Abstract
Information on spatial and temporal patterns of genetic diversity is a prerequisite to understanding the demography of populations, and is fundamental to successful management and conservation of species. In the sea, it has been observed that oceanographic and other physical forces can constitute barriers to gene flow that may result in similar population genetic structures in different species. Such similarities among species would greatly simplify management of genetic biodiversity. Here, we tested for shared genetic patterns in a complex marine area, the Baltic Sea. We assessed spatial patterns of intraspecific genetic diversity and differentiation in seven ecologically important species of the Baltic ecosystem—Atlantic herring (Clupea harengus), northern pike (Esox lucius), European whitefish (Coregonus lavaretus), three-spined stickleback (Gasterosteus aculeatus), nine-spined stickleback (Pungitius pungitius), blue mussel (Mytilus spp.), and bladderwrack (Fucus vesiculosus). We used nuclear genetic data of putatively neutral microsatellite and SNP loci from samples collected from seven regions throughout the Baltic Sea, and reference samples from North Atlantic areas. Overall, patterns of genetic diversity and differentiation among sampling regions were unique for each species, although all six species with Atlantic samples indicated strong resistence to Atlantic-Baltic gene-flow. Major genetic barriers were not shared among species within the Baltic Sea; most species show genetic heterogeneity, but significant isolation by distance was only detected in pike and whitefish. These species-specific patterns of genetic structure preclude generalizations and emphasize the need to undertake genetic surveys for species separately, and to design management plans taking into consideration the specific structures of each species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic variation is a prerequisite for species to adapt to a changing environment (Redford and Richter 1997; Reusch et al. 2005). Consequently, the importance of conserving genetic biodiversity is recognized both scientifically (e.g. Amos and Balmford 2001; Laikre et al. 2009), and politically. For example, in the new strategic plan of the United Nations Convention on Biological Diversity (CBD 1992), adopted in 2011, Target 13 explicitly addresses the conservation of genetic diversity (www.cbd.int/sp).
Identifying population genetic structures of species, describing the distribution of genetic diversity, and understanding the causes for these structures are important for effective management and conservation (Laikre et al. 2005a, 2008; Schmitt 2007). As all ecosystems contain large numbers of species, multi-species population genetic studies have been suggested as a useful first step in genetic surveys of separate geographic areas (Kelly and Palumbi 2010; Sivasundar and Palumbi 2010). Such multi-species assessments can be a valuable asset for conservation and management as they can shed light on whether or not similar management strategies may be appropriate for different species. However, these assessments are rarely implemented in practice, presumably due to the massive sampling and analytical efforts required for the simultaneous study of multiple species.
In the absence of evidence for strong selection, it is commonly assumed that the detected genetic variation is selectively neutral, reflecting the evolutionary processes of mutation, migration and drift (Utter 1991; McCusker and Bentzen 2010). Under this assumption, converging patterns among species, such as similar degrees of genetic differentiation and isolation by distance, may be shaped by similar dispersal patterns, common colonization histories, and life-history characteristics (Wares et al. 2001; Schmitt 2007; Pelc et al. 2009; Kelly and Palumbi 2010). In marine habitats, common locations of genetic discontinuities indicating shared barriers to dispersal have been found e.g. along the North American coasts (Pelc et al. 2009; Kelly and Palumbi 2010), in the Mediterranean (Patarnello et al. 2007), in the Caribbean (Taylor and Hellberg 2006), and at the entrance of the Baltic Sea (Johannesson and André 2006). Genetic similarities among species would be useful for management and conservation, for instance when marine reserves are established (Palumbi 2003). Alternatively, contrasting patterns of genetic differentiation among species could suggest that differences in life history or colonization history are major components in shaping the genetic structure of a species in a region (Kelly and Palumbi 2010). In such a situation, separate management for different groups of species, or even species-specific management would be required.
In this study we focus on the Baltic Sea, which is a sub-basin of the Atlantic Ocean formed less than 10,000 years ago as a postglacial marine environment (Zillén et al. 2008). The Baltic Sea is a highly suitable aquatic system to evaluate the presence or absence of common genetic diversity and differentiation patterns in multiple species. Environmental variation and potential barriers to dispersal possibly affecting different species in similar manner include a temperature and salinity gradient (spanning 3–30 per mille; HELCOM 2010) reaching from the entrance of the Baltic Sea to the north of the Bothnian Bay (Gabrielsen et al. 2002), and several sub-basins between which water circulation is partially restricted by submarine sills (HELCOM 2010). Species with both freshwater and marine origin have established populations which in many cases have undergone adaptations to the brackish water environment over the very short evolutionary history of the sea (Andersen et al. 2009; Gaggiotti et al. 2009; Papakostas et al. 2012).
Marginal ecosystems such as the Baltic Sea can be of great conservation value because they may harbor unique genetic variation and even novel species (Lesica and Allendorf 1995; Johannesson et al. 2011). Indeed, a new species of macroalgae has evolved inside the Baltic Sea (Pereyra et al. 2009). At the same time, the dense human population of the Baltic drainage area imposes threats to its aquatic biota via eutrophication, habitat destruction, and overfishing (Ducrotoy and Elliott 2008). These factors indicate that high priority should be given to the management of genetic diversity as the eradication of locally adapted wild populations may result in severe effects to the ecosystem (Johannesson et al. 2011). Although a reasonable number of genetic studies have been carried out on Baltic species (see Johannesson et al. 2011 for a recent review), diverse sampling designs have prevented rigorous among-study comparisons regarding patterns of populations genetic structure (but see DeFaveri et al. 2012).
Previous genetic comparisons involving several marine species have shown that most Baltic populations contain lower levels of variation than conspecific Atlantic ones (reviewed in Laikre et al. 2005a; Johannesson and André 2006; Johannesson et al. 2011). In addition, several species show large genetic differences at the entrance of the Baltic Sea (Johannesson and André 2006). Further, a genetic barrier near to the Islands of Åland has been identified in both herring (Clupea harengus; Jørgensen et al. 2005) and perch (Perca fluviatilis; Olsson et al. 2011), separating northern populations from southern ones. An important question is whether this and other barriers are consistent across taxa. Testing the hypothesis of shared overall genetic structures is of high relevance to management.
