Abstract
Tick-borne diseases like Rickettsia, Anaplasma and Ehrlichia are widespread infectious zoonoses that threaten the health of both humans and animals worldwide. Ticks and their hosts, such as hedgehogs, can play a crucial role in transmitting tick-borne diseases and the cycle of Rickettsia. To investigate the presence and identity of Rickettsia in hedgehogs and hedgehog-attached ticks in Xuyi County, Southeast China, 114 ticks were collected from 45 hedgehogs captured totally. Via morphological and molecular methods, all these ticks were identified as two species: Haemaphysalis flava (110/114, 96.5%) and Haemaphysalis longicornis (4/114, 3.5%). Rickettsia spp. were genotypically characterized by PCR targeting rrs, gltA, ompA, ompB, and sca4 gene fragments. The prevalence of spotted fever group rickettsiae (SFGR) infection found in hedgehogs and ticks was 17.8% (8/45) and 78.1% (89/114), respectively. Phylogenetic analyses demonstrated that those Rickettsia spp. belong to two species: Rickettsia heilongjiangensis (R. heilongjiangensis XY-1) and a potential new species, Candidatus Rickettsia xuyiensis XY-2. The present study gave the first evidence of R. heilongjiangensis and Candidatus R. xuyiensis in ticks and hedgehogs of Southeast China. Our findings suggest that hedgehogs might be involved in the natural transmission cycle of Rickettsia species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tick-borne intracellular bacteria, including Coxiella burnetti, Anaplasma spp., Ehrlichia spp. and Rickettsia spp., cause emergent or re-emergent infectious diseases among all continents (Ben Said et al. 2018; Boulanger et al. 2019; Eisen 2018; Fang et al. 2021). The genus Rickettsia (family Rickettsiaceae, order Rickettsiales) comprise tiny obligate intracellular bacteria capable of infecting humans and animals with mild to severe symptoms (Merhej et al. 2014; Kho et al. 2019; Shpynov et al. 2018).
Ticks are the primary vector and reservoir of Rickettsia. The spotted fever group rickettsiae (SFGR), including pathogenic and nonpathogenic species found worldwide, are transmitted mainly by hard ticks (Ixodidae) to vertebrate hosts (Parola et al. 2000; Socolovschi et al. 2009). In China, > 110 species of hard ticks have been identified, of which Haemaphysalis longicornis and H. flava are the most common species throughout China (Zhang et al. 2019). A variety of Rickettsia species—including R. japonica, R. heilongjiangensis, R. raoultii, Candidatus Rickettsia tarasevichiae and Candidatus Rickettsia principis—have been screened out from H. longicornis and H. flava (Fang et al. 2021; Jiang et al. 2018; Liu et al. 2016).
Hedgehogs mainly inhabit natural open and green spaces as well as artificial, rural and urban areas, including farmlands, parks, gardens, scrubby habitats at the edge of forests, and shrubby vegetation. They feed on a broad spectrum, including caterpillars, earthworms, small vertebrates, bird eggs, and berries and fruits (Reuter et al. 2019). Hedgehogs are crucial wild animal hosts for various ticks, including Ixodes hexagonus, H. flava, H. longicornis, H. erinacei, H. aegyptium, H. marginatum and Rhipicephalus sanguineus (Jahfari et al. 2017; Khaldi et al. 2012; Marié et al. 2012; Szekeres et al. 2019; Orkun et al. 2019; Barradas et al. 2021). Hedgehogs’ ecological and feeding habits, along with high population densities, resulting in their frequent contact with either human or domestic and wild animals, implicates the possibility of tick-borne diseases (Delogu et al. 2020). Therefore, hedgehogs may be involved in the ecology of several potential emerging pathogens.
A wide range of tick-borne bacteria has been reported in hedgehogs and their attached ticks (Skuballa et al. 2010; Szekeres et al. 2019; Bolanos-Rivero et al. 2017; Gong et al. 2020). Therefore, from a public health perspective it is of great importance to understand the local tick species, tick hosts, and SFG rickettsiae carried by them. Despite the previous extensive efforts of clarifying this problem, the knowledge about the circulation of SFG rickettsiae in areas of Southeast China, such as Xuyi County, Jiangsu Province, is still unclear (Jiang et al. 2010; Tan et al. 2012; Li et al. 2018a, b). Therefore, to evaluate the prevalence of SFG rickettsiae within the Southeast China region, the present study collected free-ranging hedgehogs and ticks from Xuyi County, Southeast China, and investigated their diversity and related SFG rickettsiae, in order to provide a scientific basis for the prevention and control of SFGR.
