Abstract
The fisheries and aquaculture sectors are growing rapidly, reflecting their importance in meeting the ever-increasing human population’s demands for animal protein. Production progress in this sector, however, is challenging as a result of increased deaths from epidemics caused by bacterial infectious diseases in aquaculturally important species. In order to minimize the impact of such diseases, quick and reliable diagnosis of pathogens, timely intervention, and control of the disease are essential to ensure the health of aquaculture and fisheries stocks. Thus, high-throughput proteomics-based matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) has been used as a rapid, accurate, and species-specific tool for the identification of bacterial fish pathogens and antibiotic resistance. The aim of this article is to review and evaluate the results of nearly three hundred papers published from 2010 to 2023 on the application of MALDI-TOF MS in aquaculture, showing that this method has been increasingly used over the years for the species-level identification and antibiotic resistance of fifty different gram-positive and negative bacterial aquatic pathogens encountered in wild or cultured fish, shellfish, invertebrates, and crustaceans. In addition, the history and general principles of the MALDI-TOF MS method are also mentioned in the article so that fish disease researchers interested in the use of this technology can see all aspects of the topic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aquaculture, a rapidly growing global industry, suffers tremendous financial losses each year as a result of bacterial, viral, fungal, and parasitic disease outbreaks (Woo and Bruno 2011; Fazio 2019). There are a number of bacterial pathogens that have been detected in wild or farmed fish, shellfish, and other aquatic organisms (Alderman 1996; Austin and Austin 2016). It is predicted that bacterial pathogens are responsible for about 35% of the diseases which damage the tissues of juvenile and adult fish, making them more vulnerable to infection (Dar 2022). Infection is particularly prevalent in fish under stress. Stressors include high stock densities, poor nutrition, the accumulation of hazardous toxic substances, contaminated water, sudden temperature changes, and low oxygenation (Stickney 2009; Natnan et al. 2021). Since bacterial infections are a major contributor to fish mortality in aquaculture, mitigating their impact is of great importance to the aquacultural and fisheries industry.
Bacterial pathogens commonly seen in inland and marine fish, invertebrates, and crustaceans are as follows: Aeromonas spp. (A. hydrophila, A. salmonicida, A. veronii, A. caviae, A. sobria), Edwardsiella spp. (E. ictaluri, E. tarda, E. piscicida), Flavobacterium psychrophilum, F. columnare, Lactococcus garvieae, Mycobacterium spp. (M. fortuitum, M. marinum), Pseudomonas spp. (P. anguilliseptica, P. fluorescens, P. plecoglossicida), Photobacterium damselae, Renibacterium salmoninarum, Streptococcus spp. (S. iniae, S. agalactiae), Staphylococcus warneri, Vagacoccus salmoninarum, Vibrio spp. (V. anguillarum, V. splendidus, V. harveyi, V. parahaemolyticus, V. alginolyticus, V. vulnificus, V. coralliilyticus), Yersinia ruckeri, and Tenacibaculum maritinum (Toranzo 2004; Noga 2010; Austin and Austin 2016; Vanamala 2022). It is crucial to monitor aquatic animal diseases through surveillance and rapid bacterial detection so that the appropriate interventions can be undertaken so that the diseases are diagnosed and managed before they present a serious threat (Adams and Thompson 2011; Crumlish 2017; Ashfaq et al. 2022) to both animal welfare and national economies that rely on marine and inland aquaculture. Accurate and rapid diagnostic techniques are, therefore, crucial.
Traditionally, bacterial fish diseases and aquatic pathogens have been identified using conventional microbiological, immunological, and molecular biological approaches. Microbiological methods are comparatively slow and require bacterial isolation and culturing and subsequent morphological observation and biochemical characterization of potential pathogens (Buller 2014; Smith 2019; Austin 2019; Duman et al. 2022). Bacterial isolates can be differentiated and classified either manually or using a number of kits and systems for biotyping based on the biochemical properties of bacteria, including the API (Biomerieux, USA), Vitek (Biomerieux, France), Biolog (Biolog, USA), and BD BBL (Becton Dickinson Microbiology Systems, USA) (Santos et al. 1993; Taylor et al. 1995; Popović et al. 2007). Serological and histopathological assays are also utilized in the diagnosis of bacterial fish diseases (Popovic et al. 2007; Timur et al. 2009; Austin 2019). Enzyme-linked immunosorbent assays (ELISA) have also been prominent in diagnosing bacterial fish diseases. ELISA detects specific antibodies or antigens associated with pathogens in fish samples, allowing for early disease identification and continuous monitoring of infection levels within fish populations (Tanrikul 2007; González and Santos 2009; Duman et al. 2022). In the late 1990s and early 2000s, molecular biological methods began to be used to identify bacterial fish diseases in aquaculture (Zlotkin et al. 1998a; Hassan et al. 2001; Altinok and Kurt 2003; González et al. 2004; Ruiz-Zarzuela et al. 2005; Altinok et al. 2008; Jung et al. 2010; Kayış et al. 2015). The most common and frequently used basic molecular techniques are standard polymerase chain reaction (PCR), PCR-based genotyping, and Sanger sequencing (SS). The basis of these techniques is the sensitive amplification of bacterial DNA, the identification of target genetic markers and their sequences as a tool for the identification of aquatic bacterial pathogens (del Cerro et al. 2002; Cunningham 2002; Rhodes et al. 2004; Beaz-Hidalgo et al. 2008; Onuk et al. 2010; Buller 2014; Kim et al. 2017; Çağatay 2022). The use of next-generation sequencing (NGS) technique based on targeted amplicon or whole-genome sequencing (Hambuch and Mayfield 2014; Lefterova et al. 2015) has started a new era in aquaculture disease diagnosis since this basic PCR-based simple sequencing technique is not sensitive enough to distinguish some closely related aquatic bacterial species (Kumar and Kocour 2017; De Bruijn et al. 2018; Natnan et al. 2021; Jaies et al. 2024; Bohara et al. 2024). Although it is very beneficial to perform species-specific sequencing using an NSG device, the high cost and the time requirement are disadvantages for the users. In this regard, one of the alternative high throughput methods, the fast, highly sensitive, specific, and less demanding matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) method has become the new focus.
MALDI-TOF MS involves analyzing the protein profiles of bacterial strains, enabling rapid and definitive identification (Seng et al. 2009; Stevenson et al. 2017). This new and powerful proteomic approach can be used as a complement and validator of other techniques for microbial identification and allowing early recognition of bacterial populations in water and effective animal health management. Beyond microbial identification, it is also highly useful for characterizing pathogen life cycles and virulence components and investigating antibiotic resistance (Lauková et al. 2018; Moreira et al. 2021; Duman et al. 2022).
In this review article, unlike the articles in the literature on the application of the MALDI-TOF MS method in the diagnosis of bacterial diseases in aquaculture and fisheries, the findings of approximately three hundred studies from 2010 to 2023 are examined in detail. In addition, the general application areas and history of the method are briefly mentioned for the readers to better understand the topic. The results of incremental studies on the species identification of fifty different gram-negative and gram-positive aquatic bacterial pathogens using MALDI-TOF MS methods are presented and evaluated. This review also aims to assess the advantages, disadvantages, and potential future directions of the method in aquaculture.
History and applications of MALDI-TOF MS
Matrix-assisted laser desorption/ionization (MALDI) mass spectrometry (MS) was initially developed by Karas et al. (1985; 1987). Around the same period, Tanaka et al. (1988) reported a “soft desorption ionization” technique that allowed for the mass spectrometric study of biological macromolecules by integrating mass analyzer time of flight (TOF) with MALDI. MALDI was named after Karas and Hillenkamp’s (1988) research on soft desorption ionization utilizing an organic compound matrix. Koichi Tanaka won the Chemistry Nobel Prize in 2002, and since then, MALDI-TOF MS has advanced significantly in proteomics and metabolomics research, allowing analysis of many different metabolites in biomedical research (Tanaka 2003). The American Food and Drug Administration (FDA) has approved several MALDI systems for commercial use in clinical microbiology laboratories globally, including the Clin-TOF (Bioyong Technology, China), the MassARRAY System (Agena Bioscience, USA), the VitekMS (bioMérieux Clinical Diagnostics, France), and the MALDI Biotyper (Bruker, Germany). These systems offer special combinations of databases, software, and mass spectrometers (Li et al. 2022). In addition, inclusive mass spectrum libraries are available for MALDI-TOF MS analysis (Table 1), which offer public access to mass spectra of numerous microorganisms (Böhme et al. 2012; Wieser et al. 2012; Torres-Sangiao et al. 2021; Zuffa et al. 2023), and users can select these libraries according to their available current system. Studies have shown that one or more libraries can be used depending on the type of microorganism being investigated.
Among the many benefits of MALDI-TOF MS, which include ease of use, high throughput, and resistance to various contaminants, the equipment is extensively utilized in regular analysis and research in clinical microbiology laboratories throughout the world, making it a versatile investment. Uses of MALDI-TOF (Fig. 1) include microbial pathogen identification (Seng et al. 2010; Croxatto et al. 2012; De Marco and Burnham 2014; Cañas et al. 2015; Erler et al. 2015; Welker et al. 2019; Anwer et al. 2022), toxic substances and allergen detection, the diagnosis of human bloodstream infection, cancer diagnosis (Royo-Cebrecos et al. 2017), determination of antibiotic resistance (Wieser et al. 2012; Singhal et al. 2015; Patel 2015; Anwer et al. 2022; Özcan 2023), genetic disorders screening (Huang et al. 2016; Li et al. 2022), pharmaceutical analysis, and drug discovery (Koh et al. 2003; Kafka et al. 2011).
Although primarily developed for clinical purposes, MALDI-TOF MS technology has been applied to many different fields to identify a broad range of microorganisms (including viruses, bacteria, fungus, and yeast), parasites, and protozoa. It has also proven to be an effective tool for rapidly and precisely identifying aerobic, anaerobic, and fastidious (low growth rate or non-cultivatable) microorganisms (Seng et al. 2009; Biswas and Rolain 2013; Singhal et al. 2015; Patel 2015; Alcala et al. 2021; Popović et al. 2022). In fact, the Google Scholar search engine retrieved 18,500 research papers published between 2003 and 2023, based on the question “Other than clinical use, what are the applications of MALDI-TOF mass spectrometry?” Applications of MALDI-TOF MS included the following: environmental microbiology for the identification of soil and waterborne pathogens and aquatic microorganisms (Singhal et al. 2015; Popović et al. 2017; Ashfaq et al. 2022; Freitas et al. 2022), veterinary microbiology to identify zoonotic pathogens (Thompson 2022; Popović et al. 2021; Arafa et al. 2023), food microbiology for the identification of foodborne pathogens (Böhme et al. 2013; Akimowicz et al. 2020; Brauge et al. 2021; Nissa et al. 2021; Haider et al. 2023), agricultural microbiology (Gandhi et al. 2013; Drissner et al. 2017; Tamaru 2023; Surányi et al. 2023), identification of plant pathogens (Wang et al. 2012; Ahmad et al. 2012; Chalupová et al. 2014; Chun et al. 2022; Sivanesan et al. 2023), and most recently in aquatic microbiology for identification of fish pathogens (Singhal et al. 2015; Assis et al. 2017; López-Cortés et al. 2017; Jansson et al. 2020; Piamsomboon et al. 2020; Moreira et al. 2021; Duman et al. 2022; Saticioglu et al. 2023).
General principles and methodology of MALDI-TOF MS
MALDI-TOF MS works by ionizing and measuring the mass-to-charge ratios (m/z) of ribosomal proteins in a microbial sample resulting in a mass spectrum that is a unique fingerprint (Singhal et al. 2015; Piamsomboon et al. 2020; Popović et al. 2022) which can be used to identify the microorganism by matching to existing bio-markers (López-Cortés et al. 2017; Duman et al. 2022). The microbial peptide mass fingerprints that are obtained are compared to the mass spectral library database of pre-existing reference samples (Brauge et al. 2021).
The MALDI-TOF MS process for microbial identification from different sample types can be summarized in three steps (Fig. 2). The first step is sample preparation from either a direct sample or culture, the second step is the analysis of the samples in the MALDI-TOF instrument, and the third step is the identification process (Patel 2015; Singhal et al. 2015; Sandalakis et al. 2017; Kazazić et al. 2019a; Kazazić et al. 2019b; Mishra et al. 2020; Dare 2006). In step one, a single colony from a cultured sample is spread onto the target plate, which can be either “direct sample spotting,” “on-target extraction,” or “full extraction” (Popović et al. 2021). One of these three methods is chosen according to the strains of microorganism, the cell wall structure, spore form, and biosafety requirements. It has been shown that the full extraction method has a higher efficacy rate in the identification of difficult bacteria, while in gram-negative bacteria, it is suitable for direct sample spotting and on-target extraction methods (Popović et al. 2021). The sample target plate is then coated with a matrix solution (such as 1 μL of 70% formic acid solution in 50% acetonitrile or 2.5% trifluoroacetic acid, or cyano 4-hydroxycinnamic acid-CHCA) and allowed to dry at room temperature. As the matrix dries, it crystallizes together with the sample, which is then analyzed in the MALDI-TOF instrument (Kazazić et al. 2019a; Kazazić et al. 2019b). The matrix plate is inserted into the instrument and exposed to a laser beam operating at a frequency of 60 Hz in auto-shot, which causes desorption and ionization. The matrix absorbs most of the energy and the microbial peptides are converted into protonated ions which are accelerated by an electrical field towards a detector with the time required to move under vacuum across the flight tube and measured according to the mass to charge ratio (m/z). Smaller analytes arrive at the detector first, followed by increasingly larger analytes. Information on TOF peaks is recorded by the instrument and used to generate a characteristic spectrum, called “a peptide mass fingerprint,” allowing for the identification of the molecules present in the samples based on the unique mass fingerprints. The distinctive calibrated spectrum acquired for each species/strain is recorded and compared with the reference spectrum database. The mass spectra of the recorded proteins, peptides, and ribosomal peptides of the sample can range from 2000 to 21,000 Da depending on the type of microorganism. The generated peak lists are used directly for matches to the reference library using the software's integrated pattern-matching algorithm. The results of the pattern-matching process are expressed in scores ranging from 0 to 3.00 on a logarithmic scale as recommended by the manufacturer. Scores below 1.70 are considered unreliable identification, while a score ≥ 1.70 is considered a genus identification, and a score ≥ 2.00 indicates species identification (De Marco and Burnham 2014; Puk et al. 2018; Surányi et al. 2023).
Identification of bacterial fish pathogens with MALDI-TOF MS
In terms of aquatic health and fisheries management, MALDI-TOF MS serves as a state-of-the-art diagnostic tool for rapid and accurate identification of pathogens affecting aquatic organisms. The proteomics-based MALDI-TOF MS method described in the “General principles and methodology of MALDI-TOF MS” section has two different applications in the diagnosis of fish pathogens. One is the direct identification of the whole-cell bacterial pathogen (Jansson et al. 2020; Piamsomboon et al. 2020; Popović et al. 2022, and the other is the identification of antibiotic resistance biomarkers (for example to beta-lactam, carbapenem, methicillin, and vancomycin) in bacterial pathogens, enabling the instant detection of resistant isolates in real time during the standard routine identification process (Zhu et al. 2002; Fernández-Álvarez et al. 2017; Sandalakis et al. 2017; Popović et al. 2017; Cordovana et al. 2019; Florio et al. 2020). This contributes to early disease detection and helps to develop timely and effective intervention strategies, promoting sustainable practices in aquaculture and healthier fish populations.
