Abstract
Environmental managers require a sensitive and reliable means to prove, with the highest level of confidence possible, where non-native fish species exist and where they do not. Therefore, a nested PCR (nPCR) protocol was developed to detect the environmental DNA (eDNA) of a case-study species, topmouth gudgeon Pseudorasbora parva, which was recently the subject of a national eradication campaign in the UK. The nPCR protocol was tested in the laboratory and in the field in a series of coordinated surveys (eDNA and conventional sampling with traps) at a commercial angling venue in southern England where an initial eDNA survey, based on conventional PCR (cPCR), found P. parva to be present in one of the seven ponds. In the laboratory, the nPCR protocol was on average 100× more sensitive than cPCR, providing a 100% detection rate at DNA concentrations of 3 × 10−8 ng/µL (8 DNA copies per µL). In the field, nPCR and conventional trapping both detected P. parva in only one of the seven angling ponds, the same infested pond as in the previous cPCR-based study. Following eradication work on the infested pond, no eDNA of P. parva was detected using nPCR in either the formerly-infested pond or the adjacent pond, which had been used to quarantine large commercially-valuable fishes. In management applications where the veracity of negative results may be of equal importance as confirmation of positive detections, nPCR protocols provide a useful addition to the analytical toolkit available to inform decision makers responsible for non-native species management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A major challenge in the conservation and management of aquatic ecosystems to combat biological invasions is the detection of non-native species (NNS), both as an early warning after their initial introduction and as a means of determining where to apply management procedures to eradicate or contain the unwanted NNS (Simberloff et al. 2005). To address this challenge, molecular techniques are being developed to detect species, even when present in very low abundance, from the environmental DNA (eDNA) that these organisms shed into the aquatic environment (Darling and Mahon 2011; Rees et al. 2014; Davison et al. 2016). These eDNA detection techniques have so far been used primarily to determine distributions (Takahara et al. 2013; Adrian-Kalchhauser and Burkhardt-Holm 2016) or, when next-generation sequencing methods are used, in studies of biodiversity (Taberlet et al. 2012; Hänfling et al. 2016; Keskin et al. 2016; Valentini et al. 2016). Indeed, virtually all applications of eDNA in aquatic environments to date, whether single-species or meta-barcoding based, have aimed to prove species presence. However, conservation management decisions to address biological invasions, such as whether to attempt an eradication, can have immense resource implications and therefore require highly sensitive analytical techniques with which to prove with the highest possible level of confidence where the undesirable NNS is absent, in addition to where it is present.
Another application where eDNA surveys can be particularly valuable is in assessing the efficacy of NNS eradication attempts (Dunker et al. 2016; Davison et al. 2017). A species of particular concern is topmouth gudgeon Pseudorasbora parva, a small cyprinid fish that first arrived in Europe as a contaminant of Asian carp consignments to Romania in the 1960s and is now widespread throughout the continent (Gozlan et al. 2010). In the UK, where it arrived via the same vector in the mid-1980s, P. parva has been the subject of a nationwide eradication campaign due to the potential threats the species poses to native fish species (Gozlan et al. 2005; Britton et al. 2007; Great Britain Non-native Species Secretariat 2015). Recent research has demonstrated the value of eDNA techniques, as a complement to conventional sampling/capture methods, for determining P. parva distribution within a given area to inform management decisions on eradication procedures (Davison et al. 2017). In that study, at a recreational angling venue in southeast England, conventional PCR-based eDNA analysis provided evidence, of sufficiently high degree of confidence to form the basis of management decisions, that P. parva had survived an attempted eradication in one pond but was not present in six adjacent but unconnected ponds (Davison et al. 2017).
Surveys based on eDNA must consider the risk of errors due to both ‘false positives’ caused by contamination, and ‘false negatives’ resulting from failure to capture eDNA in the collected sample, or from limitations of the laboratory tests used (Guillera-Arroita et al. 2017). Conservation management decisions can have serious consequences on the allocation of resources (personnel, consumables, travel) and potential collateral (environmental, socio-economic, ecosystem services) damage, and so must be based on correct information. In terms of economic costs alone, eradication operations for P. parva at three UK sites cost between £1.90 and £7.90 per m2 of water surface (Britton et al. 2008). Monitoring of eradication success, by any survey technique, also provides a challenge for managers; if success is declared prematurely and management operations ceased, the invasive species can re-establish, resulting in continued ecological impacts and increased management costs (Rout et al. 2009; Davies and Britton 2015). Field and laboratory eDNA protocols of the highest possible accuracy and reliability are necessary in order to provide a high degree of confidence in the survey results so that they can be used to inform management decisions. This is effectively a transition from ‘proof of presence’ to ‘proof of absence’, because a positive eDNA detection can normally be substantiated through conventional sampling, albeit with greater effort when extremely rare (infrequent) species are concerned. Whereas, in the case of a negative detection for a species’ eDNA, it may be impossible, or practically impossible, to prove that the target species is absent, except where the water body can be drained down in a manner that allows all specimens to be captured (again there remains the possibility of specimens of the target species being missed; e.g. Pot et al. 1984). Therefore, a more accurate and reliable eDNA approach is needed, even if this involves increased financial cost.
