Abstract
The introduction of species into new environments provides the opportunity for the evolution of new forms through admixture and novel selection pressures. The common brushtail possum, Trichosurus vulpecula vulpecula from the Australian mainland and T.v.fuliginosus from Tasmania, were introduced multiple times to New Zealand from Australia to become one of New Zealand’s most significant pests. Although derived from two subspecies, possums in New Zealand are generally considered to be a single entity. In a previous analysis, we showed that possums in the Hawkes Bay region of New Zealand appeared to consist of at least two overlapping populations. Here, we extend that analysis using a genotype-by-sequencing approach to examine the origins and population structure of those possums and compare their genetic diversity to animals sampled from Australia. We identify two populations of each subspecies in Hawkes Bay and provide clear evidence of a contact zone between them in which a hybrid form is evident. Our analysis of private alleles shows higher rates of dispersal into the contact zone than away from it, suggesting that the contact zone functions as a sink (and hence as a barrier) between the two subspecies. Given the widespread and overlapping distribution of the two subspecies across both large islands in New Zealand, it is possible that many such contact zones exist. These results suggest an opportunity for a more targeted approach to controlling this pest by recognising sub-specific differences and identifying the contact zones that may form between them.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Introductions of species into novel environments can provide the opportunity for the evolution of new forms as they naturalise (Szűcs et al. 2017). In some cases, alien species have been introduced to the same location on multiple occasions from different sources, creating genetic combinations of individuals that have not previously existed (Kolbe et al. 2004). Such admixture can enhance fitness through hybrid vigour or increased genetic diversity (Barker et al. 2019; Dlugosch, Parker 2008; Facon et al. 2008; Hirsch et al. 2017; Rius, Darling 2014) or reduce fitness through outbreeding depression (Rhymer, Simberloff 1996). It is unclear how frequently genetic factors are intrinsic to the success or otherwise of invasive species (Barker et al. 2019; Rius, Darling 2014) although propagule pressure (something that will affect the levels of genetic diversity within an invader) is one of the few acknowledged general characteristics of successful invaders (Bock et al. 2015). Identifying the role that such factors can play in successful establishment, may offer novel perspectives or pathways for the identification of high-risk invaders or provide opportunities for enhanced control of invasive pests (Szűcs et al. 2017).
The introduction of common brushtail possums (Trichosurus vulpecula; hereafter called possums) to New Zealand from Australia is one well documented instance where different forms of the same species have been brought together into a new environment. Introduced to establish a fur trade (Montague, 2000), possums were shipped to New Zealand from Tasmania and several other undocumented locations on mainland Australia, most likely in New South Wales and Victoria, between 1858 and the 1920s. All introductions ceased after that time but the transfer of possums within New Zealand continued into the 1940s. Possums are endemic to Australia with five currently recognised subspecies distributed across the continent and on several offshore islands (How, Hillcox 2000; Kerle et al. 1991). New Zealand has no naturally occurring marsupials and possums are now one of the most serious threats to biodiversity and agriculture in New Zealand damaging vegetation (Cowan 1991; Nugent et al. 2001); preying on birds and their eggs (Brown et al. 1993); and spreading bovine tuberculosis (Innes, Barker 1999). New Zealand management agencies have conducted extensive possum control for decades (Donnell 1995; Morgan 1990; Nugent, Cousins 2014; Nugent, Morriss 2013) and now aim, with the support of conservation groups, to eradicate brushtail possums as part of the “Predator-free New Zealand” campaign (Russell et al. 2015).
It is well known that the different colour forms seen in New Zealand possums are representative to some extent of their different ancestry (Kean 1971; Pracy 1974; Triggs, Green 1989), but it is generally assumed (although seldom stated) that New Zealand possums breed indiscriminately with respect to those origins (Montague 2000). However, a recent microsatellite DNA analysis of possums in Hawkes Bay, on the North Island of New Zealand, identified at least two genetically distinct groups of possums, separated by a zone of putatively introgressed animals (Sarre et al. 2014). That work implied that there were at least two breeding forms in the region, probably representing the Tasmanian subspecies and the south-eastern mainland subspecies (T.v.fuliginosus and T.v.vulpecula respectively), rather than one continuous population (Sarre et al. 2014). That proposition is supported by historical records which show that there were three recorded introductions of possums into the area (Fig. 1): possums taken directly from Tasmania and released in the Lake Waikaremoana area in 1898; “black furred” possums (also presumed to be Tasmanian in origin but translocated from elsewhere in New Zealand) released on the Mahia Peninsula a short time after the first introduction; and possums from an unknown location on mainland Australia released in the Mohaka river area in the early 1900s (Pracy 1974; Sarre et al. 2014).
