Abstract
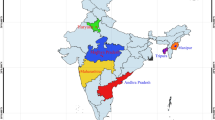
Twenty-seven enzyme systems, six random amplified polymorphic DNA (RAPD) primers, and two microsatellite loci were tested to determine intraspecific divergence in the natural population of the endangered Indian featherback fish, Chitala chitala, for the first time. The 262 samples of C. chitala were collected from six riverine locations in India: the Satluj, Ganga (Ghagra, Bhagirathi, and Brahmaputra), Mahanadi, and Narmada river systems. The analysis revealed population subdivisions, with an FST value from 0.1235 (95% confidence 0.0868–0.1621) for RAPD and a combined FST of 0.0344 (95% confidence 0.0340–0.0350) for microsatellite loci. An analysis of 38 allozyme loci did not reveal any polymorphism in the samples from any of the riverine localities; a possible explanation for this could be that the ancestors of Chitala could have faced a population reduction in prehistoric periods, as low allozyme variation is also reported for other species of Chitala from south Asia.



Similar content being viewed by others
References
Alarcon JA, Magoulas A, Georgakopoulos T, Zouros E, Alvarez MC (2004) Genetic comparison of wild and cultivated European populations of the gilthead sea bream (Sparus aurata). Aquaculture 230(1–4):65–80
Ali BA, Huang TH, Qin DN, Wang XM (2004) A review of random amplified polymorphic DNA (RAPD) markers in fish research. Rev Fish Biol Fish 14:443–453
Belkhir K, Borsa P, Goudet J, Chikhi L, Bonhomme F (1997) Genetix ver. 4.05, Genetics logiciel sous Windows pour la genetique des populations. http://www.genetix.univ-montp2.fr/genetix/genetix.htm. Accessed November 2005
BDA (2002) Biological Diversity Act, 2002, with statement of objects and reasons & notes on clauses. Professional Book Publishers, New Delhi, p 27. Download available at site of National Biodiversity Authority of India. http://www.nbaindia.org/act/act_english.htm. Accessed January 2009
Black WCIV (1993) PCR with arbitrary primers: approach with care. Insect Mol Biol 2:1–6
Bovo D, Rugge M, Shiao YH (1999) Origin of spurious multiple bands in the amplification of microsatellite sequences. Mol Pathol 52:50–51
Brown LD, Gunter PT, Waters MJ, Epifanio MJ (2000) Evaluating genetic diversity associated with propagation assisted restoration of American shad. Conserv Biol 14:294–303
CAMP (1997) Conservation assessment and management plan (CAMP) for freshwater fishes of India. Workshop report. Zoo Outreach Organization, Coimbatore/CBSG and NBFGR, Lucknow, India, pp 1–158
Chauhan T, Lal KK, Mohindra V, Singh RK, Punia P, Gopalakrishnan A, Sharma PC, Lakra WS (2007) Genetic differentiation in the population of Indian major carp, Cirrhinus mrigala (Hamilton-Buchanan, 1882): evidence from allozyme and microsatellite markers. Aquaculture 269:135–149
Chenuil A, Glazier N, Berrebi P (1999) A test of the hypothesis of an autopolyploid vs allopolyploid origin of a tetraploid lineage: Application of genus Barbus (Cyprinidae). Heredity 82:373–380
Chistiakov DA, Hellemans B, Volckaert FAM (2006) Microsatellites and their genomic distribution, evolution, function and applications: a review with special reference to fish genetics. Aquaculture 255(1–4):1–29
Croojimans RPMA, Bierbooms VAF, Komer J, Poel JJV, Groenen MAM (1997) Microsatellite markers in common carp (Cyprinus carpio). Anim Genet 28:129–134
Dever JA, Densmore LD (2001) Microsatellites in Morelet’s crocodile (Crocodylus moreletii) and their utility in addressing crocodilian population genetics questions. J Herpetol 35:541–544
DeWoody JA, Avise JC (2000) Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J Fish Biol 56:461–473
ECAFE (1966) A compendium of major international rivers in the ECAFE region. Water Resources series, United Nation publication, No. 29
