. 2003; Hammond et al. 2008a; Shmueli et al. 2009; Abosch et al. 2010; Eapen et al. 2011; Schäfer et al. 2012; Kerl et al. 2012; Kerl 2013; Deistung et al. 2013a, b; Gizewski et al. 2013; Tourdias et al. 2014; Zeineh et al. 2014; Saranathan et al. 2014; Kanowski et al. 2014; Khabipova et al. 2015). Such T2* sequences have been used in PD patients to investigate pathological alterations occurring in the SN dopaminergic system [e.g., (Cho et al. 2010b, 2011b; Kwon et al. 2012)]. Particularly at high UHF MRI the use of a T2* weighted sequence for a volumetric study is however not trivial. Pronounced B0 inhomogeneities lead to additional dephasing which may result in signal dropouts especially in regions with high iron content. Additionally, a major difficulty in interpreting T2*-weighted gradient-echo data is that the dependence of the signal on the tissue susceptibility is a non-local effect, i.e., the signal within a voxel is not only from effected by sources within but also from neighboring sources outside that voxel. Therefore, T2* hypointensity and phase contrast in gradient-echo techniques are not directly reflective of local tissue properties (Schäfer et al. 2009) which can effect volumetric measurements (Chandran et al. 2015). Shorter TE acquisition are preferable for volumetric measurements in terms of edge fidelity, but do not have the high contrast associated with midrange TE’s. What the optimal sequence is for the other subcortical structures is unclear from the current available publications and will probably differ from the SN, STN, and Th due to differences in tissue properties, most notably the lower concentrations of iron.
It should also be noted that these comparison studies should be viewed with the ongoing development of MRI contrasts such as QSM in mind (Marques and Norris 2017). QSM is a novel post-acquisition processing technique where the susceptibility of the tissue is quantified by estimating the magnetic field distribution and solves the inverse problem from field perturbation to magnetic susceptibility, while removing the background field contribution (Schweser et al. 2011, 2016). As such the QSM suffers less from non-local effects as described above which makes it an interesting contrast for volumetric studies of iron rich nuclei [e.g. (Liu et al. 2013; Alkemade et al. 2017)].
Quantitative Maps
Most of the included UHF MRI studies use standard MRI sequences that are (mainly) weighted for a certain contrast mechanism as opposed to a quantitative map, of, e.g., T1 or T2* relaxation. This is unfortunate as there are several clear advantages to quantitative MRI (qMRI) over standard weighted sequences (Weiskopf et al. 2015). One of the benefits of qMRI is that the quantitative maps can be used to generate bias-free weighted images [e.g. (Renvall et al. 2016)]. Another benefit of quantitative maps is the possibility of assigning a physical meaning to the intensity value of the image and therefore being able to provide biologically and spatially specific information (Weiskopf et al. 2015; Ropele and Langkammer 2016). For instance, T1, the parameter describing the spin–lattice relaxation, has been used as a proxy for myelin content (Koenig 1991; Stüber et al. 2014; Lutti et al. 2014; Dinse et al. 2015), whereas T2*, the parameter describing the spin–spin relaxation in combination with field inhomogeneity, and especially QSM are thought to be informative for iron concentration (Fukunaga et al. 2010; Lee et al. 2010; Cohen-Adad et al. 2012; Stüber et al. 2014).
One of the downsides of qMRI is that the acquisition time of a quantitative map is usually longer than standard weighted MRI. However, this can be solved by combining different contrast mechanisms into one data acquisition enabling quantification of multiple MRI parameters within a clinically acceptable time (Weiskopf et al. 2013). The advantage of having multiple contrasts is that each contrast contains complimentary anatomical information that can be used to inform segmentation algorithms, such as the multimodal image segmentation tool [MIST, (Visser et al. 2016a, b)].
Reporting the Demographic and MRI Protocol Values
A critical note needs to be made regarding the lack of details reported in the included papers. A substantial number of studies fail to report basic demographic information of the measured subjects. At times information regarding the exact age, gender ratio, and whether the participant is healthy is missing. This is problematic as age and disease can have substantial effects on the biological properties of the brain (Minati et al. 2007; Aquino et al. 2009; Fritzsch et al. 2014; Lorio et al. 2014; Visser et al. 2016b). In other cases, essential information regarding the MRI protocol such as field of view, matrix size, or voxel size is missing or incomplete. This hinders the reproducibility of these studies and makes it challenging to implement their sequences and protocols. As such it should be recommended that groups adhere to the guidelines on reporting neuroimaging studies (Poldrack et al. 2008; Nichols et al. 2016).
Challenges of UHF MRI
An obvious limitation of UHF MRI is the limited accessibility. Of the approximately 36,000 MRI scanners available worldwide, only ± 0.2% are UHF MRI scanners (Rinck 2016). Given the advantages for visualizing clinically relevant subcortical nuclei, this calls for an increase of UHF MRI scanner sites but we acknowledge the substantial higher purchasing and running costs of a UHF MRI scanner. A more technical challenge with UHF MRI are the B0 and B1 field inhomogeneities which increase with field strength resulting in local signal intensity variations and signal dropout (Truong et al. 2006a; van der Zwaag et al. 2015). While B0 and B1 field inhomogeneity remains an active field of research, substantial progress has already been made in overcoming these problems (van der Zwaag et al. 2015; Yarach et al. 2016; Sclocco et al. 2017). For the subcortex, the absence of nearby air–water interfaces for most of the subcortical structures means that B0 inhomogeneities are a relatively minor problem. B1 inhomogeneities are more problematic. While the standard single-channel transmit/32-channel receive coils have a relatively favorable transmit B1 pattern with highest achieved flip angles in the middle of the brain, the receive profile of the array coils means that SNR is rather lower in the midbrain than in the cortex.
While the spatial resolutions achieved by in vivo UHF-MRI are impressive, on its own, it is not able to deliver the anatomical resolution needed to visualize all structures known to be present in the human brain. At present, the combination of neuroimaging and post mortem staining’s are still needed to create a complete and comprehensive picture of the human brain in its entirety (Yang et al. 2013; Amunts et al. 2013; Forstmann et al. 2017a). An example of such a combination has been given by Ding and colleagues (Ding et al. 2016). Here they used a single post mortem brain, which was structurally scanned with 7.0T and subsequently further processed using various staining techniques. A staggering 862 cortical and subcortical areas were manually segmented and aligned to the structural MRI scans. Given that it is not yet possible to fully automatize such a pipeline nor translate it directly to the individual in vivo brain, these efforts will not quickly result in a tool to identify the structures per individual brain. However, what such a multi-modal atlas could do is to provide shape, intensity, and spatial relationship priors for automatic segmentation methods (Bogovic et al. 2013; Kim et al. 2014; Visser et al. 2016a, b).
A final limitation of UHF MRI utility is that until recently the standard FDA approval for clinical scanning only went up to 3.0T (van Osch and Webb 2014). This restriction does not seem to be based on safety concerns, as the risks associated with UHF MRI up to 8.0T are similar to 1.5 and 3.0T (Administration 2003; van Osch and Webb 2014). This limitation has hindered the use of UHF MRI in standard clinical practice which, given the clear clinical advantages, is unfortunate (Kraff et al. 2014; Trattnig et al. 2015). This limitation has been recently been resolved as the newest generation of 7.0T systems (e.g., the Siemens 7.0T MAGNETOM Terra system) has both CE and FDA clinical approval (Heimbach 2015; Healthineers 2017a, b). This might result in more institutes having a larger interest in investing in UHF MRI scanners, increasing the accessibility for clinical and non-clinical research.
Future Development
As the voxel sizes continue to decrease, involuntary subject motion becomes an increasing challenge, to the extent that muscle relaxation, cardiac pulsation, respiratory motion and swallowing have a measurable effect on the image quality (Herbst et al. 2013; Stucht et al. 2015). A possible solution for this would be prospective motion correction (PMC), where the MR gradient system is adjusted in real time to ensure that the brain remains in the same location in the imaged volume (Maclaren et al. 2012). PMC has been used in combination with UHF MRI and results of whole brain MP2RAGE scans with an isotropic resolution of 0.44 mm have been presented (Stucht et al. 2015). One of the downsides of PMC is that for the currently commercially available systems additional hardware is necessary to track the motion of the brain (Maclaren et al. 2012). Another possibility would be to use MR-based motion measures such as fat image navigators (fat-navs) (Gallichan et al. 2015; Federau and Gallichan 2016). Fat-navs are interleaved acquired high contrast images of the sub-cutaneous fat and bone marrow of the skull and can be used to estimate and correct head motion. Using these fat-navs, whole brain MP2RAGE scans with an isotropic resolution of 0.35 mm have been acquired at 7T (Stucht et al. 2015). The advantage of such high spatial resolution is that certain anatomical details such as the grey matter islands between the putamen and caudate become much more visible [see Fig. 9 for a visual comparison between two whole brain MP2RAGE datasets of which one used fat-Navs and higher spatial resolution. Data is provided by (Forstmann et al. 2014; Stucht et al. 2015; Federau and Gallichan 2016)].
Structural MP2RAGE whole brain volumes with or without retrospective motion correction. a A MP2RAGE whole brain volume that was acquired with 0.35 mm isotropic voxel resolution using Fat-Navs for retrospective motion correction. The image is based on an average of 4 scans which were registered using trilinear interpolation. The MRI data is made freely available and described in Federau and Gallichan (2016). b A single MP2RAGE whole brain volume that was acquired with 0.7 mm isotropic voxel resolution, with no motion correction. The MRI data is made freely available and described in Forstmann et al. (2014). The MP2RAGE in panel A has a voxel volume that is 8 times smaller than in panel B. This difference in voxel size results in a substantially lower PVE
Conclusion
The number of UHF MRI sites are steadily increasing as there are several advantages over lower field MRI such as intrinsic higher SNR and increased CNR. With the increase of field strength, it becomes possible to visualize small subcortical structures and their subnuclei which are challenging to localize. This is illustrated in this review by the fact that UHF MRI, with a wide range of imaging approaches, has been able to identify 169 subcortical structures in the individual brain. Some of these concern subdivisions in structures that were only identifiable as a whole at lower fields. It should however be noted that most of these structures were only identified in a single publication. This is substantial progress, but also emphasizes the amount of work yet to be done to find a comprehensive imaging approach to parcellate the subcortex per individual. With the large efforts currently directed at UHF sequence development (Marques and Norris 2017) it seems especially likely that the number of identifiable structures will increase further.