The present study is based on population genetic data from seven species of key socio-economic and/or ecological importance sampled from each of seven geographic regions throughout the Baltic Sea. The key question is whether genetic divergence patterns of these different species are similar over the Baltic Sea. Despite the adaptive relevance of such ecological variables as temperature and salinity, our data sets are not designed to address levels or types of selection affecting specific loci, noting the ambiguity of interpreting such effects on outlier loci even from extensive genomic scans (Bierne et al. 2011, 2013). Rather, we assume an overall signal of neutrality as a first approximation of reality (Ihssen et al. 1981) as balanced by divergent, convergent, and nonselective forces. This interpretation has been widely validated for diverse organisms and is particularly applicable to initial comparisons among heterogeneous data sets such as those used in this study (Utter and Seeb 2010). Each species diverges uniquely from the null hypothesis of panmixia, reflecting factors including barriers to effective migration, isolation by distance, and repeated colonizations.
Genetic data of Baltic species
Genetic data were compiled or generated for each of the following seven species selected for this study: (1) Atlantic herring (C. harengus), one of the most economically important species fished in the Baltic Sea, (2) Northern pike (Esox lucius), and (3) European whitefish (Coregonus lavaretus), two ecologically important predators and popular targets for commercial and recreational fishing, (4) three-spined stickleback (Gasterosteus aculeatus), and (5) nine-spined stickleback (Pungitius pungitius), abundant mesopredators; and two important habitat forming species, (6) the blue mussel (Mytilus trossulus) including collections from populations putatively hybridized with M. edulis at the Baltic/Atlantic interface (Väinölä and Strelkov 2011; Zbawicka et al. 2012), and (7) bladderwrack (Fucus vesiculosus), a primary producer (Table 1). The recently diverged form of bladderwrack Fucus radicans endemic to the northern and eastern parts of the Baltic Sea was not included. Sampling locations for all species were chosen to cover defined regions throughout the Baltic Sea, and where possible, adjacent Atlantic regions (Fig. 1). Sample sizes per locality varied in the range 12–48 (Table S1).
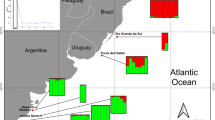
Sampling regions for the empirical material of the seven Baltic species of the present study. Definition of regions similar to Ojaveer et al. (2010) and Olsson et al. (2012b). These regions largely constitute sub-basins between which water circulation is partially restricted by submarine sills. There is a sharp salinity gradient at the Danish Belts at the entrance of the Baltic Sea located at the indicated border between the Baltic Sea and Outside the Baltic (HELCOM 2010)
Individual genotypes for 7–23 microsatellites, or in the case of the blue mussel 10 single nucleotide polymorphisms (SNPs), were generated. Detailed genotyping procedures for each separate species are provided in the Supplementary material.
We used two sets of comparative data; one included only Baltic samples for all seven species. The second set also included samples from outside the Baltic Sea (Fig. 1), and such samples were available for all species except for northern pike which lacks Atlantic (fully marine) populations (Table 2). The Atlantic sample for the whitefish, which is also a non-marine species, was collected from a fjord with brackish water on the border between Sweden and Norway (Fig. 2).
Diversity-divergence patterns and the three strongest barriers to gene flow. Diversity is shown in left part of the circles; dark higher diversity than average, light lower diversity. Divergence is shown in the right part of the figures; dark higher divergence than average, light lower divergence. Populations sampled outside the Baltic Sea were not included in diversity-divergence analyses and are shown as white circles with a dot. Barriers supported by more than half of the investigated loci are indicated with solid lines, and barriers supported by less than half of the loci are indicated with dotted lines. Barriers indicated here are supported also by traditional F ST statistics (cf. Table S2a–g). For bladderwrack there is also an indication of a barrier to gene flow at the entrance to the Baltic Sea, but it is not included among the three strongest barriers depicted here (cf. Table S2g). For herring no statistically significant differences were found among populations in the Baltic Sea, and for northern pike no sample was available outside of the Baltic as this species does not exist there
Statistical analyses
Genetic diversity, Hardy–Weinberg proportions, differentiation between samples
All loci and all separate samples were checked for deviations from Hardy–Weinberg expectation using Genepop 3.4 (Raymond and Rousset 1995) and Microchecker (van Oosterhout et al. 2004). Loci with likely null alleles or allelic dropout were removed (Supplementary material). We investigated remaining loci that might be under selection using an F ST outlier method based on the expected distribution of F ST and gene diversity (H e) using the software Lositan, simulating a neutral distribution of F ST under the stepwise mutation and infinite allele model respectively, and identifying loci falling outside of the 95 % quartiles after 100,000 simulations (Antao et al. 2008). Inclusion or exclusion of loci under potential selection affected the results only slightly, and never affected statistical significances or major conclusions. Therefore, loci potentially affected by selection were kept in all subsequent analyses.
Observed and expected heterozygosities as well as the number of alleles were estimated using Microsatellite Toolkit 3.1 (Park 2001), and allelic richness was estimated using Fstat 2.9.3.2 (Goudet 1995). For each species differences in allelic richness between the sampled regions were tested with a median test. Each locus in each sampled region was assigned to one of two groups—higher or lower allelic richness than the median allelic richness for all samples in that particular locus. A χ 2 test was used to determine whether the observed frequencies of loci with high or low allelic richness for each region differed from expected equal frequencies under the hypothesis of no difference in genetic variation among sampled regions. The degree of population differentiation, measured as F ST, was assessed using GenePop 3.4 (Raymond and Rousset 1995), and tests for genetic heterogeneity were made using ChiFish (Ryman 2006). Because data for both microsatellites and SNPs were used, some caution is warranted in among-species interpretations of estimated parameters, particularly between the blue mussel and the other species. Large numbers of alleles and high heterozygosities, typical of microsatellite loci, impose low limits on F ST values (Hedrick 1999). Conversely, SNPs are commonly limited to two alleles, thus limiting the range of possible values for heterozygosity and allelic richness. In addition to F ST we also applied G ′ST a measurement of genetic differentiation corrected for heterozygosity using the software Smogd (Crawford 2010). We note, however, that in situations that are not characterized by steady state conditions and very low migration rates, G ′ST in many cases may be difficult to interpret (Ryman and Leimar 2008, 2009).