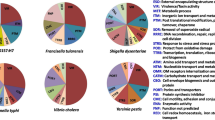
Studies have demonstrated that the MALDI-TOF MS method can identify a single bacterial disease agent or co-infectors which are of significance in marine and inland aquaculture fish species (Table 2) (Nissa et al. 2021; Natnan et al. 2021; Moreira et al. 2021; Parker-Graham et al. 2023). Many of the fish species listed in Table 2 are susceptible to disease, and some have high mortality rates resulting in substantial economic losses (Noga 2010; Austin and Austin 2016; Dar et al. 2022). Several bacterial species in the families of Aeromonadaceae and Pseudomonadaceae (Anagnostopoulos et al. 2022), Mycobacteriaceae (Ziarati et al. 2022), Streptococcaceae (Cañas et al. 2015; Kalimuddin et al. 2017), Vibrionaceae (Burbick et al. 2018), and Enterobacteriaceae (Boylan 2011; Zakrzewski et al. 2022) not only affect aquaculture production but they are also zoonoses transmitted between animal species to humans and are therefore also human public health concerns. The following sections review research publications from 2010 to 2023 retrieved using “MALDI-TOF MS” and “bacterial agents” as keywords or phrases in the Google Scholar search engine (Fig. 3). The bar graph in Fig. 3 shows the number of research studies that utilized MALDI-TOF MS for the identification of bacterial agents of aquatic diseases during this period has been gradually increasing. Table 2 summarizes the applications of MALDI-TOF MS reported in the academic literature to identify 50 major aquatic pathogens (37 g-negative and 13 g-positive) that cause common bacterial diseases in egg, larvae, juvenile, adult fish, shellfish, shrimp, and other aquatic organisms. In these publications, MALDI-TOF was either used alone or in combination with other methodologies in the identification of aquaculture pathogens. The following section summarizes the use of MALDI-TOF MS for the use of identification of both gram-negative and gram-positive fish pathogens.
Identification of gram-negative fish pathogens with MALDI-TOF MS
Aeromonadaceae
Aeromonas spp. are gram-negative, facultatively anaerobic, rod-shaped, non-spore-forming bacteria that morphologically resemble members of the family Aeromonadaceae. They are major infectious agents that cause significant financial losses and high mortality rates in many marine and freshwater fish, especially salmonids under stress (Doukas et al. 1998; Janda and Abbott 2010; Buller 2014; Austin 2017; Pekala-Safińska 2018). Well-known non-motile A. salmonicida causes furunculosis characterized by ulcers on fish skin (Balcazar et al. 2007; Beaz-Hidalgo et al. 2008), while motile aeromonads such as A. hydrophila, A. bestiarum, A. veronii, A. media, A. caviae, and A. sobria are the causative agents of motile aeromonas septicaemia (MAS) which causes abdominal edema, dermatitis, deep ulcers on skin and fins, exophthalmos, and septicemia in fish (Abreu et al. 2018; Beaz-Hidalgo et al. 2013). A number of studies have reported the identification of specific Aeromonas spp. from Salmo trutta, S. gairdnerii (Lauková et al. 2018; Jung-Schroers et al. 2019), Oncorhynchus mykiss (Popović et al. 2022; Tütmez et al. 2023), Acipenser baerii (Vázquez-Fernández et al. 2023), ornamental fish (Cardoso et al. 2021), Danio rerio, Carassius auratus (Walczak et al. 2017), and as a water-borne pathogens (Donohue et al. 2006) (Table 2). However, Pérez‐Sancho et al. (2018) have demonstrated that the accuracy of the genus-level identification of Aeromonas using MALDI-TOF MS is inconsistent when it comes to correctly identifying some of the most prevalent species linked to fish infections. In addition to bacterial identification studies, antibiotic resistance profiles of A. salmonicida (Jung-Schroers et al. 2019), A. hydrophila (Ozbey et al. 2023), A. media (Özcan 2023), A. bestiarum (Jansson et al. 2020), and A. veronii (Özcan 2022) also been determined with MALDI-TOF MS (Table 2).
Flavobacteriaceae
Numerous bacterial species (Flavobacterium spp., Tenacibaculum spp., and Chryseobacterium spp.) belonging to the Flavobacteriaceae family cause flavobacterial diseases in fish, which have devastating impacts on fish stocks in both the wild and farms around the world. Species from flavobacterium cause worldwide economic losses in the aquaculture and fisheries due to infections with high mortality in young fish and morbidity in adult fish (Austin 2017; Crumlish 2017; Wahli and Madsen 2018). The yellow-pigmented, gram-negative F. columnare is the causative agent of one of the oldest known fish diseases, Columnaris, and has been identified by MALDI-TOF MS along with F. psychrophilum the causative agent of bacterial cold water disease (BCWD) and rainbow trout fry syndrome (RTFS) (Dumpala et al. 2010; Li et al. 2015; Pérez-Sancho et al. 2017b; Fernández-Álvarez et al. 2018; Jansson et al. 2020; Neidorf and Morozova 2021; LaFrentz et al. 2022; Chinchilla et al. 2023) (Table 2). In addition, the antimicrobial resistance profiles in F. bernardetii sp. nov. (Saticioglu et al. 2021a), fatty acid composition profiles of F. kayseriense and F. turcium (Saticioglu et al. 2021b), and bacterial protein profiles of F. collinsii (Kondo et al. 2023) have been identified by using the MALDI-TOF MS method (Table 2). In the studies summarized in Table 2, it was evident that the different isolates could be accurately identified by examining the mass score of bacterial protein or mass spectral profiles of other biomarkers from MALDI-TOF MS and confirmed to be Flavobacterium spp. by distinguishing them from other species.
The second genus of bacteria in the family Flavobacteriaceae identified with the MALDI-TOF MS is Tenacibaculum maritimum (Bridel et al. 2020; Fernández-Álvarez et al. 2017). This is a strictly aerobic, gram-negative, rod, and filamentous bacteria with pale yellow flat, thin colonies. It has been identified as the etiological agent of Tenacibaculosis in farmed and wild marine fish species (Toranzo et al. 2005; Avendaño-Herrera et al. 2006; Piñeiro-Vidal et al. 2007). The pathology of tenacibaculosis has been associated with characteristic gross lesions on the body surface of fish such as ulcers, necrosis, eroded mouth, frayed fins, tail rot, and sometimes necrosis on the gills and eyes (Austin and Austin 2016; Le Breton 2020; Mabrok et al. 2023). Bridel et al. (2020) evaluated the variability among T. maritimum isolates based on genome sequence data along with the mass spectral peak characteristics and values of several ribosomal proteins detected by MALDI-TOF MS. In another study by Fernández-Álvarez et al. (2017), the protein mass profiles of 53 Tenacibaculum strains were compared using mass-up software, and the genus-specific peak mass (m/z) values were 2273.50, 2433.48, 2661.82, 4789.03, 5048.15, 5314.34, 10,507.39, and 13,148.28 and were shown to be 100% specific to T. maritimum with score values between 2.048 and 2.258.
Chryseobacterium species belonging to the order flavobacteriales are chemoorganotrophic, rod-shaped, gram-negative fish pathogens forming characteristic yellow-orange colored colonies. This genus contains more than 100 described species from a variety of habitats, including freshwater sources, soil, humans, and marine fish (Loch and Faisal 2015; Austin and Austin 2016). Chryseobacterium species are generally isolated from external lesions, gills, kidneys, and fins of diseased fish. Pérez-Sancho et al. (2017a) reported on the first analysis of Chryseobacterium species and distinguished between the species such as C. balustinum (2.866), C. oncorhynchi (2.897), C. piscicola (2.881), C. shigense (2.790), C. tructae (2.802), C. viscerum (2.941), C. chaponense (2.832), and C. piscium (2.852) using the MALDI-TOF MS technique. Popović et al. (2022) compared the results of MALDI-TOF MS and API 20E identification for gram-negative strains isolated from freshwater, sediment, and fish tissues. Among 190 isolates, two isolates were identified as C. indogenes with a mass score between 1.768 and 2.058, and the other two isolates were identified as C. scophtalmum with a mass score between 1.870 and 2.058.
Vibrionaceae
The pathogenic Vibrionaceae family, which has multiple members, consists of gram-negative, mostly halophilic, flagellated, and facultative anaerobic bacteria (Austin and Austin 2016). Some species of this family, such as Vibrio vulnificus, V. parahaemolyticus, V. mimicus, and V. cholerae, cause enteric pathogenicity such as diarrheal disease, septicemia, and serious wound infections in humans (Sanjuán et al. 2007; Malainine et al. 2013; Burbick et al. 2018; Boonstra et al. 2023). V. anguillarum, V. alginolyticus, V. vulnificus, V. parahaemolyticus, V. ordalii, V. splendidus, V. tubiashii, V. campbellii, V. harveyi, V. owensii, V. rotiferianus, V. azureus, and V. jasicida are also known to be the causative agent of Vibriosis that causes symptoms of hemorrhagic septicemia in various marine fish and freshwater fish (salmon, rainbow trout) (Silva-Rubio et al. 2008; Mougin et al. 2020; Sanches-Fernandes et al. 2022) as well as shellfish, crustaceans, and bivalves (Tanrikul 2007; Mougin et al. 2020; Liu et al. 2004). Lastly, the bacterium V. tapetis causes brown ring disease (BRD) in Manila clams (Paillard et al. 2006).
Many studies (Table 2) have demonstrated that Vibrio species isolated from a wide range of aquatic organisms such as molluscs, seahorse, shrimp, oyster, and clam (Dieckmann et al. 2010; Bauer et al. 2018; Burbick et al. 2018) and cod, char, halibut, eel, Japanese flounder, grouper, seabass, seabream, turbot, ayu, and dreamfish (Low et al. 2014; Ashfaq et al. 2022; Kazazić et al. 2019b; Jansson et al. 2020; Yavuzcan et al. 2022; Boonstra et al. 2023; Haider et al. 2023) can be accurately identified either by evaluating mass score values or peptide mass spectrum peaks. Dieckmann et al. (2010) compared the MALDI-TOF MS-based method with the RNA polymerase beta subunit gene (rpoB) sequence and showed that a total of 83 Vibrio strains were accurately identified by MALDI-TOF MS and species-specific score mass data were between 4200 and 6500 Da. In the study by Cho et al. (2017), four different MALDI-TOF preparation procedures were tested on fifty Vibrio isolates and five reference strains for the identification of Vibrio species. Formic acid and trifluoroacetic acid (TFA) extraction methods applied in the pretreatment of bacterial colonies plated on MALDI-TOF MS plates were considered to be the most suitable methods for the identification of Vibrio species. In a study conducted by Burbick et al. (2018), the MALDI-TOF MS method was compared with rpoB and rpoD gene sequence analysis to identify Vibrio species in isolates obtained from different aquatic organisms such as Paralichthys californicus (California halibut), Hippocampus abdominalis (big-bellied seahorse), Seriola lalandi (yellowtail kingfish), Atractoscion nobilis (white perch), and Pterapogon kauderni (Banggai cardinalfish). It was reported that 29 out of 35 Vibrio species (83%) were correctly identified at the species level using MALDI-TOF MS. Bauer et al. (2018) used MALDI-TOF MS analysis with score values between 1.700 and 2.410 and protein spectrum results to identify eleven pathogenic Vibrio species from Litopenaeus vannamei. These results were further compared with the 16S rRNA sequence and the sequence of the pyrH gene. Erler et al. (2015) carried out the identification of Vibrios in 997 environmental samples by comparing the MALDI-TOF MS and rpoB gene. The mass score value of V. alginolyticus was reported as 2.542, V. cholerae as 2.628, V. parahaemolyticus as 2.609, and V. vulnificus as 2.644. In order to identify Vibrio isolates, Vidal et al. (2020) compared the results of 16S rRNA, pyrH gene sequences, and MALDI-TOF MS analyses. Wu et al. (2020) identified 28 of 74 isolates obtained from seawater, plankton, marine plants, and animals, and Liu et al. (2022) identified isolates from clinical and environmental samples by the MALDI-TOF MS method. V. vulnificus, V. fluvalis, and V. splendidus were identified by analyzing peptide mass score values between 1.750 and 2.410 obtained from seawater (Haider et al. 2023) and Anguilliformes species (eels) (Boonstra et al. 2023), freshwater, ornamental fish, and moribund fish (Joansson et al. 2020; Janampa-Sarmiento et al. 2024; Pastuszka et al. 2024), respectively. V. anguillarum, another important bacterium causing vibriosis, was identified by MALDI-TOF MS from Dicentrarchus labrax and Sparus aurata with an average mass score of 2.120 to 2.500 (Kazazić et al. 2019b; Jansson et al. 2020; Mougin et al. 2021). V. alginolyticus, one of the most important causative bacteria causing vibriosis, was identified by Low et al. (2014) in Epinephelus fuscoguttatus (brown marble grouper) and by Ashfaq et al. (2019) in seawater by analyzing peptide mass values obtained from MALDI-TOF method. Bronzato et al. (2018) identified V. alginolyticus in 49 of 209 isolates obtained from Perna perna mussels using MALDI-TOF and rpoA/pyrH sequencing methods together. Wang et al. (2024) identified 35 of a total of 55 different bacterial isolates as V. alginolyticus by MALDI-TOF MS and confirmed the same isolates with a new method, rapid visual nucleic acid detection (RPA-CRISPR/Cas13a). Another bacterial species identified by the MALDI-TOF method was V. harveyi, which was also identified from Sarpa salpa (dreamfish) by Yavuzcan et al. (2022) and Culot et al. (2021) from shrimp. V. parahaemolyticus, the other causative agent of vibriosis, has been reported to be identified from D. labrax, Haliotis tuberculata (green ormer) (Malainine et al. 2013), Crassostrea gigas (Pacific oyster) (Mougin et al. 2020), and seafood samples (Fasulkova et al. 2023) and seawater (Khandeparker et al. 2023) using the MALDI-TOF method. In addition, Rahmani et al. (2021) concluded that V. tapetis isolated from Manila clams was pathogenic based on protein profiles indicating the presence of a virulence gene. Moussa et al. (2021) reported a study on marine molluscs to identify V. tapetis.
Although many studies listed above have reported the identification of Vibrio species by MALDI-TOF MS, a study by Sanches-Fernandes et al. (2022) showed that isolates belonging to V. tubiashii/V. europaeus and V. owensii/ V. jasicida/V. campbellii species could not be correctly differentiated using some commonly available databases for MALDI-TOF MS-based classification. Similarly, Mougin et al. (2020) successfully identified several strains belonging to V. campbellii, V. owensii, V. azureus, V. jasicida, V. sinaloensis, and V. rotiferianus using an in-house database called Luvibase, which contains the master spectral profiles (MSPs) of 23 strains in combination with Bruker v.9.0.0.0 database, since Bruker v.9.0.0.0 could not distinguish some Vibrio species.