Materials and methods
Ethical approval
All procedures and protocols for sample collection and processing were approved by the Administrative Committee on Animal Welfare of the Institute of Jiangsu CDC Veterinary and the Ethics Committee of the CDC of Eastern Theater (approval nrs. 2017011 and 2018012; approval dates 26-10-2017 and 15-08-2018).
Ticks and animal collection
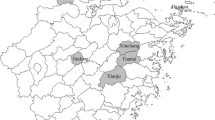
Between November 2017 and April 2019, 45 hedgehogs were captured from forest sites near Tieshan Temple in Xuyi County, Jiangsu Province, China (Fig. 1). After careful examination, ectoparasitic ticks were removed from the hedgehogs using fine forceps and placed individually into 1.5-mL tubes with 70% cleaning ethanol. After cleaning, all ticks were marked with the collection date and stored at −80 °C. The tick species were identified based on morphological criteria (Deng and Jiang 1991) and molecular biology tools (Liu et al. 2016). After being anesthetized with diethyl ether, all hedgehogs were sacrificed to collect muscle tissue, hearts, livers, spleens, lungs, kidneys, brains, and intestines, which were all stored at −80 °C.
DNA extraction
Ticks and hedgehog tissues were homogenized with a stroke-physiological saline solution individually. Homogenates were centrifuged for 10 min at 1000×g and 4 °C, and pellets were collected for DNA extraction. Genomic DNA was extracted from each specimen by using the MiniBEST Universal Genomic DNA Extraction Kit (Takara, Beijing, China) according to the manufacturer’s instructions and subsequently stored at −20 °C before use.
PCR amplification and sequencing
To identify the species of each hedgehog, a partial sequence of the mitochondrial 16S rRNA gene—approximately 201–211 nucleotides (nt) in length—was PCR-amplified using genomic DNA from hedgehog muscle tissues, based on primers (HedF and HedR) as described by Sarri et al. (2014). To identify the species of each tick, the mitochondrial 16S rRNA gene from the genomic DNA of each tick was PCR-amplified using the forward and reverse primers TickHF and TickHR (Liu et al. 2016).
The rickettsial citrate synthase (gltA) gene was chosen as the target for its genus specificity and conservativeness (Mediannikov et al. 2004). All samples were screened for the presence of gltA by nested PCR using two sets of primers, RpCS877F and RpCS1258R, and approximately a 380 bp fragment of the gltA gene was amplified. The second PCR round will be performed if no product was visible by agarose gel electrophoresis. The full-length of gltA gene was amplified in 22 tick samples using primers CS2d and CSEndr. To further characterize SFGR strains, each positive sample for the gltA gene was tested for four other genes: the 16S ribosomal RNA gene (16S rRNA), outer membrane protein A gene (ompA), outer membrane protein B gene (ompB), and surface cell antigen-4 gene (sca4). The primer sets used in each of these assays are listed in Table 1. Sterile distilled water and a previously determined rickettsial-positive tick sample were used as negative and positive controls in each run, respectively. All positive amplicons were purified with PCR Clean-Up Kit (Beyotime, Shanghai, China). Sanger dideoxy DNA sequencing was performed using the BigDye Terminator v.3.1 Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA) and an ABI Prism 3130 × genetic analyzer.