Brushtail possum introduction sites (Black: Lake Waikaremoana; Mahia Peninsula; Grey: Mohaka River) and sample collection sites (numbered from 1 to 35) in the Hawkes Bay study site in New Zealand. See Sarre et al. (2014) for further information
Resolving complex introduction histories and patterns of admixture in invasive species can be difficult, but access to population level genomic data can reveal how species integrate into new environments and whether diverse origins have enhanced or inhibited their invasion success. Here, we use genotype-by-sequencing to examine the origins and contemporary population structure of possums in Hawkes Bay, and compare their genetic diversity to animals sampled from across their native range in Australia. We test the proposition that the sub-populations, as previously defined by the distribution of microsatellite DNA variation (Sarre et al. 2014), are representative of the ancestral mainland and ancestral Tasmanian origins, and that the Tasmanian and mainland forms have established a hybrid swarm in the Hawkes Bay region that affects the direction and rate of gene flow. We also use asymmetries in the number of private alleles between pairs of sub-populations to determine the direction of that geneflow. A symmetrical distribution of private alleles among sub-populations indicates symmetry in the rates of gene flow between sub-populations and subspecies and would argue against the formation of genetic barriers between the two subspecies. We demonstrate that one sub-population is genetically identifiable as the Tasmanian subspecies T.v. fuliginosus, that possums from that sub-population are distinct from animals that are most likely mainland in origin, and that they have maintained that distinctiveness even after more than 80 years of contact. We also show that the sub-population believed to originate from mainland Australia is indeed largely T.v.vulpecula in origin, although we could not identify the mainland region from which those possums were obtained. Our data support the proposition that brushtail possums in Hawkes Bay do not behave as a single panmictic population, but rather as two subspecies with a narrow zone of contact. Gene flow into and out of that contact zone is asymmetrical and indicative of a genetic barrier between the two subspecies. The formation of similar hybrid barriers elsewhere in New Zealand could have important implications for the control of this invasive species.
Methods
Sample selection
In order to determine the Australian origins of the invasive possums at our study site in Hawkes Bay, New Zealand, we obtained samples from a broad range of locations in eastern mainland Australia (n = 19) and Tasmania (n = 11). Specifically, we sampled tissue (mainly ear tissue) from museum specimens and road killed animals, from six states and territories (Northern Territory, Australian Capital Territory, Queensland, New South Wales, Victoria and Tasmania) and Flinders Island (located in the Bass Strait between Tasmania and mainland Australia; Fig. 2). These samples encompass the four commonly recognised subspecies of Trichosurus vulpecula in eastern and northern Australia (Kerle et al. 1991): T.v vulpecula (south-eastern Australia), T.v. arnhemensis (Northern Territory), T.v. johnstonii (Queensland) and T.v.fuliginosus (Tasmania and Flinders Island) and represent the subspecies most likely to include the source populations for New Zealand possums (Fig. 2). Possums from the Hawkes Bay region of New Zealand were sampled from 24 sites (n = 253) and were selected as a subset of the samples previously genotyped using microsatellite DNA (Sarre et al. 2014). Specifically, we applied New Hybrid analysis (Anderson, Thompson 2002) to the microsatellite data previously reported by Sarre et al. (2014) to estimate the percentage of ancestry for each individual possum and indicate whether an individual is an F1 or F2 cross between subspecies. We then selected individual possums that represented all possible combinations of F1, F2 and ‘pure’ ancestry, as well as an even number of males and females, and used the NewHybrid analysis to sort all 1,605 individuals into three genetic groups: (1) largely Tasmanian in origin, (2) largely Australian mainland in origin, and (3) of mixed ancestry. We then stratified for collection site within those three groups, focussing most on samples from the sites closest to the three known sites of introduction and the previously identified contact zone (Fig. 2), and then randomly chose individuals across those strata for inclusion in the present study using genotyping-by-sequencing.
Genotype-by-sequencing
DNA was extracted from each 2mm2 piece of ear tissue sample by Diversity Arrays Technology Pty Ltd (DArT) using a salting out method as reported by (Couch et al. 2016). We used the DArTseq™ genotype-by-sequencing approach, which uses a combination of restriction enzyme complexity reduction and high throughput sequencing (Courtois et al. 2013; Cruz et al. 2013; Kilian et al. 2012) to simultaneously identify and genotype SNP markers in the absence of a reference genome. A double digest approach with the restriction enzymes PstI and SphI was used for complexity reduction before the addition of custom adapters (Kilian et al. 2012) to restriction site overhangs. Fragments that included both a PstI and an SphI adapter were amplified using primers complementary to the adapters. These also incorporated molecular identifier barcode tags, to allow multiplexing of up to 96 samples per sequencing run. PCR conditions consisted of: denaturation at 94 0C for 1 min; 30 cycles of 94 0C for 20 s, 58 0C for 30 s and 72 0C for 45 s; and a final extension period of 72 0C for 7 min. PCR products were pooled for sequencing on an Illumina HiSeq 2000 platform using 77 cycles of single end sequencing. Raw sequence reads were filtered and processed using a proprietary DArT analytical pipeline. Poor quality sequences were removed, with more stringent criteria being placed upon the barcode region than the rest of the sequence. Approximately 2,000,000 sequences per barcode were identified and used in marker calling, during which identical sequences were collapsed and filtered before screening to identify variable markers using DArT proprietary SNP and SilicoDArT algorithms (DArTsoft14).
Analysis of genotype-by-sequencing data
The R package dartR (Gruber et al. 2018) was used for population genomic analyses of the DArTseq SNP data, including F-statistics, principal coordinates analysis (PCoA) and fixed difference analysis. SNPs were filtered using the RepAvg function (Reproducibility ≥ 0.95), CallRate (0.95), with lower and upper read depth thresholds of 5 and 90 respectively, and to exclude monomorphic loci and secondary SNPs.
We estimated ancestry proportions using SNP data and geographic location in tess3r (Caye et al. 2016), for K values of between two and eight with the default settings (code supplied on request). Fixed differences were identified between the New Zealand and Australian (mainland and Tasmanian combined) possum populations and through a second analysis in which we considered only the sub-populations at the Hawkes Bay study site identified using PCoA and tess3r (Caye et al. 2016).
