Abstract
Objectives
To display a recombinant avidin fused to the autotransporter ShdA to bind biotinylated molecules on the surface of Escherichia coli.
Results
Two chimeric protein constructs containing avidin fused to the autotransporter ShdA were expressed on the surface of Escherichia coli DH5α. One fusion protein contained 476 amino acids of the ShdA α and β domains, whereas the second consisted of a 314 amino acid from α and truncated β domains. Protein production was verified by SDS-PAGE using an antibody to the molecular FLAG-tag. The surface display of the avidin-shdA fusion protein was confirmed by confocal microscopy and flow cytometry analysis, and the biotin-binding activity was evaluated by fluorescence microscopy and flow cytometry using biotin-4-fluorescein and biotinylated-ovalbumin (OVA).
Conclusions
Expression of a recombinant avidin with biotin-binding activity on the surface of E. coli was achieved using the autotransporter ShdA. This system is an alternative to bind biotinylated molecules to E. coli.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
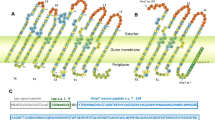
The display of heterologous proteins on the surface of engineered Enterobacteria strains has several biotechnological applications, including the evaluation of protein secretion pathways, and live vaccine development (Daugherty 2007). Expression of recombinant proteins on the surface of transformed bacteria can be achieved using their natural secretion systems; autotransporters, which comprise the type V secretion system in Gram-negative bacteria, are especially attractive due to their simplicity. In general, these proteins contain three functional domains: a N-terminal signal peptide that mediates the translocation of the protein to the bacterial periplasm through the Sec machinery; a passenger α domain, which is transported to the bacterial surface and may be trimmed and secreted into the milieu and, an outer membrane-anchored C-terminal β-domain that mediates the translocation of the α-domain to the bacterial surface (Nicolay et al. 2015). The autotransporters have been extensively used for the expression and secretion of recombinant proteins. This method is often referred to as autodisplay, indicating the surface expression of heterologous proteins by autotransporters and the substitution of the natural passenger α by a heterologous (passenger) protein through genetic engineering. Thus, the use of autotransporters to translocate heterologous proteins opens up the interesting possibility of coating bacterial surfaces with different peptides or proteins, which may be advantageous for the development of recombinant bacterial vector vaccines.
However, efficient autodisplay depends on several properties of the passenger protein, such as molecular weight, amino acid composition, tendency to form complex tertiary structures and hydrophobicity. Moreover, the recombinant protein may be toxic or constitute a metabolic burden to the transformed bacteria. Therefore, inadequate expression or poor biological function of the passenger protein is not an infrequent occurrence.
ShdA is a large outer membrane protein (OMP) of the autotransporter family from Salmonella enterica, involved in the colonization of the cecum and the Peyer’s patches of terminal ileum in mice (Kingsley et al. 2003). In addition, ShdA functions as adhesine, in virtue of its passenger domain ability to bind fibronectin and collagen I (Kingsley et al. 2004; Wagner and Hensel 2011). The translocation domain of ShdA is composed by a transmembrane β-barrel domain embedded in the outer membrane, and a linker region (connecting the ß-barrel to the passenger domain). Both have been used for in cis surface display of fusion proteins (Pompa-Mera et al. 2011, 2014). However, the use of ShdA for in trans surface display technology has not been explored.
The interaction of biotin and avidin or streptavidin has been used in many medical and biotechnological applications due to its strong affinity and specificity, allowing the bridging between biotinylated molecules to a substrate in cell surfaces (Liao et al. 2015).
In the present study, a recombinant avidin with biotin-binding activity was fused to the autotransporter ShdA This strategy allowed the in trans surface display of biotin-4-fluorescein and biotinylated ovalbumin (OVA-biotin) on the surface of Escherichia coli DH5α. This system represents a new tool for display of exogenous biotinylated molecules on cell surface of E. coli.