Phylogenetic analysis
Partial nucleotide sequences of rrs, gltA, ompA, ompB, and sca4 obtained from ticks, and hedgehog organs were compared to known sequences using the BLAST program from the NCBI website (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The best-fit evolution model for each gene was calculated via MrModeltest v.2.3 in cooperation with BEAST v.1.10.4 using the Bayesian information criterion (BIC). The K81u (K3Pu), TVM, TPM3u, TN(TN93) and K81u (K3Pu) models were selected for the gltA, ompA, ompB, rrs, and sca4 gene, respectively. Substitution rates at polymorphic sites in both genes followed a gamma distribution with a large proportion of invariable sites. Maximum Likelihood (ML) methods of phylogeny inference on the individual were conducted in BEAST in the ‘Ultrafast’ bootstrap model with the 5000 bootstrap samples, maximum interaction value 1000, and minimum correlation coefficient of 0.90. Nodal support was evaluated by bootstrap resampling for the ML trees from posterior probabilities (PP) for Bayesian inferences. Bootstrap values of 70% or more and Bayesian support values of 0.95 and higher were considered significant nodal support.
Results
The amplified partial sequences (203–215 nt) of the hosts’ 16S rRNA gene (GenBank acc. nr. of the hedgehogs: OM865773) showed the highest nucleotide similarity (96.0–97.0%) to those of Erinaceus amurensis from the GenBank database (KX9646061). Therefore, all hedgehogs (n = 45) were identified as E. amurensis. 17.8% (8/45) of the hedgehogs were PCR positive for rickettsial gltA. The highest infection rate of SFGR in the eight positive hedgehogs was seen in the brain, whereas other organs varied in their presence of rickettsial gltA (Table 2).
A total of 114 adult ticks were collected from 45 hedgehogs and identified as H. flava and H. longicornis through their morphological characteristics and partial 16S rRNA gene (Table 2). 110 of the 114 tested Haemaphysalis ticks (96.5%, GenBank acc. nr. MH520707.1) were identified as H. flava (GenBank acc. nr. OM865774), with a similarity of 99.2%. The other four (3.5%, GenBank acc. nr. KX083342.1) were identified as H. longicornis (GenBank acc. nr. OM865775), with a likeness of 99.2%. Overall, 89 of 114 ticks (78.1%) were tested positive for SFGR, with the infection rate as 80.9% (89/110) in H. flava and 0% (0/4) in H. longicornis, respectively.
An 1183 bp rrs, 1153 bp gltA, 602 bp ompA, 784 bp ompB, and 861 bp sca4 gene fragment of Rickettsia spp. was amplified and sequenced from our partial positive samples, which showed 100% identity to R. heilongjiangensis isolate Xinxian-HL9 (China), with the GenBank acc. nrs. MG9066701, MG9066691, MG9066651, MG9066671 and MG9066681, respectively. Meanwhile, phylogenetic trees (Fig. 2A–E, respectively), inferred from these genes, also showed one isolated strain formed a distinct cluster with R. heilongjiangensis in all trees, which also confirmed the identification of R. heilongjiangensis (Rickettsia heilongjiangensis XY-1).
Phylogenetic tree of Rickettsia spp. detected in ticks and hedgehogs from Southeast China with other rickettsial strains based on partial (A) rrs, (B) gltA, (C) ompA, (D) ompB and (E) sca4 sequences. Rickettsial strains in red ( ) were detected in ticks and hedgehogs of this study. Maximum likelihood (ML) methods of phylogeny inference on individuals were conducted in BEAST v.1.10.4 under the ‘Ultrafast’ Bootstrap model with 5000 bootstrap samples, maximum interaction value 1000 and minimum correlation coefficient of 0.90
) were detected in ticks and hedgehogs of this study. Maximum likelihood (ML) methods of phylogeny inference on individuals were conducted in BEAST v.1.10.4 under the ‘Ultrafast’ Bootstrap model with 5000 bootstrap samples, maximum interaction value 1000 and minimum correlation coefficient of 0.90
Another 1230 bp rrs, 1156 bp gltA, 540 bp ompA, 789 bp ompB, and 886 bp sca4 gene fragment of Rickettsia spp. was amplified and sequenced from the positive ticks. The rrs sequence showed 99.7% nucleotide identity with Candidatus Rickettsia principis (MG5172531), the gltA sequence showed 99.8% nucleotide identity with Candidatus R. principis (AY5781151), the ompB sequence showed 96.9% nucleotide identity with Candidatus R. principis isolate (MG5449911), ompA sequence showed 98.6% nucleotide identity with Rickettsia sp. NGT116-2016-Hfla (LC4610751) and sca4 showed 99.2% nucleotide identity with an uncultured Rickettsia sp. Hme_2021 (LC3794771). The isolated strain could not be classified into specific species due to a lack of consensus between the phylogenetic trees (Fig. 2). According to the gene sequence-based criteria proposed by Fournier et al. (2003), this Rickettsia isolate can therefore be classified as a potentially novel SFGR, named Candidatus Rickettsia xuyiensis-XY2.