Photobacterium damsale is a gram-negative, halophilic, rod-shaped bacteria belonging to the family Vibrionaceae. This pathogen is an emerging threat in marine aquaculture and also occasionally causes disease in poikilotherms and mammals, including humans. P. damselae subsp. piscicida (formerly known as Pasteurella piscicida) is responsible for photobacteriosis (fish pasteurellosis) causing outbreaks characterized by multifocal necrosis in the spleen, liver, and kidneys leading to high mortality rates in larvae and juveniles fish (Austin and Austin 2016), and P. damselae subsp. damselae (formerly known as Vibrio damsela) which causes hemorrhagic ulcerative skin lesions in fish has also been isolated from humans and has been recognized as a zoonotic pathogen (Andreoni and Magnani 2014). Table 2 shows that both subspecies were identified and confirmed by the spp. names by Kazazić et al. (2019a) by trying longer incubation times and three MALDI-TOF methodologies such as direct bacterial sample spotting, on-target extraction, and full extraction. Pérez-Sancho et al. (2016) also showed that all tested isolates from D. labrax, S. aurata, and M. saxatilis were identified as P. damselae subsp. damselae with an average score value of 2.256 (2.114–2.353) and P. damselae subsp. piscicida with an average of 2.286 (2.250–2.331) by MALDI-Biotyper. Yavuzcan et al. (2022) identified Photobacterium damselae with MALDI-TOF with high identification score values (Table 2).
Pseudomonadaceae
Pseudomonas is a genus of gram-negative, rod-shaped, and non-spore-forming bacteria belonging to the family Pseudomonadaceae in the class Gammaproteobacteria. Due to their extensive metabolic diversity, the 313 members of the genus (Parte et al. 2020) can live in a wide range of habitats, including water, soil, plants, insects, fish, aquatic creatures, and animals (Khan et al. 2007; Duman et al. 2021). P. putida, P. aeruginosa, P. lundensis, P. fluorescens, and P. baetica are opportunistic human pathogens (Sadikot et al. 2005; De Bentzmann and Plesiat 2011; Tohya et al. 2022). A large number of Pseudomonas spp. (P. aeruginosa, P. anguilliseptica, P. proteolytica, P. baetica, P. chlororaphis, P. fluorescens, P. koreensis, P. luteola, P. plecoglossicida, P. pseudoalcaligenes, P. lundensis, P. tructae, P. extremaustralis, and P. putida) have been reported as the most common bacterial causative agent of stress-related infections in rainbow trout and tench (strawberry disease), sea bream, sea bass, ayu, and ornamental fish. Although the symptoms of the disease vary according to the fish, they are generally described as the distended abdomen and hemorrhaging on the body surface, fin or tail rot, hemorrhagic septicemia, hemorrhagic ascites, exophthalmitis with extensive skin lesions, and external ulceration and Sekiten byo (red spot) (Altinok et al. 2006, 2007; von Siebenthal et al. 2009; Nishimori et al. 2000; López et al. 2012; Austin and Austin 2016; Walczak et al. 2017; Oh et al. 2019; Aydin et al. 2023). The MALDI-TOF MS method has been used to detect previously identified Pseudomonas spp. (Kačániová et al. 2019; Klūga et al. 2019; Popović et al. 2022) as well as the novel species P. Sivasensis, P. anatoliensis sp. nov., and P. iridis sp. nov. (Duman et al. 2020; 2021), P. haemolytica (Saticioglu et al. 2022).
Enterobacterales
Yersinia ruckeri is a gram-negative, non-spore-forming, round-tipped, rod-shaped, and facultative anaerobe bacteria belonging to the Yersiniaceae family of the Enterobacterales order. This bacterium is the causative agent of yersiniosis or enteric red mouth disease (ERM) in salmonids, especially in O. mykiss (Tobback et al. 2007; Huang et al. 2013; Austin and Austin 2016). Recent literature shows the successful identification of Y. ruckeri from O. mykiss (Jansson et al. 2020), S. salar (Ojasanya et al. 2022), and Cyprinus carpio (Kazarnikova et al. 2019) using the MALDI-TOF MS method (Table 2).
Edwardsiella tarda, a gram-negative, facultative anaerobic, small, motile, flagellated, and straight rod bacterium belonging to the order Enterobacterales and the family Hafniaceae, is a serious fish pathogen causing Edwardsiella septicemia (ES), Edwardsiellosis, fish gangrene, and red disease in both farmed and wild fish species (Plumb 1993; Thune et al. 1993; Park et al. 2012; Austin and Austin 2016). E. tarta is zoonotic and can infect fish as well as amphibians, reptiles, birds, and mammals (Xu and Zhang 2014; Verma et al. 2022). Table 2 shows two studies on the use of MALDI-TOF MS for the identification of E. tarda. The first study by Piamsomboon et al. (2020) showed that isolates from O. niloticus, a hybrid catfish, were E. tarda. Reis et al. (2023) also used MALDI-TOF MS for the identification of E. tarda isolated from Colossoma macropomum, as well as dnaJ gene sequencing to confirm the accuracy of the species identification.
Piscirickettsiaceae
Piscirickettsia salmonis is a gram-negative, recalcitrant, and psychotropic bacterium that is generally non-motile, aerobic, not encapsulated, highly fastidious, and very slow-growing, coccoid, and found in pairs or ring-shaped structures (Fryer et al. 1992; Rozas and Enríquez 2014; Makrinos and Bowden 2017). It is reported to be the causative agent of Piscirickettsiosis, also known as Salmonid Rickettsial Septicaemia (SRS) and Salmon Rickettsia Syndrome (Branson and Nieto Diaz-Munoz 1991; Lannan and Fryer 1993). It is an intracellular fish pathogen that can infect and reproduce in monocyte and macrophage cell lines. It is an economically important disease that greatly affects the production of cold-water salmonids and non-salmonids including sea bass grouper and Nile tilapia in many countries from the United States, North America, Canada, and Europe to Asia (Austin and Austin 2016). P. salmonis is very difficult to isolate, and culture from the fish cell, MALDI-TOF MS, and molecular identification have been used in combination as a powerful tool for accurate and early detection to facilitate disease control in the salmon farming industry (López-Cortés et al. 2017; Olate et al. 2016; Zrnčić et al. 2021).
Identification of gram-positive fish pathogens with MALDI-TOF MS
Renibacterium salmoninarum
R. salmoninarum, a member of the Micrococcaceae family, is a small, gram-positive, rod-shaped, fastidious, slow-growing, extracellular/intracellular pathogen causing bacterial kidney disease (BKD), especially in farmed and wild salmonid fish, reported from many countries around the world including Scotland, England, Germany, Spain, Turkey, Italy, France, Iceland, Balkan Peninsula, Canada, United States of America, Chile, and Japan (Fryer and Lannan 1993; Daly et al. 2001; Vardić et al. 2007). These bacteria can be transmitted vertically and horizontally via eggs (Evelyn et al. 1986) and primarily accumulate in the kidney and other internal organs, causing edema, granuloma lesions, and resulting in morbidity and mortality (Gutenberger et al. 1997). Clinical signs of BKD include loss of appetite, lethargy, dark colouration, exophthalmia, bloody ascites, and hemorrhages around the cloaca and fins (Rhodes et al. 2004; Austin and Austin 2016; Elliott 2017). Jansson et al. (2020) successfully identified bacterial isolates of Renibacterium at the species level from rainbow trout, Atlantic salmon, chinook salmon, coho salmon, and arctic char (Table 2). Conversely, Jia et al. (2023) compared different swab transport methods, along with molecular detection (qPCR) and MALDI-TOF methods for the identification of R. salmoninarum from bacterial colonies on selective kidney disease medium (SDKM) agar plates, but when samples from contaminated colonies on SDKM agar plates were tested with MALDI-TOF MS, the results obtained could not determine whether this isolate was R. salmoninarum because it was not taken from a single colony.
Streptococcaceae
Lactococcus garvieae is a gram-positive, facultative anaerobic, non-motile, non-spore-forming, ovoid cocci, pair, and short-chain bacterium (Ravelo et al. 2001; Vendrell et al. 2006). L. garvieae is the bacterial agent of lactococcosis, an important zoonotic disease affecting fish, other animals, and humans. It causes peritonitis, urinary tract infections, and infective endocarditis in humans (Ferrario et al. 2013; Cañas et al. 2015; Gibello 2016). In fish, it has been reported to cause lesions in the vascular endothelium, causing hemorrhages and petechiae on the surface of internal organs in rainbow trout, yellowtail, cobia, and mullet and causing large-scale deaths (Zlotkin et al. 1998b; Eldar and Ghittino 1999; Diler et al. 2002; Meyburgh et al. 2017). L. garvieae was successfully identified at the species level from salmonids by MALDI-TOF MS (Assis et al. 2017; Radosavljević et al. 2020; Saticioglu et al. 2023) and has a strong score at m/z 2.35 that corresponded to the nisin leader peptide mass peak (Sequeiros et al. 2015) (Table 2). Torres-Corral and Santos (2021) also showed that the two mass peaks belonging to the small (30S) subunit of the species-specific DNA-binding ribosomal protein were m/z 4717.04 and 9431.98 Da. They also found that a high intraspecies similarity (83% similarity) was observed among L. garvieae strains, regardless of the host species or geographical region from which the strains were isolated. Torres-Corral and Santos (2022) also showed that the identification of L. garvieae by determining the mass peaks associated with the antibiotic resistance pattern is a simple and rapid technique using protein mass patterns or biomarkers for oxytetracycline and florfenicol resistance that were associated with the expression of a peptide directly or indirectly related to phenotypic resistance.
Another bacterial species belonging to the Streptococcaceae family, which can be identified by MALDI-TOF MS, is a novel pathogen, gram-positive L. petauri, which is known as the second causative agent of lactococcosis in fish (Goodman 2017; Kotzamanidis et al. 2020; Altinok et al. 2022; Egger et al. 2023; Littman et al. 2023; Saticioglu et al. 2023).
MALDI-TOF MS can also be used to identify the gram-positive, cocci Streptococcus iniae, S. agalactiae, S. dysgalactiae, and S. parauberis belonging to the family Streptococcaceae (Mata et al. 2004; Buller 2014; Austin and Austin 2016). The disease known as warm water streptococcosis (mortalities at temperatures above 15 °C) caused by these species has signs of septicemia, suppurative exophthalmia (pop-eye), and meningoencephalitis in several aquatic species in marine and freshwater fish including rainbow trout (O. mykiss) (Lahav et al. 2004), olive flounder (Paralichthys olivaceus) (Nho et al. 2009; Kim et al. 2015), turbot (Scophthalmus maximus), flatfish (Domeénech et al. 1996), barramundi (Lates calcalifera), tilapia (Oreochromis spp.), and sturgeon (Acipenser spp.) (Deng et al. 2017). These Streptococcus species have been also recognized as zoonotic pathogens (Facklam et al. 2005; Gauthier 2015; Kim et al. 2017). Tavares et al. (2018) showed fifteen Streptococcus spp. isolates from Amazon catfish (Leiarius marmoratus × Pseudoplatystoma corruscans) were identified as with other diagnostic methods but also confirmed by MALDI-TOF MS. Table 2 summarizes the successful identification of S. agalactiae were from O. niloticus (Assis et al. 2017), S. iniae, and S. parauberis isolated from P. olivaceus (Kim et al. 2015; 2017) and from S.maximus and O. mykiss (Torres-Corral et al. 2019), S. agalactiae from O. niloticus, O. aureus and O. mossambicus (Piamsomboon et al. 2020), and S. iniae from Acipenser transmontanus (Pierezan et al. 2020).
Enterococcaceae
The gram-positive and rod-shaped Vagococcus salmoninarum (Wallbanks 1990; Schmidtke and Carson 1994) has been reported to be the causative agent of vagococcosis or “Cold Water Streptococcosis” with peritonitis in Atlantic salmon, rainbow trout, and brown trout (Michel et al. 1997; Ruiz-Zarzuela et al. 2005; Salogni et al. 2007). Although there are many articles in the literature showing the identification of V. salmoninarum by classical methods, the application of the MALDI-TOF method has been limited due to a lack of reference data (Buller and Hair 2016). Nevertheless, Torres-Corral and Santos (2019) recently reported that V. salmoninarum strains can be successfully identified by MALDI-TOF MS methods.
Mycobacteriaceae
Four Mycobacterium species from fish identified by the MALDI-TOF MS method are known as pleomorphic, acid-fast, gram-positive, aerobic, non-motile, and rods form (Gauthier and Rhodes 2009). These species are known to be the causative agents of mycobacteriosis, also known as fish tuberculosis, which develops chronically and rapidly in many fish species (Kaattari et al. 2005; Whipps et al. 2007; Zanoni et al. 2008; Jacob et al. 2009; Mugetti et al. 2021) and progresses with external symptoms such as emaciation, skin swelling, exophthalmos, open lesions, and ulceration symptoms in several organs (Austin and Austin 2016). Some Mycobacterium species are also known to be zoonotic (Bhatty et al. 2000; Ucko and Colorni 2005). The slow-growing M. marinum was first identified by Aronson in (1926), and it has been observed and identified from many fish worldwide (Lansdell et al. 1993; Puk et. al. 2018; Elgendy et al. 2023). Kurokawa et al. (2013) demonstrated that M. marinum and M. avium could be reliably identified with the MALDI-TOF MS, and Puk et al. (2018) showed accurate detection of M. marinum, M. peregrinum, M. fortuitum, and M. abscessus in aquarium fish with MALDI-TOF MS in combination with two other techniques (16S rRNA and Hsp65) to maximize the identification results.
Conclusion
In conclusion, the proteomics-based MALDI-TOF MS method appears to be an effective and reliable microbial identification approach for rapid, accurate, and specific identification and classification of many bacteria at the species level with unique fingerprints (spectral protein peaks) and mass score values generated by cellular ribosomal proteins or antimicrobial resistance markers present in bacteria compared to reference mass spectral databases. In this review, it is shown that the number of published studies that utilized MALDI-TOF MS between 2010 and 2023 has increased rapidly, reflecting the effectiveness and efficiency of the technique.
Although the use of this method in aquatic studies has been validated with numerous outstanding achievements, its use for the identification of pathogens has a number of advantages. Firstly, in addition to speeding up the detection process of diseases that damage aquatic organisms, it can efficiently screen a large number of field samples at one time due to its high throughput and low sample preparation needs. It is also a beneficial approach as it eliminates the need for expensive reagents, labor-intensive steps, and highly trained operators that are part of traditional techniques.
There are a few limitations to consider, nevertheless. The main disadvantage that may discourage users from using this technique is the high initial cost of acquiring a MALDI-TOF MS instrument, which has a low unit sample cost, although not as expensive as the NGS platforms that perform the next-generation sequencing technique. Another limitation of current devices is that bacteria cannot be sampled and identified directly from the diseased tissue sample on the instrument. Although it can accurately identify most bacterial species, the inability to detect some species not found in global proteomics databases may be one of the limitations of this method. Therefore, the potential for in-house databases such as the EnviBase library and Luvibase database, which have been developed to identify Vibrio species, is needed to identify other aquatic pathogens.
Overall, MALDI TOF MS technology has ongoing innovations and system refinement for future user convenience. Efforts to optimize sample preparation protocols and streamline data analyses can make MALDI-TOF MS more accessible and user-friendly for aquaculture practitioners and hold great promise for advancing disease diagnosis and promoting sustainable aquaculture practices in the future. Nevertheless, more fish pathogens’ score values and peak data should be uploaded to global protein databases for accurate discrimination and identification of fish pathogens. The use of this method in the aquaculture disease sector promotes sustainable practices and healthier fish populations in aquaculture by contributing to the early detection of disease and the development of timely and effective intervention strategies. Therefore, more research is crucial to facilitating the routine use of MALDI-TOF MS for the identification of pathogens of importance to the aquaculture and fisheries sectors.
Data availability
No datasets were generated or analyzed during the current study.