To further define and visualise the population genetics of animals in Hawkes Bay, we ran a STRUCTURE analysis (Pritchard et al. 2000) using all loci with identical filters (n = 502) for each value of K (assumed number of ancestral populations) between 2 and 5 with 10 repeats each of 100,000 iterations. The first 50,000 iterations were disregarded as burnin and then each of the 10 repeats were summarised using the programs Evanno (Evanno et al. 2005) and Clumpp (Jakobsson, Rosenberg 2007).
We ran STRUCTURE using both ancestral models (No Admixture and Admixture) because they differ in their underlying assumptions about the origins of the population under study (Porras-Hurtado et al. 2013). The No Admixture scenario assumes no prior knowledge about the origin of populations under study. In our case, we know something about the origins of the populations drawn from the introduction history and we have an hypothesis arising from a previous microsatellite DNA study (mainland, Tasmanian and Hybrid) so for this analysis we ignored that knowledge and checked to see if the three groups still emerged. We then ran the Admixture model using our prior knowledge of the origin of the populations to provide insight into the proportion of individuals that can be attributed to each of the presumed ancestral populations. In order to do this, we combined individuals from those sites closest to the mainland introduction sites, those from the previously identified hybrid zone (Sarre et al. 2014) and those closest to the sites populated by possums from Tasmania into sub-populations A, B and C respectively. We expected that A and C (Mainland and Tasmanian origin) would each have a unique signature, while individuals from population B should show a mixture of both A and C.
We also used a NewHybrids analysis to examine population structure among Hawkes Bay possums. While broadly similar to that provided by STRUCTURE, NewHybrids analysis differs in that it uses a detailed inheritance model to check to see if individuals are indeed hybrids when two potential source populations mainly comprising of pure individuals can be identified (Anderson, Thompson 2002). As NewHybrids can only be applied to a maximum of 200 loci, we reduced the available loci to 15,523 by filtering on call rate (≥ 0.95), and then selected the most informative (based on their polymorphic information content as implemented in dartR) 200 SNPs for NewHybrids analysis, using 30,000 sweeps with a burn in of approximately 100 sweeps, and using the Jefferies prior for both Pi and Theta. To further validate the result and its independence from the 200 loci identified as most informative, we re-ran the analysis with 200 randomly selected loci 30 times.
Directionality of gene flow through asymmetry in private alleles
To further determine the direction of interactions within the Hawkes Bay study site, we estimated the levels of asymmetry of gene flow between sub-populations using a novel approach based on the numbers of private alleles present in each sub-population. Specifically, we used the idea that a sub-population that has a net loss of individuals towards another sub-population through emigration will lose private alleles over time as it shares them with the recipient sub-population. Under this process, the sub-population that has fewer emigrants will lose private alleles (through sharing) at a slower rate that its paired sub-population while also accumulating private alleles through genetic drift. We used this logic to explore the direction of gene flow by counting the number of private alleles between sub-populations across the study area while accounting for differences in sample size. This approach differs from those developed for microsatellite DNA markers (Kennington et al. 2003; Sundqvist et al. 2016) by using the raw counts of private alleles rather than comparisons between observed and simulated asymmetries. Under our approach, we consider that a detectable asymmetry in the number of private alleles between neighbouring sub-populations is indicative of asymmetric geneflow between those two sub-populations. Conversely, symmetry in the numbers of private alleles is indicative of equal rates of gene flow in both directions between two sub-populations.
To study the gene flow into and out of the contact zone using private alleles we clustered adjacent sample sites with the aim of obtaining an approximately even number of individuals per cluster. We used the proximity of individuals as the main criterion for clustering, while also considering the genetic similarity of individuals as indicated by the NewHybrid approach (see above) using microsatellite DNA data gathered previously by Sarre et al. (2014). Using this approach, we generated six clusters (A1, B1, B2, B3, C1, C2) within the three sub-populations and confirmed their validity by running a Discriminant Analysis of Principal Component (Jombart et al. 2010). We then counted the number of private alleles for each pairwise comparison among the 6 sub-populations. Sample sizes of individuals in each sub-population varied between 20 and 36. To account for the uneven number of individuals, we sampled all sub-populations to 20 individuals (with replacement) and counted the number of private alleles in each pair of sub-samples. To reduce noise in the data set, we filtered the identified markers by minor allele frequency with a threshold of > 0.1 (based on 40*0.1 > 4 alleles) so that a cluster contained at least 5 alleles before an allele could be called private. This resulted in a data set of 506 SNPs, with a minimum minor allele frequency of 0.1 and mean minor allele frequency of 0.23. The mean number of private alleles between each resampled pair of sub-populations was then calculated from a thousand repetitions of the procedure. Finally, we calculated the mean and 95% confidence intervals of the number of private alleles and the pairwise ratio of private alleles. Sub-population pairs that had a private allele ratio larger than one, in more than 975 out of 1000 cases, were considered significantly asymmetric and hence regarded as exhibiting asymmetric gene flow.
Results
We genotyped 283 individual possums from Australia and New Zealand using the DArTseq approach, generating 27,339 SNPs, 6,596 of which were retained after filtering. Ten individuals were also removed by filtering because of high numbers of missing data leaving a total of 273 possum genotypes for analysis. We identified far fewer private alleles in the T.v.fuliginosus samples from Tasmania (mean = 25.3; n = 11) than in T.v.vulpecula sampled from mainland Australia (mean = 296.6; n = 12). Possums sampled in Tasmania also exhibited lower levels of expected (0.04) and observed (0.03) heterozygosity relative to their mainland counterparts (0.13 and 0.11 respectively). There was strong support (17.3% weight for PCoA Axis 1) for differentiation between Tasmanian individuals of T.v.fuliginosus (including those from Flinders Island) and all other individuals sampled from mainland Australia (Fig. 3). These findings are indicative of the Tasmanian history of geographical isolation, following separation of the island from mainland Australia by rising sea levels more than 9,000 years ago (Veevers 1991). There was also good support for differentiation between T. v. vulpecula and two other mainland subspecies (T. v. arnhemensis and T. v. johnsoni) although there was overlap between the latter two subspecies (Fig. 3).