References
Abduljalil AM, Schmalbrock P, Novak V, Chakeres DW (2003) Enhanced gray and white matter contrast of phase susceptibility-weighted images in ultra-high-field magnetic resonance imaging. J Magn Reson Imaging 18:284–290. https://doi.org/10.1002/jmri.10362
Abosch A, Yacoub E, Ugurbil K, Harel N (2010) An assessment of current brain targets for deep brain stimulation surgery with susceptibility-weighted imaging at 7 T. Neurosurgery 67:1745–1756. https://doi.org/10.1227/NEU.0b013e3181f74105
Administration UFAD. (2003) Guidance for industry and FDA staff: criteria for significant risk investigations of magnetic resonance diagnostic devices. Washington DC
Aggarwal M, Zhang J, Pletnikova O et al (2013) Feasibility of creating a high-resolution 3D diffusion tensor imaging based atlas of the human brainstem: a case study at 11.7T. NeuroImage 74:117–127. https://doi.org/10.1016/j.neuroimage.2013.01.061
Alarcon C, de Notaris M, Palma K et al (2014) Anatomic study of the central core of the cerebrum correlating 7-T magnetic resonance imaging and fiber dissection with the aid of a neuronavigation system. Neurosurgery 10:294–304. https://doi.org/10.1227/NEU.0000000000000271
Alexander G, Crutcher M (1990) Functional architecture of basal ganglia circuits: neural substrates of parallel processing. Trends Neurosci 13:266–271
Alexander GE, Crutcher MD, DeLong MR (1990) Basal ganglia-thalamocortical circuits: parallel substrates for motor, oculomotor, “prefrontal” and “limbic” functions. Progr Brain Res 85:119–146
Al-Helli O, Thomas DL, Massey L et al (2015) Deep brain stimulation of the subthalamic nucleus: histological verification and 9.4-T MRI correlation. Acta Neurochir 157:2143–2147. https://doi.org/10.1007/s00701-015-2599-x
Alho EJL, Grinberg L, Heinsen H, Fonoff ET (2011) Review of printed and electronic stereotactic atlases of the human brain. In Neuroimaging for clinicians-combining research and practice, 1st edn. InTech, Rijeka, pp 145–172
Alkemade A, Keuken MC, Forstmann BU (2013) A perspective on terra incognita: uncovering the neuroanatomy of the human subcortex. Front Neuroanat. https://doi.org/10.3389/fnana.2013.00040
Alkemade A, de Hollander G, Keuken MC et al (2017) Comparison of T2*-weighted and QSM contrasts in Parkinson’s disease to visualize the STN with MRI. PLoS ONE 12:e0176130–e0176113. https://doi.org/10.1371/journal.pone.0176130
Al-Radaideh AM, Wharton SJ, Lim SY et al (2013) Increased iron accumulation occurs in the earliest stages of demyelinating disease: an ultra-high field susceptibility mapping study in Clinically Isolated Syndrome. Multiple Sclerosis J 19:896–903. https://doi.org/10.1177/1352458512465135
Amunts KK, Schleicher AA, Zilles KK (1999) Broca’s region revisited: cytoarchitecture and intersubject variability. J Comp Neurol 412:319–341
Amunts K, Lepage C, Borgeat L et al (2013) BigBrain: an ultrahigh-resolution 3D human brain model. Science 340:1472–1475. https://doi.org/10.1126/science.1235381
Aquino D, Bizzi A, Grisoli M et al (2009) Age-related iron deposition in the Basal Ganglia: quantitative analysis in healthy subjects. Radiology 252:165–172. https://doi.org/10.1148/radiol.2522081399
Augustinack JC, van der Kouwe AJW, Salat DH et al (2014) H.M.’s contributions to neuroscience: a review and autopsy studies. Hippocampus 24:1267–1286. https://doi.org/10.1002/hipo.22354
Bao L, Li X, Cai C et al (2017) Quantitative susceptibility mapping using structural feature based collaborative reconstruction Pub _newline (SFCR) in the human brain. IEEE Trans Med Imag 35:2040–2050. https://doi.org/10.1109/TMI.2016.2544958
Barry RL, Coaster M, Rogers BP et al (2013) On the origins of signal variance in FMRI of the human midbrain at high field. PLoS ONE 8:e62708–e62714. https://doi.org/10.1371/journal.pone.0062708
Barth M, Poser BA (2011) Advances in high-field BOLD fMRI. Materials 4:1941–1955. https://doi.org/10.3390/ma4111941
Batson MA, Petridou N, Klomp DWJ et al (2015) Single session imaging of cerebellum at 7 T: obtaining structure and function of multiple motor subsystems in individual subjects. PLoS ONE 10:e0134933–e0134925. https://doi.org/10.1371/journal.pone.0134933
Beisteiner R, Robinson S, Wurnig M et al (2011) Clinical fMRI: Evidence for a 7T benefit over 3T. NeuroImage 57:1015–1021. https://doi.org/10.1016/j.neuroimage.2011.05.010
Benjamin P, Viessmann O, MacKinnon AD et al (2015) 7 T MRI in cerebral small vessel disease. Int J Stroke 10:659–664. https://doi.org/10.1111/ijs.12490
Betts MJ, Acosta-Cabronero J, Cardenas-Blanco A et al (2016) High-resolution characterisation of the aging brain using simultaneous quantitative susceptibility mapping (QSM) and R2* measurements at 7T. NeuroImage 138:43–63. https://doi.org/10.1016/j.neuroimage.2016.05.024
Beuls E, Gelan J, Vandersteen M et al (1993) Microanatomy of the excised human spinal cord and the cervicomedullary junction examined with high-resolution MR imaging at 9.4 T. AJNR Am J Neuroradiol 14:699–707
Beuls E, Vanormelingen L, van Aalst J et al (2003) The Arnold-Chiari type II malformation at midgestation. Pediatr Neurosurg 39:149–158. https://doi.org/10.1159/000071653
Bianciardi M, Toschi N, Edlow BL et al (2015) Toward an in vivoneuroimaging template of human brainstem nuclei of the ascending arousal, autonomic, and motor systems. Brain Connect 5:597–607. https://doi.org/10.1089/brain.2015.0347
Bianciardi M, Strong C, Toschi N et al (2017) A probabilistic template of human mesopontine tegmental nuclei from in vivo 7T MRI. NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.04.070
Blazejewska AI, Schwarz ST, Pitiot A, Stephenson MC (2013) Visualization of nigrosome 1 and its loss in PD pathoanatomical correlation and in vivo 7 T MRI. Neurology 81:534–540. https://doi.org/10.1212/wnl.0b013e31829e6fd2
Blazejewska AI, Al-Radaideh AM, Wharton S et al (2014) Increase in the iron content of the substantia nigra and red nucleus in multiple sclerosis and clinically isolated syndrome: A 7 T MRI study. J Magn Reson Imag 41:1065–1070. https://doi.org/10.1002/jmri.24644
Bogovic JA, Prince JL, Bazin P-L (2013) A multiple object geometric deformable model for image segmentation. Comput Vis Image Underst 117:145–157. https://doi.org/10.1016/j.cviu.2012.10.006
Bourekas EC, Christoforidis GA (1999) High resolution MRI of the deep gray nuclei at 8 T. J Comput Assist Tomogr 23:867–874. https://doi.org/10.1097/00004728-199911000-00009
Bouvy WH, Biessels GJ, Kuijf HJ, Kappelle LJ (2014) Visualization of perivascular spaces and perforating arteries with 7 T magnetic resonance imaging. Invest Radiol 49:307–313. https://doi.org/10.1097/rli.0000000000000027
Bouvy WH, Zwanenburg JJ, Reinink R et al (2016) Perivascular spaces on 7 T brain MRI are related to markers of small vessel disease but not to age or cardiovascular risk factors. J Cereb Blood Flow Metab 36:1708–1717. https://doi.org/10.1177/0271678X16648970
Budde J, Shajan G, Hoffmann J et al (2010) Human imaging at 9.4 T using T2*-, phase-, and susceptibility-weighted contrast. Magn Reson Med 65:544–550. https://doi.org/10.1002/mrm.22632
Budde J, Shajan G, Scheffler K, Pohmann R (2014) Ultra-high resolution imaging of the human brain using acquisition-weighted imaging at 9.4T. NeuroImage 86:592–598. https://doi.org/10.1016/j.neuroimage.2013.08.013
Budinger TF, Bird MD, Frydman L et al (2016) Toward 20 T magnetic resonance for human brain studies: opportunities for discovery and neuroscience rationale. Magn Reson Mater Phy 29:617–639. https://doi.org/10.1007/s10334-016-0561-4
Cabezas M, Oliver A, Lladó X et al (2011) A review of atlas-based segmentation for magnetic resonance brain images. Comput Methods Programs Biomed 104:e158–e177. https://doi.org/10.1016/j.cmpb.2011.07.015
Cakmakli GY, Oruckaptan H, Saka E, Elibol B (2009) Reversible acute cognitive dysfunction induced by bilateral STN stimulation. J Neurol 256:1360–1362. https://doi.org/10.1007/s00415-009-5103-9
Calamante F, Oh S-H, Tournier J-D et al (2012) Super-resolution track-density imaging of thalamic substructures: comparison with high-resolution anatomical magnetic resonance imaging at 7.0T. Hum Brain Mapp 34:2538–2548. https://doi.org/10.1002/hbm.22083
Chalifoux JR, Perry N, Katz JS, Wiggins GC (2013) The ability of high field strength 7-T magnetic resonance imaging to reveal previously uncharacterized brain lesions in patients with tuberous sclerosis complex. J Neurosurg 11:268–273. https://doi.org/10.3171/2012.12.peds12338