To date, single-species eDNA surveys have typically used either conventional PCR (cPCR) or quantitative PCR (qPCR) detection protocols (Goldberg et al. 2016). Although qPCR is generally considered to be more sensitive than cPCR (Wilcox et al. 2013; Tréguier et al. 2014; Biggs et al. 2015), some recent studies have reported little difference in their detection ability, e.g. for fish at low density in a mesocosm experiment (Nathan et al. 2014), and for invasive freshwater mussels in a river system (De Ventura et al. 2017). A potentially more sensitive approach is nested PCR (nPCR), which consists of two steps: (1) a cPCR, followed by (2) a qPCR performed on the product of the cPCR. This protocol is expected to increase detection sensitivity but has so far been little used in aquatic eDNA studies, with one exception being for the detection of salmonid fish in rivers (Clusa et al. 2017).
The aim of the present study was to develop an approach, based on conventional and molecular detection methods, to determine the presence or absence of P. parva and inform management decisions on where eradication efforts are warranted. The specific objectives of this study were to: (1) directly compare the sensitivity of cPCR and qPCR protocols in laboratory trials; (2) develop a more sensitive eDNA protocol, based on a nPCR approach, and evaluate its sensitivity; (3) undertake coordinated sampling (conventional trapping and eDNA surveys) of ponds at a known P. parva site to determine the species presence/absence in each water body prior to an eradication attempt; and (4) undertake coordinated sampling of any infested water body after eradication to check for continued persistence.
Materials and methods
Overview and study site
The laboratory and field investigations took place in three stages: (1) development and laboratory testing of PCR protocols and their eDNA detection sensitivity; (2) a coordinated, pre-eradication survey of the seven water bodies at a commercial fishery in south-east England using conventional and eDNA approaches; and (3) a post-eradication survey of two of these water bodies, the angling pond subjected to eradication measures (henceforth the ‘infested pond’) and the adjacent ‘holding’ (i.e. quarantine) pond where the rescued (i.e. large, commercially-valuable) fishes were held during and after the eradication work (details given here below).
The commercial fishery (latitude 51°N, longitude 0°E) is the same angling venue where a previous eDNA survey, using cPCR (Davison et al. 2017), demonstrated P. parva to be present in only one of the venue’s seven human-made angling ponds (areas of 0.5 to 2.4 ha). These ponds are surface-water fed only, i.e. not connected with each other nor with an adjacent stream that flows along the eastern side of the venue (for a map, see Fig. 1 in Davison et al. 2017), and any outflows from the ponds discharge into their own gravel and reed bed filters that do not retain surface water. An invasive population of P. parva was discovered in the infested pond at least as early as April 2004. An eradication attempt was conducted by the fishery owners (intensive trapping combined with introduction of a piscivorous fish species), but the persistence of a low-density population of P. parva was confirmed by cPCR of water samples and focused intensive trapping at the locations where DNA of P. parva was found (Davison et al. 2017).
Protocol sensitivity testing
Sensitivity tests were conducted using DNA extracted from P. parva dorsal muscle tissue (DNeasy Blood and Tissue Kit, Qiagen, Hilden, Germany) collected from a population in southern England; the sequence has been deposited in the open-source database Genbank (www.ncbi.nlm.nih.gov/genbank) with the accession number KR092385 (Davison et al. 2016). Several different approaches to defining limit of detection (LOD) and limit of quantification (LOQ) have been suggested, as reviewed by Hunter et al. (2017). In the present study, LOD is defined as the minimum amount of target DNA at which positive detections were recorded in one or more replicates (following the definition used in other eDNA studies, e.g. Takahara et al. 2013; Tréguier et al. 2014; Biggs et al. 2015). LOQ is generally defined as the lowest amount of target DNA that yields an acceptable level of precision and accuracy (IUPAC 1995; Tréguier et al. 2014). In the present study, LOQ has been specifically defined as detection in 100% of replicates as per Agersnap et al. (2017). Tests to determine LOD and LOQ were applied to two sources of DNA, referred to hereafter as “total DNA” and “plasmid DNA”. “Total DNA” refers to DNA extracted directly from muscle tissue, and therefore comprises both genomic and mitochondrial DNA. “Plasmid DNA” refers to targeted mitochondrial DNA obtained using cloning to create a plasmid solution for use as a standard, enabling calculation of DNA copy numbers. Concentrations of both total and plasmid DNA were measured using a Nanodrop® ND1000 Spectrophotometer (Thermo Scientific, Waltham, MA, USA) and calculated with the software ND-1000 v3.8.1 (Thermo Scientific).
To obtain the plasmid DNA, a preliminary cPCR using total DNA from P. parva was performed to amplify the 350 base-pair target region (Table 1). Cloning was performed using a TOPO® TA Cloning® Kit for Sequencing (Invitrogen, Carlsbad, CA, U.S.A.) with PCR 4-TOPO® vector including competent cells (Escherichia coli), following the manufacturer’s recommended protocol. Bacterial colonies were grown on agar plates with ImMedia™ Amp Blue (Invitrogen). Colonies not displaying blue colouration were selected and inoculated in a liquid medium containing 40 mL of LB-Medium (MP Biomedicals, Santa Ana, CA) and 50 μg/mL of Ampicillin. The plasmids were isolated using QIAprep® Spin Miniprep Kit (Qiagen) and tested with a cPCR to verify the success of the incorporation of the mitochondrial cytochrome c oxidase subunit I (mtCOI) target gene sequence into the plasmid. Copy numbers for plasmid DNA standards were calculated from DNA concentrations and base-pair lengths using the equation of Godornes et al. (2007).