PCoA of SNP genotypes from Australian brushtail possums, showing clear differences between T.v.fuliginosus (blue–Tasmania and Flinders Island) and T.v.vulpecula (purple–Australian mainland) along axis 1 but with some overlap between the other two subspecies T.v. johnsoni (green) and T.v. arnhemensis (red). Mainland possums show greater diversity than the tightly clustered Tasmanian possums
Within the Hawkes Bay study area, we identified the three distinct sub-populations (A, B, and C) seen previously using microsatellite DNA analysis (Sarre et al. 2014). Sub-population A, genetically the closest to the introduction site of animals from mainland Australia, is discrete from sub-populations B and C, but does not cluster with any mainland Australian animals sampled here (Fig. 4). Sub-populations B and C cluster most closely together and have a lower pairwise FST (0.03; Table 1) than the other pairwise comparisons (A-B: FST = 0.06; A-C: FST = 0.15). The latter FST is equivalent to that observed for T. v. vulpecula and T. v. fuliginosus in their native ranges (FST = 0.16). The highest pairwise FST values were recorded between mainland T. v. vulpecula and groups B and C (FST = 0.25 and 0.40 respectively; Table 1). No fixed allelic differences were observed between any pairwise combination of the Hawkes Bay sub-populations A, B, or C, or between any of those three groups and the two Australian subspecies (Table 2). Private alleles were observed in all pairwise combinations of groups. Overall, these data suggest that that both B and C are composed largely of descendants of Tasmanian animals. All three Hawkes Bay genetic sub-populations exhibit private alleles when compared to each other (Table 2) suggesting that gene flow is restricted between them. The smallest number of private alleles were seen between sub-populations B and C, suggesting that gene flow is higher between those two than between sub-populations A and B or between A and C.
PCoA analysis of SNP genotypes from brushtail possums sampled from the Hawkes Bay region of New Zealand (n = 253), mainland Australia (n = 19) and Tasmania and Flinders Island (n = 11). Australian animals: blue = T.v. johnsoni; aqua = T.v. arnhemensis; pink = T.v. vulpecula (all mainland); purple = T.v. fuliginosus (Tasmania). New Zealand animals: red (A) = samples closest to the introduction site for possums of mainland origin; brown (B) = sites previously identified by Sarre et al. (2014) as likely to contain animals of mixed Tasmanian and mainland ancestry; and olive green (C) = samples closest to the first release site of Tasmanian animals
The PCoA analysis including all mainland Australian, Tasmanian, and Hawkes Bay samples (Fig. 4) shows that the majority of mainland Australian possums are genetically different from both New Zealand and Tasmanian individuals. In contrast, the native Tasmanian possums cluster very closely with the Hawkes Bay possums of putative Tasmanian ancestry (sub-populations B and C), providing additional genetic evidence for Tasmanian ancestry for those Hawkes Bay animals. The possums from sub-population A appear to be distinct from both Hawkes Bay sub-populations B and C and from the native Tasmanian and mainland Australian samples. We identified two mainland Australian individuals, one sampled from around the Sydney area of New South Wales and the other from the Australian Capital Territory, that cluster closest to the Hawkes Bay individuals of putatively mainland ancestry.
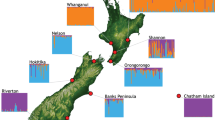
We also used SNP genotype data combined with geographic information on each sample location to generate a visual representation of population genetic structure among the Hawkes Bay possums only using tess3r (Fig. 5; Fig. S1). Here, we examined three scenarios for the ancestral makeup of the possums. (1) Two ancestral sub-populations (mainland Australia and Tasmania origins; k = 2); (2) Three ancestral sub-populations (T.v.vulpecula–mainland, T. v. fuliginosus–Tasmania, and the contact zone population; k = 3); and (3) Four ancestral sub-populations (T.v.vulpecul–mainland, two different introductions of T.v.fuliginosus–Tasmania, and the contact zone population; k = 4). Our estimates of ancestry proportions and ancestral allele frequencies show a clear disjunction between the two subspecies at their contact zone (Fig. 5; Panel 1), with the contact zone being genetically closer to the populations descended from T.v.fuliginosus than to those descended from T.v.vulpecula. The contact zone itself seems to run to the west of the Mohaka River that divides the two release points (Fig. 5; Panel 2). We find no evidence of two distinct ancestral groups of T.v.fuliginosus arising from the dual introductions of possums from Tasmania (Fig. 5; Panel 3) and it appears that this subspecies is relatively homogenous at its source within Tasmania (Fig. 4). These results were similar to those obtained using the STRUCTURE analysis (Fig. 6) which showed that the most appropriate number of groups to emerge from both No Admixture and Admixture models was three (Fig. S2) and, as predicted, demonstrated that the sub-population designated as “B” most closely resembled a combination of sub-populations A and C. Similarly, the NewHybrid plots (Fig. S3; S4) show that sub-population B consists largely of F2 hybrid individuals along.