Chandran AS, Bynevelt M, Lind CRP (2015) Magnetic resonance imaging of the subthalamic nucleus for deep brain stimulation. J Neurosurg 124:96–105. https://doi.org/10.3171/2015.1.JNS142066
Chavhan GB, Babyn PS, Thomas B et al (2009) Principles, techniques, and applications of T2*-based MR imaging and its special applications. RadioGraphics 29:1433–1449. https://doi.org/10.1148/rg.295095034
Chen Z, Johnston LA, Kwon D-H et al (2010) An optimised framework for reconstructing and processing MR phase images. NeuroImage 49:1289–1300. https://doi.org/10.1016/j.neuroimage.2009.09.071
Chilla GS, Tan CH, Xu C, Poh CL (2015) Diffusion weighted magnetic resonance imaging and its recent trend—a survey. Quant Imag Med Surg. https://doi.org/10.3978/j.issn.2223-4292.2015.03.01
Cho Z-H (2016) Review of recent advancement of ultra high field magnetic resonance imaging: from anatomy to tractography. Investig Magn Reson Imag 20:11–141. https://doi.org/10.13104/imri.2016.20.3.141
Cho Z-H, Kim Y-B, Han J-Y et al (2008a) New brain atlas—mapping the human brain in vivo with 7.0 T MRI and comparison with postmortem histology: Will these images change modern medicine? Int J Imaging Syst Technol 18:2–8. https://doi.org/10.1002/ima.20143
Cho ZH, Kim YB, Han JY et al (2008b) New brain atlas—mapping the human brain in vivo with 7.0 T MRI and comparison with postmortem histology: will these images change modern medicine? Int J Imag Syst Technol 18:2–8
Cho Z-H, Han J-Y, Hwang S-I et al (2010a) Quantitative analysis of the hippocampus using images obtained from 7.0 T MRI. NeuroImage 49:2134–2140. https://doi.org/10.1016/j.neuroimage.2009.11.002
Cho ZH, Min HK, Oh SH et al (2010b) Direct visualization of deep brain stimulation targets in Parkinson disease with the use of 7-tesla magnetic resonance imaging. J Neurosurg 113:1–9
Cho Z-H, Choi S-H, Chi J-G, Kim Y-B (2011a) Classification of the venous architecture of the pineal gland by 7T MRI. J Neuroradiol 38:238–241. https://doi.org/10.1016/j.neurad.2011.02.010
Cho ZH, Kim JM, Park SY et al (2011b) Direct visualization of Parkinson’s disease by in vivo human brain imaging using 7.0T magnetic resonance imaging. Mov Disord 26:713–718. https://doi.org/10.1002/mds.23465
Cho ZH, Son YD, Kim HK et al (2011c) Observation of glucose metabolism in the thalamic nuclei by fusion PET/MRI. J Nucl Med 52:401–404. https://doi.org/10.2967/jnumed.110.081281
Christoforidis GA, Bourekas EC, Baujan M (1999) High resolution MRI of the deep brain vascular anatomy at 8 T: susceptibility-based enhancement of the venous structures. J Comput Assist Tomogr 23:857–866. https://doi.org/10.1097/00004728-199911000-00008
Cock PJA, Antao T, Chang JT et al (2009) Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25:1422–1423. https://doi.org/10.1093/bioinformatics/btp163
Cohen-Adad J, Polimeni JR, Helmer KG et al (2012) T2* mapping and B0 orientation-dependence at 7T reveal cyto- and myeloarchitecture organization of the human cortex. NeuroImage 60:1006–1014. https://doi.org/10.1016/j.neuroimage.2012.01.053
Cosottini M, Frosini D, Pesaresi I et al (2014) MR imaging of the Substantia Nigra at 7 T enables diagnosis of Parkinson disease. Radiology 271:831–838. https://doi.org/10.1148/radiol.14131448
Cosottini M, Frosini D, Pesaresi I et al (2015) Comparison of 3T and 7T susceptibility-weighted angiography of the substantia nigra in diagnosing Parkinson disease. Brain 36:461–466. https://doi.org/10.3174/ajnr.A4158
Costagli M, Symms MR, Angeli L et al (2015) Assessment of silent T1-weighted head imaging at 7 T. Eur Radiol 26:1879–1888. https://doi.org/10.1007/s00330-015-3954-2
Daniluk S, Davies G, Ellias K SA, et al (2009) Assessment of the variability in the anatomical position and size of the subthalamic nucleus among patients with advanced Parkinson’s disease using magnetic resonance imaging. Acta Neurochir 152:201–210. https://doi.org/10.1007/s00701-009-0514-z
De Reuck J, Caparros-Lefebvre D (2014) Prevalence of small cerebral bleeds in patients with progressive supranuclear palsy: a neuropathological study with 7.0-Tesla magnetic resonance imaging correlates. Folia Neuropathol. https://doi.org/10.5114/fn.2014.47843
De Martino F, Moerel M, van de Moortele P-F et al (2013) Spatial organization of frequency preference and selectivity in the human inferior colliculus. Nat Commun 4:1386. https://doi.org/10.1038/ncomms2379
De Reuck JL, Deramecourt V, Auger F et al (2014) Iron deposits in post-mortem brains of patients with neurodegenerative and cerebrovascular diseases: a semi-quantitative 7.0 T magnetic resonance imaging study. Eur J Neurol 21:1026–1031. https://doi.org/10.1111/ene.12432
de Rotte AAJ, van der Kolk AG, Rutgers D et al (2014) Feasibility of high-resolution pituitary MRI at 7.0 T. Eur Radiol 24:2005–2011. https://doi.org/10.1007/s00330-014-3230-x
De Reuck JL, Deramecourt V, Auger F et al (2015) The significance of cortical cerebellar microbleeds and microinfarcts in neurodegenerative and cerebrovascular diseases. Cerebrovasc Dis 138–143. https://doi.org/10.1159/000371488
de Rotte AAJ, Groenewegen A, Rutgers DR et al (2015) High resolution pituitary gland MRI at 7.0 T: a clinical evaluation in Cushing’s disease. Eur Radiol 26:271–277. https://doi.org/10.1007/s00330-015-3809-x
de Hollander G, Keuken MC, van der Zwaag W et al (2017) Comparing functional MRI protocols for small, iron-rich basal ganglia nuclei such as the subthalamic nucleus at 7 T and 3 T. Hum Brain Mapp 38:3226–3248. https://doi.org/10.1002/hbm.23586
De Reuck J, Auger F, Durieux N et al (2017) Frequency and topography of small cerebrovascular lesions in vascular and in mixed dementia: a post-mortem 7-tesla magnetic resonance imaging study with neuropathological correlates. Folia Neuropathol 1:31–37. https://doi.org/10.5114/fn.2017.66711
Deistung A, Schäfer A, Schweser F et al (2013a) High-resolution MR imaging of the human brainstem in vivo at 7 T. Front Hum Neurosci. https://doi.org/10.3389/fnhum.2013.00710
Deistung A, Schäfer A, Schweser F et al (2013b) Toward in vivo histology: a comparison of quantitative susceptibility mapping (QSM) with magnitude-, phase-, and R2*-imaging at ultra-high magnetic field strength. 65:299–314. https://doi.org/10.1016/j.neuroimage.2012.09.055
Denison RN, Vu AT, Yacoub E et al (2014) Functional mapping of the magnocellular and parvocellular subdivisions of human LGN. Neuroimage 102:358–369. https://doi.org/10.1016/j.neuroimage.2014.07.019
Derix J, Yang S, Lüsebrink F et al (2014) Visualization of the amygdalo-hippocampal border and its structural variability by 7T and 3T magnetic resonance imaging. Hum Brain Mapp 35:4316–4329. https://doi.org/10.1002/hbm.22477
Devlin JT, Poldrack RA (2007) In praise of tedious anatomy. NeuroImage 37:1033–1041. https://doi.org/10.1016/j.neuroimage.2006.09.055
Dezortova M, Herynek V, Krssak M et al (2012) Two forms of iron as an intrinsic contrast agent in the basal ganglia of PKAN patients. Contrast Media Mol Imag 7:509–515. https://doi.org/10.1002/cmmi.1482
Di Ieva A, Tschabitscher M, Galzio RJ et al (2011) The veins of the nucleus dentatus: anatomical and radiological findings. NeuroImage 54:74–79. https://doi.org/10.1016/j.neuroimage.2010.07.045
Diedrichsen J, Maderwald S, Küper M et al (2011) Imaging the deep cerebellar nuclei: a probabilistic atlas and normalization procedure. NeuroImage 54:1786–1794. https://doi.org/10.1016/j.neuroimage.2010.10.035
Ding L, Gold JI (2013) The basal ganglia’s contributions to perceptual decision making. Neuron 79:640–649. https://doi.org/10.1016/j.neuron.2013.07.042
Ding S-L, Royall JJ, Sunkin SM et al (2016) Comprehensive cellular-resolution atlas of the adult human brain. J Comp Neurol. https://doi.org/10.1002/cne.24080
Dinse J, Härtwich N, Waehnert MD et al (2015) A cytoarchitecture-driven myelin model reveals area-specific signatures in human primary and secondary areas using ultra-high resolution in-vivo brain MRI. NeuroImage 114:71–87. https://doi.org/10.1016/j.neuroimage.2015.04.023
Doan NT, Orban de Xivry J, Macq B (2010) Effect of inter-subject variation on the accuracy of atlas-based segmentation applied to human brain structures. In: Dawant BM, Haynor DR (eds) SPIE, 76231S–76231S11
Dortch RD, Moore J, Li K et al (2013) Quantitative magnetization transfer imaging of human brain at 7T. NeuroImage 64:640–649. https://doi.org/10.1016/j.neuroimage.2012.08.047