Conventional PCR (350 bp)
The cPCR used in all field surveys, and in sensitivity testing (referred to hereafter as cPCR 350) used specific primers to amplify P. parva DNA, designed to amplify a 350 base-pair sequence of the mtCOI gene (Table 1). Specificity of these primers was tested in silico against all sequences in the NCBI Genbank database using the NCBI Primer Blast software (www.ncbi.nlm.nih.gov/tools/primer-blast/; Ye et al. 2012). The primers were also tested experimentally in cPCRs against 0.1 ng genomic DNA extracts from Cyprinidae species which are likely to occur at the study site: common carp Cyprinus carpio, common bream Abramis brama, roach Rutilus rutilus and rudd Scardinius erythrophthalmus, with none of the triplicate cPCRs showing amplification for any of these species (Davison et al. 2016).
A further pair of cPCR primers, referred to hereafter as cPCR 101, were designed to amplify a target region of 101 base pairs. The purpose of this primer pair was to enable a direct comparison in sensitivity tests with the pair targeting a longer region (cPCR 350) to assess whether length of target region affected sensitivity. This primer pair was used only for comparative sensitivity testing in the laboratory, and it was not used in the field surveys.
Conventional PCRs were performed with a total reaction mixture of 20 μL containing 2 μL of DNA samples (total DNA, plasmid or eDNA), 0.5 μM of each specific primer, 10 μL of HotStar Taq® Plus DNA polymerase 2× (Qiagen Fast Cycling PCR Kit) and 2 μL of Coral Load Fast Cycling Dye 10× (Qiagen). De-ionised water was added to obtain the total mixture volume. The cycling conditions were 95 °C for 5 min, followed by 35 cycles of 96 °C for 5 s, 62 °C for 5 s and 68 °C for 12 s, with a final extension at 72 °C for 1 min. PCR products were visualised after 60 min of electrophoresis migration on 2% agarose gel, stained with SYBR™ Gold Nucleic Acid Gel Stain (Invitrogen). For both laboratory validation trials and eDNA field samples, five cPCR replicates were analysed in each machine run, on three discrete machine runs (i.e. 15 replicates in total).
Quantitative and nested PCR
Specific P. parva primers and probes were designed for qPCR to amplify a 101 base-pair sequence of the mitochondrial cytochrome c oxidase subunit I (COI) gene, occurring within the sequence amplified by the cPCR 350 primers (Table 1). The primers were successfully tested for specificity in silico against all sequences in the Genbank database using NCBI Primer-BLAST software, in which no species likely to be present at UK freshwater sites corresponded to the primer pair to within four base-pair mismatches (a level of mismatch within that used for assessing specificity by recent fish eDNA studies, e.g. Harper et al. 2019). The primers were also tested experimentally against genomic DNA of C. carpio, A. brama, R. rutilus and S. erythrophthalmus, with no amplification observed. Real-time qPCRs were performed using an Applied Biosystems™ Step One™ system (Applied Biosystems, Foster City, CA, USA) using the following thermocycling profile: 2 min at 50 °C, 10 min at 95 °C, followed by 35 cycles of 15 s denaturation at 95 °C, and 60 s annealing-extension at 60 °C.
PCRs were performed in a 20 μL reaction mixture containing 2 μL of total DNA, plasmid or eDNA, 1 μL of assay mix (18 μM forward and reverse primers and 5 μM probe) for the targeted species (Applied Biosystems™), 10 μL of TaqMan® Genotyping Master Mix (Applied Biosystems™) and 7 μL of de-ionised water. Samples and standards were analysed in triplicate. The standard curve comprised a range of five or six dilutions of a selected standard (plasmid or total DNA), acting as positive samples to confirm reaction efficacy. The dilution series was constructed from the standard on the day of analysis. Finally, the lengths of the qPCR products were checked using 2% agarose gel electrophoresis after addition of DNA Gel Loading Dye (6×) (Invitrogen).
The nPCR protocol consisted of two steps: (1) a cPCR, using the cPCR 350 primer pair and the protocol described above; and if the initial cPCR produced a negative result, then (2) a qPCR was performed on 2 μL from each completed cPCR. Five cPCR replicates were performed on each sample. Each cPCR replicate that produced a negative result was then subjected to qPCR in triplicate.
Coordinated pre- and post-eradication surveys
Water samples were collected on 6 and 7 September 2016 from 12 littoral zone locations spread at approximately equal distances from each other around the shores of all ponds, using the same statistically-designed sampling protocol developed specifically for these ponds (Davison et al. 2017). Water samples were collected at about 1.5 m distance from the bank using a 183-cm-long sampling pole fitted with a 500 mL polypropylene sampling cup (Camlab Ltd, Cambridge, UK), which, between samples, was disinfected thoroughly with Microsol 3+ sterilising solution (Anachem Ltd, Luton, UK) and washed with de-ionised water. New sampling poles and cups were used for each pond to ensure no contamination risk. The sampling cup was moved in a standardised manner from the bank, to the greatest extent reached by the pole, ensuring no contact with the bottom sediment. At each sampling location, three replicates of 300 mL water, obtained using the sampling cup, were injected through a Sterivex-GP 0.22 μm sterile filter cartridge (EMD Millipore, Billerica, MA, USA) using a 50 mL sterile syringe (Thermo Scientific) that is designed to attach directly onto the cartridge’s input opening. Cartridges from each location were sealed in individual plastic bags and immediately frozen (− 20 °C) for transportation back to the laboratory. On each sampling day, water from a sterilized bottle of de-ionised water from the laboratory was also filtered, handled and transported in the same manner as the pond samples, and analysed in the laboratory to test for contamination.