Estimates of ancestry proportions and ancestral allele frequencies in the Hawkes Bay region of New Zealand identified using tess3r (Caye et al., 2016). This figure is generated using SNP genotype data and geographic information on sample locations. Three ancestral groupings were tested to visualise the likely ancestry of the Hawkes Bay possums: Two ancestral populations (left-hand panel: mainland Australia and Tasmania; k = 2); Three ancestral populations (middle panel: T.v.vulpecula—mainland, T. v. fuliginosus–Tasmania, and the contact zone population; k = 3); and Four ancestral populations (right-hand panel: T.v.vulpecula—mainland, two different introductions of T.v.fuliginosus—Tasmania, and the contact zone population; k = 4). See Fig. S1 for a graphical representation of the cross-validation scores
with small numbers of both parental genotypes while sub-populations A and C are comprised largely of parental individuals with mainland and Tasmanian origins respectively. There is little evidence of F1 individuals in any of the Hawkes Bay animals genotyped.
Finally, asymmetric geneflow was identified between several pairs of the six genetic clusters using our private allele approach (Fig. 7, Table S1). The number of private alleles was consistently lower in sub-population A1 (mainland origin individuals) than in the three sub-populations within the contact zone (B1, B2 and B3). The same was true for sub-population C1 (Tasmanian origin individuals) relative to sub-populations within the contact zone (B1, B2 and B3). We also identified a significant asymmetry in the geneflow between sub-populations C1 and C2. There was no apparent effect of distance on the asymmetries between clusters (Fig. S6). In summary, the asymmetry in private alleles indicates that the direction of geneflow is into the contact zone, with only minor geneflow outwards.
STRUCTURE analysis plot of Hawkes Bay brushtail possums with admixture for all loci (following filtering). The blue vertical lines in the structure plots represent the putative boundaries between the three groups with the left-hand side including samples from sites closest to the introduction sites for mainland origin animals and those on the right including animals closest to the introduction sites for animals from Tasmania
Asymmetry in geneflow estimated using the number of private alleles. Black lines indicate significant asymmetries in geneflow based on 95% confidence intervals obtained through 1000 replicates of resamples of 20 individuals; the level of significance is indicated by the thickness of the lines; arrows indicate the predominant direction of gene flow. Grey arrows indicate non-significant asymmetries between pairs of sub-populations
Discussion
New Zealand possums are the subject of enormous national pest animal control effort and considerable scientific endeavour (Byrom et al. 2016; Clout, Sarre 1997; Eason et al. 2017; Forsyth et al. 2018; Goldson et al. 2015; Livingstone et al. 2015; Montague 2000; Rouco et al. 2018; Rouco et al. 2017). While the history of their introduction from Tasmania and the Australian mainland is extremely well documented, New Zealand possums are considered almost universally to be representatives of a single lineage, albeit one with colour variants. This view is consistent with the prevailing taxonomy that discriminates between them only at the subspecific level (Kerle et al. 1991). Here, we demonstrate that this expectation of possums mixing indiscriminately with respect to subspecies, does not hold in practice in the introduced New Zealand population, at least within the Hawkes Bay study area. Specifically, possums in Hawkes Bay that are derived from two Australian subspecies are more closely resembling their two ancestral entities separated by a contact zone than one integrated population. This is despite having ample opportunity to disperse, mingle, and interbreed for over 80 years (between 40 and 80 generations). We found that new genetic forms, representing individuals of mixed ancestry (F2 +) between the two subspecies, occur mainly in the intermediate contact zone between the sites of the original introductions. We also observe that gene flow tends to move inwards, into that contact zone more commonly than outwards and found no evidence of parental genotypes moving across the contact zone and into the opposite parental sub-population. As such, the contact zone that has formed appears to act as a sink and limits dispersal.
Our results provide clear evidence that possums in Hawkes Bay behave as three distinct groups rather than a single panmictic population—a result that reflects previous findings of genetic structure at the site (Etherington et al. 2014; Sarre et al. 2014). We show that Hawkes Bay possums with putative Tasmanian ancestry have high genetic similarity to modern Tasmanian populations (T.v.fuliginosus), with only modest levels of introgression of alleles from T.v.vulpecula. In contrast, Hawkes Bay possums with putatively Australian mainland origin are genetically distinct from all Australian mainland individuals genotyped. As such, we were unable to identify the mainland Australian source for this group. These genome-level observations from the Hawkes Bay region mirror genetic patterns found more broadly in New Zealand possums using allozyme electrophoresis (Triggs, Green 1989). In that study, populations consisting predominantly of black possums were shown to be genetically similar to possums in Tasmania, while predominantly grey populations were genetically closer to Victorian and New South Wales possums.
There are several possible explanations for the genetic differences observed between the putative T.v.vulpecula animals in Hawkes Bay and the T.v.vulpecula animals sampled from mainland Australia. One possibility is that hybridisation between individuals of mainland Australian and Tasmanian origin masks the genetic signature of the mainland origins of New Zealand possums. Alternatively, the possums that derive from mainland Australia may be descended from a mixture of animals from several mainland sources, that were brought together in Hawkes Bay, and therefore do not resemble any single extant Australian population. Another possibility is that genetic drift—through founder effects and 80 years of isolation and/or selection imposed by a New Zealand environment—may have altered the genetic makeup of the Hawkes Bay population of T.v.vulpecula sufficiently to diverge from mainland Australian possums. This may even have resulted in the retention of genetic variants that are now rare (or even lost) in mainland Australia—a phenomenon seen in stoats introduced to New Zealand from England (Veale et al. 2015). Each of these three factors may contribute to our observations, although the fact that possums from sub-population A are genetically more similar to T.v.fuliginosus and to sub-populations B and C (Fig. 4), than they are to T.v.vulpecula, suggests that there has been leakage of T.v.fuliginosus alleles into sub-population A. A more detailed genomic haplotypic study, combined with mitochondrial DNA analysis, may resolve this issue. Additional sampling closer to the original release site for T.v.vulpecula in the Hawkes Bay may also help to identify the mainland location of origin for these possums.