Duchin Y, Abosch A, Yacoub E et al (2012) Feasibility of using ultra-high field (7 T) MRI for clinical surgical targeting. PLoS ONE 7:e37328–e37310. https://doi.org/10.1371/journal.pone.0037328
Dula AN, Welch EB, Creasy JL et al (2010) Challenges and opportunities of ultra-high field MRI. In: Van Toi V, Khoa TQD (eds) The Third International Conference on the Development of Biomedical Engineering in Vietnam. Springer Berlin Heidelberg, Berlin, Heidelberg, pp 1–5
Dumoulin SO, Fracasso A, van der Zwaag W et al (2017) Ultra-high field MRI_ Advancing systems neuroscience towards mesoscopic human brain function. NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.01.028
Dunbar RIM (1992) Neocortex size as a constraint on group size in primates. J Hum Evol 22:469–493. https://doi.org/10.1016/0047-2484(92)90081-j
Dunnen Den WF, Staal MJ (2005) Anatomical alterations of the subthalamic nucleus in relation to age: a postmortem study. Mov Disord 20:893–898. https://doi.org/10.1002/mds.20417
Duyn JH (2010) Study of brain anatomy with high-field MRI: recent progress. Magn Reson Imaging 28:1210–1215. https://doi.org/10.1016/j.mri.2010.02.007
Duyn JH (2012) The future of ultra-high field MRI and fMRI for study of the human brain. NeuroImage 62:1241–1248. https://doi.org/10.1016/j.neuroimage.2011.10.065
Eapen M, Zald DH, Gatenby JC et al (2011) Using high-resolution MR imaging at 7T to evaluate the anatomy of the midbrain dopaminergic system. AJNR Am J Neuroradiol 32:688–694. https://doi.org/10.3174/ajnr.A2355
Emir UE, Tuite PJ, Öz G (2012) Elevated pontine and putamenal GABA levels in mild-moderate parkinson disease detected by 7 T proton MRS. PLoS ONE 7:e30918–e30918. https://doi.org/10.1371/journal.pone.0030918
Faull OK, Jenkinson M, Clare S, Pattinson KTS (2015) Functional subdivision of the human periaqueductal grey in respiratory control using 7 T fMRI. NeuroImage 113:356–364. https://doi.org/10.1016/j.neuroimage.2015.02.026
Federative Committee on Anatomical Terminology (1998) Terminologia Anatomica, Thieme Stuttgart
Federau C, Gallichan D (2016) Motion-correction enabled ultra-high resolution in-vivo 7T-MRI of the brain. PLoS ONE 11:e0154974–e0154912. https://doi.org/10.1371/journal.pone.0154974
Foroutan P, Murray ME, Fujioka S et al (2013) Progressive supranuclear palsy: high-field-strength MR microscopy in the human Substantia Nigra and globus pallidus. Radiology 266:280–288. https://doi.org/10.1148/radiol.12102273
Forstmann BU, Anwander A, Schäfer A et al (2010) Cortico-striatal connections predict control over speed and accuracy in perceptual decision making. Proc Natl Acad Sci 107:15916–15920. https://doi.org/10.1073/pnas.1004932107
Forstmann BU, Keuken MC, Jahfari S et al (2012) Cortico-subthalamic white matter tract strength predict interindividual efficacy in stopping a motor response. NeuroImage 60:370–375
Forstmann BU, Keuken MC, Schäfer A et al (2014) Multi-modal ultra-high resolution structural 7-Tesla MRI data repository. Sci Data 1:140050–140058. https://doi.org/10.1038/sdata.2014.50
Forstmann B, de Hollander G, van Maanen L et al (2017a) Towards a mechanistic understanding of the human subcortex. Nat Rev 18(1):57
Forstmann BU, Isaacs BR, Temel Y (2017b) Ultra-high field MRI guided deep brain stimulation. Trends Biotechnol 35(10):904–907
Fracasso A, van Veluw SJ, Visser F et al (2016) Lines of Baillarger in vivo and ex vivo: Myelin contrast across lamina at 7T MRI and histology. NeuroImage 133:163–175. https://doi.org/10.1016/j.neuroimage.2016.02.072
Francis S, Panchuelo RS (2014) Physiological measurements using ultra-high field fMRI: a review. Physiol Meas 35:R167–R185. https://doi.org/10.1088/0967-3334/35/9/R167
Fritzsch D, Reiss-Zimmermann M, Trampel R (2014) Seven-tesla magnetic resonance imaging in Wilson disease using quantitative susceptibility mapping for measurement of copper accumulation. Invest Radiol 49:299–306. https://doi.org/10.1097/rli.0000000000000010
Frosini D, Ceravolo R, Tosetti M et al (2017) Nigral involvement in atypical parkinsonisms: evidence from a pilot study with ultra-high field MRI. J Neural Transm 123:509–513. https://doi.org/10.1007/s00702-016-1529-2
Fujioka S, Murray ME, Foroutan P et al (2011) Magnetic resonance imaging with 21.1 T and pathological correlations-diffuse Lewy body disease. Rinsho Shinkeigaku 51:603–607. https://doi.org/10.5692/clinicalneurol.51.603
Fukunaga M, Li TQ, van Gelderen P et al (2010) Layer-specific variation of iron content in cerebral cortex as a source of MRI contrast. Proc Natl Acad Sci 107:3834–3839. https://doi.org/10.1073/pnas.0911177107
Gallichan D (2017) Diffusion MRI of the human brain at ultra-high field (UHF)_ A review. NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.04.037
Gallichan D, Marques JP, Gruetter R (2015) Retrospective correction of involuntary microscopic head movement using highly accelerated fat image navigators (3D FatNavs) at 7T. Magn Reson Med 75:1030–1039. https://doi.org/10.1002/mrm.25670
Ghaznawi R, de Bresser J, van der Graaf Y et al (2017) Detection and characterization of small infarcts in the caudate nucleus on 7 T MRI: The SMART-MR study. J Cereb Blood Flow Metab. https://doi.org/10.1177/0271678X17705974
Giuliano A, Donatelli G, Cosottini M et al (2017) Hippocampal subfields at ultra high field MRI: An overview of segmentation and measurement methods. Hippocampus 27:481–494. https://doi.org/10.1002/hipo.22717
Gizewski ER, de Greiff A, Maderwald S et al (2007) fMRI at 7 T: Whole-brain coverage and signal advantages even infratentorially? NeuroImage 37:761–768. https://doi.org/10.1016/j.neuroimage.2007.06.005
Gizewski ER, Maderwald S, Linn J et al (2013) High-resolution anatomy of the human brain stem using 7-T MRI: improved detection of inner structures and nerves? Neuroradiology 56:177–186. https://doi.org/10.1007/s00234-013-1312-0
Gizewski ER, Mönninghoff C, Forsting M (2015) Perspectives of ultra-high-field MRI in neuroradiology. Clin Neuroradiol. https://doi.org/10.1007/s00062-015-0437-4
Gorka AX, Torrisi S, Shackman AJ et al (2017) Intrinsic functional connectivity of the central nucleus of the amygdala and bed nucleus of the stria terminalis. NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.03.007
Grabner G, Poser BA, Fujimoto K et al (2014) A study-specific fMRI normalization approach that operates directly on high resolution functional EPI data at 7 T. NeuroImage 100:710–714. https://doi.org/10.1016/j.neuroimage.2014.06.045
Grossman RI, GOMORI JM, RAMER KN et al (1994) Magnetization-transfer—theory and clinical-applications in neuroradiology. RadioGraphics 14:279–290. https://doi.org/10.1148/radiographics.14.2.8190954
Haacke EM, Mittal S, Wu Z et al (2008) Susceptibility-weighted imaging: technical aspects and clinical applications, part 1. Am J Neuroradiol 30:19–30. https://doi.org/10.3174/ajnr.A1400
Haber SN, Calzavara R (2009) The cortico-basal ganglia integrative network: the role of the thalamus. Brain Res Bull 78:69–74
Hallgren B, Sourander P (1958) The effect of age on the non-haemin iron in the human brain. J Neurochem 3:41–51
Hammond KE, Lupo JM, Xu D et al (2008a) Development of a robust method for generating 7.0 T multichannel phase images of the brain with application to normal volunteers and patients with neurological diseases. NeuroImage 39:1682–1692. https://doi.org/10.1016/j.neuroimage.2007.10.037
Hammond KE, Metcalf M, Carvajal L et al (2008b) Quantitative in vivo magnetic resonance imaging of multiple sclerosis at 7 T with sensitivity to iron. Ann Neurol 64:707–713. https://doi.org/10.1002/ana.21582
Healthineers S (2017a) With 7 T scanner Magnetom Terra, Siemens Healthineers introduces new clinical field strength in MR imaging. pp 1–4
Healthineers S (2017b) FDA Clears MAGNETOM Terra 7T MRI Scanner From Siemens Healthineers. pp 1–2
Heimbach S (2015) New 7 T MRI research system ready for future clinical use. pp 1–3
Herbst M, Maclaren J, Lovell-Smith C et al (2013) Reproduction of motion artifacts for performance analysis of prospective motion correction in MRI. Magn Reson Med 71:182–190. https://doi.org/10.1002/mrm.24645
Hollander G, Keuken MC, Bazin P-L et al (2014) A gradual increase of iron toward the medial-inferior tip of the subthalamic nucleus. Hum Brain Mapp 35:4440–4449. https://doi.org/10.1002/hbm.22485
Johansen-Berg H (2013) Human connectomics—what will the future demand? NeuroImage 80:541–544. https://doi.org/10.1016/j.neuroimage.2013.05.082