Contribution of separate sampling regions to total genetic diversity
For each species separately, the relative contribution of each sampled region to the total genetic diversity of the global Baltic Sea population was estimated in terms of diversity and divergence using the methods described by Petit et al. (1998), implemented in the software MolKin 2.0 (Gutiérrez et al. 2005). Briefly, for each sample we estimated (1) within-sample diversity measured as allelic richness of the sample relative to the allelic richness of the other samples of the same species, and (2) genetic differentiation of the sample in relation to the other samples of the same species using a measure related to Nei’s D ST and G ST (Gutiérrez et al. 2005). Positive values of relative diversity and/or differentiation for a particular sampled region indicate that the sample of that region contributes positively to total genetic diversity of the global Baltic population. Negative values correspondingly indicate that the relative diversity or divergence of the sample in question is low and does not contribute to total genetic diversity (Petit et al. 1998).
The values for relative diversity and differentiation were used to categorize each sample into one of four categories, as identified by Swatdipong et al. (2009) including (i) higher diversity-higher divergence, (ii) higher diversity-lower divergence, (iii) lower diversity-higher divergence, and (iv) lower diversity-lower divergence. Samples in each category can be expected to be characterized by the differing roles of migration and genetic drift affecting the genetics of populations. Categories i and ii are considered to have the largest potential of containing unique genetic material and should potentially be prioritized in conservation (Swatdipong et al. 2009). The observed strong divergence of Baltic populations from Atlantic conspecifics (Johannesson and André 2006) prompted the exclusion of Atlantic samples from these analyses to amplify the diversity-divergence classification within the Baltic Sea. The difference in the distribution of observed frequencies of the four diversity-divergence categories in different geographic regions relative to the expected frequencies under the null hypothesis of random distribution of diversity-divergence was tested with a χ 2 test for independence.
Areas of genetic discontinuities
We used the software Barrier 2.2 (Manni et al. 2004) to locate areas of major genetic discontinuities. Barrier applies Monmonier’s algorithm to detect the areas of highest genetic change on a map (genetic barriers) where the samples are represented by their geographic coordinates and connected by Delauney triangulation. The software produces as many barriers as the user defines, regardless of how strong these barriers are, i.e. if they are supported by significant F ST values or not. For example in the case of the Atlantic herring in this study, there is no significant differentiation among populations within the Baltic Sea, but Barrier still identifies genetic breaks if asked to do so. On the contrary, there are cases where most samples are significantly differentiated from each other, in these cases many identified barriers are supported by significant F ST values. For each species we assessed several barriers from pairwise F ST values over all loci and compared their relative location among species. We discarded all barriers not supported by F ST values significant after Bonferroni correction. We illustrate the three major barriers identified by Barrier within each separate species. The strength of each of these barriers was quantified from the number of loci supporting the barrier. For each separate species, we differentiated between barriers supported by more or less than half of the loci as suggested by LeClerc et al. (2008).
Association between geographical distance and genetic divergence
We examined the association between geographical distance and genetic divergence (isolation by distance, IBD) with a Mantel test using the package Ecodist 1.1.3 (Goslee and Urban 2007) in the software R 2.12.2 (R Development Core Team 2011), using 10,000 permutations, and bootstrapping confidence limits with 1,000 iterations. Genetic divergence was measured as F ST/(1 − F ST), and geographic distances between sample sites were calculated as shortest waterway distance using ArcGIS 10 (ESRI 2010, Redlands, CA, USA). Both raw and log transformed distances were used (Rousset 1997), but only results based on raw distances are presented, since the two measurements of geographic distance gave very similar results. Two Mantel tests were conducted for each species including (1) all samples, and (2) only Baltic Sea samples.
Results
We found few deviations from Hardy–Weinberg proportions. Observed and expected heterozygosity varied in the range 0.073–0.832 and allelic richness in the range 1.400–14.115. Overall F ST values ranged from <0.01 to 0.47. As expected G ′ST values were higher, but the relative difference in magnitude among species were the same for F ST and G ′ST (Table 2; details for separate species and localities are provided in Table S1).
Distinct signatures of genetic variation among sampling locations existed for each species based on various measurements. All species except the Atlantic herring exhibit significant allele frequency differences among sampling regions within the Baltic Sea, although for three-spined stickleback only one pairwise F ST value remained significant after Bonferroni correction (Table 2; Pairwise F ST values between all samples for each species are found in Tables S2 a–g). Allelic richness also varies significantly among regions. However, the patterns of this within-species variability over the Baltic Sea vary widely among species (Table 3; Figs. 2, 3) as reflected by a lack of tendency for higher- or lower-divergence samples from different species to occur in the same geographic region (Table 3; χ 2 = 7.80, df = 6, p = 0.25; Fig. 2). Similarly, samples with high or low relative genetic diversity do not cluster in any particular region (Table 3; χ 2 = 3.42, df = 6, p = 0.76; Fig. 2). A similar disparity is evident for specific diversity-divergence categories to cluster in a specific region even if only the most extreme samples that have the highest relative diversity or divergence in each species are included (χ 2 = 25.19, df = 18, p = 0.12).
Association between geographic and genetic distance (isolation by distance, IBD). Correlation coefficients for line equation and significance level of Mantel test (*0.05 > p > 0.01, *0.01 > p > 0.001, ***0.001 > p). Two Mantel tests were performed, one for the total material (all points, dotted line) and one for Baltic only samples (filled points, full line)
Four of the species: Northern pike, whitefish, nine-spined stickleback and bladderwrack show significant pairwise differentiation between almost all samples (Table S2a–g). Although overall values of F ST are moderate in the three first species, the significant values imply limited gene flow among most sampling areas. We observe isolation by distance in both species of freshwater origin (pike and whitefish), but apart from that there are few similarities between these two species regarding location of barriers and samples of high diversity or divergence. Isolation by distance was also present for herring when the Atlantic sample was included, but was not detectable in any other species in this study (Fig. 3). Among the originally marine species (herring, blue mussel, bladderwrack, and three-spined stickleback) we find few shared patterns, except for the common genetic barrier reported by the software Barrier that separates Baltic samples from Atlantic ones (one of the strongest barriers in all species with Atlantic samples, except bladderwrack, although the Atlantic-Baltic transition is also supported by significant F ST values in that species) and the lack of major genetic barriers supported by significant F ST values within the Baltic Sea for herring and three-spined stickleback. Additional barriers occur at different locations for all seven species. For each species we illustrate the location of the three most important barriers identified by the software Barrier, that are also supported by significant F ST values. The locations of these three major barriers are almost unique for each species (Fig. 2).