Our results provide strong evidence that animals in sub-population B are a hybrid form of T.v. vulpecula and T.v. fuliginosus—the product of interbreeding between sub-populations A and C. The formation of a stable contact zone between two colonisers can arise when competition between the two is stronger than intraspecific competition or when hybrids are inviable or less fit than the parent taxa (Goldberg, Lande 2007). To our knowledge, interactions between the two subspecies or the viability of their offspring in nature have not been reported. We cannot therefore distinguish between these two possibilities. The positioning of the contact zone between two colonisers is likely to be attracted to, but not necessarily centred upon, a partial dispersal barrier (Goldberg, Lande 2007). In the case of the possums of Hawkes Bay, the presence of the Mohaka River between the release sites of the mainland Australian origin T.v.vulpecula and the Tasmanian origin T.v. fuliginosus provides such a partial barrier, and may have influenced the positioning of the contact zone near the eastern edge of this river.
The distribution of private alleles among the possum sites suggests that there is more immigration into the contact zone than out of it, from both the T.v.fuliginosus dominated population in the east and north of the zone, and from the T.v.vulpecula dominated population to the south west. The net result is that very few individuals with predominantly T.v.vulpecula ancestry (sub-population A) migrate completely through the contact zone (sub-population B) and integrate into the predominantly T.v.fuliginosus population (sub-population C), or vice versa. The contact zone thus appears to act as a sink for the two subspecies and, as such, represents a non-topographical (invisible) barrier to dispersal.
The existence of non-topographical or invisible barriers have implications for management, in this case particularly for the national program aimed at a predator free New Zealand (Russell et al. 2015). Populations with a mix of both grey (T.v.vulpecula) and black (T.v.fuliginosus) individuals occur in many parts of New Zealand (Cowan 2001; Kean 1971; Triggs, Green 1989) and hybrid animals have been reported previously in the Orongorongo valley on the North Island (Kean 1971). It is therefore possible that many such contact zones occur on the main islands of New Zealand. These zones could inhibit local dispersal and provide barriers that were previously invisible, but which could now be exploited for control. Conversely, possums can disperse quickly into depopulated habitats (Efford et al. 2000; Ji et al. 2001), so poisoning or trapping across contact zones could inadvertently break down existing subspecific barriers and increase local rates of dispersal. Documentation of the existence of additional contact zones on the main islands of New Zealand would provide greater opportunities to take advantage of these invisible barriers, with potential to produce faster localised eradication results.
Possums are the principal wildlife host of bovine tuberculosis in New Zealand (Morgan 1990), and the main cause of Tb persistence in cattle and deer herds in New Zealand, although the disease has not been recorded in Australian possums (Nugent, Cousins 2014; Tweddle, Livingstone 1994). If hybrid possums are less fit than either of the parental subspecies then the risk that possums are susceptible to bovine tuberculosis may be exacerbated. Moreover, there is some evidence that black coated animals, presumably T.v.fuliginosus, exhibit higher resistance to 1080 poison (Henderson et al. 1999; McIlroy 1982) than T.v.vulpecula–a phenomenon that has previously been hypothesised for marsupials more generally (Deakin et al. 2013). Thus, baiting with 1080 (a common approach used to control possums in New Zealand) could affect disproportionately the viability of one subspecies over the other, or even the viability of certain hybrids over others. Further research in this area, particularly through genomic analyses (Eldridge et al. 2020) will be particularly important if total eradication of possums is to be a key management objective.
In conclusion, we have identified clear evidence of a contact zone or border between two subspecies of introduced brushtail possums in Hawkes Bay, New Zealand, that produces a hybrid form. More work is required to determine whether this zone has formed because of inter-sub-specific competition or reduced hybrid fitness. The contact zone functions as a sink, with an asymmetry of gene flow tending towards the contact zone and away from both subspecies. It is therefore likely that the zone functions as a barrier to dispersal between the subspecies. Greater sampling would be required to determine the extent of the contact zone. Given the widespread overlapping distribution of the two subspecies in New Zealand, it is possible that there are many such hybrid zones that could function as barriers to dispersal across New Zealand. These hybrid zones could provide opportunities for managers to adopt a more nuanced approach to control of this introduced pest species, by accounting for differences between the subspecies and by taking advantage of previously unrecognised borders at contact zones.
Code availability
All genetic data are available on https://figshare.com/; https://doi.org/10.6084/m9.figshare.13040522 (embargoed for 12 months).