Jones DK, Knösche TR, Turner R (2013) White matter integrity, fiber count, and other fallacies: the do“s and don”ts of diffusion MRI. NeuroImage 73:239–254. https://doi.org/10.1016/j.neuroimage.2012.06.081
Kanowski M, Voges J, Buentjen L et al (2014) Direct visualization of anatomic subfields within the superior aspect of the human lateral thalamus by MRI at 7T. Am J Neuroradiol 35:1721–1727. https://doi.org/10.3174/ajnr.A3951
Kemper VG, De Martino F, Emmerling TC et al (2017) High resolution data analysis strategies for mesoscale human functional MRI at 7 and 9.4T. NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.03.058
Keren NI, Taheri S, Vazey EM et al (2015) Histologic validation of locus coeruleus MRI contrast in post-mortem tissue. 113:235–245. https://doi.org/10.1016/j.neuroimage.2015.03.020
Kerl HU (2013) Imaging for deep brain stimulation: The zona incerta at 7 T. WJR 5:5–12. https://doi.org/10.4329/wjr.v5.i1.5
Kerl HU, Gerigk L, Pechlivanis I et al (2012) The subthalamic nucleus at 7.0 T: evaluation of sequence and orientation for deep-brain stimulation. Acta Neurochir 154:2051–2062. https://doi.org/10.1007/s00701-012-1476-0
Keuken MC, Bazin PL, Schäfer A et al (2013) Ultra-high 7T MRI of structural age-related changes of the subthalamic nucleus. J Neurosci 33:4896–4900. https://doi.org/10.1523/JNEUROSCI.3241-12.2013
Keuken MC, Bazin PL, Crown L et al (2014) Quantifying inter-individual anatomical variability in the subcortex using 7T structural MRI. NeuroImage 94:40–46. https://doi.org/10.1016/j.neuroimage.2014.03.032
Keuken MC, van Maanen L, Bogacz R et al (2015) The subthalamic nucleus during decision-making with multiple alternatives. Hum Brain Map 36:4041–4052. https://doi.org/10.1002/hbm.22896
Keuken MC, Bazin PL, backhouse K et al (2017) Effects of aging on T1, T2*, and QSM MRI values in the subcortex. Brain Struct Funct. https://doi.org/10.1007/s00429-016-1352-4
Khabipova D, Wiaux Y, Gruetter R, Marques JP (2015) A modulated closed form solution for quantitative susceptibility mapping—a thorough evaluation and comparison to iterative methods based on edge prior knowledge. NeuroImage 107:163–174. https://doi.org/10.1016/j.neuroimage.2014.11.038
Kim NR, Chi JG, Choi SH, Kim YB (2011) Identification and morphologic assessment of mesocoelic recess by in vivo human brain imaging with 7.0-T magnetic resonance imaging. J Comput Assist Tomogr 35:486–491. https://doi.org/10.1097/rct.0b013e31821de1cc
Kim J, Lenglet C, Duchin Y et al (2014) Semiautomatic segmentation of brain subcortical structures from high-field MRI. IEEE J Biomed Health Inform 18:1678–1695. https://doi.org/10.1109/JBHI.2013.2292858
Kim J-H, Son Y-D, Kim J-H et al (2015a) Self-transcendence trait and its relationship with in vivo serotonin transporter availability in brainstem raphe nuclei_ An ultra-high resolution PET-MRI study. Brain Res 1629:63–71. https://doi.org/10.1016/j.brainres.2015.10.006
Kim J-H, Son Y-D, Kim J-H et al (2015b) Serotonin transporter availability in thalamic subregions in schizophrenia_ A study using 7.0-T MRI with [11C]DASB high-resolution PET. Psychiatr Res 231:50–57. https://doi.org/10.1016/j.pscychresns.2014.10.022
Kim J-M, Jeong H-J, Bae YJ et al (2016) Loss of substantia nigra hyperintensity on 7 T MRI of Parkinson’s disease, multiple system atrophy, and progressive supranuclear palsy. Parkinsonism Relat Disord 26:47–54. https://doi.org/10.1016/j.parkreldis.2016.01.023
Kim J-H, Kim J-H, Son Y-D et al (2017a) Altered interregional correlations between serotonin transporter availability and cerebral glucose metabolism in schizophrenia: a high-resolution PET study using [11C]DASB and [18F]FDG. Schizophr Res 182:55–65. https://doi.org/10.1016/j.schres.2016.10.020
Kim JH, Son YD, Kim JM et al (2017b) Interregional correlations of glucose metabolism between the basal ganglia and different cortical areas: an ultra-high resolution PET/MRI fusion study using 18F-FDG. Braz J Med Biol Res 51:a009621–7. https://doi.org/10.1590/1414-431x20176724
Kirov II, Hardy CJ, Matsuda K et al (2013) In vivo 7 T imaging of the dentate granule cell layer in schizophrenia. Schizophr Res 147:362–367. https://doi.org/10.1016/j.schres.2013.04.020
Kitajima M, Korogi Y, Kakeda S et al (2008) Human subthalamic nucleus: evaluation with high-resolution MR imaging at 3.0 T. Neuroradiology 50:675–681. https://doi.org/10.1007/s00234-008-0388-4
Kleiner-Fisman G, Herzog J, Fisman DN et al (2006) Subthalamic nucleus deep brain stimulation: summary and meta-analysis of outcomes. Mov Disord 21:S290–S304. https://doi.org/10.1002/mds.20962
Koenig SH (1991) Cholesterol of myelin is the determinant of gray-white contrast in MRI of brain. Magn Resonan Med 20:285–291. https://doi.org/10.1002/mrm.1910200210
Kollia K, Maderwald S, Putzki N et al (2009) First clinical study on ultra-high-field MR imaging in patients with multiple sclerosis: comparison of 1.5T and 7T. AJNR Am J Neuroradiol 30:699–702. https://doi.org/10.3174/ajnr.A1434
Kraff O, Fischer A, Nagel AM et al (2014) MRI at 7 T and above: demonstrated and potential capabilities. J Magn Reson Imaging 41:13–33. https://doi.org/10.1002/jmri.24573
Küper M, Dimitrova A, Thürling M et al (2011a) Evidence for a motor and a non-motor domain in the human dentate nucleus—an fMRI study. NeuroImage 54:2612–2622. https://doi.org/10.1016/j.neuroimage.2010.11.028
Küper M, Thürling M, Stefanescu R et al (2011b) Evidence for a motor somatotopy in the cerebellar dentate nucleus-an FMRI study in humans. Hum Brain Mapp 33:2741–2749. https://doi.org/10.1002/hbm.21400
Küper M, Wünnemann MJS, Thürling M et al (2013) Activation of the cerebellar cortex and the dentate nucleus in a prism adaptation fMRI study. Hum Brain Map 35:1574–1586. https://doi.org/10.1002/hbm.22274
Kwon D-H, Kim J-M, Oh S-H et al (2012) Seven-tesla magnetic resonance images of the substantia nigra in Parkinson disease. Ann Neurol 71:267–277. https://doi.org/10.1002/ana.22592
Larkman DJ (2007) The g-Factor and Coil Design. In: Parallel imaging in clinical MR applications. Springer Berlin Heidelberg, Berlin, pp 37–48
Lee J, Shmueli K, Fukunaga M et al (2010) Sensitivity of MRI resonance frequency to the orientation of brain tissue microstructure. Proc Natl Acad Sci USA 107:5130–5135. https://doi.org/10.1073/pnas.0910222107
Lee JY, Jeong H-J, Lee JH et al (2014) An investigation of lateral geniculate nucleus volume in patients with primary open-angle glaucoma using 7 T magnetic resonance imaging. Invest Ophthalmol Vis Sci 55:3468–3469. https://doi.org/10.1167/iovs.14-13902
Lenglet C, Abosch A, Yacoub E et al (2012) Comprehensive in vivo mapping of the human basal ganglia and thalamic connectome in individuals using 7T MRI. PLoS ONE 7:e29153. https://doi.org/10.1371/journal.pone.0029153
Lenroot RK, Giedd JN (2008) The changing impact of genes and environment on brain development during childhood and adolescence: Initial findings from a neuroimaging study of pediatric twins. Dev Psychopathol 20:1120–1161. https://doi.org/10.1017/S0954579408000552
Liem MK, Lesnik Oberstein SAJ, Versluis MJ et al (2012) 7T MRI reveals diffuse iron deposition in putamen and caudate nucleus in CADASIL. J Neurol Neurosurg Psychiat 83:1180–1185. https://doi.org/10.1136/jnnp-2012-302545
Limousin P, Pollak P, Benazzouz A et al (1995) Effect of parkinsonian signs and symptoms of bilateral subthalamic nucleus stimulation. Lancet 345:91–95
Liu F, Zhang Z, Lin X et al (2011) Development of the human fetal cerebellum in the second trimester: a post mortem magnetic resonance imaging evaluation. J Anat 219:582–588. https://doi.org/10.1111/j.1469-7580.2011.01418.x
Liu T, Eskreis-Winkler S, Schweitzer AD, Chen W (2013) Improved subthalamic nucleus depiction with quantitative susceptibility mapping. Radiology. https://doi.org/10.1148/radiol.13121991/-/DC1
Liu C, Li W, Tong KA et al (2014) Susceptibility-weighted imaging and quantitative susceptibility mapping in the brain. J Magn Reson Imaging 42:23–41. https://doi.org/10.1002/jmri.24768
Lorio S, Lutti A, Kherif F et al (2014) Disentangling in vivo the effects of iron content and atrophy on the ageing human brain. NeuroImage 103:280–289. https://doi.org/10.1016/j.neuroimage.2014.09.044
Lotfipour AK, Wharton S, Schwarz ST et al (2011) High resolution magnetic susceptibility mapping of the substantia nigra in Parkinson’s disease. J Magn Reson Imaging 35:48–55. https://doi.org/10.1002/jmri.22752
Lüsebrink F, Wollrab A, Speck O (2013) Cortical thickness determination of the human brain using high resolution 3T and 7T MRI data. NeuroImage 70:122–131. https://doi.org/10.1016/j.neuroimage.2012.12.016