References
Abreu REF, Magalhaes TC, Souza RC, Oliveira STL, Ibelli AMG, Demarqui FN, Gouveia GV (2018) Environmental factors on virulence of Aeromonas hydrophila. Aqua Inter 26:495–507. https://doi.org/10.1007/s10499-017-0230-2
Adams A, Thompson KD (2011) Development of diagnostics for aquaculture: challenges and opportunities. Aqua Res 42:93–102. https://doi.org/10.1111/j.1365-2109.2010.02663.x
Ahmad F, Babalola OO, Tak HI (2012) Potential of MALDI-TOF mass spectrometry as a rapid detection technique in plant pathology: identification of plant-associated microorganisms. Anal Bioanal Chem 404:1247–1255. https://doi.org/10.1007/s00216-012-6091-7
Akimowicz M, Bucka-Kolendo J (2020) MALDI-TOF MS-application in food microbiology. Acta Bioch Polonica 67(3):327–332. https://doi.org/10.18388/abp.2020_5380
Alcalá L, Marín M, Ruiz A, Quiroga L, Zamora-Cintas M, Fernández-Chico MA, Muñoz P, Rodríguez-Sánchez B (2021) Identifying anaerobic bacteria using MALDI-TOF mass spectrometry: a four-year experience. Front Cell Infec Micro 11:521014. https://doi.org/10.3389/fcimb.2021.521014
Alderman DJ (1996) Geographical spread of bacterial and fungal diseases of crustaceans. Rev Sci Tec OIE 603–632. https://doi.org/10.20506/rst.15.2.943
Altinok I, Kurt I (2003) Molecular diagnosis of fish diseases: a review. Turk J Fish Aqua Sci 3(2):131–138
Altinok I, Kayis S, Capkin E (2006) Pseudomonas putida infection in rainbow trout. Aqua 261:850–855. https://doi.org/10.1016/j.aquaculture.2006.09.009
Altinok I, Balta F, Capkin E, Kayis S (2007) Disease of rainbow trout caused by Pseudomonas luteola. Aqua 273:393–397. https://doi.org/10.1016/j.aquaculture.2007.10.025
Altinok I, Capkin E, Kayis S (2008) Development of multiplex PCR assay for simultaneous detection of five bacterial fish pathogens. Vet Micro 131:332–338. https://doi.org/10.1016/j.vetmic.2008.04.014
Altinok I, Ozturk RC, Ture M (2022) NGS analysis revealed that Lactococcus garvieae Lg-per was Lactococcus petauri in Türkiye. J Fish Dis 45:1839–1843. https://doi.org/10.1111/jfd.13708
Anagnostopoulos DA, Parlapani FF, Natoudi S, Syropoulou F, Kyritsi M, Vergos I, Hadjichristodoulou C, Kagalou I, Boziaris IS (2022) Bacterial communities and antibiotic resistance of potential pathogens involved in food safety and public health in fish and water of Lake Karla, Thessaly. Greece Pathogens 11(12):1473. https://doi.org/10.3390/pathogens11121473
Andreoni F, Magnani M (2014) Photobacteriosis: prevention and diagnosis. J Immun Res 793817:1–7. https://doi.org/10.1155/2014/793817
Anwer R, Darami H, Almarri FK, Albogami MA, Alahaydib F (2022) MALDI-TOF MS for rapid analysis of bacterial pathogens causing urinary tract infections in the Riyadh Region. Dis 10:78. https://doi.org/10.3390/diseases10040078
Arafa SH, Elbanna K, Osman GE, Abulreesh HH (2023) Candida diagnostic techniques: a review. J Umm Al-Qura Uni Appl Sci 9:360–367. https://doi.org/10.1007/s43994-023-00049-2
Aronson JD (1926) Spontaneous tuberculosis in salt water fish. J Infec Dis 39(4):315–320. https://doi.org/10.1093/infdis/39.4.315
Ashfaq MY, Al-Ghouti MA, Qiblawe H, Rodrigues DF, Hu Y, Zouari N (2019) Isolation, identification and biodiversity of antiscalant degrading seawater bacteria using MALDI-TOF-MS and multivariate analysis. Sci Total Environ 656:910–920. https://doi.org/10.1016/j.scitotenv.2018.11.477
Ashfaq MY, Da’na DA, Al-Ghouti MA (2022) Application of MALDI-TOF MS for identification of environmental bacteria: a review. J Environ Man 305:114359. https://doi.org/10.1016/j.jenvman.2021.114359
Assis GB, Pereira FL, Zegarra AU, Tavares GC, Leal CA, Figueiredo HC (2017) Use of MALDI-TOF mass spectrometry for the fast identification of gram-positive fish pathogens. Front Micro 8:1492. https://doi.org/10.3389/fmicb.2017.01492
Austin B (2017) Diagnosis and control of diseases of fish and shellfish, 1st edn. Published by John Wiley and Sons Ltd., Chennai, India, Diagnosis and control of diseases of fish and shellfish. https://doi.org/10.1002/9781119152125
Austin B (2019) Methods for the diagnosis of bacterial fish diseases. Mar Life Sci Tech 1:41–49. https://doi.org/10.1007/s42995-019-00002-5
Austin B, Austin DA (2016) Bacterial Fish Pathogens, 6th Ed. Springer, Chichester, UK. https://doi.org/10.1007/978-3-319-32674-0
Avendaño-Herrera R, Toranzo AE, Magariños B (2006) Tenacibaculosis infection in marine fish caused by Tenacibaculum maritimum: a review. Dis Aqua Org 71(3):255–266. https://doi.org/10.3354/dao071255
Aydin A, Sudagidan M, Mamatova Z, Yurt MNZ, Ozalp VC, Zornu J, Tavornpanich S, Brun E (2023) Bacterial skin microbiota of seabass from Aegean fish farms and antibiotic susceptibility of psychrotrophic Pseudomonas. Foods 12(10):1956. https://doi.org/10.3390/foods12101956
Balcazar JL, Vendrell D, de Blas I, Ruiz-Zarzuela I, Girones O, Muzquiz JL (2007) Quantitative detection of Aeromonas salmonicida in fish tissue by real-time PCR using self-quenched, fluorogenic primers. J Med Micro 56(3):323–328. https://doi.org/10.1099/jmm.0.46647-0
Bauer J, Teitge F, Neffe L, Adamek M, Jung A, Peppler C, Jung-Schroers V (2018) Recommendations for identifying pathogenic Vibrio spp. as part of disease surveillance programmes in recirculating aquaculture systems for Pacific white shrimps (Litopenaeus vannamei). J Fish Dis 41(12):1877–1897. https://doi.org/10.1111/jfd.12897
Beaz-Hidalgo R, López-Romalde S, Toranzo AE, Romalde JL (2008) Polymerase chain reaction amplification of repetitive intergenic consensus and repetitive extragenic palindromic sequences for molecular typing of Pseudomonas anguilliseptica and Aeromonas salmonicida. J Aqua Anim Health 20:75–85. https://doi.org/10.1577/H07-007.1
Beaz-Hidalgo R, Martinez-Murcia A, Figueras MJ (2013) Reclassification of Aeromonas hydrophila subsp. dhakensis Huys et al. 2002 and Aeromonas aquariorum Martinez-Murcia et al. 2008 as Aeromonas dhakensis sp. nov. comb nov. and emendation of the species Aeromonas hydrophila. Syst Appl Micro 36:171–176. https://doi.org/10.1016/j.syapm.2012.12.007
Bhatty MA, Turner DP, Chamberlain ST (2000) Mycobacterium marinum hand infection: case reports and review of literature. British J Plastic Sur 53(2):161–165. https://doi.org/10.1054/bjps.1999.3245
Biswas S, Rolain JM (2013) Use of MALDI-TOF mass spectrometry for identification of bacteria that are difficult to culture. J Micro Met 92(1):14–24. https://doi.org/10.1016/j.mimet.2012.10.014
Bohara K, Joshi P, Acharya KP, Ramena G (2024) Emerging technologies revolutionising disease diagnosis and monitoring in aquatic animal health. Rev Aqua 16(2):836–854. https://doi.org/10.1111/raq.12870
Böhme K, Fernández-No IC, Barros-Velázquez J, Gallardo JM, Cañas B, Calo-Mata P (2012) SpectraBank: an open access tool for rapid microbial identification by MALDI-TOF MS fingerprinting. Electrophoresis 33(14):2138–2142. https://doi.org/10.1002/elps.201200074
Böhme K, Fernández-No IC, Pazos M, Gallardo JM, Barros-Velázquez J, Cañas B, Calo-Mata P (2013) Identification and classification of seafood borne pathogenic and spoilage bacteria: 16S rRNA sequencing versus MALDI-TOF MS fingerprinting. Electrophoresis 34(6):877–887. https://doi.org/10.1002/elps.201200532
Boonstra M, Fouz B, van Gelderen B, Dalsgaard I, Madsen L, Jansson E, Amaro C, Haenen O (2023) Fast and accurate identification by MALDI-TOF of the zoonotic serovar E of Vibrio vulnificus linked to eel culture. J Fish Dis 46(4):445–452. https://doi.org/10.1111/jfd.13756
Boylan S (2011) Zoonoses associated with fish. Vet Clin Exo Anim Prac 14(3):427–438. https://doi.org/10.1016/j.cvex.2011.05.003
Branson EJ, Nieto Diaz-Munoz D (1991) Description of a new disease condition occurring in farmed coho salmon, Oncorhynchus kisutch (Walbaum) in South America. J Fish Dis 14:147–156. https://doi.org/10.1111/j.1365-2761.1991.tb00585.x
Brauge T, Trigueros S, Briet A, Debuiche S, Leleu G, Gassilloud B, Wilhelm A, Py JS, Midelet G (2021) MALDI-TOF mass spectrometry fingerprinting performance versus 16S rDNA sequencing to identify bacterial microflora from seafood products and sea water samples. Front Mar Sci 8:650116. https://doi.org/10.3389/fmars.2021.650116
Bridel S, Bourgeon F, Marie A, Saulnier D, Pasek S, Nicolas P, Bernardet JF, Duchaud E (2020) Genetic diversity and population structure of Tenacibaculum maritimum, a serious bacterial pathogen of marine fish: from genome comparisons to high throughput MALDI-TOF typing. Vet Res 51(1):1–17. https://doi.org/10.1186/s13567-020-00782-0
Bronzato GF, Oliva MS, Alvin MG, Pribul BR, Rodrigues DP, Coelho SM, Souza M (2018) MALDI-TOF MS as a tool for the identification of Vibrio alginolyticus from Perna perna mussels (Linnaeus, 1758). Pesquisa Vet Brasileira 38:1511–1517. https://doi.org/10.1590/1678-5150-PVB-5233
Buller BN (2014) Bacteria and fungi from fish and other aquatic animals. 2nd Ed. Bentley, CABI Publishing. Western Australia p.1–423. 978–0851997384
Buller N, Hair S (2016) Identification of bacteria from aquatic animals. Micro Australia 37(3):129–131
Burbick CR, Nydam SD, Hendrix GK, Besser TE, Diaz D, Snekvik K (2018) Use of matrix-assisted laser desorption ionization time-of-flight mass spectrometry for the identification of pathogenic Vibrio in fish. J Aqua Anim Health 30(4):332–338. https://doi.org/10.1002/aah.10044
Çağatay IT (2022) FTA® card tool for sampling and rapid diagnosis of bacterial diseases from rainbow trout (Oncorhynchus mykiss) tissue. Aqua Inter 30:419–428. https://doi.org/10.1007/s10499-021-00810-6
Cañas VH, Ramirez MP, Jiménez FB, Martin MR, Casas CM, Arriaza MM, Marí JN (2015) Lactococcus garvieae endocarditis in a native valve identified by MALDI-TOF MS and PCR-based 16s rRNA in Spain: a case report. New Micro New Infec 5:13–15. https://doi.org/10.1016/j.nmni.2015.02.00
Cardoso PH, Moreno LZ, de Oliveira CH, Gomes VT, Silva APS, Barbosa MR, Sato MIZ, Balian SC, Moreno AM (2021) Main bacterial species causing clinical disease in ornamental freshwater fish in Brazil. Folia Micro 66:231–239. https://doi.org/10.1007/s12223-020-00837-x
Chalupová J, Raus M, Sedlářová M, Šebela M (2014) Identification of fungal microorganisms by MALDI-TOF mass spectrometry. Biotec Advan 32(1):230–241. https://doi.org/10.1016/j.biotechadv.2013.11.002
Chinchilla B, Vázquez-Fernández E, Rebollada-Merino A, Pérez-Sancho M, Domínguez L, Rodríguez-Bertos A (2023) First detection of Flavobacterium psychrophilum in juvenile Siberian sturgeons (Acipenser baerii) and description of the pathological findings. J Fish Dis 46:887–894. https://doi.org/10.1111/jfd.13801
Cho Y, Kim E, Han SK, Yang SM, Kim MJ, Kim HJ, Kim HY (2017) Rapid identification of vibrio species isolated from the southern coastal regions of Korea by MALDI-TOF mass spectrometry and comparison of MALDI sample preparation methods. J Micro Biotech 27(9):1593–1601. https://doi.org/10.4014/jmb.1704.04056
Chun S, Gopal J, Muthu M (2022) A consolidative synopsis of the MALDI-TOF MS accomplishments for the rapid diagnosis of microbial plant disease pathogens. TrAC Trends Anal Chem 156:116713. https://doi.org/10.1016/j.trac.2022.116713
Cordovana M, Pranada AB, Ambretti S, Kostrzewa M (2019) MALDI-TOF bacterial subtyping to detect antibiotic resistance. Clin Mass Spectr 14:3–8. https://doi.org/10.1016/j.clinms.2019.06.002
Croxatto A, Prodhom G, Greub G (2012) Applications of MALDI-TOF mass spectrometry in clinical diagnostic microbiology. FEMS Micro Rev 36(2):380–407. https://doi.org/10.1111/j.1574-6976.2011.00298.x
Crumlish M (2017) Bacterial diagnosis and control in fish and shellfish. Diagnosis and control of diseases of fish and shellfish. 1st ed. Edited by Brian Austin and Aweeda Newaj-Fyzul. Published by John Wiley and Sons Ltd. p. 5–18. https://doi.org/10.1002/9781119152125.ch2
Culot A, Grosset N, Bruey Q, Auzou M, Giard JC, Favard B, Gautier M (2021) Isolation of Harveyi clade Vibrio spp. collected in aquaculture farms: how can the identification issue be addressed? J Microb Met 180:106106. https://doi.org/10.1016/j.mimet.2020.106106
Cunningham CO (2002) Molecular diagnosis of fish and shellfish diseases: present status and potential use in disease control. Aqua 206(1–2):19–55. https://doi.org/10.1016/S0044-8486(01)00864-X
Daly JG, Griffiths SG, Kew AK, Moore AR, Olivier G (2001) Characterization of attenuated Renibacterium salmoninarum strains and their use as live vaccines. Dis Aqua Org 44(2):121–126. https://doi.org/10.3354/dao044121
Dar GH, Bhat RA, Qadri H, Al-Ghamdi KM, Hakeem KR. (Eds.) (2022) Bacterial fish diseases. Academic Press. Elsiver Inc. UK. p. 118–144.
Dare D (2006) Rapid bacterial characterization and identification by MALDI-TOF mass spectrometry. In: Tang YW, Stratton CW. (Eds.), Advanced techniques in diagnostic microbiology. Springer Science Business Media, LLC, NY. p. 117–134.