References
Anderson EC, Thompson EA (2002) A model-based method for identifying species hybrids using multilocus genetic data. Genetics 160:1217–1229
Barker BS, Cocio JE, Anderson SR et al (2019) Potential limits to the benefits of admixture during biological invasion. Mol Ecol 28:100–113
Bock DG, Caseys C, Cousens RD et al (2015) What we still don’t know about invasion genetics. Mol Ecol 24:2277–2297
Brown K, Innes J, Shorten R (1993) Evidence that possums prey on and scavenge birds' eggs, birds and mammals. Notornis 1
Byrom AE, Innes J, Binny RN (2016) A review of biodiversity outcomes from possum-focused pest control in New Zealand. Wildl Res 43:228–253
Caye K, Deist TM, Martins H et al (2016) TESS3: Fast inference of spatial population structure and genome scans for selection. Mol Ecol Resour 16:540–548
Clout MN, Sarre SD (1997) Model marsupial or menace? A review of research on brushtail possums in Australia and New Zealand. Wildl Soc Bull 25:168–172
Couch AJ, Unmack PJ, Dyer FJ, et al. (2016) Who's your mama? Riverine hybridisation of threatened freshwater trout cod and murray cod. PeerJ. 2016
Courtois B, Audebert A, Dardou A, et al. (2013) Genome-wide association mapping of root traits in a japonica rice panel. PloS one. 8
Cowan PE (1991) Effects of introduced Australian brushtail possums (Trichosurus vulpecula) on the fruiting of the endemic New Zealand nikau palm (Rhopalostylis sapida). NZ J Bot 29:91–93
Cowan PE (2001) Advances in New Zealand mammalogy 1990–2000: Brushtail possum. J R Soc N Z 31:15–29
Cruz VMV, Kilian A, Dierig DA (2013) Development of DArT Marker Platforms and Genetic Diversity Assessment of the U.S. Collection of the New Oilseed Crop Lesquerella and Related Species. PloS one. 8
Deakin JE, Cooper DW, Sinclair JJ et al (2013) Towards an understanding of the genetic basis behind 1080 (sodium fluoroacetate) tolerance and an investigation of the candidate gene ACO2. Aust J Zool 61:69–77
Dlugosch KM, Parker IM (2008) Invading populations of an ornamental shrub show rapid life history evolution despite genetic bottlenecks. Ecol Lett 11:701–709
Donnell CFJO (1995) Possums as conservation pests. In: Donnell CFJO (ed) Workshop on Possums as Conservation Pests Department of Conservation, Christchurch, New Zealand,
Eason CT, Shapiro L, Ogilvie S et al (2017) Trends in the development of mammalian pest control technology in New Zealand. New Zealand J Zool 44:267–304
Efford M, Warburton B, Spencer N (2000) Home-range changes by brushtail possums in response to control. Wildl Res 27:117–127
Eldridge MDB, Deakin JE, MacDonald AJ et al (2020) The Oz Mammals Genomics (OMG) initiative: developing genomic resources for mammal conservation at a continental scale. Australian Zool 40:505–509
Etherington TR, Perry GLW, Cowan PE, et al. (2014) Quantifying the direct transfer costs of common brushtail possum dispersal using least-cost modelling: A combined cost-surface and accumulated-cost dispersal kernel approach. PLoS ONE 9
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Facon B, Pointier J-P, Jarne P et al (2008) High genetic variance in life-history strategies within invasive populations by way of multiple introductions. Curr Biol 18:363–367
Forsyth DM, Ramsey DSL, Perry M et al (2018) Control history, longitude and multiple abiotic and biotic variables predict the abundances of invasive brushtail possums in New Zealand forests. Biol Invasions 20:2209–2225
Goldberg EE, Lande R (2007) Species’ borders and dispersal barriers. Am Nat 170:297–304
Goldson SL, Bourdot GW, Brockerhoff EG et al (2015) New Zealand pest management: current and future challenges. J R Soc N Z 45:31–58
Gruber B, Unmack PJ, Berry OF et al (2018) dartr: an r package to facilitate analysis of SNP data generated from reduced representation genome sequencing. Mol Ecol Resour 18:691–699
Henderson RJ, Frampton CM, Morgan DR et al (1999) The efficacy of Baits containing 1080 for control of Brushtail Possums. J Wildl Manag 63:1138–1151
Hirsch H, Brunet J, Zalapa JE et al (2017) Intra- and interspecific hybridization in invasive Siberian elm. Biol Invasions 19:1889–1904
How RA, Hillcox SJ (2000) Brushtail possum, Trichosurus vulpecula, populations in south-western Australia: demography, diet and conservation status. Wildl Res 27:81–89
Innes J, Barker G (1999) Ecological consequences of toxin use for mammalian pest control in New Zealand-an overview. N Z J Ecol 23:111–127
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Ji W, Sarre SD, Aitken N et al (2001) Sex-biased dispersal and a density-independent mating system in the Australian brushtail possum, as revealed by minisatellite DNA profiling. Mol Ecol 10:1527–1537
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC genetics. 11
Kean RI (1971) Selection for melanism and for low reproductive rate in Trichosurus vulpecula (Marsupialia). Proc New Zealand Ecol Soc 18:42–47
Kennington WJ, Gockel J, Partridge L (2003) Testing for asymmetrical gene flow in a Drosophila melanogaster body-size cline. Genetics 165:667–673
Kerle JA, McKay GM, Sharman GB (1991) A systematic analysis of the brushtail possum, Trichosurus-vulpecula (Kerr, 1792)(Marsupialia, Phalangeridae). Aust J Zool 39:313–331
Kilian k, Wenzl P, Huttner E, et al (2012) Diversity Arrays Technology: A Generic Genome Profiling Technology on Open Platforms. In: Paompanon F, Bonin A (eds) Data Production and Analysis in Population Genomics. Springer, New York, pp 67–90
Kolbe JJ, Kolbe JJ, Glor RE et al (2004) Genetic variation increases during biological invasion by a Cuban lizard. Nature 431:177–181