Samples from the northern and southern extremes of the Baltic showed high relative divergence in most species, coupled with high diversity in some of the species (herring and pike in the north, bladderwrack and blue mussel in the south). However, a signal of a major genetic break in these areas was seen only in the two species; pike and blue mussel. Except for the barrier at the entrance of the Baltic Sea the locations of the three most important genetic breaks were unique for each species (Fig. 2). Genetic patterns for each species in this study are briefly described below as illustrated in Figs. 2 and 3, and fine scale structuring for each species is provided in Table S2a–g.
Atlantic herring
There were low and non significant levels of differentiation among sampling sites of Baltic herring (F ST = 0.0009; Table 2). We found the largest genetic divergences between Baltic and Atlantic samples (average F ST = 0.0075) and this difference was also statistically significant. Consistently lower relative diversity and higher relative differentiation were observed in the southern and eastern regions. These patterns were reversed in adjacent northwestern regions, and both higher diversity and divergence occurred in northernmost Bothnian Bay.
Northern pike
All pairwise comparisons among pike samples were significantly differentiated from each other, with an overall moderate F ST-value of 0.03 (Tables 2, S2b) and a significant isolation by distance. Major genetic discontinuities distinguish the Bothnian Bay and Baltic Proper East samples.
European whitefish
Baltic whitefish samples were notable for mostly well differentiated samples with moderate overall differentiation (F ST = 0.04; Tables 2, S2c) and significant isolation by distance. The strongest barrier is located between the southernmost Baltic samples and the rest of the Baltic Sea with a fairly homogenous area of lower differentiation in the northern Bothnian Bay.
Three-spined stickleback
The low but statistically significant F ST of <0.001 within the Baltic Sea and the lack of isolation by distance suggests very weak genetic structuring or genetic uniformity in the region (Tables 2, S2d). The lower diversities in the northern and eastern regions contrasted with the generally higher values in the western samples.
Nine-spined stickleback
Baltic samples were characterized by a moderate overall differentiation, although almost all samples were significantly differentiated from each other (F ST = 0.03; Tables 2, S2e) and lack of isolation by distance, though the test for isolation by distance approached significance (p = 0.095 when the Atlantic sample was included in the analysis).
Blue mussel
Overall F ST is 0.47 (Table 2) with a strong barrier separating two southwestern samples and a second barrier distinguishing island and mainland samples in the Baltic Proper West. High diversity at southern sampling sites contrasted with lower diversity and higher divergence in northern samples. The strikingly high F ST might reflect species mixture and introgression. M. trossulus is indigenous to the Baltic Sea but is closely related to M. edulis (common name also blue mussel), native to the North Sea. These taxa are known to hybridize and it is possible that our southern samples include very rare M. edulis specimens. The two species are difficult to distinguish even by genetic techniques, and geographic distribution and genetic characteristics of these species are continuously subject to revision (Riginos and Cunningham 2005; Steinert et al. 2012).
Bladderwrack
The three strongest barriers to gene flow occur in the northern part of the Baltic, although the high overall F ST (0.14; Table 2) indicated strong genetic structuring overall, with all sampling locations being significantly differentiated from each other (Table S2g).
Discussion
We conducted the first multi-species study in the Baltic Sea where a large number of individuals and loci were collected from the same areas covering the full Baltic Sea. Surprisingly, we detected few shared genetic patterns in the seven species analyzed with respect to location of the three major genetic barriers to gene flow and diversity-divergence patterns (Fig. 2). An exception to this general lack of consistence is the genetic break between the Atlantic and the Baltic Sea.
We observe a variety of genetic patterns ranging from large and significant differences among sampling regions in both genetic variation and divergence, to very little differentiation within the Baltic Sea. The most pronounced, genetic breaks occurred almost individually for each species in different regions of the Baltic Sea, although several species showed significant pairwise differentiation between the majority of the samples (Table S2a–g). At the northern extreme, five of six samples from the Bothnian Bay showed high diversity, but no shared major genetic barrier was present in this region (Table 3; Fig. 2). Unlike previous studies of herring and perch (Jørgensen et al. 2005; Olsson et al. 2011) we found few shared major genetic breaks associated with the different sub-basins of the Baltic Sea, e.g. around the Åland Islands.
Potential causes of variability patterns
The species-specific genetic patterns in the Baltic Sea, including relative amount of genetic variation, location of major genetic breaks, and isolation by distance are likely dependent on a multitude of factors including salinity tolerance, oceanographic features, life history, and population history (Table 1). For example, salinity has been correlated with genetic differentiation in Atlantic herring and three-spined stickleback, using both selected and neutral loci (André et al. 2011; Lamichhaney et al. 2012; Limborg et al. 2012; DeFaveri et al. 2013); ocean connectivity has been correlated with genetic divergence in herring (Teacher et al. 2013) as has temperature for herring and three-spined stickleback (Limborg et al. 2012; DeFaveri et al. 2013).
Additional factors that have been demonstrated to affect genetic structure include larval development and dispersal (Kyle and Boulding 2000). For example, the free-floating larval stage in Atlantic herring and a later pelagic life stage mediate potential for long distance dispersal and is a likely explanation for the lack of genetic structuring for herring within the Baltic Sea shown here, as well as in previous studies using neutral genetic markers (Bekkevold et al. 2005; Jørgensen et al. 2005). Genetic divergence among herring populations has indeed been shown to be affected more by ocean currents than geographic distance (Teacher et al. 2013). Ocean currents are more likely to affect species with freefloating life stages, such as herring, or bladderwrack, for which dispersal of eggs are limited, but detached adults have the potential for dispersal by means of rafting (Tatarenkov et al. 2007). Species with stationary development on the other hand, such as European whitefish and Northern pike, which are both associated with freshwater spawning, are likely to have more limited dispersal. The observed pattern of isolation by distance found in whitefish and pike in the present study as well as previous studies (Laikre et al. 2005b; Olsson et al. 2012a) is consistent with such limited dispersal and suggests that migration predominantly takes place between geographically proximate populations. It should be noted that recent studies have detected isolation by distance also in herring (Teacher et al. 2013) and three-spined and nine-spined stickleback (DeFaveri et al. 2012). Those studies included larger sample sizes and/or more genetic markers than examined here, however, and may thus have been characterized by higher statistical power for detection of isolation by distance.