Lutti A, Dick F, Sereno MI, Weiskopf N (2014) Using high-resolution quantitative mapping of R1 as an index of cortical myelination. NeuroImage. https://doi.org/10.1016/j.neuroimage.2013.06.005
Maclaren J, Herbst M, Speck O, Zaitsev M (2012) Prospective motion correction in brain imaging: a review. Magn Reson Med 69:621–636. https://doi.org/10.1002/mrm.24314
Makris N, Swaab DF, van der Kouwe A et al (2013a) Volumetric parcellation methodology of the human hypothalamus in neuroimaging: normative data and sex differences. NeuroImage 69:1–10. https://doi.org/10.1016/j.neuroimage.2012.12.008
Marques JP, Gruetter R (2013) New developments and applications of the MP2RAGE sequence—focusing the contrast and high spatial resolution R1 mapping. PLoS ONE 8:e69294–e69211. https://doi.org/10.1371/journal.pone.0069294
Marques JP, Norris DG (2017) How to choose the right MR sequence for your research question at 7T and above? NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.04.044
Marques JP, van der Zwaag W, Granziera C et al (2010) Cerebellar cortical layers: in vivo visualization with structural high-field-strength MR imaging 1. Radiology 254:942–948. https://doi.org/10.1148/radiol.09091136
Marques JP, Khabipova D, Gruetter R (2017) Studying cyto and myeloarchitecture of the human cortex at ultra-high field with quantitative imaging_ R1, R2* and magnetic susceptibility. NeuroImage 147:152–163. https://doi.org/10.1016/j.neuroimage.2016.12.009
Marrakchi-Kacem L, Vignaud A, Sein J et al (2016) Robust imaging of hippocampal inner structure at 7T: in vivo acquisition protocol and methodological choices. Magn Reson Mater Phys 29:475–489. https://doi.org/10.1007/s10334-016-0552-5
Massey LA, Miranda MA, Zrinzo L, Al-Helli O (2012) High resolution MR anatomy of the subthalamic nucleus: imaging at 9.4 T with histological validation. NeuroImage 59:2035–2044. https://doi.org/10.1016/j.neuroimage.2011.10.016
Mavridis I, Boviatsis E, Anagnostopoulou S (2014) Stereotactic anatomy of the human subthalamic nucleus: providing coordinates for accurate electrode placement. J Neurol Surg A Cent Eur Neurosurg 75:289–298. https://doi.org/10.1055/s-0034-1368093
Mazziotta JC, Toga AW, Evans A et al (1995) A probabilistic atlas of the human brain: theory and rationale for its development the international consortium for brain mapping (ICBM). NeuroImage 2:89–101
McRobbie DW, Moore EA, Graves MJ, Prince MR (2006) MRI from Picture to Proton, 1st edn. Cambridge University Press, Cambridge
Meijer FJA, Steens SC, van Rumund A et al (2016) Nigrosome-1 on susceptibility weighted imaging to differentiate Parkinson’s disease From Atypical Parkinsonism: an in vivo and ex vivo pilot study. Pol J Radiol 81:363–369. https://doi.org/10.12659/PJR.897090
Meng H, Zhang Z, Geng H et al (2012) Development of the subcortical brain structures in the second trimester: assessment with 7.0-T MRI. Neuroradiology 54:1153–1159. https://doi.org/10.1007/s00234-012-1069-x
Mestres-Missé A, Bazin P-L, Trampel R et al (2014) Dorsomedial striatum involvement in regulating conflict between current and presumed outcomes. NeuroImage 98:159–167. https://doi.org/10.1016/j.neuroimage.2014.05.002
Middleton FA, Strick PL (2000a) Basal ganglia and cerebellar loops: motor and cognitive circuits. Brain Res Rev 31:236–250. https://doi.org/10.1016/s0165-0173(99)00040-5
Middleton FA, Strick PL (2000b) Basal ganglia output and cognition: evidence from anatomical, behavioral, and clinical studies. Brain Cogn 42:183–200. https://doi.org/10.1006/brcg.1999.1099
Miller MI, Younes L, Ratnanather JT, Brown T (2015) Amygdalar atrophy in symptomatic Alzheimer’s disease based on diffeomorphometry: the BIOCARD cohort. Neurobiol Aging 36:S3–S10. https://doi.org/10.1016/j.neurobiolaging.2014.06.032
Minati L, Grisoli M, Carella F (2007) Imaging degeneration of the substantia nigra in Parkinson disease with inversion-recovery MR imaging. Am J Neuroradiol 28:309–313. https://doi.org/10.1016/j.acra.2006.01.049
Mitsumori F, Watanabe H, Takaya N et al (2011) Toward understanding transverse relaxation in human brain through its field dependence. Magn Reson Med 68:947–953. https://doi.org/10.1002/mrm.23301
Moenninghoff C, Kraff O, Schlamann M et al (2010) Assessing a dysplastic cerebellar gangliocytoma (Lhermitte-Duclos disease) with 7T MR imaging. Korean J Radiol 11:244–245. https://doi.org/10.3348/kjr.2010.11.2.244
Moerel M, De Martino F, Ugurbil K et al (2015) Processing of frequency and location in human subcortical auditory structures. Sci Rep. https://doi.org/10.1038/srep17048
Mollink J, Baarsen KM, Dederen PJWC. et al (2016) Dentatorubrothalamic tract localization with postmortem MR diffusion tractography compared to histological 3D reconstruction. Brain Struct Funct 1–15. https://doi.org/10.1007/s00429-015-1115-7
Nichols TE, Das S, Eickhoff SB et al (2016) Best practices in data analysis and sharing in neuroimaging using MRI. Nature Neurosci. https://doi.org/10.1101/054262
Novak P, Novak V, Kangarlu A et al (2001a) High resolution MRI of the brainstem at 8 T. J Comput Assist Tomogr 25:242–246. https://doi.org/10.1097/00004728-200103000-00016
Novak V, Abduljalil A, Kangarlu A, Slivka A (2001b) Intracranial ossifications and microangiopathy at 8 T MRI. Magn Reson Imag 19:1133–1137. https://doi.org/10.1016/s0730-725x(01)00447-7
O’Brien KR, Kober T, Hagmann P et al (2014) Robust T1-weighted structural brain imaging and morphometry at 7T using MP2RAGE. PLoS ONE 9:e99676–e99677. https://doi.org/10.1371/journal.pone.0099676
Paek SH, Lee J-Y, Kim H-J et al (2011) Electrode position and the clinical outcome after bilateral subthalamic nucleus stimulation. J Korean Med Sci 26:1344. https://doi.org/10.3346/jkms.2011.26.10.1344
Pereira JLB, Furie S, Sharim J et al (2016) Lateralization of the subthalamic nucleus with age in Parkinson’s disease. Basal Ganglia 6:83–88. https://doi.org/10.1016/j.baga.2016.01.003
Peters AM, Brookes MJ, Hoogenraad FG et al (2007) T2* measurements in human brain at 1.5, 3 and 7 T. Magn Reson Imag 25:748–753. https://doi.org/10.1016/j.mri.2007.02.014
Plantinga BR, Temel Y, Roebroeck A et al (2014) Ultra-high field magnetic resonance imaging of the basal ganglia and related structures. Front Hum Neurosci 8:1–22. https://doi.org/10.3389/fnhum.2014.00876/abstract
Plantinga BR, Roebroeck A, Kemper VG et al (2016a) Ultra-high field MRI post mortem structural connectivity of the human subthalamic nucleus, substantia nigra, and globus pallidus. Front Neuroanat 10:110–117. https://doi.org/10.3389/fnana.2016.00066
Plantinga BR, Temel Y, Duchin Y et al (2016b) Individualized parcellation of the subthalamic nucleus in patients with Parkinson’s disease with 7T MRI. NeuroImage. https://doi.org/10.1016/j.neuroimage.2016.09.023
Pohmann R, Speck O, Scheffler K (2015) Signal-to-noise ratio and MR tissue parameters in human brain imaging at 3, 7, and 9.4 T using current receive coil arrays. Magn Reson Med 75:801–809. https://doi.org/10.1002/mrm.25677
Poldrack R, Fletcher P, Henson R et al (2008) Guidelines for reporting an fMRI study. NeuroImage 40:409–414
Priovoulos N, Jacobs HIL, Ivanov D et al (2017) High-resolution in vivo imaging of human locus coeruleus by magnetization transfer MRI at 3T and 7T. NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.07.045
Raz N (2005) Regional brain changes in aging healthy adults: general trends, individual differences and modifiers. Cereb Cortex 15:1676–1689. https://doi.org/10.1093/cercor/bhi044
Renvall V, Witzel T, Wald LL, Polimeni JR (2016) Automatic cortical surface reconstruction of high-resolution T1 echo planar imaging data. NeuroImage 134:338–354. https://doi.org/10.1016/j.neuroimage.2016.04.004
Rijkers K, Temel Y, Visser-Vandewalle V et al (2007) The microanatomical environment of the subthalamic nucleus. J Neurosurg 107:198–201
Rinck P (2016) Magnetic resonance in medicine, 9 edn. Wiley-Blackwell, Hoboken
Robitaille P-M, Berliner L (2007) Ultra high field magnetic resonance imaging. Springer Science & Business Media, NewYork
Robitaille P, Kangarlu A (1999) RF penetration in ultra high field MRI: challenges in visualizing details within the center of the human brain. J Comput Assist Tomogr 23:845–849. https://doi.org/10.1097/00004728-199911000-00006
Romanzetti S, Mirkes CC, Fiege DP et al (2014) Mapping tissue sodium concentration in the human brain: a comparison of MR sequences at 9.4 T. NeuroImage 96:44–53. https://doi.org/10.1016/j.neuroimage.2014.03.079