Conventional trapping in each of the angling ponds consisted of ten, previously-unused, rectangular minnow traps of 45 cm length and 25 cm width and height with 3 mm mesh, which were deployed on 7 September 2016 (i.e. same date as eDNA surveying, the use of new traps to avoid potential DNA contamination). Traps were baited using fishmeal pellets (21 mm diameter) and exposed for 12 h, with the numbers of fish captured recorded for P. parva only. Only five traps were used in the pond known to contain P. parva due to a periodical check of the traps revealing high numbers of P. parva captured. Once P. parva presence was confirmed, the traps were retrieved.
Post-eradication surveys (eDNA, trapping) were completed approximately six months after the fishery undertook procedures to eradicate P. parva. This consisted of complete drain-down of the infested (i.e. treatment) pond during which the larger and more commercially valuable fish were collected, passed through a salt bath (≈ 30 ppt) and placed into one of the adjacent ponds, henceforth the ‘quarantine’ pond. On 8 June 2017, three replicate water samples of 300 mL were collected (as described above) at 12 littoral zone locations from the treatment and the quarantine ponds. These samples were collected and analysed in the same manner as described above.
Laboratory processing of the pond-water samples
In the laboratory, DNA was extracted from the cartridges using a PowerWater Sterivex™ DNA Isolation Kit (MoBio, Carlsbad, CA, USA), with a final elution volume of 100 mL. The extracted sample was then diluted 1:5 in deionised water to dilute potential inhibitors (McKee et al. 2015), and a nPCR then performed using the conditions described above on 2 µL of diluted sample. To confirm that negative results in the qPCR were not detection errors (‘false negatives’) caused by PCR inhibition, five replicate samples from four locations per pond were spiked with 0.01 ng of P. parva total DNA and compared against controls of deionised water spiked with the same DNA quantity.
Sample extraction, PCR preparation and post-PCR analyses were each undertaken in separate rooms of a laboratory dedicated to molecular biology, observing strict anti-contamination procedures (no transfer of equipment between rooms; changing of lab coats when moving between rooms; thorough cleaning of all equipment and surfaces before and after use, and treating of equipment under UV light; use of sterile filter tips for pipettes). Increased risk of contamination is an important consideration with nested PCR protocols, due to the increased handling of amplified DNA. This risk was minimised by placing prepared reagents into well plates in a fume cabinet in a separate room from where the completed cPCR template was added, using different pipettes and gloves.
Statistical analysis
Differences between treatments in the sensitivity testing (plasmid DNA cPCR 350 vs. plasmid DNA cPCR 101; plasmid DNA cPCR 350 vs. plasmid DNA qPCR; total DNA cPCR 350 vs. total DNA qPCR) were tested by Permutational (Univariate) Analysis of Variance (PERANOVA). This was based on a one fixed-factor design consisting of Detection rate at two levels. PERANOVA was carried out in PERMANOVA + v1.0.8 for PRIMER v6.1.18 (Anderson et al. 2008), using a Euclidean distance, 9999 permutations of the residuals under a full model (Anderson and Robinson 2001), and with statistical effects evaluated at α = 0.05. Notably, the advantage of PERANOVA compared to ‘traditional’ (fully parametric) ANOVA is that the stringent assumptions of normality and homoscedasticity, which often prove unrealistic when dealing with biological data sets, are ‘relaxed’ considerably.
Results
Sensitivity testing
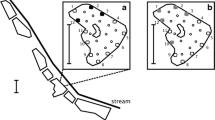
No statistical difference (permutational ANOVA) in sensitivity was observed between cPCR 350 and qPCR, in laboratory trials using plasmid DNA (F #1,20 = 0.924, P = 0.415; # = permutational) or total DNA (F #1,20 = 0.569, P = 0.480; # = permutational). Nested PCR proved the most sensitive of the three protocols (Fig. 1a, b; Table 2). In trials on plasmid DNA (Fig. 1b), the LOQ was 3.34 × 10−8 ng/µL using nPCR, corresponding to 8 DNA copies per µL, compared to 3.34 × 10−6 ng/µL (764 copies per µL) using both cPCR 350 and qPCR. At target DNA concentrations below LOQ, cPCR 350 produced more positive detections than qPCR for a given concentration. The LOD for qPCR was 3.34 × 10−8 ng/µL (76 copies per µL), with no positive detections at 3.34 × 10−9 ng/µL, whereas both cPCR 350 and nPCR produced detections at the lowest DNA concentration tested, 3.34 × 10−10 ng/µL (< 1 copy per µL), in 20% and 27% of replicates, respectively.
When total DNA was tested (Fig. 1a), nPCR again proved more sensitive than cPCR 350 or qPCR in terms of LOQ. In all three protocols, a more sensitive LOQ (i.e. more detections at low concentrations) was achieved when using plasmid DNA than when using total DNA. Although the LOQ showed less sensitivity using cPCR than using qPCR, at concentrations below qPCR LOQ there was a higher probability of detection using cPCR 350 (e.g. 31% of replicates at 9.79 × 10−5 ng/µL) than using qPCR (11% of replicates at 9.79 × 10−5 ng/µL). As with plasmid DNA, the number of detections using qPCR declined to zero before cPCR and nPCR (qPCR no detections at or below 9.79 × 10−6 ng/µL, nPCR detections in 15% of replicates at 9.79 × 10−8 ng/µL, cPCR 350 detections in 8% of replicates at 9.79 × 10−8 ng/µL).