GenBank acc. nrs. of partial sequences obtained in the study are: MZ646340-MZ646341 (rrs genes of Rickettsia spp.), MZ646342-MZ646345 (gltA, sca4, ompB, and ompA gene of R. heilongjiangensis XY-1), MZ646346-MZ646349 (gltA, ompA, sca4, and ompB gene of Candidatus R. xuyiensis XY-2), OM865773(16S rRNA gene of hedgehog), OM865774 and OM865775 (16S rRNA gene of H. flava and H. longicornis).
Discussion
This study reports Rickettsia sp.'s finding in hedgehogs and ticks from Jiangsu Province, Southeast China. In addition, this is the first report of R. heilongjiangensis and a novel potential species of Rickettsia (Candidatus R. xuyiensis XY-2) in H. flava and hedgehogs of Xuyi, Southeast China.
SFGRs are widely distributed throughout China and tend to have regional characteristics. Previous studies have revealed the extensive diversity of rickettsiae among tick species and geographic areas (El-Mahallawy et al. 2015; Fang et al. 2021). There are many hills and low mountains in the Xuyi area, and it has a developed presence of animal husbandry. Many tick bites have been reported among fever cases with thrombocytopenia syndrome in Jiangsu Province (Li et al. 2017). Therefore, improving the knowledge on the prevalence of Rickettsia in ticks and hosts from this region can identify potential rickettsioses in the population and reduce the risk for tick-borne Rickettsia transmission.
Small mammals and ticks are intermediate hosts or vectors of many zoonoses. Hedgehogs, one of the most important hosts of ticks, can play an essential role in the natural foci of tick-borne pathogens (Orkun et al. 2019). Our results demonstrate that the dominant tick species carried by hedgehogs in the Xuyi area is H. flava, followed by H. longicornis, which agrees with the findings of both Sun et al. (2009) and Lan et al. (2016). The number of ticks carried by each hedgehog in this study may vary significantly due to the sampling season or the activity tracking of hedgehogs. Additionally, an initial screening test using gltA nested PCR revealed that 78.1% of the ticks and 17.8% of the hedgehogs were infected with SFG rickettsiae. This percentage was significantly higher than previous work from the Sichuan (33.5%), Yunnan (12.1%), and Zhejiang (7.5%) provinces (Liu et al. 2020; Sun et al. 2015; Zhang et al. 2018). We found that the prevalence of SFGR infection of ticks in eight positive hedgehogs was 76.9 ~ 100%, and most SFGR infection of ticks collected from negative hedgehogs was 0–100%. Of 89 ticks infected with SFG rickettsiae, 63 positive ticks were carried by eight positive hedgehogs (Table 2). As the ticks may suck the blood of these hedgehogs, they had a high positive rate of spotted fever.
Furthermore, we determined partial sequences of the gltA gene of SFG rickettsiae by conventional PCR in 45 hedgehogs organs (heart, brain, intestine, spleen, lung, liver, kidney). Based on the sequences of the gltA gene obtained from 45 hedgehogs’ organs, the highest infection rate of SFG rickettsiae was detected in the brain (5/45); other organs varied in their presence of rickettsial gltA (Table 2). Therefore, the brain of the hedgehog may be particularly susceptible to Rickettsia. Our findings also indicate that hedgehogs and their carrying ticks can serve as the animal host and vector for SFG rickettsiae.