De Bentzmann S, Plesiat P (2011) The Pseudomonas aeruginosa opportunistic pathogen and human infections. Environ Micro 13:1655–1665. https://doi.org/10.1111/j.1462-2920.2011.02469.x
De Marco ML, Burnham CAD (2014) Diafiltration MALDI-TOF mass spectrometry method for culture-independent detection and identification of pathogens directly from urine specimens. American J Clin Patho 141(2):204–212. https://doi.org/10.1309/AJCPQYW3B6JLKILC
De Bruijn I, Liu Y, Wiegertjes GF, Raaijmakers JM (2018) Exploring fish microbial communities to mitigate emerging diseases in aquaculture. FEMS Micro Eco 94(1):fix161. https://doi.org/10.1093/femsec/fix161
del Cerro A, Marques I, Guijarro JA (2002) Simultaneous detection of A. salmonicida, F.psychrophilum and Y. ruckeri three major fish pathogens, by multiplex PCR. Appl Environ Micro 68:5177–5180. https://doi.org/10.1128/AEM.68.10.5177-5180.2002
Deng ML, Yu ZH, Geng Y, Wang KY, Chen DF, Huang XL, Lai WM (2017) Outbreaks of Streptococcosis associated with Streptococcus iniae in Siberian sturgeon (Acipenser baerii) in China. Aqua Res 48(3):909–919. https://doi.org/10.1111/are.12934
Dieckmann R, Strauch E, Alter T (2010) Rapid identification and characterization of Vibrio species using whole-cell MALDI-TOF mass spectrometry. J Appl Micro 109(1):199–211. https://doi.org/10.1111/j.1365-2672.2009.04647.x
Diler O, Altun S, Adiloglu AK, Kubilay A, Isıklı B (2002) First occurrence of Streptococcosis affecting farmed rainbow trout (Oncorhynchus mykiss) in Turkey. Bull Assoc Fish Pathol 22:21–26
Domeénech A, Derenaáandez-Garayzábal J, Pascual C, Garcia J, Cutuli M, Moreno M, Collins M, Dominguez L (1996) Streptococcosis in cultured turbot, Scopthalmus maximus (L.) associated with Streptococcus parauberis. J Fish Dis 19:33–38. https://doi.org/10.1111/j.1365-2761.1996.tb00117.x
Donohue MJ, Smallwood AW, Pfaller S, Rodgers M, Shoemaker JA (2006) The development of a matrix-assisted laser desorption/ionization mass spectrometry-based method for the protein fingerprinting and identification of Aeromonas species using whole cells. J Micro Metho 65(3):380–389. https://doi.org/10.1016/j.mimet.2005.08.005
Doukas V, Athanassopoulou F, Karagouni E, Dotsika E (1998) Aeromonas hydrophila infection in cultured sea bass, Dicentrarchus labrax L. and Puntazzo puntazzo Cuvier from the Aegean Sea. J Fish Dis 21:317–320. https://doi.org/10.1046/j.1365-2761.1998.00105.x
Drissner D, Freimoser FM (2017) MALDI-TOF mass spectroscopy of yeasts and filamentous fungi for research and diagnostics in the agricultural value chain. Chem Bio Techno Agricul 4(1):1–12. https://doi.org/10.1186/s40538-017-0095-7
Duman M, Mulet M, Saticioglu IB, Altun S, Gomila M, Lalucat J, Garcia-Valdes E (2020) Pseudomonas sivasensis sp. nov. isolated from farm fisheries in Turkey. Syste Appl Micro 43(4):126103. https://doi.org/10.1016/j.syapm.2020.126103
Duman M, Mulet M, Altun S, Saticioglu IB, Ozdemir B, Ajmi N, Lalucat J, García-Valdés E (2021) The diversity of Pseudomonas species isolated from fish farms in Turkey. Aqua 535:736369. https://doi.org/10.1016/j.aquaculture.2021.736369
Duman M, Altun S, Saticioglu IB (2022) General assessment of approaches to the identification of aquatic bacterial pathogens: a methodological review. North Am J Aqua 84(4):405–426. https://doi.org/10.1002/naaq.10260
Dumpala PR, Gülsoy N, Lawrence ML, Karsi A (2010) Proteomic analysis of the fish pathogen Flavobacterium columnare. Proteome Sci 8:1–1. http://www.proteomesci.com/content/8/1/26
Egger RC, Rosa JCC, Resende LFL, de Padu SB, de Oliveira-Barbosa F, Zerbini MT, Tavares GC, Figueiredo HCP (2023) Emerging fish pathogens Lactococcus petauri and L. garvieae in Nile tilapia (Oreochromis niloticus) farmed in Brazil. Aqua 565:739093. https://doi.org/10.1016/j.aquaculture.2022.739093
Eldar A, Ghittino C (1999) Lactococcus garvieae and Streptococcus iniae infections in rainbow trout (Oncorhynchus mykiss): similar but different diseases. Dis Aqua Org 36:227–231. https://doi.org/10.3354/dao036227
Elgendy MY, Ali SE, Abbas WT, Algammal AM, Abdelsalam M (2023) The role of marine pollution on the emergence of fish bacterial diseases. Chemosphere 6:140366. https://doi.org/10.1016/j.chemosphere.2023.140366
Elliott DG (2017) Renibacterium salmoninarum. In fish viruses and bacteria: pathobiology and protection. P. 286–297. Wallingford, UK. CABI Publishing. https://doi.org/10.1079/9781780647784.028
Erler R, Wichels A, Heinemeyer EA, Hauk G, Hippelein M, Reyes NT, Gerdts G (2015) VibrioBase: a MALDI-TOF MS database for fast identification of Vibrio spp. that are potentially pathogenic in humans. Syst Appl Micro 38(1):16–25. https://doi.org/10.1016/j.syapm.2014.10.009
Evelyn TPT, Prosperi-Porta L, Ketcheson JE (1986) Persistence of the kidney-disease bacterium, Renibacterium salmoninarum, in coho salmon, Oncorhynchus kisutch (Walbaum), eggs treated during and after water-hardening with providone-iodine. J Fish Dis 9:461–464. https://doi.org/10.1111/j.1365-2761.1986.tb01040.x
Facklam R, Elliott J, Shewmaker L, Reingold A (2005) Identification and characterization of sporadic isolates of Streptococcus iniae isolated from humans. J Clin Micro 43(2):933–937. https://doi.org/10.1128/jcm.43.2.933-937.2005
Fasulkova R, Orozova P, Stratev D (2023) Identification of Vibrio parahemolyticus isolated from seafood via matrix-assisted laser desorption/ıonization time of flight mass spectrometry. J Food Qual Haz Cont 10:135–141. https://doi.org/10.18502/jfqhc.10.3.13644
Fazio F (2019) Fish hematology analysis as an important tool of aquaculture: a review. Aqua 500:237–242. https://doi.org/10.1016/j.aquaculture.2018.10.030
Fernández-Álvarez C, Torres-Corral Y, Saltos-Rosero N, Santos Y (2017) MALDI-TOF mass spectrometry for rapid differentiation of Tenacibaculum species pathogenic for fish. Appl Micro Biotec 101:5377–5390. https://doi.org/10.1007/s00253-017-8324-3
Fernández-Álvarez C, Torres-Corral Y, Santos Y (2018) Use of ribosomal proteins as biomarkers for identification of Flavobacterium psychrophilum by MALDI-TOF mass spectrometry. J Proteomics 170:59–69. https://doi.org/10.1016/j.jprot.2017.09.007
Ferrario C, Ricci G, Milani C, Lugli GA, Ventura M, Eraclio G, Borgo F, Fortina MG (2013) Lactococcus garvieae: where is it from? A first approach to explore the evolutionary history of this emerging pathogen. PLoS One 8:e84796. https://doi.org/10.1371/journal.pone.0084796
Florio W, Baldeschi L, Rizzato C, Tavanti A, Ghelardi E, Lupetti A (2020) Detection of antibiotic resistance by MALDI-TOF mass spectrometry: an expanding area. Front Cell Infec Micro 10:572909. https://doi.org/10.3389/fcimb.2020.572909
Freitas J, Perestrelo R, Vaz-Pires P, Câmara JS (2022) Bacterial diversity analysis of coastal superficial seawaters near aquaculture facilities, using MALDI-TOF approach and Ribopeaks database. Aqua 556:738263. https://doi.org/10.1016/j.aquaculture.2022.738263
Fryer JL, Lannan CN (1993) The history and current status of Renibacterium salmoninarum, the causative agent of bacterial kidney disease in Pacific salmon. Fish Res 17:15–33. https://doi.org/10.1016/j.vetmic.2004.11.007
Fryer JL, Lannan C, Giovannoni SJ, Wood ND (1992) Piscirickettsia salmonis gen. nov.; sp. nov. the causative agent of an epizootic disease in salmonid fishes. Inter J Syst Bac 42:120–126. https://doi.org/10.1099/00207713-42-1-120
Gandhi K, Kumar A, Sarkar P, Aghav A, Lal D (2013) MALDI-TOF MS: applications in dairy and related sectors. Res Rev J Dairy Sci Tech 2:2319–3409
Gauthier DT (2015) Bacterial zoonoses of fishes: a review and appraisal of evidence for linkages between fish and human infections. Vet J 203:27–35. https://doi.org/10.1016/j.tvjl.2014.10.028
Gauthier DT, Rhodes MW (2009) Mycobacteriosis in fishes: a review. Vet J 180(1):33–47. https://doi.org/10.1016/j.tvjl.2008.05.012
Gibello A, Galán-Sánchez F, Blanco MM, Rodríguez-Iglesias M, Domínguez L, Fernández-Garayzábal JF (2016) The zoonotic potential of Lactococcus garvieae: an overview on microbiology, epidemiology, virulence factors and relationship with its presence in foods. Res Vet Sci 109:59–70. https://doi.org/10.1016/j.rvsc.2016.09.010
González SF, Krug MJ, Nielsen ME, Santos Y, Call DR (2004) Simultaneous detection of marine fishpathogens by using multiplex PCR and a DNA microarray. J Clin Micro 42:1414–1419. https://doi.org/10.1128/jcm.42.4.1414-1419.2004
González SF, Santos Y (2009) Serological methods for the detection of pathogenic bacteria in aquaculture: present status and prospects. Fisheries, Aquaculture and Biotechnology: Agrobios, India. p. 131–144.
Goodman LB, Lawton MR, Franklin-Guild RJ, Anderson RR, Schaan L, Thachil AJ, Wiedmann M, Miller CB, Alcaine SD, Kovac J (2017) Lactococcus petauri sp. nov. isolated from an abscess of a sugar glider. Int J Syst Evol Micro 67:4397–4404. https://doi.org/10.1099/ijsem.0.002303
Gutenberger SK, Duimstra JR, Rohovec JS, Fryer JL (1997) Intracellular survival of Renibacterium salmoninarum in trout mononuclear phagocytes. Dis Aquat Org 28:93–106. https://doi.org/10.3354/dao028093
Haider A, Ringer M, Kotroczó Z, Mohácsi-Farkas C, Kocsis T (2023) The importance of protein fingerprints in bacterial identification: the MALDI-TOF technique. J Environ Geo 16(1–4):38–45. https://doi.org/10.14232/jengeo-2023-44716
Hambuch TM, Mayfield J (2014) Next generation sequencing. 4131–4139. https://doi.org/10.1016/B978-0-12-386456-7.07717-0
Hassan AA, Khan IU, Abdulmawjood A, Lämmler C (2001) Evaluation of PCR methods for rapid identification and differentiation of Streptococcus uberis and Streptococcus parauberis. J Clin Micro 39:1618–1621. https://doi.org/10.1128/JCM.39.4.1618
Huang Y, Runge M, Michael GB, Schwarz S, Jung A, Steinhagen D (2013) Biochemical and molecular heterogeneity among isolates of Yersinia ruckeri from rainbow trout (Oncorhynchus mykiss, Walbaum) in northwest Germany. BMC Vet Res 9:215. https://doi.org/10.1186/1746-6148-9-215
Huang S, Xu Y, Liu X, Zhou M, Wu X, Jia Y (2016) Molecular newborn screening of four genetic diseases in Guizhou Province of South China. Gene 591(1):119–122. https://doi.org/10.1016/j.gene.2016.07.019
Jacobs JM, Stine CB, Baya AM, Kent ML (2009) A review of mycobacteriosis in marine fish. J Fish Dis 32(2):119–130. https://doi.org/10.1111/j.1365-2761.2008.01016.x
Jaies I, Shah FA, Qadiri SSN, Qayoom I, Bhat BA, Dar SA, Bhat FA (2024) Immunological and molecular diagnostic techniques in fish health: present and future prospectus. Mol Biol Rep 51(1):551. https://doi.org/10.1007/s11033-024-09344-5
Janampa-Sarmiento PC, Reis FY, Egger RC, de Pádua SB, Marcelino SA, Cunha JL, Pierezan F, Figueiredo HC, Tavares GC (2024) First report of Vibrio vulnificus outbreak in farm-raised sorubim (Pseudoplatystoma sp.) from Brazil. Fishes 9(2):54. https://doi.org/10.3390/fishes9020054
Janda JM, Abbott SL (2010) The genus Aeromonas: taxonomy, pathogenicity, and infection. Clin Micro Rev 23:5–73. https://doi.org/10.1128/CMR.00039-09
Jansson E, Haenen O, Nonnemann B, Madsen L, Van Gelderen E, Aspán A, Säker E, Gulla S, Colquhoun DJ, Roozenburg-Hengst I (2020) MALDI-TOF MS: a diagnostic tool for identification of bacterial fish pathogens. Bull Euro Associ Fish Patho 40(6):240–248. https://eafp.org/download/2020-volume40/issue_6/40-6-240-jansson.pdf.