Materials and methods
Strains and plasmids
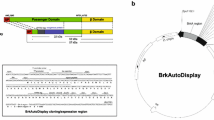
Escherichia coli DH5α (supE44 Δlac169 φ80lacZΔM15 hsdR17 recA1 endA1 gyrA96 thi-1 relA1) was used in this study. The vectors pENP (3415 bp) (Pompa-Mera et al. 2011), pAH3545 (3545 bp) (kindly donated by Dr. ML Penichet, UCLA, USA), pENP-Avi (3794 bp), pJEAY (3727 bp) and pJEAY-Avi (4094 bp) were used to express a recombinant avidin from Gallus gallus. All plasmids contain the nirB promoter (inducible under anaerobic conditions), with the exception of pAH3545, a 12,526 bp plasmid that encodes the LTB signal peptide for translocation of the fusion protein to the periplasmic space, as previously reported (Ruiz-Pérez et al. 2002). A FLAG-tag was included to trace the fusion proteins as reported previously (Pompa-Mera et al. 2011). Two variants of ShdA were constructed, one with the complete α-β-domains (wShdA) (476 amino acid residues) and one a previously reported Δα-β-domain variant of ShdA (Pompa-Mera et al. 2011, 2014) containing a shorter version of ShdA (sShdA) (314 amino acid residues). All recombinant bacterial strains were handled according to national biosafety regulations for genetically modified organisms that was published and updated in DOF 06-03-2009.
Plasmid construction
The expression vector pJEAY-Avidin, encoding the LTB-Avidin-FLAG-ShdA fusion protein, was constructed from the plasmid pENP as follows: a full-length sequence coding for the complete α-β domains of ShdA (wShdA) was amplified by PCR using Salmonella enterica serovar Typhimurium SL3261 genomic DNA as a template. Based on the GenBank sequence accession number AF140550.2, the primers sense shda-1 and antisense Tshda-2 were designed, containing the restriction sites NheI and BamHI (underlined), respectively. The resulting 1428 bp amplicon encodes 476 amino acids (positions 1560–2035). After restriction enzyme digestion, the amplicon was ligated to plasmid pENP. This strategy allowed the short ShdA (sShdA) sequence to be replaced and generated a complete ShdA fused to avidin (wShdA-Avi). The sequence coding for a Gallus gallus avidin was amplified by PCR from plasmid pAH3545 using primers avi-1 and avi-2 with HindIII and XhoI restriction sites incorporated at the 5′- and 3′-ends, respectively. The primer sequences used to construct plasmids are listed in Supplementary Table 1. The avidin encoding sequence (379 bp) was inserted in plasmid pJEAY that had been digested with the same enzymes (Supplementary Fig. 1). The resulting plasmid, pENP-Avi, contains the sShdA domain fused to avidin (sShdA-Avi). Competent E. coli DH5α cells were transformed with the pENP, pENP-Avi, pJEAY and, pJEAY-Avi plasmids. All restriction enzymes were purchased from New England Biolabs (Beverly, MA, USA). PCR reagents and plasmid preparation kits were purchased from Invitrogen (Palo Alto, CA, USA) and Qiagen (Qiagen, Mexico), respectively. DNA manipulation was performed according to the methods described by Sambrook and Russell (1989). The constructs engineered in this study were verified by nucleotide sequencing using the Big Dye Terminator Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA) and the pNirl primer (5′-TTCAGGTAAATTTGATACATCAAA-3′). Sequencing reactions were analysed on an ABI Prism 310 genetic analyser (applied biosystems).
Culture and induction conditions
Escherichia coli DH5α transformants were cultured on Luria–Bertani (LB) agar plates supplemented with 100 μg/ml of ampicillin at 37 °C. For inducing conditions, bacterial strains were incubated at 37 °C in 45 ml of thioglycolate medium (Difco Laboratories, Detroit, MI, USA) under an anaerobic atmosphere using a GasPak Anaerobic System (Becton–Dickinson Microbiology Systems, Cockeysville, MD, USA) until OD540nm = 1.0 was reached.
Analysis of expression of ShdA-Avidin fusion proteins by immunoblotting
Expression of the ShdA-Avidin fusion proteins was demonstrated by sodium dodecylsulfate-polyacrylamide gel electrophoresis (SDS–PAGE) of whole-cell lysates and outer membrane fraction from Escherichia coli DH5α strains transformed with plasmids pENP-Avi, pJEAY-Avi (encoding sShdA-Avi and wShdA-Avi, respectively), pJEAY and pENP (controls). Outer membrane proteins were prepared as previously described (Park et al. 2013). The samples were electrophoresed under reducing conditions (1% SDS, 140 mM 2-mercaptoethanol, 95 °C, 10 min) in 12% gels and then transferred onto nitrocellulose membrane 0.45 μm (BioRad®). Immunoblotting of protein extracts was performed with a mouse monoclonal anti-FLAG® M2 antibody (F3165, Sigma®) diluted to 1:1500 in PBS 5% skim milk (BioRad®). After washing, a secondary goat anti-mouse IgG antibody (H + L)-HRP conjugate (Zymed®) that was diluted to 1:3000 was applied. The immune reaction was revealed by 4-chloro-1-α-naphthol-30% H2O2 in PBS (pH 7.4). Expression analysis was performed with the BioRad® ChemiDocTM XRS System and Image LabTM Software v. 2.0 (Hercules, CA), using the protein ladder PageRuler™ Plus Prestained Protein Ladder (Thermo ScientificTM) to estimate the fusion proteins molecular size.