For the molecular classification of SFG rickettsiae that were obtained in the study, partial sequences of rrs, gltA, ompA, ompB, and sca4 were analyzed. Phylogenetic trees inferred from rrs, gltA, ompA, ompB, and sca4 analysis are shown in Fig. 2. One isolated strain formed a distinct cluster with R. heilongjiangensis in all trees (Fig. 2) and thus were identified as R. heilongjiangensis (XY-1). Nucleotide sequence analysis of five genes of R. heilongjiangensis XY-1 showed 97–100% similarity with R. heilongjiangensis isolate Xinxian-HL9 and R. japonica YH_M. The isolated strain could not be classified into specific species due to a lack of consensus between the phylogenetic trees. It shares a branch with the previously reported Candidatus R. principis isolate Kh-81, uncultured Rickettsia sp. Hme_2021, uncultured Rickettsia sp. clone 7–17, whereas it forms a separate branch in the ompB and sca4 phylogenetic tree (Fig. 2). Moreover, in the isolated strain, the sequence nucleotide identity to recognized Rickettsia species was < 99.8, 99.9, 98.8, 99.2 and 99.3% for rrs, gltA, ompA, ompB, and sca4, respectively, which suggests that this agent is novel potential SFG Rickettsia according to Fournier et al. (2003). Therefore, this species was provisionally named Candidatus R. xuyiensis-XY2, concerning the location where it was found. Our findings indicate that hedgehogs and H. flava collected from hedgehogs in Southeast China were infected with R. heilongjiangensis and Candidatus R. xuyiensis. The diseases caused by these pathogens should therefore be monitored in Southeast China. Further isolation and identification are needed to obtain morphological characteristics and the entire genome of these species. Previous studies have reported that H. longicornis is an essential vector of R. heilongjangensis (Jiang et al. 2019; Liu et al. 2020; Zhuang et al. 2018). However, no rickettsiae were detected in any of the H. longicornis ticks collected in our study, which may be due to the limited sample size of H. longicornis ticks.
There are some limitations of this study worth noting. Firstly, our investigation is biased because the infection rates were calculated using ticks collected from the infected hedgehogs, where we collected fewer ticks from uninfected hedgehogs. Therefore, the actual infection rates might be lower than those determined by this research. Secondly, we mainly focused on the infection rates and tick species collected from hedgehogs, and we did not identify ticks carried by other small mammals in the Xuyi area. Thus, it is crucial to find Rickettsiales infection among other local animals and humans in a subsequent study.
References
Anstead CA, Chilton NB (2013) A novel Rickettsia species detected in vole ticks (Ixodes angustus) from Western Canada. Appl Environ Microbiol 79:7583–7589. https://doi.org/10.1128/AEM.02286-13
Barradas PF, Mesquita JR, Mateus TL, Ferreira P, Amorim I, Gartner F, de Sousa R (2021) Molecular detection of Rickettsia spp in ticks and fleas collected from rescued hedgehogs (Erinaceus europaeus) in Portugal. Exp Appl Acarol 83:449–460
Ben Said M, Belkahia H, Messadi L (2018) Anaplasma spp in North Africa: a review on molecular epidemiology, associated risk factors and genetic characteristics. Ticks Tick Borne Dis 9:543–555
Bolanos-Rivero M, Carranza-Rodriguez C, Rodriguez NF, Gutierrez C, Perez-Arellano JL (2017) Detection of Coxiella burnetii DNA in peridomestic and wild animals and ticks in an endemic region (Canary Islands, Spain). Vector Borne Zoonotic Dis 17:630–634. https://doi.org/10.1089/vbz.2017.2120