Jia B, Burnley H, Gardner IA, Saab ME, Doucet A, Hammell KL (2023) Diagnosis of Renibacterium salmoninarum infection in harvested Atlantic salmon (Salmo salar L.) on the east coast of Canada: Clinical findings, sample collection methods and laboratory diagnostic tests. J Fish Dis 46(5):575–589. https://doi.org/10.1111/jfd.13770
Jung MY, Chang YH, Kim W (2010) A real-time PCR assay for detection and quantification of Lactococcus garvieae. J Appl Micro 108:1694–1701. https://doi.org/10.1111/j.1365-2672.2009.04568.x
Jung-Schroers V, Jung A, Ryll M, Bauer J, Teitge F, Steinhagen D (2019) Diagnostic methods for identifying different Aeromonas species and examining their pathogenicity factors, their correlation to cytotoxicity and adherence to fish mucus. J Fish Dis 42(2):189–219. https://doi.org/10.1111/jfd.12917
Kaattari IM, Rhodees MW, Kator H, Kaattari SL (2005) Comparative analysis of mycobacterial infections in wild striped basss Morone saxatilis from Chesapeake Bay. Dis Aquat Organ 67:125–132. https://doi.org/10.3354/dao067125
Kačániová M, Klūga A, Kántor A, Medo J, Žiarovská J, Puchalski C, Terentjeva M (2019) Comparison of MALDI-TOF MS Biotyper and 16S rDNA sequencing for the identification of Pseudomonas species isolated from fish. Micro Patho 132:313–318. https://doi.org/10.1016/j.micpath.2019.04.024
Kafka AP, Kleffmann T, Rades T, McDowell A (2011) The application of MALDI TOF MS in biopharmaceutical research. Inter J Pharm 417(1–2):70–82. https://doi.org/10.1016/j.ijpharm.2010.12.010
Kalimuddin S, Chen SL, Lim CT, Koh TH, Tan TY, Kam M, Wong CW, Mehershahi KS, Chau ML, Ng LC (2017) 2015 epidemic of severe Streptococcus agalactiae sequence type 283 infections in Singapore associated with the consumption of raw freshwater fish: a detailed analysis of clinical, epidemiological and bacterial sequencing data. Clin Infec Dis 64(2):145–152. https://doi.org/10.1093/cid/cix021
Karas M, Hillenkamp F (1988) Laser desorption ionization of proteins with molecular masses exceeding 10000 Daltons. Anal Chem 60:2299–2301
Karas M, Bachmann D, Hillenkamp F (1985) Influence of the wavelength in high-irradiance ultraviolet laser desorption mass spectrometry of organic molecules. Anal Chem 57(14):2935–2939
Karas M, Bachman D, Bahr U, Hillenkamp F (1987) Matrix-assisted ultraviolet laser desorption of non volatile compounds. Int J Mass Spect Ion Proc 78:53–68. https://doi.org/10.1016/0168-1176(87)87041-6
Kayis S, Er A, Yılmaz C, Düzgün A, Köse Ö, Kurtoğlu IZ (2015) Aeromonas hydrophila as a causative agent of blue sac fry syndrome in different trout species. J Fish Dis 38(12):1069–1071. https://doi.org/10.1111/jfd.12326
Kazarnikova A, Trishina A, Galeotti M, Manzano M, Abrosimova N, Ermakov A (2019) Detecting Yersinia ruckeri on the death of carp (Cyprinus carpio) farmed in southern Russia. In IOP Conference Series: Earth Environ Sci. 403(1):012039. IOP Publishing. https://doi.org/10.1088/1755-1315/403/1/012039
Kazazić SP, Popović NT, Strunjak-Perović I, Babić S, Florio D, Fioravanti M, Bojanić K, Čož-Rakovac R (2019a) Matrix-assisted laser desorption/ionization time of flight mass spectrometry identification of Vibrio (Listonella) anguillarum isolated from sea bass and sea bream. PLoS One 14(11):e0225343. https://doi.org/10.1371/journal.pone.0225343
Kazazić Pečur S, Popović NT, Strunjak-Perović I, Florio D, Fioravanti M, Babić S, Čož-Rakovac R (2019b) Fish photobacteriosis-the importance of rapid and accurate identification of Photobacterium damselae subsp. piscicida. J Fish Dis 42(8):1201–1209. https://doi.org/10.1111/jfd.13022
Khan NH, Ishii Y, Kimata-Kino N, Esaki H, Nishino T, Nishimura M, Kogure K (2007) Isolation of Pseudomonas aeruginosa from open ocean andcomparison with freshwater, clinical, and animal isolates. Micro Ecol 53:173–186. https://doi.org/10.1007/s00248-006-9059-3
Khandeparker L, Gardade L, Anil AC (2024) Influence of environmental settings in geographically distinct ports on the protein profiles of cultivable bacteria using MALDI–TOF mass spectrometry. Mar Eco 45(1):e12777. https://doi.org/10.1111/maec.12777
Kim SW, Jang HB, Lee JS, Im SP, Lazarte JMS, Seo JP, Lee WJ, Kim JS, Jung TS (2015) Comparison of proteome typing and serotyping of Streptococcus parauberis isolates from olive flounder (Paralichthys olivaceus). J Micro Met 118:168–172. https://doi.org/10.1016/j.mimet.2015.09.015
Kim A, Nguyen TL, Kim DH (2017) Modern Methods of Diagnosis. B. Austin, A. Newaj-Fyzul, Editors. Diagnosis and control of diseases of fish and shellfish. Published by Wiley and Son Inc., Chichester, UK. p.109–137. https://doi.org/10.1002/9781119152125.ch5
Klūga A, Kačániová M, Terentjeva M (2019) Identification and antibiotic susceptibility of bacterial microbiota of freshwater fish. Potravinarstvo 13(1):208–414. https://doi.org/10.5219/1063
Koh HL, Yau WP, Ong PS, Hegde A (2003) Current trends in modern pharmaceutical analysis for drug discovery. Drug Disco Today 8(19):889–897. https://doi.org/10.1016/S1359-6446(03)02846-0
Kondo Y, Ohara K, Fujii R, Nakai Y, Sato C, Naito M, Tsukuba T, Kadowaki T, Sato K (2023) Transposon mutagenesis and genome sequencing identify two novel, tandem genes involved in the colony spreading of Flavobacterium collinsii, isolated from an ayu fish. Plecoglossus Altivelis Front Cell Infec Micro 13:60. https://doi.org/10.3389/fcimb.2023.1095919
Kotzamanidis C, Malousi A, Bitchava K, Vafeas G, Chatzidimitriou D, Skoura L, Papadimitriou E, Chatzopoulou F, Zdragas A (2020) First report of isolation and genome sequence of L. petauri strain from a rainbow trout lactococcosis outbreak. Curr Micro 77:1089–1096. https://doi.org/10.1007/s00284-020-01905-8
Kumar G, Kocour M (2017) Applications of next-generation sequencing in fisheries research: a review. Fisheries Res 186:11–22. https://doi.org/10.1016/j.fishres.2016.07.021
Kurokawa S, Kabayama J, Fukuyasu T, Hwang SD, Park CI, Park SB, del Castillo CS, Hikima JI, Jung TS, Kondo H (2013) Bacterial classification of fish-pathogenic Mycobacterium species by multigene phylogenetic analyses and MALDI Biotyper identification system. Mar Biotec 15:340–348. https://doi.org/10.1007/s10126-012-9492-x
LaFrentz BR, Králová S, Burbick CR, Alexander TL, Phillips CW, Griffin MJ, Waldbieser GC (2022) The fish pathogen Flavobacterium columnare represents four distinct species: F. columnare, F. covae sp. nov., F. davisii sp. nov. and F. oreochromis sp. nov., and emended description of F. columnare. Syst Appl Micro 45(2):126293. https://doi.org/10.1016/j.syapm.2021.126293
Lahav D, Eyngor M, Hurvitz A, Ghittino C, Lublin A, Eldar A (2004) Streptococcus iniae type II infections in rainbow trout Oncorhynchus mykiss. Dis Aqua Org 62(1):177–180. https://doi.org/10.3354/dao062177
Lannan CN, Fryer JL (1993) Piscirickettsia salmonis, a major pathogen of salmond fish in Chile. Fish Res 17:115–121. https://doi.org/10.1016/0165-7836(93)90011-U
Lansdell W, Dixon B, Smith N, Benjamin L (1993) Communications: isolation of several mycobacterium species from fish. J Aqua Anim Health 5(1):73–76. https://doi.org/10.1577/1548-8667(1993)
Lauková A, Kubašová I, Kandričáková A, Strompfová V, Žitňan R, Simonová MP (2018) Relation to enterocins of variable Aeromonas species isolated from trouts of Slovakian aquatic sources and detected by MALDI-TOF mass spectrometry. Folia Micro 63:749–755. https://doi.org/10.1007/s12223-018-0616-1
Le Breton A (2020) 11. Tenacibaculum group. In: Zrncic S. (ed.). Diagnostic manual for the main pathogens in European seabass and Gilthead seabream aquaculture. Zaragoza: CIHEAM, Options Méditerranéennes: Série B. Etudes et Recherches. 75:p. 97–106.
Lefterova MI, Suarez CJ, Banaei N, Pinsky BA (2015) Next-generation sequencing for infectious disease diagnosis and management: a report of the association for molecular pathology. J Mol Diagnos 17(6):623–634. https://doi.org/10.1016/j.jmoldx.2015.07.004
Li N, Qin T, Zhang XL, Huang B, Liu ZX, Xie HX, Zhang J, McBride MJ, Nie P (2015) Development and use of a gene deletion strategy to examine the two chondroitin lyases in virulence of Flavobacterium columnare. Appl Environ Micro 81:7394–7402. https://doi.org/10.1128/AEM.01586-15
Li D, Yi J, Han G, Qiao L (2022) MALDI-TOF mass spectrometry in clinical analysis and research. ACS Measure Sci Au 2(5):385–404. https://doi.org/10.1021/acsmeasuresciau.2c00019
Littman EM, Heckman TI, Yazdi Z, Veek T, Mukkatira K, Adkison M, Powell A, Camus A, Soto E (2023) Temperature-associated virulence, species susceptibility and interspecies transmission of a Lactococcus petauri strain from rainbow trout. Dis of Aqua Org 155:147–158. https://doi.org/10.3354/dao03747
Liu CH, Cheng W, Hsu JP, Chen JC (2004) Vibrio alginolyticus infection in the white shrimp Litopenaeus vannamei confirmed by polymerase chain reaction and 16S rDNA sequencing. Dis Aqua Org 61(1–2):169–174. https://doi.org/10.3354/dao061169
Liu T, Kang L, Xu J, Wang J, Gao S, Li Y, Li J, Yuan Y, Yuan B, Wang J, Zhao B (2022) PVBase: a MALDI-TOF MS database for fast identification and characterization of potentially pathogenic Vibrio species from multiple regions of China. Fron Micro 13:872825. https://doi.org/10.3389/fmicb.2022.872825
Loch TP, Faisal M (2015) Emerging flavobacterial infections in fish: a review. J Advan Res 6(3):283–300. https://doi.org/10.1016/j.jare.2014.10.009
López JR, Dieguez AL, Doce A, de la Roca E, de la Herran R, Navas JI, Toranzo AE, Romalde JL (2012) Pseudomonas baetica sp. nov., a fish pathogen isolated from wedge sole, Dicologlossa cuneata (Moreau). Int J Syst Evol Micro 62:874–882. https://doi.org/10.1099/ijs.0.030601-0
López-Cortés XA, Nachtigall FM, Olate VR, Araya M, Oyanedel S, Diaz V, Jakob E, Ríos-Momberg M, Santos LS (2017) Fast detection of pathogens in salmon farming industry. Aqua 470:17–24. https://doi.org/10.1016/j.aquaculture.2016.12.008
Low CF, Shamsudin MN, Chee HY, Aliyu-Paiko M, Idrus ES (2014) Putative apolipoprotein A-I, natural killer cell enhancement factor and lysozyme g are involved in the early immune response of brown marbled grouper, Epinephelus fuscoguttatus, Forskal, to Vibrio alginolyticus. J Fish Dis 37(8):693–701. https://doi.org/10.1111/jfd.12153
Mabrok M, Algammal AM, Sivaramasamy E, Hetta HF, Atwah B, Alghamdi S, Fawzy A, Avendaño-Herrera R, Rodkhum C (2023) Tenacibaculosis caused by Tenacibaculum maritimum: updated knowledge of this marine bacterial fish pathogen. Front Cell Infec Micro 12:1068000. https://doi.org/10.3389/fcmb.2022.1068000
Makrinos DL, Bowden TJ (2017) Growth characteristics of the intracellular pathogen, Piscirickettsia salmonis, in tissue culture and cell-free media. J Fish Dis 40(8):1115–1127. https://doi.org/10.1111/jfd.12578
Malainine SM, Moussaoui W, Prévost G, Scheftel JM, Mimouni R (2013) Rapid identification of Vibrio parahaemolyticus isolated from shellfish, sea water and sediments of the Khnifiss lagoon, Morocco, by MALDI-TOF mass spectrometry. Lett Appl Micro 56(5):379–386. https://doi.org/10.1111/lam.12060
Mata AI, Gibello A, Casamayor A, Blanco MM, Domínguez L, Fernández-Garayzábal JF (2004) Multiplex PCR assay for detection of bacterial pathogens associated with warm-water streptococcosis in fish. Appl Environ Micro 70(5):3183–3187. https://doi.org/10.1128/AEM.70.5.3183-3187.2004
Mellmann A, Cloud J, Maier T, Keckevoet U, Ramminger I, Iwen P, Harmsen D (2008) Evaluation of matrix-assisted laser desorption ionization-time-of-flight mass spectrometry in comparison to 16S rRNA gene sequencing for species identification of nonfermenting bacteria. J Clin Micro 46(6):1946–1954. https://doi.org/10.1128/jcm.00157-08
Meyburgh CM, Bragg RR, Boucher CE (2017) Lactococcus garvieae: an emerging bacterial pathogen of fish. Dis Aqua Org 123(1):67–79. https://doi.org/10.3354/dao03083
Michel C, Nougayrede P, Eldar A, Sochon E, De Kinkelin P (1997) Vagococcus salmoninarum, a bacterium of pathological significance in rainbow trout Oncorhynchus mykiss farming. Dis Aquatic Org 30(3):199–208. https://doi.org/10.3354/dao030199
Mishra SS, Das R, Sahoo SN, Swain P (2020) Biotechnological tools in diagnosis and control of emerging fish and shellfish diseases. In Genomics and Biotechnological Advances in Veterinary, Poultry, and Fisheries. Academic Press. p. 311–360. https://doi.org/10.1016/B978-0-12-816352-8.00014-X
Moreira M, Schrama D, Farinha AP, Cerqueira M, Raposo de Magalhaes C, Carrilho R, Rodrigues P (2021) Fish pathology research and diagnosis in aquaculture of farmed fish; a proteomics perspective. Animals 11(1):125. https://doi.org/10.3390/ani11010125
Mougin J, Flahaut C, Roquigny R, Bonnin-Jusserand M, Grard T, Le Bris C (2020) Rapid identification of Vibrio species of the harveyi clade using MALDI-TOF MS profiling with main spectral profile database implemented with an in-house database: luvibase. Front Micro 11:586536. https://doi.org/10.3389/fmicb.2020.586536
Mougin J, Roquigny R, Flahaut C, Bonnin-Jusserand M, Grard T, Le Bris C (2021) Abundance and spatial patterns over time of Vibrionaceae and Vibrio harveyi in water and biofilm from a seabass aquaculture facility. Aqua 542:736862. https://doi.org/10.1016/j.aquaculture.2021.736862
Moussa M, Cauvin E, Le Piouffle A, Lucas O, Bidault A, Paillard C, Garcia C (2021) A MALDI-TOF MS database for fast identification of Vibrio spp. potentially pathogenic to marine mollusks. Appl Micro Biotech 105:2527–2539. https://doi.org/10.1016/j.aquaculture.2021.73686210.1007/s00253-021-11141-0
Mugetti D, Varello K, Pastorino P, Tomasoni M, Menconi V, Bozzetta E, Dondo A, Prearo M (2021) Investigation of potential reservoirs of non-tuberculous mycobacteria in a European sea bass (Dicentrarchus labrax) farm. Pathogens 10(8):1014. https://doi.org/10.3390/pathogens10081014
Natnan ME, Mayalvanan Y, Jazamuddin FM, Aizat WM, Low CF, Goh HH, Azizan KA, Bunawan H, Baharum SN (2021) Omics strategies in current advancements of infectious fish disease management. Biology 10(11):1086. https://doi.org/10.3390/biology10111086