Rooney WD, Johnson G, Li X et al (2007) Magnetic field and tissue dependencies of human brain longitudinal1H2O relaxation in vivo. Magn Reson Med 57:308–318. https://doi.org/10.1002/mrm.21122
Ropele S, Langkammer C (2016) Iron quantification with susceptibility. NMR Biomed. https://doi.org/10.1002/nbm.3534
Rudko DA, Solovey I, Gati JS et al (2014) Multiple sclerosis: improved identification of disease-relevant changes in gray and white matter by using susceptibility-based MR imaging. Radiology 272:851–864. https://doi.org/10.1148/radiol.14132475
Saranathan M, Tourdias T, Bayram E et al (2014) Optimization of white-matter-nulled magnetization prepared rapid gradient echo (MP-RAGE) imaging. Magn Reson Med 73:1786–1794. https://doi.org/10.1002/mrm.25298
Satpute AB, Wager TD, Cohen-Adad J (2013) Identification of discrete functional subregions of the human periaqueductal gray. Proc Natl Acad Sci 110(42):17101–17106
Schäfer A, Wharton S, Gowland P, Bowtell R (2009) Using magnetic field simulation to study susceptibility-related phase contrast in gradient echo MRI. NeuroImage 48:126–137. https://doi.org/10.1016/j.neuroimage.2009.05.093
Schäfer A, Forstmann BU, Neumann J et al (2012) Direct visualization of the subthalamic nucleus and its iron distribution using high-resolution susceptibility mapping. Hum Brain Mapp 33:2831–2842
Schindler S, Schönknecht P, Schmidt L et al (2013) Development and evaluation of an algorithm for the computer-assisted segmentation of the human hypothalamus on 7-Tesla magnetic resonance images. PLoS ONE 8:e66394–e66398. https://doi.org/10.1371/journal.pone.0066394
Schindler S, Schreiber J, Bazin P-L et al (2017) Intensity standardisation of 7T MR images for intensity-based segmentation of the human hypothalamus. PLoS ONE 12:e0173344–e0173322. https://doi.org/10.1371/journal.pone.0173344
Schmidt FM, Schindler S, Adamidis M et al (2017a) Habenula volume increases with disease severity in unmedicated major depressive disorder as revealed by 7T MRI. Eur Arch Psychiatry Clin Neurosci 267:107–115. https://doi.org/10.1007/s00406-016-0675-8
Schmidt MA, Engelhorn T, Marxreiter F et al (2017b) Ultra high-field SWI of the substantia nigra at 7T: reliability and consistency of the swallow-tail sign. BMC Neurol. https://doi.org/10.1186/s12883-017-0975-2
Schreiner SJ, Liu X, Gietl AF, Wyss M (2014) Regional Fluid-Attenuated Inversion Recovery (FLAIR) at 7 T correlates with amyloid beta in hippocampus and brainstem of cognitively normal elderly. Front Aging Neurosci 6:529. https://doi.org/10.3389/fnagi.2014.00240
Schweser F, Deistung A, Lehr BW, Reichenbach JR (2011) Quantitative imaging of intrinsic magnetic tissue properties using MRI signal phase: an approach to in vivo brain iron metabolism? NeuroImage 54:2789–2807. https://doi.org/10.1016/j.neuroimage.2010.10.070
Schweser F, Deistung A, Reichenbach JR (2016) Foundations of MRI phase imaging and processing for quantitative susceptibility mapping (QSM). Zeitschrift für medizinische Physik 26:6–34. https://doi.org/10.1016/j.zemedi.2015.10.002
Sclocco R, Beissner F, Bianciardi M et al (2017) Challenges and opportunities for brainstem neuroimaging with ultrahigh field MRI. NeuroImage. https://doi.org/10.1016/j.neuroimage.2017.02.052
Setsompop K, Feinberg DA, Polimeni JR (2016) Rapid brain MRI acquisition techniques at ultra-high fields. NMR Biomed 29:1198–1221. https://doi.org/10.1002/nbm.3478
Shmueli K, de Zwart JA, van Gelderen P et al (2009) Magnetic susceptibility mapping of brain tissue in vivo using MRI phase data. Magn Reson Med 62:1510–1522. https://doi.org/10.1002/mrm.22135
Sladky R, Baldinger P, Kranz GS et al (2013) High-resolution functional MRI of the human amygdala at 7T. Eur J Radiol 82:728–733. https://doi.org/10.1016/j.ejrad.2011.09.025
Solano-Castiella E, Schäfer A, Reimer E et al (2011) Parcellation of human amygdala in vivo using ultra high field structural MRI. NeuroImage 58:741–748. https://doi.org/10.1016/j.neuroimage.2011.06.047
Solbach K, Kraff O, Minnerop M et al (2014) Cerebellar pathology in Friedreich’s ataxia: atrophied dentate nuclei with normal iron content. NeuroImage 6:93–99. https://doi.org/10.1016/j.nicl.2014.08.018
Soria G, de Notaris M, Tudela R et al (2011) Improved assessment of ex vivo brainstem neuroanatomy with high-resolution MRI and DTI at 7 T. Anat Rec 294:1035–1044. https://doi.org/10.1002/ar.21383
Stefanescu MR, Thürling M, Maderwald S et al (2013) A 7T fMRI study of cerebellar activation in sequential finger movement tasks. Exp Brain Res 228:243–254. https://doi.org/10.1007/s00221-013-3558-5
Stefanescu MR, Dohnalek M, Maderwald S et al (2015) Structural and functional MRI abnormalities of cerebellar cortex and nuclei in SCA3, SCA6 and Friedreich’s ataxia. Brain 138:1182–1197. https://doi.org/10.1093/brain/awv064
Strotmann B, Heidemann RM, Anwander A et al (2013a) High-resolution MRI and diffusion-weighted imaging of the human habenula at 7 T. J Magn Reson Imaging 39:1018–1026. https://doi.org/10.1002/jmri.24252
Strotmann B, Kögler C, Bazin PL (2013b) Mapping of the internal structure of human habenula with ex vivo MRI at 7T. Front Hum Neurosci. https://doi.org/10.3389/fnhum.2013.00878
Stüber C, Morawski M, Schäfer A et al (2014) Myelin and iron concentration in the human brain: a quantitative study of MRI contrast. NeuroImage 93:95–106. https://doi.org/10.1016/j.neuroimage.2014.02.026
Stucht D, Danishad KA, Schulze P et al (2015) Highest resolution in vivo human brain mri using prospective motion correction. PLoS ONE 10:e0133921. https://doi.org/10.1371/journal.pone.0133921.s003
Tang X, Holland D, Dale AM et al (2014) Shape abnormalities of subcortical and ventricular structures in mild cognitive impairment and Alzheimer’s disease: detecting, quantifying, and predicting. Hum Brain Mapp 35:3701–3725. https://doi.org/10.1002/hbm.22431
Temel Y, Blokland A, Steinbusch H, Visser-Vandewalle V (2005) The functional role of the subthalamic nucleus in cognitive and limbic circuits. Prog Neurobiol 76:393–413
Thayyil S, Cleary JO, Sebire NJ et al (2009) Post-mortem examination of human fetuses: a comparison of whole-body high-field MRI at 9·4 T with conventional MRI and invasive autopsy. Lancet 374:467–475. https://doi.org/10.1016/S0140-6736(09)60913-2
Thomas BP, Welch EB, Niederhauser BD et al (2008) High-resolution 7T MRI of the human hippocampus in vivo. J Magn Reson Imaging 28:1266–1272. https://doi.org/10.1002/jmri.21576
Thompson PM, Cannon TD, Narr KL et al (2001) Genetic influences on brain structure. Nat Neurosci 4:1253–1258
Thulborn K, Lui E, Guntin J et al (2015) Quantitative sodium MRI of the human brain at 9.4 T provides assessment of tissue sodium concentration and cell volume fraction during normal aging. NMR Biomed 29:137–143. https://doi.org/10.1002/nbm.3312
Thürling M, Küper M, Stefanescu R et al (2011) Activation of the dentate nucleus in a verb generation task: a 7T MRI study. NeuroImage 57:1184–1191. https://doi.org/10.1016/j.neuroimage.2011.05.045
Thürling M, Hautzel H, Küper M et al (2012) Involvement of the cerebellar cortex and nuclei in verbal and visuospatial working memory: a 7T fMRI study. NeuroImage 62:1537–1550. https://doi.org/10.1016/j.neuroimage.2012.05.037
Thürling M, Kahl F, Maderwald S et al (2015) Cerebellar cortex and cerebellar nuclei are concomitantly activated during eyeblink conditioning: a 7T fMRI study in humans. J Neurosci 35:1228–1239. https://doi.org/10.1523/JNEUROSCI.2492-14.2015
Tona K-D, Keuken MC, Rover M et al (2017) In vivo visualization of the locus coeruleus in humans: quantifying the test–retest reliability. Brain Struct Funct. https://doi.org/10.1007/s00429-017-1464-5
Tourdias T, Saranathan M, Levesque IR et al (2014) Visualization of intra-thalamic nuclei with optimized white-matter-nulled MPRAGE at 7T. NeuroImage 84:534–545. https://doi.org/10.1016/j.neuroimage.2013.08.069
Trampel R, Reimer E, Huber L et al (2013) Anatomical brain imaging at 7T using two-dimensional GRASE. Magn Reson Med 72:1291–1301. https://doi.org/10.1002/mrm.25047
Trattnig S, Bogner W, Gruber S et al (2015) Clinical applications at ultrahigh field (7 T). Where does it make the difference? NMR Biomed. https://doi.org/10.1002/nbm.3272
Trattnig S, Springer E, Bogner W et al (2016) Key clinical benefits of neuroimaging at 7T. NeuroImage. https://doi.org/10.1016/j.neuroimage.2016.11.031