There were no statistically significant differences in sensitivity between cPCR 350 and cPCR 101 (F #1,20 = 0.569, P = 0.480; # = permutational).
Field surveys
In pre-eradication surveys, both eDNA and trapping, P. parva was detected in one pond only, the known infested (i.e. treatment) pond, with eDNA detection at the first (cPCR) stage of the nPCR protocol, thus confirming the previous cPCR results of Davison et al. (2017). All water samples from the other six ponds of this angling venue were negative for DNA of P. parva, and this included all replicates at both the cPCR and qPCR stages of the nPCR protocol. No inhibition was detected in any sample (following the dilution steps undertaken to reduce inhibition). All positive controls (total DNA controls in the cPCR stage, and plasmid DNA standard curves in the qPCR) demonstrated successful amplification.
In the post-eradication surveys, all water and trappings samples from both the treatment pond and the quarantine pond yielded negative results, including all replicates at both the cPCR stage and the final qPCR stage of the nPCR protocol. No inhibition of eDNA was detected, and all positive controls demonstrated successful amplification.
Discussion
All three protocols (cPCR, qPCR and nPCR) proved to be applicable for evaluating the success of the P. parva eradication operation. The greater sensitivity of nPCR in the present study, relative to cPCR or qPCR approaches, confirms the results of two similar studies (Clusa et al. 2017; Jackson et al. 2017). The LOQ of the nPCR was lower than the detection limit of 1 × 10−5 ng/µL reported by Clusa et al. (2017), whereas Agersnap et al. (2017) demonstrated a higher level of sensitivity (LOD and LOQ of one copy per µL of extracted sample) using qPCR than was recorded in this study. A LOD of less than one copy number per reaction has been reported in other studies using qPCR (Hunter et al. 2017; Serrao et al. 2017).
The increased sensitivity demonstrated by this nPCR protocol is likely to be largely a consequence of the increased number of cycles, with 70 in the two rounds of amplification compared to 35 in the cPCR or qPCR alone. Increased sensitivity could potentially have been obtained from the cPCR or qPCR protocols by increasing the number of cycles to 45–55 as used in some other eDNA studies (e.g. Tréguier et al. 2014; Biggs et al. 2015). However, one potential advantage of the nPCR approach is that it uses refreshed (new) reagents after 35 cycles. Length of DNA amplicon targeted by the primers is another factor that could conceivably affect sensitivity (Deagle et al. 2006), but comparative testing of cPCR primers for two different fragment lengths (101 bp and 350 bp) showed no statistical difference in the present study. Piggott (2016) similarly found no evidence that target amplicon size was a limiting factor in eDNA detectability.
In comparative tests using plasmid DNA, qPCR was not found to be more sensitive than cPCR (Fig. 1), although it should be noted that there were differences between the two protocols, e.g. different reagents and fragment target lengths. This contradicts some other studies (e.g. Tréguier et al. 2014; Biggs et al. 2015; Piggott 2016) but is consistent with others (e.g. Nathan et al. 2014; De Ventura et al. 2017). Indeed, at concentrations below the 100% detection limit, more detections were obtained using cPCR than with qPCR. In such circumstances, where both positive and negative results are obtained from a set of replicates, it would be important in a management context to set a threshold limit on the number of positive replicates required before assigning presence/absence status. De Ventura et al. (2017) similarly found cPCR to provide a more robust protocol than qPCR in a direct comparison, producing fewer false negatives at very low target DNA concentrations. This is despite the inherent challenge of interpretation of weak bands, which in cPCR may be on the borderline of visual acuity of the observer. These findings, particularly when the lower financial costs are considered, demonstrate that cPCR remains an effective tool, particularly for routine monitoring and/or survey applications.
Plasmid DNA provided a much lower value for sensitivity (LOQ) than total DNA, using all three PCR protocols. This demonstrates one of the difficulties in comparing protocol sensitivity between different studies, with some reporting values for DNA sensitivity based on total tissue-extracted DNA (e.g. Tréguier et al. 2014; Biggs et al. 2015) whereas others base these values on plasmid DNA (e.g. Takahara et al. 2012; Jane et al. 2015). The use of plasmid DNA enables the calculation of copy numbers for reporting relative sensitivities of protocols as recommended by Goldberg et al. (2016).
‘Proving a negative’, i.e. declaring a species to be absent using any survey method, is difficult due to the inherent uncertainty that is associated with any form of field sampling approach (Rout et al. 2009; Britton et al. 2011). Detection rates using conventional methods are well known to vary according to the gear used (e.g. Jackson and Harvey 1997), in the use of citizen science (e.g. Ruiz-Gutierrez et al. 2016), and the spatial scale used (Barry and Elith 2006). Even a moderate variation in the technique used to apply a sampling method, such as how the dip net is handled in the water during point-abundance sampling by electrofishing (Copp and Garner 1995), will affect the likelihood of capturing a species and consequently its ‘detection’ for purposes of calculating species richness. However, any increase in the sensitivity of a sampling protocol, such as seen here with the nPCR protocol, provides increased confidence in a negative detection result. For example, the nPCR protocol presented here was up to 100× more sensitive at detecting plasmid DNA than the cPCR protocol used previously to survey these same water bodies (Davison et al. 2017). The increased sensitivity did not, in this instance, result in detections that would not have been achieved by cPCR or conventional trapping, and further field trials on water bodies containing smaller populations of the target species would be beneficial to demonstrate the merit of the technique. The extent to which this increased sensitivity translates into increased detections in environmental samples (i.e. to assess the level of sensitivity that is required from the laboratory protocol to detect eDNA from a small fish population in a water body of a given size) requires further study. False negatives can derive from several stages in sampling design, in both field and laboratory (Darling and Mahon 2011). In field applications, it is conceivable that modifications to sampling strategy, such as sampling location (within a water body) or quantity of water filtered, may have as great an effect on the overall result (i.e. positive/negative) as improvements to the sensitivity of the laboratory protocol.