Other factors potentially affecting genetic diversity in the Baltic Sea include postglacial colonization of the area by different phylogenetic lineages. Nine-spined stickleback in the Baltic Sea has been shown to consist of one western and one eastern lineage meeting roughly at the entrance of the Baltic Sea (Shikano et al. 2010; Teacher et al. 2011), as previously also shown for cod (Nielsen et al. 2003) and the bivalve Macoma balthica (Luttikhuizen et al. 2012). A more extreme example of transition zones is represented by the blue mussel, where the species M. trossulus, native to the Baltic Sea is hybridized with M. edulis (Riginos and Cunningham 2005). The high diversity of the southwestern-most sample of blue mussel potentially reflects increased hybridization in this area, but does not seem to affect the general results of the study as relative diversity-divergence or locations of barriers do not change if this sample is excluded from analyses (data not shown). A combination of ecological and demographic aspects and selective forces is probably important for each species in the Baltic Sea. These potential forces apparently do not affect the different species in the Baltic Sea in the same manner, thus, there is no generalization to be made among species.
The majority of the species in this study are sampled in most of the defined sampling areas, but there is some heterogeneity among species regarding the exact sample sites (Fig. 2). The exact location of each genetic barrier cannot be defined without even more detailed sampling. However, relative barriers among major areas within the Baltic Sea should be possible to detect for all species.
The potential role of selection
The initial neutral expectations of our data do not exclude the influence of selective forces affecting the observed patterns. Indeed, such influences commonly enhance rather than reduce the observed population structures of such data sets (see e.g. Utter and Seeb 2010), which has been documented in herring of the Baltic-Atlantic including the temporal stability of such selective patterns (Larsson et al. 2007, 2010). Selection most likely plays an important role in shaping genetic patterns in the Baltic Sea that are usually not detectable using neutral genetic markers because of migration rates so high that allele frequencies at selectively neutral loci are homogenized. Recent studies of three-spined stickleback, one of the focal species for this study with the lowest levels of genetic structuring, show evidence of considerable divergence in phenotypic traits and selected loci giving direct evidence of adaptive divergence (DeFaveri et al. 2013; DeFaveri and Merilä 2013). Further studies on selected loci will likely extend and complement the knowledge based on presumed neutral markers. For management purposes this addition will be of particular interest since management and conservation units can be identified more precisely using both selected and neutral loci (Allendorf et al. 2010; Funk et al. 2012).
Genetic divergence between the Atlantic and the Baltic Sea
The generally strong genetic distinctions observed between Baltic and Atlantic samples (Fig. 2; Table S2a–g) coincide with a sharp salinity gradient and reduced water circulation in the Danish belts (HELCOM 2010; Johannesson and André 2006; Johannesson et al. 2011). This shared genetic barrier is now supported by a wide range of fish species, such as the sand goby (Larmuseau et al. 2009), sprat (Limborg et al. 2009), herring (Limborg et al. 2012; Lamichhaney et al. 2012), whitefish (Olsson et al. 2012a) and sticklebacks (Shikano et al. 2010; DeFaveri et al. 2013). The relatively large genetic differentiation in this region has generally been attributed to selection for the low saline environment followed by adaptive divergence and subsequent reduced gene flow which affect selected as well as putatively neutral genetic markers, in species with a marine background (Limborg et al. 2012; Teacher et al. 2013; DeFaveri et al. 2013). However, it is important to note that demographic rather than non-adaptive forces, such as secondary contact between divergent lineages, or the formation of hybrid zones, have also generated similar patterns of genetic discontinuities in this region.
Relating our findings to previous studies
Our findings augment previous investigations within the Baltic Sea. For separate species within the Baltic Sea the magnitude and geographic pattern of genetic divergence were similar to previous results for herring using putatively neutral genetic markers (Bekkevold et al. 2005; Jørgensen et al. 2005), three-spined stickleback (Mäkinen et al. 2006; DeFaveri et al. 2012), Northern pike (Laikre et al. 2005b), and European whitefish (Olsson et al. 2012a).
Genetic biodiversity has been studied more or less extensively in several other species in addition to those of our study. Baltic populations that are genetically isolated from populations outside the Baltic are found in cod (Gadus morhua; Nielsen et al. 2003) and flounder (Platichtys flesus; Hemmer-Hansen et al. 2007). Isolation by distance patterns in the Baltic has been observed both for marine species, e.g. eelpout (Zoarces viviparus; Kinitz et al. 2013) and freshwater species, e.g. perch (Olsson et al. 2011), but also lack thereof e.g. in turbot (Psetta maxima; Florin and Höglund 2007). Genetic diversity has previously been both positively and negatively correlated with latitude within the Baltic Sea (Olsson et al. 2011; Kinitz et al. 2013).
Management implications
The apparent lack of shared genetic patterns in the Baltic Sea has consequences both for management and future research. Scientists, as well as managers, should be cautious regarding generalizing genetic patterns among species in the Baltic region, and this lack of a general pattern challenges conservation management of gene level biodiversity. For instance, common indicators of genetic biodiversity will be difficult to find, and optimal procedures for implementing the Strategic Plan of the Convention on Biological Diversity adopted in 2010 (www.cbd.int) are not obvious. Different biological traits, possibly unique to each species, are likely to shape genetic patterns and therefore need to be identified and taken into account in management. Similarly, the species-specific patterns might increase identified problems of institutional uncertainty regarding genetic variation (cf. Sandström 2010, 2011). For instance, creating an understanding for the gene level of biodiversity and communicating easily understandable advice from researchers to managers in the Baltic Sea region is a complex task, and there is an obvious risk that the existing lag of implementing existing conservation policy to genetic biodiversity (Laikre 2010; Laikre et al. 2010) remains difficult to overcome. It is clear that Baltic populations are genetically distinct from North Atlantic populations and should be actively conserved as unique genetic and biological resources.
Future comparisons among species with more extensive sampling including both additional species and sampling sites seem likely to reveal more subtle shared genetic patterns than detected in this study. However, at present when genetics is used as a base for sound management, recommendations should be made on a species-by-species basis. Clearly, providing means for adaptive management of Baltic Sea genetic biodiversity is complex and challenging for both scientists and managers.
Conclusions
Each species in the environmentally heterogeneous Baltic Sea that was included in our study displayed a unique genetic pattern of diversity and divergence. Genetic differences among Baltic Sampling sites were present among most of the seven species (except for Atlantic herring, and very small differences for three-spined stickleback), as was the barrier to gene flow at the entrance of the Baltic Sea. Our main conclusion is that in the Baltic Sea ecosystem where environmental gradients occur and where separate species have different origins (freshwater or marine), genetic patterns of variation and divergence are not shared among species. In order to infer management and conservation units, each species of interest must be investigated separately. These findings stress the overall need for genetic surveys of high spatial resolution, in particular in areas of high environmental complexity such as the Baltic Sea.