Confocal immunofluorescence microscopy and flow cytometry analysis
Surface expression of the sShdA-Avi and wShdA-Avi fusion proteins was evaluated by fluorescence confocal microscopy. After induction, the bacterial strains were harvested, washed thrice with PBS and overnight fixed in 4% (w/v) paraformaldehyde. After washing with PBS, cells were incubated with PBS supplemented with 3% (w/v) bovine serum albumin (BSA; Sigma-Aldrich). Then, cells were incubated with the monoclonal anti-FLAG® M2 antibody (F3165, Sigma®) at dilution of 1:50 in PBS, overnight at 25 °C with agitation. The cells were washed thrice with PBS and incubated with a goat anti-mouse IgG (H + L)-FITC conjugate (Zymed®), diluted to 1:100 in PBS for 2 h at 25 °C with agitation in the dark. The samples were suspended in 50 μl PBS, and 5 μl of the bacterial suspensions was placed on a poly-l-lysine coated microscopic slides and examined on a Nikon Ti Eclipse inverted confocal microscope. Imaging was performed using a 100× (oil immersion, NA 1.45) objective lens. The FITC-label was excited using a built-in 488 nm laser line. This light source was also used to acquire the transmitted light channel. FITC fluorescence was read in the 500–550 nm range. Images were acquired and analysed using NIS Elements v.5.00.
Surface display of the ShdA-Avidin fusion proteins on the cell surface of E. coli was also confirmed by flow cytometry. Cells were labelled with monoclonal anti-FLAG antibody, diluted to 1:100 and goat anti-mouse IgG (H + L)-FITC conjugate were used (Zymed®), diluted to 1:250, and analysed in a Beckman Coulter FacScalibur using FLOWJO v.7.5 software (Tree Star, Inc., Ashland, OR).
Evaluation of the biotin-binding activity of the ShdA-avidin fusion proteins
The ability of the two recombinant avidin fusion proteins (sShdA-Avi and wShdA-Avi) to bind biotin or biotinylated molecules was assessed, using biotin-4-fluorescein and ovalbumin (biotinylated-OVA).
After induction, the transformed bacterial strains were harvested, washed twice with PBS, and adjusted to 1 × 108 CFU. Cells were harvested, washed twice with PBS, adjusted to 1 × 108 CFU and incubated at 37 °C with 50 µl of biotin-4-fluorescein (Sigma-Aldrich) at 1 × 10−12, 1 × 10−10, 1 × 10−8, 1 × 10−6, 1 × 10−4, and 1 × 10−2 M concentrations. Bacterial suspensions were analysed by flow cytometry and indirect immunofluorescence assay (IFA).
The binding of the biotinylated ovalbumin (bio-OVA) to the ShdA-Avidin displayed on the bacterial surfaces was evaluated by IFA. Briefly, ovalbumin OVA (Sigma-Aldrich, St. Louis Missouri, USA) was biotinylated using an ImmunoProbe™ Biotinylation Kit (Sigma-Aldrich) according to the manufacturer’s instructions. After culturing cells under inducing conditions, 1 × 108 CFU of each bacterial strain was incubated with 50 µl of bio-OVA at concentrations of 1 × 10−4, 1 × 10−3, and 1 × 10−2 M. Bacterial suspensions were washed twice with PBS, and pellets were suspended in 500 µl PBS. Then, samples were incubated at 37 °C with agitation with the primary antibody, anti-Ovalbumin in dilution 1:1000. After three washes with PBS, samples were incubated with goat anti-mouse IgG (H + L)-FITC conjugate (Zymed®), diluted to 1:1500. Finally, the samples were suspended in 50 μl PBS, and 5 μl of the bacterial suspensions was placed on slides and observed using a fluorescence microscope (BX-40; Olympus).