The results of the eDNA survey of the seven water bodies matched the results of the conventional sampling, with both methods detecting P. parva in the known infested pond, and neither method detecting the species in the six remaining ponds. Experimental trials (in 100 m2 mesocosms) have demonstrated that trapping is not completely effective at detecting P. parva at low densities (Britton et al. 2011). Baited traps (deployed for 1 h) showed 100% detection at densities > 0.5 m−2, but imperfect detection at densities of 0.02 and 0.1 m−2 (Britton et al. 2011). Surveys of other fish species using eDNA methods have frequently proved more effective than conventional methods at detecting species (e.g. Takahara et al. 2013; Janosik and Johnston 2015; Sigsgaard et al. 2015). Further study would be needed to determine the efficiency of eDNA sampling, relative to conventional sampling, at varying population densities, as such comparisons will be species specific (Hinlo et al. 2017) and are also likely to be site specific.
Following draw-down of the infested pond by the fishery owners, the negative eDNA detection for P. parva in the infested and quarantine ponds indicated that eradication of the infested pond had been successful and that it was highly unlikely that any P. parva were transferred into the quarantine holding pond. The six-month delay between eradication and water sampling would have provided sufficient time for: (1) any surviving P. parva to introduce more eDNA into the ponds (and possibly even reproduce, as spawning in southern England occurs between April and June; Beyer 2008), and (2) any remaining DNA from dead/removed P. parva to degrade to non-detectable levels (Dejean et al. 2011). The present study, therefore, clearly demonstrates the potential of eDNA surveying as a tool to identify which water bodies require eradication efforts and to assess the success of those eradication attempts (Dunker et al. 2016; Davison et al. 2017).
Environmental DNA surveys are often seen as a less-expensive substitute for conventional methods (Biggs et al. 2015; Sigsgaard et al. 2015), although this does not hold true for all sampling programmes under all circumstances (Smart et al. 2016). For example, to detect brook trout Salvelinus fontinalis in streams, eDNA analysis imposed 67% less cost than triple-pass electrofishing and required lower sampling effort (Evans et al. 2017). However, for simple presence/absence detection in that study, eDNA was more expensive than electrofishing, when fishing operations were halted upon first detection of the target species. Conventional fish surveying approaches can also provide information on population structure, which eDNA surveys cannot, and therefore eDNA sampling may in some applications represent a complement, rather than an alternative, to conventional sampling.
The work required to confirm definitive absence of an invasive fish species in a pond would be both costly and potentially environmentally destructive. The enhanced sensitivity of the nPCR protocol provides increased confidence that the negative results obtained were not ‘false negatives’, but this comes at increased financial cost. In the present study, the extra analysis involved with the two steps of the nPCR protocol increased the combined costs of consumables and laboratory analysis by 1.6×. Despite these considerations, nPCR protocols are recommended for any eDNA survey in which a high level of confidence is required in the declaration of a negative result, e.g. where the presence or absence of a species will form the basis of decisions for management action.
References
Adrian-Kalchhauser I, Burkhardt-Holm P (2016) An eDNA assay to monitor a globally invasive fish species from flowing freshwater. PLoS ONE 11:e0147558
Agersnap S, Larsen WB, Knudsen SW, Strand D, Thomsen PF, Hesselsøe M, Mortensen PB, Vrålstad T, Møller PR (2017) Monitoring of noble, signal and narrow-clawed crayfish using environmental DNA from freshwater samples. PLoS ONE 12:e0179261
Anderson MJ, Robinson J (2001) Permutation tests for linear models. Aust N Z J Stat 43:75–88
Anderson MJ, Gorley RN, Clarke KR (2008) PERMANOVA + for PRIMER: guide to software and statistical methods. PRIMER-E, Plymouth
Barry S, Elith J (2006) Error and uncertainty in habitat models. J Appl Ecol 43:413–423
Beyer K (2008) Ecological implications of introducing Leucaspius delineatus (Heckel, 1843) and Pseudorasbora parva (Temminck and Schlegel, 1842) into inland waters in England. Ph.D. thesis, University of Hull. https://hydra.hull.ac.uk/resources/hull:1002. Accessed 9 June 2018
Biggs J, Ewald N, Valentini A, Gaboriaud C, Dejean T, Griffiths RA, Foster J, Wilkinson JW, Arnell A, Brotherton P, Williams P (2015) Using eDNA to develop a national citizen science-based monitoring programme for the great crested newt (Triturus cristatus). Biol Conserv 183:19–28