References
Allendorf FW, Hohenlohe PA, Luikart G (2010) Genomics and the future of conservation genetics. Nat Rev Genet 11:697–709
Amos W, Balmford A (2001) When does conservation genetics matter? Heredity 87:257–265
Andersen O, Wetten OF, De Rosa MC, André C, Carelli Alinovi C, Colafranceschi M, Brix O, Colosimo A (2009) Haemoglobin polymorphisms affect the oxygen-binding properties in Atlantic cod populations. Proc R Soc B 276:833–841
André C, Larsson LC, Laikre L, Bekkevold D, Brigham J, Carvalho GR, Dahlgren TG, Hutchinson WF, Mariani S, Mudde K, Ruzzante DE, Ryman N (2011) Detecting population structure in a high gene-flow species, Atlantic herring (Clupea harengus): direct, simultaneous evaluation of neutral vs putatively selected loci. Heredity 106:270–280
Antao T, Lopes A, Lopes RJ, Beja-Pereira A, Luikart G (2008) LOSITAN: a workbench to detect molecular adaptation based on a F ST outlier-method. MBC Bioinform 9:323. doi:10.1186/1471-2105-9-323
Bekkevold D, André C, Dahlgren TG, Clausen LAW, Torstensen E, Mosegaard H, Carvalho GR, Christensen TB, Norlinder E, Ruzzante DE (2005) Environmental correlates of population differentiation in Atlantic herring. Evolution 59:2656–2668
Bierne N, Welch J, Loire E, Bonhomme F, David P (2011) The coupling hypothesis: why genome scans may fail to map local adaptation genes. Mol Ecol 20:2044–2072
Bierne N, Roze D, Welch JJ (2013) Pervasive selection or is it…? why are F ST outliers sometimes so frequent? Mol Ecol 22:2061–2064
CBD (1992) Convention on biological diversity (with annexes). Concluded at Rio de Janeiro 1760(30619):I-30619
Crawford N (2010) Smogd: software for the measurement of genetic diversity. Mol Ecol Res 10:556–557
DeFaveri J, Merilä J (2013) Evidence for adaptive phenotypic differentiation in Baltic Sea sticklebacks. J Evol Biol 26:1700–1715
DeFaveri J, Shikano T, Ab Ghani NI, Merilä J (2012) Contrasting population structures in two sympatric fishes in the Baltic Sea basin. Mar Biol 159:1659–1672
DeFaveri JP, Jonsson PR, Merilä J (2013) Heterogenous genomic differentiation in marine threespine sticklebacks: adaptation along an environmental gradient. Evolution 159:1659–1672
Ducrotoy JP, Elliott M (2008) The science and management of the North Sea and the Baltic Sea: natural history, present threats and future challenges. Mar Pollut Bull 57:8–21
ESRI (2010) ArcGIS Desktop 10.0. Environmental Systems Research Institute, Redlands
Florin A-B, Höglund J (2007) Absence of population structure of turbot (Psetta maxima) in the Baltic Sea. Mol Ecol 16:115–126
Funk WC, McKay JK, Hohenlohe PA, Allendorf FW (2012) Harnessing genomics for delineating conservation units. Trends Ecol Evol 27:489–496
Gabrielsen TM, Brochmann C, Rueness J (2002) The Baltic Sea as a model system for studying postglacial colonization and ecological differentiation, exemplified by the red alga Ceramium tenuicorne. Mol Ecol 11:2083–2095
Gaggiotti OE, Bekkevold D, Jørgensen HBH, Foll M, Carvalho GR, André C, Ruzzante DE (2009) Disentangling the effects of evolutionary, demographic, and environmental factors influencing genetic structure of natural populations: Atlantic herring as a case study. Evolution 63:2939–2951
Goslee SC, Urban DL (2007) The ecodist package for dissimilarity-based analysis of ecological data. J Stat Softw 22:1–19
Goudet J (1995) FSTAT (vers 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Gutiérrez JP, Royo LJ, Álvarez I, Goyache F (2005) MolKin v2.0: a computer program for genetic analysis of populations using molecular coancestry information. J Hered 96:718–721
Hedrick PW (1999) Variable loci and their interpretation in evolution and conservation. Evolution 53:313–318
HELCOM (2010) Atlas of the Baltic Sea. Helsinki Commission (HELCOM). ISBN 978-952-67205-2-4
Hemmer-Hansen J, Nielsen EEG, Grønkjaer P, Loeschke V (2007) Evolutionary mechanisms shaping the genetic population structure of marine fishes: lessons from the European flounder (Platichthys flesus). Mol Ecol 16:3104–3118
Ihssen PE, Booke HE, Casselman JM, McGlade JM, Payne NR, Utter FM (1981) Stock identification—material and methods. Can J Fish Aquat Sci 38:1838–1855
Johannesson K, André C (2006) Life on the margin: genetic isolation and diversity loss in a peripheral marine ecosystem, the Baltic Sea. Mol Ecol 15:2013–2029
Johannesson K, Smolarz K, Grahn M, André C (2011) The future of Baltic Sea populations: local extinction or evolutionary rescue? Ambio 40:179–190
Jørgensen HBH, Hansen MM, Bekkevold D, Ruzzante DE, Loeschke V (2005) Marine landscapes and population genetic structure of herring (Clupea harengus) in the Baltic Sea. Mol Ecol 14:3219–3234
Kelly RP, Palumbi SR (2010) Genetic structure among 50 species of the northeastern pacific rocky intertidal community. PLoS ONE 5(1):e8594. doi:10.1371/journal.pone.0008594
Kinitz T, Quack M, Paulus M, Veith M, Bergek S, Strand J, Tuvikene A, Soirinsuo A, Hochkirch A (2013) Strong isolation-by-distance in the absence of genetic population structure in the eelpout (Zoarces viviparus 1758). Ecol Indic 27:116–122
Kyle CJ, Boulding EG (2000) Comparative population genetic structure of marine gastropods (Littorina spp.) with and without pelagic larval dispersal. Mar Biol 137:835–845
Laikre L (2010) Genetic diversity is overlooked in international conservation policy implementation. Conserv Genet 11:349–354
Laikre L, Palm S, Ryman N (2005a) Genetic population structure of fishes: implications for coastal zone management. Ambio 34:111–119
Laikre L, Miller LM, Palme A, Palm S, Kapuscinski AR, Thoresson G, Ryman N (2005b) Spatial genetic structure of northern pike (Esox lucius) in the Baltic Sea. Mol Ecol 14:1955–1964