Elliott NG, Reilly A (2003) Likelihood of bottleneck event in the history of the Australian population of Atlantic salmon (Salmo salar L.). Aquaculture 215(1–4):31–44
Farias IP, Hrbek T, Brinkmann H, Sampaio I, Meyer A (2003) Characterization and isolation of DNA Microsatellite primers for Arapaima gigas, an economically important but severely over-exploited fish species of the Amazon basin. Mol Ecol Notes 3:128–130
Frentiu FD, Ovenden JR, Street R (2001) Australian lungfish (Neoceratodus forsteri: Dipnoi) have low genetic variation at allozyme and mitochondrial DNA loci: a conservation alert? Conserv Genet 2:63–67
Froese R, Pauly D (2003) Fish base. World Wide Web Electronic Publication. World Fish Centre (ICLARM). http://www.FishBase.Org. Accessed January 2009
Glenn TC, Dessauer HD, Braun MJ (1998) Characterization of microsatellite DNA loci in American alligators. Copeia 3:591–601
Goswami PK (2000) Endangered featherback Chitala chitala (Hamilton-Buchanan). In: Ponniah AG, Sarkar UK (eds) Fish biodiversity of north-east India, NBFGR-NATP Publication 2, pp 124–125
Goudet J, Raymond M, DeMeeus T, Rousset F (1996) Testing differentiation in diploid populations. Genetics 139:463–471
Hamilton-Buchanan F (1822) An account of the fishes found in the river Ganges and its branches. Edinburgh and London. Fishes Ganges i–vii:1–405
Hardwick R (2005) The clown knifefish, Chitala ornata. http://www.practicalfishkeeping.co.uk/pfk/pages/show_article.php?article_id=43. Accessed January 2009
Hartl DL, Clark AG (1997) Principles of population genetics, 3rd edn. Sinauer Associates, Inc., Sunderlands, MA, p 542
Hora SL (1953) The Satpura hypothesis. Sci Prog, London 41:245–255
Jones AG, Blouin MS, Arnold SJ (2001) Genetic variation in two population of the rough-skinned newt (Taricha granulosa) assessed using novel tetranucleotide microsatellite loci. Mol Ecol Notes 1:293–296
Kemp A (1991) Australian Mesozoic and Cainozoic lungfish. In: Vickers-Rich P, Monaghan JM, Baird RF, Rich TH (eds) Vertebrate palaeontology of Australasia. Pioneer Design Studio Pvt. Ltd, Victoria, pp 465–497
Kendall MG, Stuart A (1977) The advanced theory of statics, vol 1, 4th edn. Charles Griffin, London
Lal KK, Mandal A, Singh RK, Punia P, Kapoor D, Chauhan UK, Singh SP, Mohindra V (2006) Genetic divergence in two feather back fishes, Chitala chitala and Notopterus notopterus. J Appl Ichthyol 22(5):369–373
Lessios HA (1992) Testing electrophoretic data for agreement with Hardy–Weinberg expectations. Mar Biol 112:517–523
Lilabati H, Vishwanath W (1998) Biochemical, nutritional and microbiological quality of ice stored Notopterus chitala of Imphal market, Manipur. Indian J Fish 45(4):441–446
Lynch M, Milligan BG (1994) Analysis of population genetic structure with RAPD markers. Mol Ecol 3:91–99
Miller LM, Senanan WA (2003) Review of Northern pike population genetics research and its implications for management. North Am J Fish Mgmt 23(1):297–306
Miller MP (1997) Tools for population genetic analysis (TFPGA, V. 1.3): a windows program for the analysis of allozyme and molecular population genetic data. Department of Biological Sciences, Northern Arizona University, Flagstaff, AZ
Mohindra V, Narain L, Peyush P, Gopalakrishanan A, Mandal A, Kapoor D, Ponniah AG, Lal KK (2005) Microsatellite DNA markers for population-genetics study of Labeo dyocheilus (McClelland, 1839). J Appl Ichthyol 21:1–5
Naish KA, Skibinski DOF (1998) Tetranucleotide microsatellite loci for Indian major carps. J Fish Biol 53:886–889
Nayyar RP (1966) Karyotype studies in thirteen species of fishes. Genetica 37:78
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Punia P, Gupta HS, Singh RK, Mohindra V, Lal KK, Chauhan RVS, Lakra WS (2006) Polymorphic microsatellite markers isolated from partially enriched genomic library of Chitala chitala. Mol Ecol Notes 6(4):1263