Britton JR, Davies GD, Brazier M, Pinder AC (2007) A case study on the population ecology of a topmouth gudgeon (Pseudorasbora parva) population in the UK and the implications for native fish communities. Aquat Conserv Mar Freshw Ecosyst 17:749–759
Britton JR, Brazier M, Davies GD, Chare SI (2008) Case studies on eradicating the Asian cyprinid Pseudorasbora parva from fishing lakes in England to prevent their riverine dispersal. Aquat Conserv Mar Freshw Ecosyst 18:867–876
Britton JR, Pegg J, Gozlan RE (2011) Quantifying imperfect detection in an invasive pest fish and the implications for conservation management. Biol Conserv 144:2177–2181
Clusa L, Ardura A, Fernández S, Roca AA, García-Vázquez E (2017) An extremely sensitive nested PCR-RFLP mitochondrial marker for detection and identification of salmonids in eDNA from water samples. PeerJ 5:e3045
Copp GH, Garner P (1995) Evaluating microhabitat use of fish larvae and juveniles with Point Abundance Sampling. Folia Zool 44:145–158
Darling JA, Mahon AR (2011) From molecules to management: adopting DNA-based methods for monitoring biological invasions in aquatic environments. Environ Res 111:978–988
Davies GD, Britton JR (2015) Assessing the efficacy and ecology of biocontrol and biomanipulation for managing invasive pest fish. J Appl Ecol 52:1264–1273
Davison PI, Créach V, Liang W-J, Andreou D, Britton JR, Copp GH (2016) Laboratory and field validation of a simple method for detecting four species of non-native freshwater fish using eDNA. J Fish Biol 89:1782–1793
Davison PI, Copp GH, Créach V, Vilizzi L, Britton JR (2017) Applications of environmental DNA analysis to inform invasive fish eradication operations. Sci Natl 104:35
De Ventura L, Kopp K, Seppälä K, Jokela J (2017) Tracing the quagga mussel invasion along the Rhine river system using eDNA markers: early detection and surveillance of invasive zebra and quagga mussels. Manag Biol Invasions 8:101–112
Deagle BE, Eveson JP, Jarman SN (2006) Quantification of damage in DNA recovered from highly degraded samples—a case study on DNA in faeces. Front Zool 3:11
Dejean T, Valentini A, Duparc A, Pellier-Cuit S, Pompanon F, Taberlet P, Miaud C (2011) Persistence of environmental DNA in freshwater ecosystems. PLoS ONE 6:e23398
Dunker KJ, Sepulveda AJ, Massengill RL, Olsen JB, Russ OL, Wenburg JK, Antonovich A (2016) Potential of environmental DNA to evaluate northern pike (Esox lucius) eradication efforts: an experimental test and case study. PLoS ONE 11:e0162277
Evans NT, Shirey PD, Wieringa JG, Mahon AR, Lamberti GFA (2017) Comparative cost and effort of fish distribution detection via environmental DNA analysis and electrofishing. Fisheries 42:91–99
Godornes C, Leader BT, Molini BJ, Centurion-Lara A, Lukehart SA (2007) Quantitation of rabbit cytokine mRNA by real-time RT-PCR. Cytokine 38:1–7
Goldberg CS, Turner CR, Deiner K, Klymus KE, Thomsen PF, Murphy MA, Spear SF, McKee A, Oyler-McCance SJ, Cornman RS, Laramie MB (2016) Critical considerations for the application of environmental DNA methods to detect aquatic species. Methods Ecol Evol 7:1299–1307
Gozlan RE, St-Hilaire S, Feist SW, Martin P, Kents ML (2005) Disease threat to European fish. Nature 435:1046
Gozlan RE, Andreou D, Asaeda T, Beyer K, Bouhadad R, Burnard D, Caioloa N, Cakic P, Djikanovic V, Esmaeili HR, Falka I, Golicher D, Harka A, Jeney G, Kováč V, Musil J, Nocita A, Povž M, Poulet N, Virbickas T, Wolter C, Tarkan AS, Tricarico E, Trichkova T, Verreycken H, Witkowski A, Zhang C-G, Zweimueller I, Britton JR (2010) Pan-continental invasion of Pseudorasbora parva: towards a better understanding of freshwater fish invasions. Fish Fish 11:315–340
Great Britain Non-native Species Secretariat (2015) The Great Britain Invasive Non-native Species strategy. UK Government, York
Guillera-Arroita G, Lahoz-Monfort JJ, Van Rooyen AR, Weeks AR, Tingley R (2017) Dealing with false-positive and false-negative errors about species occurrence at multiple levels. Methods Ecol Evol 8:1081–1091
Hänfling B, Handley LL, Read DS, Hahn C, Li J, Nichols P, Blackman RC, Oliver A, Winfield IJ (2016) Environmental DNA metabarcoding of lake fish communities reflects long-term data from established survey methods. Mol Ecol 25:3101–3119
Harper LR, Griffiths NP, Handley LL, Sayer CD, Read DS, Harper KJ, Blackman RC, Li J, Hänfling B (2019) Development and application of environmental DNA surveillance for the threatened crucian carp (Carassius carassius). Freshw Biol 64:93–107
Hinlo R, Furlan E, Suitor L, Gleeson D (2017) Environmental DNA monitoring and management of invasive fish: comparison of eDNA and fyke netting. Manag Biol Invasions 8:89–100
Hunter ME, Dorazio RM, Butterfield JSS, Meigs-Friend G, Nico LG, Ferrante JA (2017) Detection limits of quantitative and digital PCR assays and their influence in presence–absence surveys of environmental DNA. Mol Ecol Resour 17:221–229