Laikre L, Larsson LC, Palmé A, Charlier J, Josefsson M, Ryman N (2008) Potentials for monitoring gene level biodiversity: using Sweden as an example. Biodiv Conserv 17:893–910
Laikre L, Nilsson T, Primmer CR, Ryman N, Allendorf FW (2009) Importance of genetics in the interpretation of favourable conservation status. Conserv Biol 23:1378–1381
Laikre L, Allendorf FW, Aroner LC, Baker CS, Gregovich DP, Hansen MM, Jackson JA, Kendall KC, McKelvey K, Neel MC, Olivieri I, Ryman N, Schwartz MK, Short Bull R, Stetz JB, Tallmon DA, Taylor BL, Vojta CD, Waller DM, Waples RS (2010) Neglect of genetic diversity in implementation of the convention on biological diversity. Conserv Biol 24:86–88
Lamichhaney S, Martinez Barrio A, Rafati N, Sundström G, Rubin CJ, Gilbert ER, Berglund J, Wetterbom A, Laikre L, Webster MT, Grabherr M, Ryman N, Andersson J (2012) Population-scale sequencing reveals genetic differentiation due to local adaptation in Atlantic herring. Proc Natl Acad Sci 109:19345–19350
Larmuseau MHD, Vad Houdt JKJ, Guelinckx J, Hellemans B, Volckairt FAM (2009) Distributional and demographic consequences of Pleistocene climate fluctuations for a marine demersal fish in the north-eastern Atlantic. J Biogeogr 36:1138–1151
Larsson LC, Laikre L, Palm S, André C, Carvalho GR, Ryman N (2007) Concordance of allozyme and microsatellite differentiation in a marine fish, but evidence of selection at a microsatellite locus. Mol Ecol 16:1135–1147
Larsson LC, Laikre L, Andre C, Dahlgren TG, Ryman N (2010) Temporally stable genetic structure of heavily exploited Atlantic herring (Clupea harengus) in Swedish waters. Heredity 104:40–51
LeClerc É, Mailhot Y, Mingelbier M, Bernatchez L (2008) The landscape genetics of yellow perch (Perca flavenscens) in a large fluvial ecosystem. Mol Ecol 17:1702–1717
Lesica P, Allendorf FW (1995) When are peripheral-populations valuable for conservation. Conserv Biol 9:753–760
Limborg MT, Pedersen JS, Hemmer-Hansen J, Tomkiewicz J, Bekkevold D (2009) Genetic population structure of European sprat Sprattus sprattus: differentiation across a steep environmental gradient in a small pelagic fish. Mar Ecol Prog Ser 379:213–224
Limborg MT, Heylar SJ, de Bruyn M, Taylor MI, Nielsen EE, Ogden R, Consortium FPT, Bekkevold D (2012) Environmental selection on transcriptome-derived SNPs in a high gene flow marine fish, the Atlantic herring (Clupea harengus). Mol Ecol 21:3686–3703
Luttikhuizen PC, Drent J, Peijnenburg KTCA, van der Veer HW, Johannesson K (2012) Genetic architecture in a marine hybrid zone: comparing outlier detection and genomic clines analysis in the bivalve Macoma balthica. Mol Ecol 21:3048–3061
Mäkinen HS, Cano JM, Merilä J (2006) Genetic relationships among marine and freshwater populations of the European three-spined stickleback (Gasterosteus aculeatus) revealed by microsatellites. Mol Ecol 15:1519–1534
Manni F, Guérard E, Hever E (2004) Geographic patterns of (Genetic, Morphologic, Linguistic) variation: how barriers can be detected by using Monmonier’s algorithm. Hum Biol 76:173–190
McCusker MR, Bentzen P (2010) Positive relationships between genetic diversity and abundance in fishes. Mol Ecol 19:4852–4862
Nielsen EE, Hansen MM, Ruzzante DE, Meldrup D, Grønkjaer P (2003) Evidence of a hybrid-zone in Atlantic cod (Gadus morhua) in the Baltic and the Danish Belt Sea revealed by individual admixture analysis. Mol Ecol 12:1479–1508
Ojaveer H, Jaanus A, MacKenzie BR, Martin G, Olenin S, Radziejewska T, Telesh I, Zettler ML, Zaiko A (2010) Status of biodiversity in the Baltic Sea. PLoS ONE 5:e12467
Olsson J, Mo K, Florin A-B, Aho T, Ryman N (2011) Genetic population structure of perch Perca fluviatilis along the Swedish coast of the Baltic Sea. J Fish Biol 79:122–137
Olsson J, Florin AB, Mo K, Aho T, Ryman N (2012a) Genetic structure of whitefish (Coregonus maraena) in the Baltic Sea. Estuar Coast Shelf S 97:104–113
Olsson J, Bergström L, Gårdmark A (2012b) Abiotic drivers of coastal fish community change during four decades in the Baltic Sea. ICES J Mar Sci 68:961–970
Østbye K, Bernatchez L, Naesje TF, Himberg K-JM, Hindar K (2005) Evolutionary history of the European whitefish Coregonus lavaretus species complex as inferred from mtDNA phylogeography and gill-raker numbers. Mol Ecol 14:4371–4387
Palumbi SR (2003) Population genetics, demographic connectivity, and the design of marine reserves. Ecol Appl 13:146–158
Papakostas S, Vasemägi A, Vähä J-P, Himberg M, Peil L, Primmer CR (2012) A proteomics approach reveals divergent molecular responses to salinity in populations of European whitefish (Coregonus lavaretus). Mol Ecol 21:3516–3530
Park SDE (2001) The Excel microsatellite toolkit, version 3.1. Animal Genomics Laboratory, University College Dublin. (http://animalgenomics.ucd.ie/sdepark/ms-toolkit/)
Patarnello T, Volckaert FAMJ, Castilho R (2007) Pollars of Hercules: is the Atlantic-Mediterranean transition a phylogeographical break? Mol Ecol 16:4426–4444
Pelc RA, Warner RR, Gaines SD (2009) Geographical patterns of genetic structure in marine species with contrasting life histories. J Biogeogr 36:1881–1890
Pereyra R, Bergström L, Kautsky L, Johannesson K (2009) Rapid speciation in a newly opened postglacial marine environment. BMC Evol Biol 9:70. doi:10.1186/1471-2148-9-70
Petit R, Mousadik A, Pons O (1998) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12:844–855
Raymond M, Rousset F (1995) GENEPOP Version 1.2: population genetics software for exact tests and ecumenicism. J Hered 86:248–249