Results
Expression of ShdA-avidin under control of the nirB promoter
Two ShdA-Avidin fusion proteins were constructed, including a short sShdA-Avidin construct (amino acids 1721–2035) containing (from the 5′-terminus) LTB-Avidin-FLAG-ΔShdA and a long wShdA-Avidin construct (amino acids 1560–2035) containing LTB-Avidin-FLAG-ShdA. The fusion genes were cloned into the pENP and pJEAY vectors under control of the inducible nirB promoter (Supplementary Fig. 1). A schematic representation of the two forms of recombinant avidin is shown in Fig. 1a. E. coli DH5α was transformed with the vectors pENP-Avi and pJEAY-Avi. The production of the fusion proteins was demonstrated in whole-cell lysates by SDS-PAGE (Fig. 1b), and immunoblotting using antibodies to the FLAG-tag (Fig. 1c).
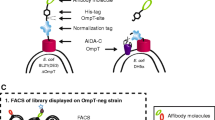
Expression of fusion proteins containing recombinant monomeric avidin in whole-cell lysates and outer membrane fraction of E. coli DH5α. a Hypothetical model for the display of recombinant monomeric avidin by ShdA autotransporter on the surface of E. coli DH5α. b SDS-PAGE analysis and c immunoblotting analysis of fusion proteins LTB-FLAG-ΔShdA (lanes 3 and 4), LTB-Avi-FLAG-ΔShdA (lanes 5 and 6), LTB-FLAG-ShdA (lanes 7 and 8) and LTB-Avi-FLAG-ShdA (lanes 9 and 10). A whole-cell lysate and outer membrane fraction derived from untransformed E. coli DH5α strain was included as a negative control (lanes 1 and 2, respectively). Lanes 1, 3, 5, 7, and 9, total proteins; lanes 2, 4, 6, 8 and 10, outer membrane fraction. All fusion proteins were tracked with an anti-FLAG monoclonal antibody
The expected bands corresponding to sShdA-Avi and wShdA-Avi fusion proteins with molecular weights of 52.2 and 70.1 kDa were observed as shown in Fig. 1c, lanes 5 and 9, respectively. In the E. coli strains transformed with the pENP and pJEAY plasmids, protein bands with molecular weights of 39 and 56.2 kDa were observed, respectively (Fig. 1c, lanes 3 and 7). A total protein extract from untransformed E. coli DH5α was included as a negative control in Fig. 1c, lane 1. Thus, the molecular weight of the recombinant avidin was consistent. Although no quantitative measurements of protein production were performed, it is noteworthy that sShdA-Avi (Fig. 1c, lane 5) showed lower expression as estimated by the 52.2 kDa band intensity when compared with the intensity of sShdA alone (39 kDa) (Fig. 1c, lane 3). In contrast, cloning of recombinant avidin in the pJEAY vector also resulted in low expression, as indicated by the intensity of the 73.5 kDa band (Fig. 1c, lane 9). These results show that E. coli successfully expressed the fusion construct of the FLAG-tagged avidin.
In addition, sShdA-Avi and wShdA-Avi fusion proteins were detected in outer membrane fractions of induced strains of E. coli carrying pENP-Avi and pJEAY-Avi plasmids (Fig. 1c, lanes 6 and 10, respectively). Similar expression of sShdA-Avi and wShdA-Avi fusion proteins, was detected in outer membrane fractions from E. coli transformed with pENP-Avi and pJEAY-Avi plasmids, respectively.
Display of ShdA-avidin fusion proteins on the E. coli cell surface
To confirm the presence and localization of avidin on the E. coli cell surface, recombinant strains were analyzed by confocal immunofluorescence microscopy. Fluorescence was observed in E. coli carrying pENP-Avi (Fig. 2c) and pJEAY-Avi plasmids (Fig. 2e). Expression of wShdA-Avi fusion protein on the cell surface of E. coli (strain pJEAY-Avi) was higher in comparison to showed by E. coli strain pENP-Avi. Histograms showed in Fig. 2f confirmed the display of recombinant avidin on the cell surface of E. coli transformed with the pENP-Avi and pJEAY-Avi plasmids. The expression of sShdA-Avi and wShdA-Avi fusion proteins, reached a 2-log-relative-light-unit displacement with respect to the untransformed E. coli DH5α control. Thus, recombinant avidin was efficiently translocated and displayed on cell surface of E. coli, through the two version of ShdA autotransporter (sShdA and wShdA).