IUPAC (1995) Nomenclature in evaluation of analytical methods including detection and quantification capabilities. Pure Appl Chem 67:1699–1723
Jackson DA, Harvey HH (1997) Qualitative and quantitative sampling of lake fish communities. Can J Fish Aquat Sci 54:2807–2813
Jackson M, Myrholm C, Shaw C, Ramsfield T (2017) Using nested PCR to improve detection of earthworm eDNA in Canada. Soil Biol Biochem 113:215–218
Jane SF, Wilcox TM, McKelvey KS, Young MK, Schwartz MK, Lowe WH, Letcher BH, Whiteley AR (2015) Distance, flow and PCR inhibition: eDNA dynamics in two headwater streams. Mol Ecol Resour 15:216–227
Janosik AM, Johnston CE (2015) Environmental DNA as an effective tool for detection of imperilled fishes. Environ Biol Fish 98:1889–1893
Keskin E, Unal EM, Atar HH (2016) Detection of rare and invasive freshwater fish species using eDNA pyrosequencing: lake Iznik ichthyofauna revised. Biochem Syst Ecol 67:29–36
McKee AM, Spear SF, Pierson TW (2015) The effect of dilution and the use of a post extraction nucleic acid purification column on the accuracy, precision and inhibition of environmental DNA samples. Biol Conserv 183:70–76
Nathan LM, Simmons M, Wegleitner BJ, Jerde CL, Mahon AR (2014) Quantifying environmental DNA signals for aquatic invasive species across multiple detection platforms. Environ Sci Technol 48:12800–12806
Piggott M (2016) Evaluating the effects of laboratory protocols on eDNA detection probability for an endangered freshwater fish. Ecol Evol 6:2739–2750
Pot W, Noakes DLG, Ferguson MM, Coker G (1984) Quantitative sampling of fishes in a simple system: failure of conventional methods. Hydrobiologia 114:249–254
Rees HC, Maddison BC, Middleditch DJ, Patmore JRM, Gough KC (2014) The detection of aquatic animal species using environmental DNA—a review of eDNA as a survey tool in ecology. J Appl Ecol 51:1450–1459
Rout TM, Thompson CJ, McCarthy MA (2009) Robust decisions for declaring eradication of invasive species. J Appl Ecol 46:782–786
Ruiz-Gutierrez V, Hooten MB, Grant EHC (2016) Uncertainty in biological monitoring: a framework for data collection and analysis to account for multiple sources of sampling bias. Methods Ecol Evol 7:900–909
Serrao NR, Reid SM, Wilson CC (2017) Establishing detection thresholds for environmental DNA using receiver operator characteristics. Conserv Genet Resour. https://doi.org/10.5061/s12686-017-0817-y
Sigsgaard EE, Carl H, Møller PR (2015) Monitoring the near-extinct European weather loach in Denmark based on environmental DNA from water samples. Biol Conserv 183:46–52
Simberloff D, Parker IM, Windle PN (2005) Introduced species policy, management, and future research needs. Front Ecol Environ 3:12–20
Smart AS, Weeks AR, Van Rooyen AR, Moore A, McCarthy MA, Tingley R (2016) Assessing the cost-efficiency of environmental DNA sampling. Methods Ecol Evol 7:1291–1298
Taberlet P, Coissac E, Pompanon F, Brochmann C, Willerslev E (2012) Towards next-generation biodiversity assessment using DNA metabarcoding. Mol Ecol 21:2045–2050
Takahara T, Minamoto T, Yamanaka H, Doi H, Kawabata Z (2012) Estimation of fish biomass using environmental DNA. PLoS ONE 7:e35868
Takahara T, Minamoto T, Doi H (2013) Using environmental DNA to estimate the distribution of an invasive fish species in ponds. PLoS ONE 7:e56584
Tréguier A, Paillisson JM, Dejean T, Valentini A, Schlaepfer MA, Roussel JM (2014) Environmental DNA surveillance for invertebrate species: advantages and technical limitations to detect invasive crayfish Procambarus clarkii in freshwater ponds. J Appl Ecol 51:871–879
Valentini A, Taberlet P, Miaud C, Civade R, Herder J, Thomsen PF, Bellemain E, Besnard A, Coissac E, Boyer F, Gaboriaud C (2016) Next-generation monitoring of aquatic biodiversity using environmental DNA metabarcoding. Mol Ecol 25:929–942
Wilcox TM, McKelvey KS, Young MK, Jane SF, Lowe WH, Whiteley AR, Schwartz MK (2013) Robust detection of rare species using environmental DNA: the importance of primer specificity. PLoS ONE 8:e59520
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden T (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13:134
Acknowledgements
This study was funded entirely by the UK Department of Environment, Food & Rural Affairs under a research contract (SF0265) awarded to GHC, and under operational funding to the Environment Agency (GDD). The travel of MF-P to the UK was funded by a travel grant from Conseil Regional de Bretagne. The authors would like to thank N. Edmonds for assistance in the field and to the owners of the angling venue in Kent for permission to access their property for this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Davison, P.I., Falcou-Préfol, M., Copp, G.H. et al. Is it absent or is it present? Detection of a non-native fish to inform management decisions using a new highly-sensitive eDNA protocol. Biol Invasions 21, 2549–2560 (2019). https://doi.org/10.1007/s10530-019-01993-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-019-01993-z