Truong T-K, Chakeres DW, Beversdorf DQ et al (2006a) Effects of static and radiofrequency magnetic field inhomogeneity in ultra-high field magnetic resonance imaging. Magn Reson Imag 24:103–112. https://doi.org/10.1016/j.mri.2005.09.013
Truong T-K, Chakeres DW, Scharre DW et al (2006b) Blipped multi gradient-echo slice excitation profile imaging (bmGESEPI) for fastT2* measurements with macroscopicB0 inhomogeneity compensation. Magn Reson Med 55:1390–1395. https://doi.org/10.1002/mrm.20916
Turner R (2011) Neuroscientific applications of high-field MRI in humans. In: High-Field MR, Imaging. Springer, New York
Turner R (2013) Where matters: new approaches to brain analysis. In: Geyer S, Turner R (eds) Microstructural parcellation of the human cerebral cortex. Springer, Heidelberg
Turner R, De Haan D (2017) Bridging the gap between system and cell: The role of ultra-high field MRI in human neuroscience. Progr Brain Res. https://doi.org/10.1016/bs.pbr.2017.05.005
Turner R, Geyer S (2014) Comparing like with like: the power of knowing where you are. Brain Connect 4:547–557. https://doi.org/10.1089/brain.2014.0261
Uylings HBM, Rajkowska G, Sanz-Arigita E et al (2005) Consequences of large interindividual variability for human brain atlases: converging macroscopical imaging and microscopical neuroanatomy. Anat Embryol 210:423–431. https://doi.org/10.1007/s00429-005-0042-4
van Osch MJP, Webb AG (2014) Safety of ultra-high field MRI: what are the specific risks? Curr Radiol Rep 2:61–68. https://doi.org/10.1007/s40134-014-0061-0
van Bergen JMG, Hua J, Unschuld PG et al (2016) Quantitative susceptibility mapping suggests altered brain iron in premanifest huntington disease. AJNR Am J Neuroradiol 37:789–796. https://doi.org/10.3174/ajnr.A4617
van den Bogaard SJA, Dumas EM, Teeuwisse WM et al (2011) Exploratory 7-Tesla magnetic resonance spectroscopy in Huntington’s disease provides in vivo evidence for impaired energy metabolism. J Neurol 258:2230–2239. https://doi.org/10.1007/s00415-011-6099-5
van der Kolk AG, Hendrikse J, Zwanenburg JJM et al (2013) European journal of radiology. Eur J Radiol 82:708–718. https://doi.org/10.1016/j.ejrad.2011.07.007
van der Zwaag W, Schäfer A, Marques JP et al (2015) Recent applications of UHF-MRI in the study of human brain function and structure: a review. NMR Biomed. https://doi.org/10.1002/nbm.3275
Vaughan JT, Griffiths JR (2012) RF Coils for MRI. Wiley, Hoboken
Verma G, Hariharan H, Nagarajan R et al (2013) Implementation of two-dimensional L-COSY at 7 T: an investigation of reproducibility in human brain. J Magn Reson Imag 40:1319–1327. https://doi.org/10.1002/jmri.24510
Visser E, Keuken MC, Douaud G et al (2016a) Automatic segmentation of the striatum and globus pallidus using MIST: multimodal image segmentation tool. NeuroImage 125:479–497. https://doi.org/10.1016/j.neuroimage.2015.10.013
Visser E, Keuken MC, Forstmann BU, Jenkinson M (2016b) Automated segmentation of the substantia nigra, subthalamic nucleus and red nucleus in 7T data at young and old age. NeuroImage 139:324–336. https://doi.org/10.1016/j.neuroimage.2016.06.039
Wang X-Y, Zhao L, Yu T et al (2016) Assessment of age-related morphometric changes of subcortical structures in healthy people using ultra-high field 7 T magnetic resonance imaging. Front Aging Neurosci 8:893–899. https://doi.org/10.3389/fnagi.2016.00224
Wargo CJ, Gore JC (2013) Localized high-resolution DTI of the human midbrain using single-shot EPI, parallel imaging, and outer-volume suppression at 7T. Magn Resonance Imag 31:810–819. https://doi.org/10.1016/j.mri.2013.01.013
Weiskopf N, Suckling J, Williams G (2013) Quantitative multi-parameter mapping of R1, PD*, MT, and R2* at 3T: a multi-center validation. Front Neurosci. https://doi.org/10.3389/fnins.2013.00095
Weiskopf N, Mohammadi S, Lutti A, Callaghan MF (2015) Advances in MRI-based computational neuroanatomy. Curr Opin Neurol 28:313–322. https://doi.org/10.1097/wco.0000000000000222
Weiss M, Alkemade A, Keuken MC et al (2015) Spatial normalization of ultrahigh resolution 7 T magnetic resonance imaging data of the postmortem human subthalamic nucleus: a multistage approach. Brain Struct Funct 220:1695–1703. https://doi.org/10.1007/s00429-014-0754-4
Wen Q, Kelley DAC, Banerjee S et al (2015) Clinically feasible NODDI characterization of glioma using multiband EPI at 7 T. YNICL 9:291–299. https://doi.org/10.1016/j.nicl.2015.08.017
Wharton S, Bowtell R (2010) Whole-brain susceptibility mapping at high field: a comparison of multiple- and single-orientation methods. NeuroImage 53:515–525. https://doi.org/10.1016/j.neuroimage.2010.06.070
Wharton S, Schäfer A, Bowtell R (2010) Susceptibility mapping in the human brain using threshold-based k-space division. Magn Reson Med 63:1292–1304. https://doi.org/10.1002/mrm.22334
Wiggins GC, Polimeni JR, Potthast A et al (2009) 96-Channel receive-only head coil for 3 T: design optimization and evaluation. Magn Reson Med 62:754–762. https://doi.org/10.1002/mrm.22028
Wonderlick JS, Ziegler DA, Hosseini-Varnamkhasti P et al (2009) Reliability of MRI-derived cortical and subcortical morphometric measures: effects of pulse sequence, voxel geometry, and parallel imaging. NeuroImage 44:1324–1333. https://doi.org/10.1016/j.neuroimage.2008.10.037
Wright PJ, Mougin OE, Totman JJ et al (2008) Water proton T 1 measurements in brain tissue at 7, 3, and 1.5T using IR-EPI, IR-TSE, and MPRAGE: results and optimization. Magn Resonance Med 21:121–130. https://doi.org/10.1007/s10334-008-0104-8
Yang S, Yang Z, Fischer K, Zhong K (2013) Integration of ultra-high field MRI and histology for connectome based research of brain disorders. Front Neuroanat. https://doi.org/10.3389/fnana.2013.00031/abstract
Yao B, Li T-Q, Gelderen PV et al (2009) Susceptibility contrast in high field MRI of human brain as a function of tissue iron content. NeuroImage 44:1259–1266. https://doi.org/10.1016/j.neuroimage.2008.10.029
Yarach U, Luengviriya C, Stucht D et al (2016) Correction of B0-induced geometric distortion variations in prospective motion correction for 7T MRI. Magn Reson Mater Phy 29:319–332. https://doi.org/10.1007/s10334-015-0515-2
Zeineh MM, Parekh MB, Zaharchuk G et al (2014) Ultrahigh-resolution imaging of the human brain with phase-cycled balanced Steady-state free precession at 7 T. Invest Radiol 49:278–289. https://doi.org/10.1097/rli.0000000000000015
Zhang Y, Brady M, Smith S (2001) Segmentation of brain MR images through a hidden Markov random field model and the expectation-maximization algorithm. IEEE Trans Med Imaging 20:45–57
Zhang Z, Liu S, Lin X et al (2011) Development of fetal brain of 20 weeks gestational age: assessment with post-mortem magnetic resonance imaging. Eur J Radiol 80:e432–e439. https://doi.org/10.1016/j.ejrad.2010.11.024
Zielman R, Teeuwisse W, Bakels F et al (2014) Biochemical changes in the brain of hemiplegic migraine patients measured with 7 T 1H-MRS. Cephalalgia 34:959–967. https://doi.org/10.1177/0333102414527016
Zrinzo L, Zrinzo LV, Massey LA et al (2011) Targeting of the pedunculopontine nucleus by an MRI-guided approach: a cadaver study. J Neural Transm 118:1487–1495. https://doi.org/10.1007/s00702-011-0639-0
Zwanenburg JJM, Hendrikse J, Takahara T et al (2008) MR angiography of the cerebral perforating arteries with magnetization prepared anatomical reference at 7T: comparison with time-of-flight. J Magn Reson Imag 28:1519–1526. https://doi.org/10.1002/jmri.21591
Zwanenburg JJM, Hendrikse J, Visser F et al (2009) Fluid attenuated inversion recovery (FLAIR) MRI at 7.0 T: comparison with 1.5 and 3.0 T. Eur Radiol 20:915–922. https://doi.org/10.1007/s00330-009-1620-2
Acknowledgements
The work was supported by a Vidi grant by the Dutch Organization for Scientific Research (NWO) (BUF) and a starter grant from the European Research Council (ERC) (BUF). We would like to thank Bob Turner, Andreas Schäfer and Pierre-Louis Bazin for helpful discussions on the use of UHF MRI and Rosie Mulray for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
The authors declare that the work complies to the ethical standards.
Additional information
Handling Editor: Christoph M. Michel.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Keuken, M.C., Isaacs, B.R., Trampel, R. et al. Visualizing the Human Subcortex Using Ultra-high Field Magnetic Resonance Imaging. Brain Topogr 31, 513–545 (2018). https://doi.org/10.1007/s10548-018-0638-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10548-018-0638-7