R Development Core Team (2011) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, URL http://www.R-project.org/
Redford KH, Richter BD (1997) Conservation of biodiversity in a world of use. Conserv Biol 13:1246–1256
Reusch TBH, Ehlers A, Hammerli A, Worm B (2005) Ecosystem recovery after climatic extremes enhanced by genotypic diversity. Proc Natl Acad Sci USA 102:2826–2831
Riginos C, Cunningham CW (2005) Local adaptation and species segregation in two mussel (Mytilus edulis x Mytilus trossulus) hybrid zones. Mol Ecol 14:381–400
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Ryman N (2006) CHIFISH: a computer program for testing for genetic heterogeneity at multiple loci using Chi square and Fisher’s exact test. Mol Ecol Notes 6:285–287
Ryman N, Leimar O (2008) Effect of mutation on genetic differentiation among nonequilibrium populations. Evolution 62:2250–2259
Ryman N, Leimar O (2009) G ST is still a useful measure of genetic differentiation—a comment on Jost’s D. Mol Ecol 18:2084–2087
Sandström A (2010) Institutional and substantial uncertainty—explaining the lack of adaptability in fish stocking policy. Mar Policy 34:357–1365
Sandström A (2011) Navigating acomplex policy system—explaining local divergences in Swedish fish stocking policy. Mar Policy 35:419–425
Schmitt T (2007) Molecular biogeography of Europe: Pleistocene cycles and postglacial trends. Front Zool 4:11. doi:10.1186/1742-9994-4-11
Shikano T, Shimada Y, Herczeg G, Merilä J (2010) History vs. habitat type: explaining the genetic structure of European nine-spined stickleback (Pungitius pungitius) populations. Mol Ecol 19:1147–1161
Sivasundar A, Palumbi SR (2010) Life history, ecology and the biogeography of strong genetic breaks among 15 species of Pacific rockfish, Sebastes. Mar Biol 157:1433–1452
Steinert G, Huelsken T, Gerlach G, Binida-Emonds ORP (2012) Species status and population structure of mussels (Mollusca: Bivalvia: Mytilus spp.) in the Wadden Sea of Lower Saxony (Germany). Org Divers Evol 12:387–402
Swatdipong A, Vasemägi A, Kosikinen MT, Piironen J, Primmer CR (2009) Unanticipated population structure of European grayling in its northern distribution: implications for conservation prioritization. Front Zool 6:6. doi:10.1186/1742-9994-6-6
Tatarenkov A, Jönsson RB, Kautsky L, Johannesson K (2007) Genetic structure in populations of Fucus vesiculosus (Phaeophuceae) over spatial scales from 10 m to 800 km. J Phycol 43:675–685
Taylor MS, Hellberg ME (2006) Comparative phylogeography in a genus of coral reef fishes: biogeography and genetic concordance in the Caribbean. Mol Ecol 15:695–707
Teacher AGF, Shikano T, Karjalainen ME, Merilä J (2011) Phylogeography and genetic structuring of European none-spined sticklebacks (Pungitius pugitius)—mitochondrial DNA evidence. PLoS ONE 6(5):e19476. doi:10.1371/journal.pone0019476
Teacher AGF, André C, Merilä J, Wheat CW (2012) Whole mitochondrial genome scan for population structure and selection in the Atlantic herring. BMC Evol Biol 12:248
Teacher AGF, André C, Jonsson PR, Merilä J (2013) Oceanographic connectivity and environmental correlates of genetic structuring in Atlantic herring in the Baltic Sea. Evol Appl 6:549–567
Utter F (1991) Biochemical genetics and fishery management: an historical perspective. J Fish Biol 39(Suppl A):1–20
Utter F, Seeb J (2010) A perspective on positive relationships between genetic diversity and abundance in fishes. Mol Ecol 19:483–3833
Väinölä R, Strelkov P (2011) Mytilus trossulus in Northern Europe. Mar Biol 158:817–833
van Oosterhout C, Hutchinson WF, Wills DP, Shipley P (2004) MICRO-Checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Wares JP, Gaines SD, Cunningham CW (2001) A comparative study of asymmetric migration events across a marine biogeography boundary. Evolution 55:295–306
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Zbawicka M, Drywa A, Smietanka B, Wenne R (2012) Identification and validation of novel SNP markers in European populations of marine Mytilus mussels. Mar Biol 159:1347–1362
Zillén L, Conley DJ, Andrén T, Andrén E, Björck S (2008) Past occurrences of hypoxia in the Baltic Sea and the role of climate variability, environmental change and human impact. Earth Sci Rev 91:77–92
Acknowledgments
This work was carried out within the framework of the BaltGene research program (Baltic Sea Genetic Biodiversity; http://www.tmbl.gu.se:16080/baltgene/index.html). BaltGene was funded from the European Community’s Framework Programme (FP/2007-2013) under Grant agreement n 217246 made with the joint Baltic Sea research and development programme BONUS. The Academy of Finland (Grants 129662 and 134728 to JM, 138043 to AGFT, Grant 141231 to CRP), the Swedish Research Council (NR and CeMEB), the Swedish Research Council for Environmental, Agricultural Sciences and Spatial Planning (Formas; LL, NR, LK, KJ, and CeMEB), The Royal Swedish Academy of Sciences, Marie Curie Intra-European Fellowship no. 327293 (AGFT), the Estonian Science Foundation (Grant No. 8215 to AV), The Gordon and Betty Moore Foundation (FU), and the Carl Trygger Foundation (LL) are gratefully acknowledged. We thank Kirsi Kähkönen and Anna-Karin Ring for help with herring genotyping, Mikhael Ozerov for data analysis advice and numerous people who helped with obtaining the samples used in this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Wennerström, L., Laikre, L., Ryman, N. et al. Genetic biodiversity in the Baltic Sea: species-specific patterns challenge management. Biodivers Conserv 22, 3045–3065 (2013). https://doi.org/10.1007/s10531-013-0570-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-013-0570-9