Neidorf A, Morozova M (2021) Diagnosis and treatment of flexibacteriosis of koi carp (Cyprinus carpio koi). In IOP Conf Ser Earth Envir Sci 937(3):032040. https://doi.org/10.1088/17551315/937/3/032040
Nho SW, Shin GW, Park SB, Jang HB, Cha IS, Ha MA, Kim YR, Park YK, Dalvi RS, Kang BJ (2009) Phenotypic characteristics of Streptococcus iniae and Streptococcus parauberis isolated from olive flounder (Paralichthys olivaceus). FEMS Micro Lett 293:20–27. https://doi.org/10.1111/j.1574-6968.2009.01491.x
Nishimori E, Kita-Tsukamoto K, Wakabayashi H (2000) Pseudomonas plecoglossicida sp. nov., the causative agent of bacterial haemorrhagic ascites of ayu, Plecoglossus altivelis. Int J Syst Evol Micro 50:83–89. https://doi.org/10.1099/00207713-50-1-83
Nissa MU, Pinto N, Parkar H, Goswami M, Srivastava S (2021) Proteomics in fisheries and aquaculture: an approach for food security. Food Cont 127:108125. https://doi.org/10.1016/j.foodcont.2021.108125
Noga EJ (2010) Fish disease: diagnosis and treatment, 2nd edn. Published by John Wiley and Sons Ltd., Hoboken, NJ, USA
Oh WT, Jun JW, Giri SS, Yun S, Kim HJ, Kim SG, Kim SW, Kang JW, Han SJ, Kwon J (2019) Pseudomonas tructae sp. Nov., novel species isolated from rainbow trout kidney. Int J Syst Evol Micro 69:3851–3856. https://doi.org/10.1099/ijsem.0.003696
Ojasanya RA, Gardner IA, Groman D, Saksida S, Saab ME, Thakur KK (2022) Development and validation of main spectral profile for rapid identification of Yersinia ruckeri isolated from Atlantic salmon using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Front Vet Sci 9:1031373. https://doi.org/10.3389/fvets.2022.1031373
Olate VR, Nachtigall FM, Santos LS, Soto A, Araya M, Oyanedel S, Díaz V, Marchant V, Rios-Momberg M (2016) Fast detection of Piscirickettsia salmonis in Salmo salar serum through MALDI-TOF-MS profiling. J Mass Spect 51(3):200–206. https://doi.org/10.1002/jms.3734
Onuk EE, Ciftci A, Findik A, Durmaz Y (2010) Development and evaluation of a multiplex PCR assay for simultaneous detection of F. psychrophilum, Y. ruckeri and A. salmonicida in culture fisheries. J Vet Sci 11(3):235–241. https://doi.org/10.4142/jvs.2010.11.3.235
Ozbey G, Tanriverdi ES, Basusta A, Lakshmanappa YS, Otlu B, Zigo F (2023) Investigation for the presence of bacteria and antimicrobial resistance genes in sea snails (Rapana venosa). Ann Agric Environ Med 30(2):235–243. https://doi.org/10.26444/aaem/163582
Özcan F (2022) Investigation of diseases caused by Aeromonas media in rainbow trout (Oncorhynchus mykiss) in commercial fish farms using MALDI-TOF and specification of antibiotic sensitivity profiles of the agent. Rev Cient e32191:1–5. https://doi.org/10.52973/rcfcv-e32191
Özcan F (2023) Observation of inherently contaminated Oncorhynchus mykiss Walbaum, 1792 by Aeromonas veronii with MALDI-TOF and culture methods and specification of antibiotic sensitivity profiles of agent in a commercial farms. Indian J Ani Res 1:4. https://doi.org/10.18805/IJAR.BF-1444
Paillard C, Gausson S, Nicolas JL, Le Pennec JP, Haras D (2006) Molecular identification of Vibrio tapetis, the causative agent of the brown ring disease of Ruditapes philippinarum. Aqua 253(1–4):25–38. https://doi.org/10.1016/j.aquaculture.2005.03.047
Park SB, Aoki T, Jung TS (2012) Pathogenesis of and strategies for preventing Edwardsiella tarda infection in fish. Vet Res 43:1–17. http://www.veterinaryresearch.org/content/43/1/67
Parker-Graham CA, Heckman TI, Griffin MJ, Soto E (2023) Infectious diseases of warm water fish in marine and brackish waters. In Climate Change on Diseases and Disorders of Finfish in Cage Culture. 3rd Ed. Eds by Woo PTK and Subasinghe RP. CABI Publishing. USA p. 163–201. https://doi.org/10.1079/9781800621640.000
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of prokaryptic names with standing in nomenclature moves to the DSMA. Inter J Syst Evo Micro 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Pastuszka A, Guz L, Michalak K, Pietras-Ożga D, Puk K (2024) Vibrio infection in freshwater fish in Poland. Polish J Vet Sci 117–125. https://doi.org/10.24425/pjvs.2024.149341
Patel R (2015) MALDI-TOF MS for the diagnosis of infectious diseases. Clin Chem 61(1):100–111. https://doi.org/10.1373/clinchem.2014.221770
Pekala-Safińska A (2018) Contemporary threats of bacterial infections in freshwater fish. J Vet Res 62(3):261. https://doi.org/10.2478/jvetres-2018-0037
Pérez-Sancho M, Ana IV, Awad M, Kostrzewa M, Domínguez L, Fernández-Garayzábal JF (2016) Differentiation of Photobacterium damselae subspecies using matrix-assisted laser-desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) in fish isolates. Aqua 464:159–164. https://doi.org/10.1016/j.aquaculture.2016.06.024
Pérez-Sancho M, Vela AI, Kostrzewa M, Zamora L, Casamayor A, Dominguez L, Fernández-Garayzábal JF (2017) First analysis by MALDI-TOF MS technique of Chryseobacterium species relevant to aquaculture. J Fish Dis 41(2):389–393. https://doi.org/10.1111/jfd.12706
Pérez-Sancho M, Vela AI, Wiklund T, Kostrzewa M, Domínguez L, Fernández-Garayzábal JF (2017) Differentiation of Flavobacterium psychrophilum from F. psychrophilum-like species by MALDI-TOF mass spectrometry. Res Vet Sci 115:345–352. https://doi.org/10.1016/j.rvsc.2017.06.022
Pérez-Sancho M, Cerdá I, Fernández-Bravo A, Domínguez L, Figueras MJ, Fernández-Garayzábal JF, Vela AI (2018) Limited performance of MALDI-TOF for identification of fish Aeromonas isolates at species level. J Fish Dis 41(10):1485–1493. https://doi.org/10.1111/jfd.12837
Piamsomboon P, Jaresitthikunchai J, Hung TQ, Roytrakul S, Wongtavatchai J (2020) Identification of bacterial pathogens in cultured fish with a custom peptide database constructed by matrix-assisted laser desorption/ ionization time-of-flight mass spectrometry (MALDI-TOF). BMC Vet Res 16:1–10. https://doi.org/10.1186/s12917-020-2274-1
Pierezan F, Shahin K, Heckman TI, Ang J, Byrne BA, Soto E (2020) Outbreaks of severe myositis in cultured white sturgeon (Acipenser transmontanus L.) associated with Streptococcus iniae. J Fish Dis 43(4):485–490. https://doi.org/10.1111/jfd.13145
Piñeiro-Vidal M, Centeno-Sestelo G, Riaza A, Santos Y (2007) Isolationof pathogenic Tenacibaculum maritimum-related organisms from diseased turbot and sole cultured in the Northwest of Spain. Bull Eur Assoc Fish Pathol 27(1):29
Plumb JA (1993) Edwardsiella septicaemia. In Bacterial Diseases of Fish ed. Inglis V, Roberts RJ, Bromage NR. p. 61–79. Cambridge, UK: Cambridge University Press. https://doi.org/10.1079/9781845935542.05
Popović NT, Kazazić SP, Strunjak-Perović I, Čož-Rakovac R (2017) Differentiation of environmental aquatic bacterial isolates by MALDI-TOF. Environ Res 152:7–16. https://doi.org/10.1016/j.envres.2016.09.020
Popović NT, Kazazić SP, Bojanić K, Strunjak-Perović I, Čož-Rakovac R (2021) Sample preparation and culture condition effects on MALDI-TOF MS identification of bacteria: a review. Mass Spect Rev 42(5):1589–1603. https://doi.org/10.1016/j.envres.2016.09.020
Popović NT, Kepec S, Kazazić SP, Strunjak-Perović I, Bojanić K, Čož-Rakovac R (2022) Identification of environmental aquatic bacteria by mass spectrometry supported by biochemical differentiation. PLoS One 17(6):e0269423. https://doi.org/10.1371/journal.pone.0269423
Popović NT, Čož-Rakovac R, Strunjak-Perović I (2007) Commercial phenotypic tests (API 20E) in diagnosis of fish bacteria. Vet Med 52(2):49–53. http://vri.cz/docs/vetmed/52-2-49.pdf
Puk K, Banach T, Wawrzyniak A, Adaszek Ł, Zietek J, Winiarczyk S, Guz L (2018) Detection of Mycobacterium marinum, M. peregrinum, M. fortuitum and M. abscessus in aquarium fish. J Fish Dis 41:153–156. https://doi.org/10.1111/jfd.12666
Radosavljević V, Radanović O, Zdravković N, Savić B, Stanković M, Zorić JM, Veljović L, Nešić K (2020) The first outbreak of lactococcosis caused by Lactococcus garvieae in Serbia. Arch Vet Med 13(1):53–68. https://doi.org/10.46784/e-avm.v13i1.78
Rahmani A, Vercauteren M, Vranckx K, Boyen F, Bidault A, Pichereau V, Decostere A, Paillard C, Chiers K (2021) MALDI-TOF MS as a promising tool to assess potential virulence of Vibrio tapetis isolates. Aqua 530:735729. https://doi.org/10.1016/j.aquaculture.2020.735729
Ravelo C, Magarinos B, Romalde JL, Toranzo AE (2001) Conventional versus miniaturized systems for the phenotypic characterization of Lactococcus garvieae strains. Bull Eur Assoc Fish Pathol 21:136–144
Reis FYT, Rocha VP, Janampa-Sarmiento PC, Costa HL, Egger RC, Passos NC, de Assis CHS, Carneiro SP, Santos ÁF, Silva BA, Dorella FA (2023) Edwardsiella tarda in Tambaqui (Colossoma macropomum): a pathogenicity, antimicrobial susceptibility and genetic analysis of Brazilian isolates. Animals 13(18):2910. https://doi.org/10.3390/ani13182910
Rhodes LD, Coady AM, Deinhard RK (2004) Identification of a msa gene in R. salmoninarum the associated virulence phenotype. Appl Environ Mic 70:6488–6494. https://doi.org/10.1128/AEM.70.11.6488-6494.2004
Royo-Cebrecos C, Gudiol C, Ardanuy C, Pomares H, Calvo M, Carratalà J (2017) A fresh look at polymicrobial bloodstream infection in cancer patients. PLoS One 12(10):e0185768. https://doi.org/10.1371/journal.pone.0185768
Rozas M, Enríquez R (2014) Piscirickettsiosis and Piscirickettsia salmonis in fish: a review. J Fish Dis 37(3):163–188. https://doi.org/10.1111/jfd.12211
Ruiz-Zarzuela I, de Bias I, Gironés O, Ghittino C, Múazquiz JL (2005) Isolation of Vagococcus salmoninarum in rainbow trout, Oncorhynchus mykiss (Walbaum), broodstocks: characterization of the pathogen. Vet Res Comm 29:553–562. https://doi.org/10.1007/s11259-005-2493-8
Rupp M, Knüsel R, Sindilariu PD, Schmidt-Posthaus H (2019) Identification of important pathogens in European perch (Perca fluviatilis) culture in recirculating aquaculture systems. Aqua Inter 27:1045–1053. https://doi.org/10.1007/s10499-019-00382-6
Sadikot RT, Blackwell TS, Christman JW, Prince AS (2005) Pathogen-host interactions in Pseudomonas aeruginosa pneumonia. American J Resp Criti Care Medic 11:1209–1223. https://doi.org/10.1164/rccm.200408-1044SO
Salogni C, Perantoni P, Pitozzi A, Loris G, Alborali GL (2007) Vagococcus salmoninarum: descrizione di un focolaio di malattia in riproduttori di trota iridea (Oncorhynchus mykiss). Ittiopatologia 4:59–66
Sanches-Fernandes GM, Sá-Correia I, Costa R (2022) Vibriosis outbreaks in aquaculture: addressing environmental and public health concerns and preventive therapies using gilthead seabream farming as a model system. Front Micro 13:904815. https://doi.org/10.3389/fmicb.2022.904815
Sandalakis V, Goniotakis I, Vranakis I, Chochlakis D, Psaroulaki A (2017) Use of MALDI-TOF mass spectrometry in the battle against bacterial infectious diseases: recent achievements and future perspectives. Exp Rev Proteomics 14(3):253–267. https://doi.org/10.1080/14789450.2017.1282825
Sanjuán E, Amaro C (2007) Multiplex PCR assay for detection of Vibrio vulnificus biotype 2 and simultaneous discrimination of serovar E strains. Appl Environ Micro 73(6):2029–2032. https://doi.org/10.1128/AEM.02320-06
Santos Y, Romalde JL, Bandín I, Magariños B, Núñez S, Barja JL, Toranzo AE (1993) Usefulness of the API-20E system for the identification of bacterial fish pathogens. Aqua 116:111–120. https://doi.org/10.1016/0044-8486(93)90002-G
Saticioglu IB, Ay H, Altun S, Sahin N, Duman M (2021) Flavobacterium bernardetii sp. nov., a possible emerging pathogen of farmed rainbow trout (Oncorhynchus mykiss) in cold water. Aqua 540:736717. https://doi.org/10.1016/j.aquaculture.2021.736717
Saticioglu IB, Ay H, Altun S, Duman M, Sahin N (2021) Flavobacterium turcicum sp. nov. and Flavobacterium kayseriense sp. nov. isolated from farmed rainbow trout in Turkey. Syst Appl Micro 44(2):126186. https://doi.org/10.1016/j.syapm.2021.126186
Saticioglu IB, Mulet M, Duman M, Altun S, Gomila M, Lalucat J, García-Valdés E (2022) First occurrence and whole-genome comparison of Pseudomonas haemolytica isolated in farmed rainbow trout. Aqua Res 53(12):4472–4486. https://doi.org/10.1111/are.15944
Saticioglu IB, Onuk EE, Ay H, Ajmi N, Demirbas E, Altun S (2023) Phenotypic and molecular differentiation of Lactococcus garvieae and Lactococcus petauri isolated from trout. Aqua 577:739933. https://doi.org/10.1016/j.aquaculture.2023.739933
Schmidtke LM, Carson J (1994) Chararacteristics of Vagococcus salmoninarum isolated from diseased salmonid fish. J Appl Bacterio 77(2):229–236. https://doi.org/10.1111/j.1365-2672.1994.tb03068.x
Seng P, Drancourt M, Gouriet F, La Scola B, Fournier PE, Rolain JM, Raoult D (2009) Ongoing revolution in bacteriology: routine identification of bacteria by matrix-assisted laser desorption ionization time of flight mass spectrometry. Clin Infec Dis 49(4):543–551. https://doi.org/10.1086/600885
Seng P, Rolain JM, Fournier PE, La Scola B, Drancourt M, Raoult D (2010) MALDI-TOF-mass spectrometry applications in clinical microbiology. Future Micro 5(11):1733–1754. https://doi.org/10.2217/fmb.10.127
Sequeiros C, Garcés ME, Vallejo M, Marguet ER, Olivera NL (2015) Potential aquaculture probiont Lactococcus lactis TW34 produces nisin Z and inhibits the fish pathogen Lactococcus garvieae. Arch Micro 197:449–458. https://doi.org/10.1007/s00203-014-1076-x
Silva-Rubio A, Avendaño-Herrera R, Jaureguiberry B, Toranzo AE, Magariños B (2008) First description of serotype O3 in Vibrio anguillarum strains isolated from salmonids in Chile. J Fish Dis 31(3):235
Singhal N, Kumar M, Kanaujia PK, Virdi JS (2015) MALDI-TOF mass spectrometry: an emerging technology for microbial identification and diagnosis. Front Mic 6:791. https://doi.org/10.3389/fmicb.2015.00791
Sivanesan I, Gopal J, Hasan N, Muthu M (2023) A systematic assessment of matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF MS) application for rapid identification of pathogenic microbes that affect food crops: delivered and future deliverables. RSC Adv 13(25):17297–17314. https://doi.org/10.1039/D3RA01633A
Smith SA (2019) Fish diseases and medicine. In Smith SA. (Ed.). Taylor and Frances Group. CRC Press.USA. p.46–85.