Display of recombinant monomeric avidin on the cell surface of E. coli strains, revealed by confocal immunofluorescence microscopy. a E. coli DH5α observed (in light field), b pENP c pENP-Avi, d pJEAY and e pJEAY-Avi. Slides observed at 100× magnification. f Histograms showing the expression of fusion proteins on the surface of E. coli DH5α transformed with different plasmids. Cells were labelled with an anti-FLAG monoclonal antibody and FITC conjugate anti-IgG antibody. Fluorescent images showed in the lower correspond to the arrows indicated in b–e. Each scale bar represents 5 μm. A representative figure of three independent experiments is shown
Biotin-binding activity of ShdA-avidin displayed on the surface of E. coli
The functional activity of the recombinant ShdA-Avidin displayed on the bacterial surface was further evaluated by flow cytometry. Biotin-binding activity was determined using serial dilutions of biotin-4-fluorescein (Supplementary Fig. 2). High concentrations of biotin-4-fluorescein [1.25 × 10−4 M] were necessary to reach an optimal biotin-binding activity. Histogram and dot-plots shown in Fig. 3 revealed different biotin-binding activities between sShdA-Avi and wSdhA-Avi when the bacteria were incubated with 2.5 × 10−4 M biotin-4-fluorescein. E. coli pENP and pENP-Avi showed the highest mean fluorescence intensity (MFI) due to the retention of the biotin-4-fluorescein on their cell surfaces (MFI = 70.62 and MFI = 87.05, respectively) compared to the fluorescence detected with E. coli-pJEAY and E. coli-pJEAY-Avi (MFI = 42.17 and MFI = 40.25, respectively) (Fig. 3a).
Biotin-binding activity of recombinant monomeric avidin on the surface of E. coli DH5α transformed with different plasmids. a Histograms corresponding to biotin-4-fluorescein bound to recombinant avidin on the surface of E. coli strains. b Representative dot plots of side scatter and mean fluorescence intensity from flow-cytometric analyses of E. coli pENP, pENP-Avi, pJEAY and pJEAY-Avi after treatment with 2.5 × 10−4 M biotin-4-fluorescein; the percentage of fluorescent cells is presented. An untransformed E. coli DH5α strain was used as a control
These results demonstrate that ShdA-Avidin showed biotin-binding activity. Both variants of the fusion protein ShdA-Avidin showed biotin-binding activity, reaching a 2-log-relative-light-unit displacement with respect to the untransformed E. coli DH5α control.
Both sShdA-Avi and wShdA-Avi showed biotin-binding activity in a concentration-dependent manner. Non-specific biotin-4-fluorescein binding activity was observed in all strains at the highest concentration tested (5 × 10−4 M). Non-specific biotin-binding was also observed in the strains transformed with pENP, pENP-Avi and pJEAY-Avi, suggesting that ShdA increases the binding of biotin-4-fluorescein on the bacterial surface (Supplementary Fig. 2).
The Fig. 3b shows dot plots of cells exhibiting a positive fluorescence side scattering signal after incubation with 2.5 × 10−4 M biotin-4-fluorescein. The percentage of fluorescent cells for E. coli transformed with pENP-Avi or pJEAY-Avi was 74 and 42. 5%, respectively. However, E. coli DH5α strains transformed with pENP or pENP-Avi showed higher MFIs, suggesting that sShdA facilitates the unspecific binding of avidin. E. coli strains not expressing avidin (DH5α and DH5α-pJEAY) also showed background fluorescence (24.62 and 42.17, respectively). A significant fluorescent signal was detected from the pENP-Avi strain incubated with 2.5 × 10−4 M biotin-4-fluorescein. Non-specific fluorescence was observed with E. coli DH5α and DH5α-pJEAY. Thus, these results demonstrated that avidin displayed on the surface of E. coli cells was indeed able to bind biotinylated molecules (Fig. 3a), with the biotin-binding activity being more efficient in strains expressing short sShdA compared to the complete wShdA (Fig. 3b).
In addition, IFA was employed to determine whether the recombinant avidin on the surface of E. coli was able to bind biotin-4-fluorescein and biotinylated ovalbumin (biotinylated-OVA). A few bacteria transformed with pJEAY showed an ability to bind biotin-4-fluorescein (Fig. 4a) and biotinylated-OVA (Fig. 4b), which demonstrates that the E. coli strain had a basal ability to bind biotin. However, E. coli pJEAY-Avi efficiently bound biotinylated-OVA (Fig. 4b). It is noteworthy that bacterial aggregates were formed, suggesting mesh formation due to multimeric binding on the bacterial surface.