Boulanger N, Boyer P, Talagrand-Reboul E, Hansmann Y (2019) Ticks and tick-borne diseases. Med Mal Infect 49:87–97. https://doi.org/10.1016/j.medmal.2019.01.007
Delogu M, Cotti C, Lelli D, Sozzi E, Trogu T, Lavazza A, Garuti G, Castrucci MR, Vaccari G, Marco MA, Moreno A (2020) Eco-virological preliminary study of potentially emerging pathogens in hedgehogs (Erinaceus europaeus) recovered at a Wildlife Treatment and Rehabilitation Center in Northern Italy. Animals 10(3):407. https://doi.org/10.3390/ani10030407
Deng GF, Jiang ZJ (1991) Economic insect fauna of Chinese, vol 39. Acari Leach Science Press, Beijing
Eisen L (2018) Pathogen transmission in relation to duration of attachment by Ixodes scapularis ticks. Ticks Tick Borne Dis 9:535–542. https://doi.org/10.1016/j.ttbdis.2018.01.002
El-Mahallawy HS, Lu G, Kelly P, Xu D, Li Y, Fan W, Wang C (2015) Q fever in China: a systematic review, 1989–2013. Epidemiol Infect 143:673–681. https://doi.org/10.1017/S0950268814002593
Fang LZ, Lei SC, Yan ZJ, Xiao X, Liu JW, Gong XQ, Yu H, Yu XJ (2021) Detection of Multiple Intracellular Bacterial Pathogens in Haemaphysalis flava Ticks Collected from Hedgehogs in Central China. Pathogens 10:8. https://doi.org/10.3390/pathogens10020115
Fournier PE, Roux V, Raoult D (1998) Phylogenetic analysis of spotted fever group rickettsiae by study of the outer surface protein rOmpA. Int J Syst Bacteriol 48(Pt 3):839–849. https://doi.org/10.1099/00207713-48-3-839
Fournier PE, Dumler JS, Greub G, Zhang J, Wu Y, Raoult D (2003) Gene sequence-based criteria for identification of new rickettsia isolates and description of Rickettsia heilongjiangensis sp. nov. J Clin Microbiol 41:5456–5465. https://doi.org/10.1128/JCM.41.12.5456-5465.2003
Reuter G, Várallyay E, Baráth D, Földvári G, Szekeres S, Boros A, Kapusinszky B, Delwart E, Pankovics P (2019) Analysis of a novel RNA virus in a wild northern white-breasted hedgehog (Erinaceus roumanicus). Arch Virol 164(12):3065–3071. https://doi.org/10.1007/s00705-019-04414-7
Gong XQ, Xiao X, Liu JW, Han HJ, Qin XR, Lei SC, Yu XJ (2020) Occurrence and genotyping of Coxiella burnetii in hedgehogs in China. Vector Borne Zoonotic Dis 20:580–585. https://doi.org/10.1089/vbz.2019.2589
Jahfari S, Ruyts SC, Frazer-Mendelewska E, Jaarsma R, Verheyen K, Sprong H (2017) Melting pot of tick-borne zoonoses: the European hedgehog contributes to the maintenance of various tick-borne diseases in natural cycles urban and suburban areas. Parasites Vectors 10:134. https://doi.org/10.1186/s13071-017-2065-0
Jiang LP, Meng Q, Gui QR, Tong F, Wang Z (2010) Detection of rOmp A and gltA genes of spotted fever group rickettsiae from tick specimens in Zhejiang province. Chin J Vector Biol Control 21:350–352
Jiang J, An H, Lee JS, O’Guinn ML, Kim HC, Chong ST, Zhang Y, Song D, Burrus RG, Bao Y, Klein TA, Richards AL (2018) Molecular characterization of Haemaphysalis longicornis-borne rickettsiae, Republic of Korea and China. Ticks Tick Borne Dis 9:1606–1613. https://doi.org/10.1016/j.ttbdis.2018.07.013
Jiang J, Choi YJ, Kim J, Kim HC, Klein TA, Chong ST, Richards AL, Park HJ, Shin SH, Song D, Park KH, Jang WJ (2019) Distribution of Rickettsia spp. in Ticks from Northwestern and Southwestern Provinces, Republic of Korea Korean. J Parasitol 57:161–166. https://doi.org/10.3347/kjp.2019.57.2.161
Khaldi M, Socolovschi C, Benyettou M, Barech G, Biche M, Kernif T, Raoult D, Parola P (2012) Rickettsiae in arthropods collected from the North African Hedgehog (Atelerix algirus) and the desert hedgehog (Paraechinus aethiopicus) in Algeria. Comp Immunol Microbiol Infect Dis 35:117–122. https://doi.org/10.1016/j.cimid.2011.11.007