Stevenson LG, Drake SK, Murray PR (2017) Rapid identification of bacteria in positive blood culture broths by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J Clin Micro 55(3):754–760. https://doi.org/10.1128/jcm.01541-09
Stickney RR (2009) Diseases of Aquaculture Species. In Stickney RR. (Ed.), Aquaculture: an ıntroductory text. 2nd Ed. CABI Publishing. Wallingford, USA. p. 148–173. https://doi.org/10.1079/9781845935894.0148
Suárez S, Ferroni A, Lotz A, Jolley KA, Guérin P, Leto J, Guiso N (2013) Ribosomal proteins as biomarkers for bacterial identification by mass spectrometry in the clinical microbiology laboratory. J Micro Meth 94(3):390–396. https://doi.org/10.1016/j.mimet.2013.07.021
Surányi BB, Taczman-Brückner A, Mohácsi-Farkas C, Engelhardt T (2023) Rapid identification of bacteria from agricultural environment using MALDI-TOF MS. Acta Aliment 52(1):113–120. https://doi.org/10.1556/066.2022.00202
Tamura H (2023) A MALDI‐TOF MS proteotyping approach for environmental, agricultural and food microbiology. microbiological identification using MALDI‐TOF and tandem mass spectrometry: industrial and environmental applications. Ed. by Shah HN, Gharbia SE, Shah AJ, Tranfield EY, Thompson, KC. Published by Wiley and Sons Inc. p.147–182. https://doi.org/10.1002/9781119814085.ch6
Tanaka K, Waki H, Ido Y, Akita S, Yoshida Y, Yoshida T, Matsuo T (1988) Protein and polymer analyses up to 100 000 by laser ionization time off light mass spectrometry. Rap Com Mass Spectrom 2(8):151–153. https://doi.org/10.1002/rcm.1290020802
Tanaka K (2003) The origin of macromolecule ionization by laser irradiation [Lecture]. In: Frangsmyr T, ed. The Nobel Prizes 2002. Stockholm, Sweden: Nobel Foundation. https://doi.org/10.1002/anie.200300585
Tanrikul TT (2007) Vibriosis as an epizootic disease of rainbow trout (Onchorynchus mykiss) in Turkey. Pakistan J Bio Sci 10(10):1733–1737. https://doi.org/10.3923/pjbs.2007.1733.1737
Tavares GC, de Queiroz GA, Assis GB, Leibowitz MP, Teixeira JP, Figueiredo HC, Leal CA (2018) Disease outbreaks in farmed Amazon catfish (Leiarius marmoratus x Pseudoplatystoma corruscans) caused by Streptococcus agalactiae, S. iniae, and S. dysgalactiae. Aqua 495:384–392. https://doi.org/10.1016/j.aquaculture.2018.06.027
Taylor PW, Crawford T, Shotts EB (1995) Comparison of two biochemical test systems with conventional methods for the identification of bacteria pathogenic to warmwater fish. J Aqua Anim Health 7:312–317. https://doi.org/10.1577/1548-8667(1995)007%3c0312:COTBTS%3e2.3.CO;2
Thompson JE (2022) Matrix-assisted laser desorption ionization-time-of-flight mass spectrometry in veterinary medicine: recent advances (2019-present). Vet World 15(11):2623. https://doi.org/10.14202/vetworld.2022.2623-2657
Thune RL, Stanley LA, Cooper RK (1993) Pathogenesis of gram negative bacterial infections in warm water fish. Annu Rev Fish Dis 3:37–68. https://doi.org/10.1016/0959-8030(93)90028-A
Timur G, Karataş S, Akayli T, Ercan MD, Yardimci RE (2009) A histopathological study of Hexamitiasis in farmed rainbow trout (Oncorhynchus mykiss) fry in Turkey. Bull Eur Ass Fish Pathol 29(3):104
Tobback E, Decostere A, Hermans K, Haesebrouck F, Chiers K (2007) Yersinia ruckeri infections in salmonid fish. J Fish Dis 30(5):257–268. https://doi.org/10.1111/j.1365-2761.2007.00816.x
Tohya M, Teramoto K, Watanabe S, Hishinuma T, Shimojima M, Ogawa M, Tada T, Tabe Y, Kirikae T (2022) Whole-genome sequencing-based re-identification of Pseudomonas putida/fluorescens clinical isolates identified by biochemical bacterial identification systems. Micro Spectrum 10(2):e02491-e2521. https://doi.org/10.1128/spectrum.02491-21
Toranzo AE (2004) Report about fish bacterial diseases. Mediterranean Aquaculture Laboratories 49–89.
Toranzo AE, Magariños B, Romalde JL (2005) A review of the main bacterial fish diseases in mariculture systems. Aqua 246:37–61. https://doi.org/10.1016/j.aquaculture.2005.01.002
Torres-Corral Y, Santos Y (2019) Identification and typing of Vagococcus salmoninarum using genomic and proteomic techniques. J Fish Dis 42(4):597–612. https://doi.org/10.1111/jfd.12967
Torres-Corral Y, Santos Y (2021) Clonality of Lactococcus garvieae isolated from rainbow trout cultured in Spain: a molecular, immunological, and proteomic approach. Aqua 545:737190. https://doi.org/10.1016/j.aquaculture.2021.737190
Torres-Corral Y, Santos Y (2022) Predicting antimicrobial resistance of Lactococcus garvieae: PCR detection of resistance genes versus MALDI-TOF protein profiling. Aqua 553:738098. https://doi.org/10.1016/j.aquaculture.2022.738098
Torres-Corral Y, Clara FÁ, Ysabel S (2019) Proteomic and molecular fingerprinting for identification and tracking of fish pathogenic Streptococcus. Aqua 498:322–334. https://doi.org/10.1016/j.aquaculture.2018.08.041
Torres-Sangiao E, Leal Rodriguez C, García-Riestra C (2021) Application and perspectives of MALDI-TOF mass spectrometry in clinical microbiology laboratories. Micro 9(7):1539. https://doi.org/10.3390/microorganisms9071539
Tütmez ÇS, Özbey Ü, Tanrıverdi ES, Aksu Ö, Otlu B, Özbey G (2023) Identification of Aeromonas species in trout in Tunceli province by MALDI-TOF MS method. J Popu Therap Clin Phar 30(11):346–354. https://doi.org/10.47750/jptcp.2023.30.11.035
Ucko M, Colorni A (2005) Mycobacterium marinum infections in fish and humans in Israel. J Clin Micro 43(2):892–895. https://doi.org/10.1128/jcm.43.2.892-895.2005
Vanamala P, Sindhura P, Sultana U, Vasavilatha T, Gul MZ (2022) Common bacterial pathogens in fish: an overview. Bacterial Fish Dis 279–306. https://doi.org/10.1016/B978-0-323-85624-9.00010-5
Vardić I, Kapetanović D, Valić D, Kurtović B, Teskeredžić Z, Teskeredžić E (2007) Detection of Renibacterium salmoninarum in tissue of brook trout (Salvelinus fontinalis) by nested RT-PCR. Croatian J Fish Ribarstvo 65(1):15–24
Vázquez-Fernández E, Chinchilla B, Rebollada-Merino A, Domínguez L, Rodríguez-Bertos A (2023) An outbreak of Aeromonas salmonicida in juvenile Siberian Sturgeons (Acipenser baerii). Animals 13(17):2697. https://doi.org/10.3390/ani13172697
Vendrell D, Balcazar JL, Ruiz-Zarzuela I, de Blas I, Girones O, Muzquiz JL (2006) Lactococcus garvieae in fish: a review. Comp Immun Micro Infect Dis 29:177–198. https://doi.org/10.1016/j.cimid.2006.06.003
Verma RK, Sankhla MS, Jadhav S, Parihar K, Gulliya S, Kumar R, Sonone SS (2022) Global status of bacterial fish diseases in relation to aquatic pollution. In Bacterial Fish Diseases. Academic Press. Elsevier Inc. p.155–182. https://doi.org/10.1016/B978-0-323-85624-9.00017-8
Vidal LM, Venas TM, Gonçalves AR, Mattsson HK, Silva RV, Nóbrega MS, Azevedo GP, Garcia GD, Tschoeke DA, Vieira VV, Thompson FL (2020) Rapid screening of marine bacterial symbionts using MALDI-TOF MS. Arch Micro 202:2329–2336. https://doi.org/10.1007/s00203-020-01917-9
von Siebenthal BA, Jacob A, Wedekind C (2009) Tolerance of whitefish embryos to Pseudomonas fluorescens linked to genetic and maternal effects, and reduced by previous exposure. Fish Shellfish Immun 26(3):531–535. https://doi.org/10.1016/j.fsi.2009.02.008
Wahli T, Madsen L (2018) Flavobacteria, a never ending threat for fish: a review. Cur Clin Micro Rep 5:26–37. https://doi.org/10.1007/s40588-018-0086-x
Walczak N, Puk K, Guz L (2017) Bacterial flora associated with diseased freshwater ornamental fish. J Vet Res 61(4):445. https://doi.org/10.1515/jvetres-2017-0070
Wallbanks S, Martinez-Murcia AJ, Fryer JL, Phillips BA, Collins MD (1990) 16S rRNA sequence determination for members of the genus Carnobacterium and related lactic acid bacteria and description of Vagococcus salmoninarum sp. nov. Inter J Syst Evol Micro 40(3):224–230. https://doi.org/10.1099/00207713-40-3-224
Wang Y, Hou Y, Liu X, Lin N, Dong Y, Liu F, Xia W, Zhao Y, Xing W, Chen J, Chen C (2024) Rapid visual nucleic acid detection of Vibrio alginolyticus by recombinase polymerase amplification combined with CRISPR/Cas13a. World J Micro Biotech 40(2):51. https://doi.org/10.1007/s11274-023-03847-2
Wang Y, Zhou Q, Li B, Liu B, Wu G, Ibrahim M, Xie G, Li H, Sun G (2012) Differentiation in MALDI-TOF MS and FTIR spectra between two closely related species Acidovorax oryzae and Acidovorax citrulli. BMC Micro 12(1):1–7. http://www.biomedcentral.com/1471-2180/12/182
Welker M, Van Belkum A, Girard V, Charrier JP, Pincus D (2019) An update on the routine application of MALDI-TOF MS in clinical microbiology. Expert Rev Proteomics 16(8):695–710. https://doi.org/10.1080/14789450.2019.1645603
Whipps CM, Butler WR, Pourahmad F, Watral VG, Kent ML (2007) Molecular systematics support the revival of Mycobacterium salmoniphilum (ex Ross 1960) sp. nov., nom. rev., a species closely related to Mycobacterium chelonei. Int J Syst Evol Micro 57:2525–2531. https://doi.org/10.1099/ijs.0.64841-0
Wieser A, Schneider L, Jung J, Schubert S (2012) MALDI-TOF MS in microbiological diagnostics identification of microorganisms and beyond (mini review). Appl Micro Biotec 93:965–974. https://doi.org/10.1007/s00253-011-3783-4
Woo PT, Bruno DW (eds) (2011) Fish diseases and disorders, vol 3. CABI Publishing. USA, Viral, Bacterial and Fungal Infections
Wu J, Zhou Y, Liu X, Cao Y, Hu C, Chen Y (2020) Extension and application of a database for the rapid identification of Vibrio using MALDI-TOF MS. Acta Oceano Sinica 39:140–146. https://doi.org/10.1007/s13131-020-1635-8
Xu T, Zhang XH (2014) Edwardsiella tarda: an intriguing problem in aquaculture. Aqua 431:129–135. https://doi.org/10.1016/j.aquaculture.2013.12.001
Yavuzcan H, Secer FS, Yilmaz BH, Tunar MA (2022) Exemplifying pathobiome concept through case study: co-infection with Vibrio harveyi, Photobacterium damsela and Cryptocaryon irritans in Salema (Sarpa salpa). J Istanbul Vet Sci 6(3):110–115. https://doi.org/10.30704/http-www-jivs-net.1128614
Zakrzewski AJ, Zarzecka U, Chajęcka-Wierzchowska W, Zadernowska A (2022) A comparison of methods for identifying Santos ales isolates from fish and prawns. Pathogens 11(4):410. https://doi.org/10.3390/pathogens11040410
Zanoni RG, Florio D, Fioravanti ML, Rossi M, Prearo M (2008) Occurrence of Mycobacterium spp. in ornamental fish in Italy. J Fish Dis 31(6):433–441. https://doi.org/10.1111/j.1365-2761.2008.00924.x
Zhu W, Zhou S, Chu W (2020) Comparative proteomic analysis of sensitive and multi-drug resistant Aeromonas hydrophila isolated from diseased fish. Micro Pathogenesis 139:103930. https://doi.org/10.1016/j.micpath.2019.103930
Ziarati M, Zorriehzahra MJ, Hassantabar F, Mehrabi Z, Dhawan M, Sharun K, Emran TB, Dhama K, Chaicumpa W, Shamsi S (2022) Zoonotic diseases of fish and their prevention and control. Vet Quart 42(1):95–118. https://doi.org/10.1080/01652176.2022.2080298
Zlotkin A, Hershko H, Eldar A (1998) Possible transmission of Streptococcus iniae from wild fish to cultured marine fish. Appl Environ Micro 64:4065–4067. https://doi.org/10.1128/AEM.64.10.4065-4067.1998
Zlotkin A, Eldar A, Ghittino C, Bercovier H (1998) Identification of Lactococcus garvieae by PCR. J Clin Micro 36:983–985. https://doi.org/10.1128/jcm.36.4.983-985.1998
Zrnčić S, Vendramin N, Boutrup TS, Boye M, Madsen L, Nonneman B, Oraić D (2021) First description and diagnostics of disease caused by Piscirickettsia salmonis in farmed European sea bass (Dicentrarchus labrax Linnaeus) from Croatia. J Fish Dis 44(7):1033–1042. https://doi.org/10.1111/jfd.13366
Zuffa S, Schmid R, Bauermeister AP, Gomes PW, Caraballo-Rodriguez AM, El Abiead Y. Dorrestein PC (2023) A taxonomically-informed mass spectrometry search tool for microbial metabolomics data. BioRxiv 2023–07. https://doi.org/10.1101/2023.07.20.549584
Acknowledgements
The authors would like to thank Andrea Hogan (Lincoln Univeristy, New Zealand) and Dr. Rachel Forrest (Massey Univeristy, New Zealand) for grammar editing.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK).
Author information
Authors and Affiliations
Contributions
The author is responsible for research, conceptualization, writing, review and editing.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Ethical approval was not required for the research in association with this manuscript.
Competing interests
The author declares no competing interests.
Additional information
Handling Editor: Brian Austin
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Çağatay, İ.T. Use of proteomic-based MALDI-TOF mass spectra for identification of bacterial pathogens in aquaculture: a review. Aquacult Int (2024). https://doi.org/10.1007/s10499-024-01544-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10499-024-01544-x