Determination of binding of biotinylated molecules to recombinant avidin displayed on the surface of E. coli DH5α by an indirect immunofluorescence assay (IFA). a IFA of E. coli strains incubated with 2.5 × 10−4 M biotin-4-fluorescein. b IFA of E. coli strains incubated with 1 × 10−4, 1 × 10−3, 1 × 10−2 M biotinylated-OVA and revealed with anti-OVA antibody and conjugate FITC-anti-IgG. Slides were observed at 100 × magnification. Each scale bar represents 5 μm
Discussion
Autotransporters are proteins that constitutes the type V secretion system of Gram-negative bacteria and have been widely used to autodisplay functional heterologous peptides or proteins on the surface of bacteria. This method is simple and has several advantages over the production of recombinant proteins from other bacterial compartments. However, autodisplay has several limitations; for instance, the folded, recombinant protein may not resemble the native protein, and this method is not suitable when glycosylation and posttranslational modifications are important for the function of the recombinant protein. An alternative to solve the problems associated with the inadequate production or improper folding of recombinant proteins displayed on the bacterial surface is the in trans surface display system, where the protein of interest is first purified from a separate source and then linked to the bacterial surface (Kalyanasundram et al. 2015).
Here, we report a similar in trans surface display system using the autodisplay of avidin on the bacterial surface. The in trans surface display of avidin was performed using the autotransporter ShdA to externally bridge the biotinylated molecules biotin-4-fluorescein and biotinylated-OVA. The binding of biotin-avidin/streptavidin is the strongest non-covalent interaction known in nature (Chivers et al. 2011), and this property has been exploited as a biological tool for a wide range of applications. The approach we describe here may represent a universal system to bind biotinylated molecules with diverse biochemical natures on the bacterial surface and is especially attractive, for rapid agglutination diagnostic tests or vaccine development. Furthermore, this system represents an efficient alternative for gene transfer approaches, in which cloning and expression of recombinant proteins have some limitations, in the expression of proteins with posttranslational modifications (Liao et al. 2015).
To achieve bacterial surface binding by the avidin–biotin interaction, the activity of the recombinant avidin must be preserved. Various systems for the heterologous expression of recombinant streptavidins in E. coli have been published (Demonte et al. 2014). Several studies designed to isolate the recombinant avidin have reported different technical difficulties, such as the intracellular accumulation of the heterologous protein, the formation of inclusion bodies and partial toxicity (Wu and Wong 2006). In addition, additional purification or renaturation steps are required when the secretion of bioactive avidin into the medium is desirable.
Although we cannot predict the proper folding of the avidin translocated to the bacterial surface as a fusion protein with ShdA and although it seems highly improbable, we found that it was indeed displayed on the surface of E. coli DH5α and was able to bind biotin. Two versions of ShdA-Avi were constructed. The short version of ShdA (ΔShdA), containing a truncated α domain, was more efficiently displayed than the complete ShdA. While a number of studies have shown that incomplete, truncated forms of the anchoring motif are better to display proteins surface than the complete α passenger domain (Han and Lee 2015); results from this study showed that complete ShdA autotransporter (wShdA) allowed a efficient expression of recombinant monomeric avidin as passenger domain. The secretion mechanism of autotransporters typically allows some passenger domain folding and the presence of disulphide bonds. The fusion protein ShdA-Avi was able to translocate avidin despite containing eight antiparallel β-strands and interconnecting loops. The surface display of a tetrameric avidin is not possible with autodisplay; however, a monomeric functional avidin was enough to bind biotinylated molecules. To increase recombinant protein production, we used the inducible promotor nirB, which offers some advantages over constitutive expression. This study shows that the nirB promoter is suitable for the expression of an avidin with bioactive function in E. coli DH5α. This approach offers some clear advantages, such as the availability of an avidin with biotin-binding activity having the potential to bridge biotinylated molecules without compromising the viability of host bacteria. Another advantage of this system is the fact that the recombinant avidin contains only 124 amino acids, maintaining its affinity for biotin. It is known that the monomeric avidin has a lower affinity than the tetrameric avidin; however, both ShdA-Avi fusion proteins were able to bind biotin at a concentration of 2.5 × 10−4 M. These recombinant avidins lack the carbohydrate side chains, avoiding the problems reported in several applications of avidin–biotin technology (Hytonen et al. 2004).
While the flow cytometry analysis showed that the avidin displayed on the E. coli surface was able to bind biotin, one limitation of this study is the inability to determine the amount of biotin-binding sites per bacterial cell.