Kho KL, Tan PE, Tay ST (2019) Diversity of rickettsiae in feeding and questing ticks collected from a Malaysian Forest Reserve Area. J Med Entomol 56:547–552. https://doi.org/10.1093/jme/tjy168
Lan C, Shen Y, You Y, Dai X, Wu X, Zhao Q (2016) Investigation of ticks on small mammals in Wuxi City, China. J Hyg Insect Equip 22:372–374
Li Z, Hu J, Cui L, Hong Y, Liu J, Li P, Guo X, Liu W, Wang X, Qi X, Wu B, Feng Z, Shen A, Liu X, Zhao H, Tan W, Zhou J, Xing Z, Bao C (2017) Increased prevalence of severe fever with Thrombocytopenia syndrome in Eastern China clustered with multiple genotypes and reasserted virus during 2010–2015. Sci Rep 7:6503. https://doi.org/10.1038/s41598-017-06853-1
Li BB, Yang PF, Liu CC, Li SS, Liu L, Jin J, Zhao HR, Yao HB, He NJ (2018a) Detection and phylogenetic analysis of rickettsiae from tick in Huai ’an. Mod Prev Med 45:2641–2646
Li J, Hu W, Wu T, Li HB, Sun Y, Chen Z, Shi Y, Zong J, Latif A, Wang L, Yu L, Yu XJ, Liu BY, Liu Y (2018b) Japanese Spotted Fever in Eastern China, 2013. Emerg Infect Dis 24:2107–2109. https://doi.org/10.3201/eid2411.170264
Liu H, Li Q, Zhang X, Li Z, Wang Z, Song M, Wei F, Wang S, Liu Q (2016) Characterization of rickettsiae in ticks in northeastern China. Parasit Vectors 9:498. https://doi.org/10.1186/s13071-016-1764-2
Liu H, Liang X, Wang H, Sun X, Bai X, Hu B, Shi N, Wang N, Zhang X, Huang L, Liao J, Huang F, Zhang H, Si X, Huang S, Jin N, Liu Q, Li L (2020) Molecular evidence of the spotted fever group Rickettsiae in ticks from Yunnan Province, Southwest China. Exp Appl Acarol 80:339–348. https://doi.org/10.1007/s10493-020-00467-5
Marié JL, Davoust B, Socolovschi C, Raoult D, Parola P (2012) Molecular detection of rickettsial agents in ticks and feas collected from a European hedgehog (Erinaceus europaeus) in Marseilles. France Comp Immunol Microbiol Infect Dis 35:77–79. https://doi.org/10.1016/j.cimid.2011.11.005
Mediannikov OY, Sidelnikov Y, Ivanov L, Mokretsova E, Fournier PE, Tarasevich I, Raoult D (2004) Acute tick-borne rickettsiosis caused by Rickettsia heilongjiangensis in Russian Far East. Emerg Infect Dis 10:810–817. https://doi.org/10.3201/eid1005.030437
Merhej V, Angelakis E, Socolovschi C, Raoult D (2014) Genotyping, evolution and epidemiological findings of Rickettsia species. Infect Genet Evol 25:122–137. https://doi.org/10.1016/j.meegid.2014.03.014
Orkun O, Cakmak A, Nalbantoglu S, Karaer Z (2019) Molecular detection of a novel Babesia sp. and pathogenic spotted fever group rickettsiae in ticks collected from hedgehogs in Turkey: Haemaphysalis erinacei, a novel candidate vector for the genus Babesia. Infect Genet Evol 69:190–198. https://doi.org/10.1016/j.meegid.2019.01.028
Parola P, Paddock CD, Socolovschi C, Labruna MB, Mediannikov O, Kernif T, Abdad MY, Stenos J, Bitam I, Roux V, Raoult D (2000) Phylogenetic analysis of members of the genus Rickettsia using the gene encoding the outer-membrane protein rOmpB (ompB). Int J Syst Evol Microbiol 50(Pt 4):1449–1455. https://doi.org/10.1099/00207713-50-4-1449
Roux V, Raoult D (2000) Phylogenetic analysis of members of the genus Rickettsia using the gene encoding the outer-membrane protein rOmpB (ompB). Int J Syst Evol Microbiol 50 Pt 4:1449–1455. https://doi.org/10.1099/00207713-50-4-1449
Sarri C, Stamatis C, Sarafidou T, Galara I, Godosopoulos V, Kolovos M, Liakou C, Tastsoglou S, Mamuris Z (2014) A new set of 16S rRNA universal primers for identification of animal species. Food Control 43:35–41. https://doi.org/10.1080/19440049.2019.1580389