Raymond M, Rousset F (1995a) Genepop version 3.1: population genetics software for exact test and ecumenicism. J Hered 86:248–249. http://www.cefe.cnrs-mop.fr/genepop.html. Accessed November 2005
Raymond M, Rousset F (1995b) An exact test for population differentiation. Evolution 49:1280–1283
Robertson BC, Gemmell NJ (2005) Microsatellite DNA markers for the study of population structure in the New Zealand fur seal Arctocephalus forsteri. Department of Conservation, Wellington. DOC Science Internal Series 196, pp 18
Rousset F, Raymond M (1995) Testing heterozygote excess and deficiency. Genetics 140:1413–1419
Ruzzante DE, Taggart C, Cook D, Goddard S (1996) Genetic differentiation between inshore and offshore Atlantic cod (Gadus morhua) of Newfoundland: microsatellite DNA variation and antifreeze level. Can J Fish Aquat Sci 53:634–645
Sarkar UK, Deepak PK, Negi RS, Singh SP, Kapoor D (2006) Captive breeding of endangered fish Chitala chitala (Hamilton-Buchanan) for species conservation and sustainable utilization. Biodivers Conserv 15(11):3579–3589
Sarkar UK, Negi RS, Deepak PK, Lakra WS, Paul SK (2008) Biological parameters of the endangered fish Chitala chitala (Osteoglossiformes: Notopteridae) from some Indian Rivers. Fish Res 90:170–177
Scribner KT, Pearce JM (2000) Microsatellites: evolutionary and methodological background and empirical applications at individual, population, and phylogenetic levels. In: Baker A (ed) Molecular methods in ecology. Blackwell Science Limited, England, pp 235–271
Shaklee JB, Phelps SR, Salini J (1990) Analysis of fish stock structure and mixed- stock fisheries by the electrophoretic characterization of allelic allozymes. In: Whitmore DH (ed) Electrophoretic and isoelectric focussing techniques in fisheries management. CRC Press, Boca Raton, FL, pp 173–196
Sodsuk PK, Sodsuk S (2000) Genetic diversity of featherback fish in Thailand. Kasetsart J (Nat Sci) 34:227–239
Sokal RR, Rohlf S (1995) Biometry. W. H. Freeman, New York
Venkatesh B, Ning Y, Brenner S (1999) Late changes in spliceosomal introns define clades in vertebrate evolution. Proc Natl Acad Sci U S A 96(18):10267–10271
Wang D, Shi J, Carlson SR, Cregan PB, Ward RW, Diers BW (2003) A low-cost, high-throughput polyacrylamide gel electrophoresis system for genotyping with microsatellite DNA markers. Crop Sci 43:1828–1832
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Whitmore DH (1990) Electrophoretic and isoelectric focussing techniques in fisheries management. CRC Press, Boca Raton, FL, pp 1–350
Williams JGK, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Wilson AJ, Hutchings JA, Ferguson MM (2004) Dispersal in a stream dwelling salmonid: inferences from tagging and microsatellite studies. Conserv Genet 5:25–37
Wright S (1951) The genetical structure of populations. Ann Eugen 15:323–354
Zardoya R, Vollmer DM, Craddock C, Streelman JT, Karl S, Meyer A (1996) Evolutionary conservation of microsatellite flanking regions and their use in resolving the phylogeny of cichlid (Pisces: Perciformes). Proc R Soc Lond B 263:1589–1598
Zhang YP, Wang XX, Ryder OA, Li MP, Zhang HM, Yong Y, Wang PY (2002) Genetic diversity and conservation of endangered animal species. Pure Appl Chem 74:575–584
Acknowledgments
Financial support received from the NATP-ICAR (subproject MM-18) is thankfully acknowledged. Excellent technical assistance provided by Mr. R. S. Sah, Mr. Akhilesh Mishra, Mr. Rajesh Kumar, and Mr. Sree Ram is also duly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mandal, A., Lal, K.K., Mohindra, V. et al. Evaluation of Genetic Variation in the Clown Knifefish, Chitala chitala, Using Allozymes, RAPD, and Microsatellites. Biochem Genet 47, 216–234 (2009). https://doi.org/10.1007/s10528-008-9219-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-008-9219-x