The non-specific bond of biotin-4-fluorescein observed at a concentration of 5 × 10−4 M can be attributed to the saturation point being reached, which occurs with high biotin-4-fluorescein concentrations. However, non-specific binding to biotin-4-fluorescein at low concentrations [3.12 × 10−5 M] was also observed, particularly with E. coli pENP. The presence of the conserved lysine residue within a consensus sequence (Ala/Val-Met-Lys-Met) found in carboxylases might contribute to the biotin binding (Healy et al. 2010). In contrast, avidin has a high level of nonspecific binding to various biological components at physiological pH as a result of its high isoelectric point (Takakura et al. 2012).
The production of soluble, recombinant streptavidin in Escherichia coli may be difficult. Therefore, some studies have focused on their expression on the bacterial surface. The production of a functional streptavidin on the bacterial surface of E. coli UT5600 has been shown using the autotransporter AIDA-I (Park et al. 2011). However, to our knowledge, this is the first report of the expression and display of avidin by an autotransporter in E. coli that maintains its biological properties.
Conclusions
Here, we report on a recombinant avidin fused to the membrane anchoring motif of ShdA in E. coli DH5α. This approach constitutes an effective in trans surface display system for exogenous biotinylated molecules. This method could provide an alternative, wide-ranging method to bind proteins and other biomolecules to bacterial surfaces.
References
Chivers CE, Koner AL, Lowe ED, Howarth M (2011) How the biotin-streptavidin interaction was made even stronger: investigation via crystallography and a chimaeric tetramer. Biochem J 435:55–63
Daugherty PS (2007) Protein engineering with bacterial display. Curr Opin Struct Biol 17:474–480
Demonte D, Dundas CM, Park S (2014) Expression and purification of soluble monomeric streptavidin in Escherichia coli. Appl Microbiol Biotechnol 98:6285–6295
Han MJ, Lee SH (2015) An efficient bacterial surface display system based on a novel outer membrane anchoring element from the Escherichia coli protein yiaT. FEMS Microbiol Lett 362:1–7
Healy S, McDonald MK, Wu X, Yue WW, Kochan G, Oppermann U, Gravel RA (2010) Structural impact of human and Escherichia coli biotin carboxyl carrier proteins on biotin attachment. Biochemistry 49:4687–4694
Hytonen VP, Laitinen OH, Airenne TT, Kidron H, Meltola NJ, Porkka EJ, Horha J, Paldanius T, Maatta JA, Nordlund HR, Johnson MS, Salminen TA, Airenne KJ, Yla-Herttuala S, Kulomaa MS (2004) Efficient production of active chicken avidin using a bacterial signal peptide in Escherichia coli. Biochem J 384:385–390
Kalyanasundram J, Chia SL, Song AA, Raha AR, Young HA, Yusoff K (2015) Surface display of glycosylated Tyrosinase related protein-2 (TRP-2) tumour antigen on Lactococcus lactis. BMC Biotechnol 15:113
Kingsley RA, Humphries AD, Weening EH, De Zoete MR, Winter S, Papaconstantinopoulou A, Dougan G, Bäumler AJ (2003) Molecular and phenotypic analysis of the CS54 island of Salmonella enterica serotype typhimurium: identification of intestinal colonization and persistence determinants. Infect Immun 71:629–640
Kingsley RA, Abi Ghanem D, Puebla-Osorio N, Keestra AM, Berghman L, Bäumler AJ (2004) Fibronectin binding to the Salmonella enterica serotype Typhimurium ShdA autotransporter protein is inhibited by a monoclonal antibody recognizing the A3 repeat. J Bacteriol 186:4931–4939
Liao TY, Lau A, Joseph S, Hytönen V, Hmama Z (2015) Improving the immunogenicity of the Mycobacterium bovis BCG vaccine by non-genetic bacterial surface decoration using the avidin-biotin system. PLoS ONE 10:e0145833
Nicolay T, Vanderleyden J, Spaepen S (2015) Autotransporter-based cell surface display in gram-negative bacteria. Crit Rev Microbiol 41:109–123
Park M, Jose J, Thommes S, Kim JI, Kang MJ, Pyun JC (2011) Autodisplay of streptavidin. Enzyme Microb Technol 48:307–311
Park TJ, Heo NS, Yim SS, Park JH, Jeong KJ, Lee SY (2013) Surface display of recombinant proteins on Escherichia coli by BclA exosporium of Bacillus anthracis. Microb Cell Fact 12:81