Sekeyova Z, Roux V, Raoult D (2001) Phylogeny of Rickettsia spp. inferred by comparing sequences of ‘gene D’, which encodes an intracytoplasmic protein. Int J Syst Evol Microbiol 51:1353–1360. https://doi.org/10.1099/00207713-51-4-1353
Shpynov SN, Fournier PE, Pozdnichenko NN, Gumenuk AS, Skiba AA (2018) New approaches in the systematics of rickettsiae. New Microbes New Infect 23:93–102. https://doi.org/10.1016/j.nmni.2018.02.012
Skuballa J, Petney T, Pfaffle M, Taraschewski H (2010) Molecular detection of Anaplasma phagocytophilum in the European hedgehog (Erinaceus europaeus) and its ticks. Vector Borne Zoonotic Dis 10:1055–1057. https://doi.org/10.1089/vbz.2009.0150
Socolovschi C, Mediannikov O, Raoult D, Parola P (2009) The relationship between spotted fever group Rickettsiae and ixodid ticks. Vet Res 40:34. https://doi.org/10.1051/vetres/2009017
Sun J, Lin J, Gong Z, Chang Y, Ye X, Gu S, Pang W, Wang C, Zheng X, Hou J, Ling F, Shi X, Jiang J, Chen Z, Lv H, Chai C (2015) Detection of spotted fever group Rickettsiae in ticks from Zhejiang Province, China. Exp Appl Acarol 65:403–411. https://doi.org/10.1007/s10493-015-9880-9
Sun B, Xue X, Hou W (2009) Background investigation of tick and acarid at Qingdao international airport. Port Health Control 14(6):39–41
Szekeres S, van Docters Leeuwen A, Toth E, Majoros G, Sprong H, Foldvari G (2019) Road-killed mammals provide insight into tick-borne bacterial pathogen communities within urban habitats. Transbound Emerg Dis 66:277–286. https://doi.org/10.1111/tbed.13019
Tan ZY, Li L, Zhang LJ (2012) Cross-sectional survey on the prevalence of antibodies to several types of Rickettsia in human and livestock in Jiangsu province. Suzhou Univ J Med Sci 32(4):445–449
Zhang X, Geng J, Du J, Wang Y, Qian W, Zheng A, Zou Z (2018) Molecular identification of Rickettsia species in Haemaphysalis ticks collected from Southwest China. Vector-Borne and Zoonotic Diseases 18(12):663–668. https://doi.org/10.1089/vbz.2017.2231
Zhang YK, Zhang XY, Liu JZ (2019) Ticks (Acari: Ixodoidea) in China: Geographical distribution, host diversity, and specificity. Arch Insect Biochem Physiol 102:e21544. https://doi.org/10.1002/arch.21544
Zhuang L, Du J, Cui XM, Li H, Tang F, Zhang PH, Hu JG, Tong YG, Feng ZC, Liu W (2018) Identification of tick-borne pathogen diversity by metagenomic analysis in Haemaphysalis longicornis from Xinyang. China. Infect Dis Poverty 7:45. https://doi.org/10.1186/s40249-018-0417-4
Acknowledgements
This study was funded by grants from projects supported by the General project of Jiangsu Health Committee (H2019015), National Natural Science Foundation of China (U1602223). We would like to thank Yu Wang and Baojun Xu for their help with sampling location and sample collection.
Author information
Authors and Affiliations
Contributions
Designed the study: CQZ, WLT, and CMS; sample collection: CQZ, WLT, LLA, YQ, and YSL; processed samples and extracted DNA: CQZ and HL; performed PCR and sequencing: CQZ; Phylogenetic analysis: CQZ, FQY, and QWW; analyzed data and wrote the manuscript: CQZ and WLT; reviewed the manuscript: CQZ, WLT, and CMS. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, C., Ai, L., Qi, Y. et al. Molecular detection of spotted fever group rickettsiae in hedgehogs (Erinaceus amurensis) and hedgehog-attached ticks in Xuyi County, Southeast China. Exp Appl Acarol 88, 97–111 (2022). https://doi.org/10.1007/s10493-022-00721-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10493-022-00721-y