Livingstone PG, Hancox N, Nugent G et al (2015) Development of the New Zealand strategy for local eradication of tuberculosis from wildlife and livestock. N Z Vet J 63:98–107
McIlroy JC (1982) The sensitivity of Australian animals to 1080. III. Marsupial and eutherian herbivores. Australian Wildlife Research 9:505–517
Montague TL (2000) The brushtail possum: Biology. Manaaki Whenua Press, Lincoln, New Zealand, Impact and Management of an Introduced Marsupial
Morgan DR (1990) Behavioural responses of possums (Trichosurus vulpecula) to baits used in aerial control operations. N Z J Ecol 179:601–613
Nugent G, Cousins DV (2014) Zoonotic tuberculosis in Australia and New Zealand. In: Thoen CO, Steele JH and Kaneene JB (eds). Wiley-Blackwell, pp. 221–23_Ch.20
Nugent G, Morriss GA (2013) Delivery of toxic bait in clusters: a modified technique for aerial poisoning of small mammal pests. N Z J Ecol 37:246–255
Nugent G, Fraser W, Sweetapple P (2001) Top down or bottom up? Comparing the impacts of introduced arboreal possums and ‘terrestrial’ ruminants on native forests in New Zealand. Biol Cons 99:65–79
Porras-Hurtado L, Ruiz Y, Santos C, et al. (2013) An overview of STRUCTURE: applications, parameter settings, and supporting software. Frontiers in Genetics 4
Pracy LT (1974) Introduction and Liberation of the Opossum into New Zealand. Information Series. New Zealand Forest Service, Wellington, Information Series Wellington, NZ, pp. 1–28
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rhymer J, Simberloff D (1996) Extinction by hybridization and introgression. Annu Rev Ecol Syst 27:83–109
Rius M, Darling JA (2014) How important is intraspecific genetic admixture to the success of colonising populations? Trends Ecol Evol 29:233–242
Rouco C, Norbury GL, Anderson DP (2017) Movements and habitat preferences of pests help to improve population control: the case of common brushtail possums in a New Zealand dryland ecosystem. Pest Manag Sci 73:287–294
Rouco C, Jewell C, Richardson KS et al (2018) Brushtail possum (Trichosurus vulpecula) social interactions and their implications for bovine tuberculosis epidemiology. Behaviour 155:621–637
Russell JC, Innes JG, Brown PH, et al. (2015) Predator-free New Zealand: Conservation country. BioScience. pp. 520–525
Sarre SD, Aitken N, Adamack AT et al (2014) Creating new evolutionary pathways through bioinvasion: the population genetics of brushtail possums in New Zealand. Mol Ecol 23:3419–3433
Sundqvist L, Keenan K, Zackrisson M et al (2016) Directional genetic differentiation and relative migration. Ecol Evol 6:3461–3475
Szűcs M, Vahsen ML, Melbourne BA et al (2017) Rapid adaptive evolution in novel environments acts as an architect of population range expansion. Proc Natl Acad Sci 114:13501–13506
Triggs SJ, Green WQ (1989) Geographic patterns of genetic distribution in brushtail possums in Trichosurus vulpecula and implications for pest control. N Z J Ecol 12:1–10
Tweddle NE, Livingstone P (1994) Bovine tuberculosis control and eradication programs in Australia and New Zealand. Vet Microbiol 40:23–39
Veale AJ, Holland OJ, McDonald RA et al (2015) An invasive non-native mammal population conserves genetic diversity lost from its native range. Mol Ecol 24:2156–2163
Veevers J (1991) Phanerozoic Australia in the changing configuration of proto-pangea through Gondwanaland and Pangea to the present dispersed continents. Aust Syst Bot 4:1–11
Acknowledgements
We thank Arthur Georges and Andrew O’Reilly Nugent for advice on data analysis and Matt Young, Sam Venables, and Llara Weaver for lab assistance. Samples were kindly contributed by Robert Palmer and Leo Joseph (Australian National Wildlife Collection), Belinda Bauer and Kathryn Medlock (Tasmanian Museum and Art Gallery), Tammy Gordon and David Maynard (Queen Victoria Museum and Art Gallery), Joanne Sumner (Museum Victoria), Sandy Ingleby (Australia Museum), Gavin Dally (Museum and Art Gallery of the Northern Territory) and Elise Dewar (Department of Primary Industries, Parks, Water and Environment). The New Zealand samples were originally curated by Nicola Aitken.
Funding
Project funding was provided to SDS from the Invasive Animals CRC (1.L.21). CDC was supported by an Australian Postgraduate Award and an Invasive Animals CRC postgraduate scholarship.
Author information
Authors and Affiliations
Contributions
Conceived project: SDS, CDC, BG, AJM, PC; Collected samples: PC (direct trapping of New Zealand samples) with SDS; CDC coordination of collection of Australian samples from roadkill and museum specimens; Conducted lab work: CDC; Conducted analyses and generated figures: CDC and BG; Wrote the manuscript: CDC and SDS with contributions from AJM, CEH, BG, PC. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
There were no conflicts of interest or competing interests identified.
Ethics approval
Possum tissues were obtained from museums, the UC tissue collection (reported previously in Sarre et al. 2014), or road killed animals in Tasmania. The latter were collected under a licence to collect roadkill in Tasmania issued by Tasmania’s Department of Primary Industries, Parks, Water and Environment (Permit No. 13240).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Campbell, C.D., Cowan, P., Gruber, B. et al. Has the introduction of two subspecies generated dispersal barriers among invasive possums in New Zealand?. Biol Invasions 23, 3831–3845 (2021). https://doi.org/10.1007/s10530-021-02609-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-021-02609-1