Pompa-Mera EN, Yepez-Mulia L, Ocana-Mondragon A, Garcia-Zepeda EA, Ortega-Pierres G, Gonzalez-Bonilla CR (2011) Trichinella spiralis: intranasal immunization with attenuated Salmonella enterica carrying a gp43 antigen-derived 30mer epitope elicits protection in BALB/c mice. Exp Parasitol 129:393–401
Pompa-Mera EN, Arroyo-Matus P, Ocana-Mondragon A, Gonzalez-Bonilla CR, Yepez-Mulia L (2014) Protective immunity against enteral stages of Trichinella spiralis elicited in mice by live attenuated salmonella vaccine that secretes a 30-mer parasite epitope fused to the molecular adjuvant C3d-P28. Res Vet Sci 97:533–545
Ruiz-Pérez F, León-Kempis R, Santiago-Machuca A, Ortega-Pierres G, Barry E, Levine M, González-Bonilla C (2002) Expression of the Plasmodium falciparum immunodominant epitope (NANP) 4 on the surface of Salmonella enterica using the autotransporter MisL. Infect Immun 70:3611–3620
Sambrook J, Russell DW (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Takakura Y, Oka N, Kajiwara H, Tsunashima M (2012) Engineering of novel tamavidin 2 muteins with lowered isoelectric points and lowered non-specific binding properties. J Biosci Bioeng 114:485–489
Wagner C, Hensel M (2011) Adhesive mechanisms of Salmonella enterica. Adv Exp Med Biol 715:17–34
Wu SC, Wong SL (2006) Intracellular production of a soluble and functional monomeric streptavidin in Escherichia coli and its application for affinity purification of biotinylated proteins. Protein Expr Purif 46:268–273
Acknowledgments
This work was approved by the Local Research and Bioethics Committee of the Instituto Mexicano del Seguro Social (IMSS) and was supported by grants from the IMSS–FIS (FIS/IMSS/PROT/566) and (UC-MEXUS-CONACYT, number CN-07-122). Héctor Daniel Pardavé-Alejandre is a PhD student from the Programa de Doctorado en Ciencias Biomédicas, Facultad de Medicina, Universidad Nacional Autónoma de México (UNAM), and received a PhD Fellowship from CONACYT (#228005) and COMECYT (12BCD0045-I). The authors acknowledge Dr. M. Penichet from the Division of Surgical Oncology, UCLA, USA for the donation of the pAH3545 plasmid and Javier Hernández Acosta for his technical assistance. We also thank Vadim Pérez Koldenkova Ph.D. from Laboratorio Nacional de Microscopía Avanzada, Centro Médico Nacional S. XXI, IMSS, by its technical assistance in confocal microscopy, and the Laboratorios San Ángel S.A. for their financial support in the publication of this manuscript.
Supporting information
Supplementary Table 1—List of primers used in the construction of the pENP-Avi, pJEAY and pJEAY-Avi plasmids.
Supplementary Figure 1—Schematic representation of the construction of the pENP-Avi and pJEAY-Avi plasmids. A plasmid containing a full-length sequence coding for the complete α-β domains of the ShdA AT (complete ShdA) was constructed using the pENP plasmid to replace the short ShdA AT (ΔShdA). A DNA fragment encoding for 124 amino acids, corresponding to monomeric avidin from Gallus gallus, was amplified by PCR using the pAH3545 plasmid as template. The resulting amplicon (379 bp) was digested with HindIII and XhoI enzymes and inserted into the pJEAY vector, creating the pJEAY-Avi plasmid.
Supplementary Figure 2—Determination of the optimal concentration of biotin-4-fluorescein bound to recombinant monomeric avidin displayed by E. coli DH5α. Strains of E. coli DH5α were cultured under inducing conditions and incubated with different concentrations of biotin-4-fluorescein.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors have no conflicts of interest to declare.
Ethical approval
The study was approved by the Ethics Committee and the Research Committee of the National Committee of Scientific Research of the Instituto Mexicano del Seguro Social with the registration number R-2012-785-064.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pardavé-Alejandre, H.D., Alvarado-Yaah, J.E., Pompa-Mera, E.N. et al. Autodisplay of an avidin with biotin-binding activity on the surface of Escherichia coli. Biotechnol Lett 40, 591–600 (2018). https://doi.org/10.1007/s10529-018-2507-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-018-2507-6