Abstract
An appealing strategy for finding novel bioactive molecules in Nature consists in exploring underrepresented and -studied microorganisms. Here, we investigated the antimicrobial and tumoral anti-proliferative bioactivities of twenty-three marine and estuarine bacteria of the fascinating phylum Planctomycetota. This was achieved through extraction of compounds produced by the Planctomycetota cultured in oligotrophic medium followed by an antimicrobial screening against ten relevant human pathogens including Gram-positive and Gram-negative bacteria, and fungi. Cytotoxic effects of the extracts were also evaluated against five tumoral cell lines. Moderate to potent activities were obtained against Enterococcus faecalis, methicillin-sensitive and methicillin-resistant Staphylococcus aureus and vancomycin-sensitive and vancomycin-resistant Enterococcus faecium. Anti-fungal effects were observed against Trichophyton rubrum, Candida albicans and Aspergillus fumigatus. The highest cytotoxic effects were observed against human breast, pancreas and melanoma tumoral cell lines. Novipirellula caenicola and Rhodopirellula spp. strains displayed the widest spectrum of bioactivities while Rubinisphaera margarita ICM_H10T affected all Gram-positive bacteria tested. LC-HRMS analysis of the extracts did not reveal the presence of any known bioactive natural product, suggesting that the observed activities are most likely caused by novel molecules, that need identification. In summary, we expanded the scope of planctomycetal species investigated for bioactivities and demonstrated that various strains are promising sources of novel bioactive compounds, which reenforces the potential biotechnological prospects offered by Planctomycetota.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Humanity is currently facing a shortage of effective drugs for many of its health challenges. Noncommunicable diseases, which include cancer, diabetes, obesity, cardiovascular diseases and chronic respiratory conditions, are presently accountable for more than 70% of the world deaths (WHO 2019a). Particularly, cancer alone was responsible for around 10 million worldwide deaths in 2020. By this date, lung and colon cancer were the most deadly, while breast cancer was the most common (WHO 2019a). Although prevention and early detections are crucial in reducing the burden of cancer, finding novel effective treatments with lower side effects is also of great importance for improving the recovery rates and the quality of life of patients undergoing therapy. On another perspective, perhaps one of the most worrying health problems today is the continuous rise of antibiotic resistance in microbial pathogens (WHO 2017, 2019b). According to the World Health Organization (WHO), deaths caused by resistant microorganisms have the tendency to increase exponentially. Bacteria classified as critical and high priority by WHO include antibiotic resistant Pseudomonas aeruginosa, Acinetobacter baumannii, Neisseria gonorrhoeae, Staphylococcus aureus, Helicobacter pylori, Campylobacter sp., Salmonella sp. and many Enterobacteriaceae (WHO 2017), as well as various fungi (WHO 2022). The still ongoing COVID-19 outbreak revealed how complex a worldwide epidemy can be regarding public health and socioeconomic impact. In the verge of the imminent possibility of new pandemics caused by resistant microorganisms, we are now in need of novel successful medications.
Nature has always been a shear unlimited source for bioactive molecules and, in particular, still untapped environments such as the Oceans have gained attention in recent years due to their rich biological and metabolic diversities (Gerwick and Moore 2012; Santos, et al. 2020b). In particular, microorganisms offer many advantages such as ease of manipulation and large potential for the production of many bioactive compounds and promising leads. In fact, many known antimicrobials come from (marine) microorganisms (Santos, et al. 2020b). Marine Actinomycetota, Pseudomonadota, Bacillota, Bacteroidota, Cyanobacteria, fungi and dinoflagellates have shown to be excellent sources of metabolite diversity (Anjum, et al. 2018; Braña, et al. 2017; Feling, et al. 2003; Harinantenaina Rakotondraibe, et al. 2015; Leao, et al. 2013; Reynolds, et al. 2018; Rodriguez, et al. 2018; Schulze, et al. 2015; Tareq, et al. 2014; Wiese, et al. 2018; Zhang, et al. 2018, 2016). Nevertheless, they have been more extensively studied in the past decades, while other microbial groups still remain underexplored. An attractive approach for finding novel effective drugs and lower rediscovery rates would be to investigate these underexplored microorganisms.
The vast majority of estimated bacterial diversity is still uncultured in laboratory (Barer and Harwood 1999; Rinke, et al. 2013), which heavily restricts what could be screened for biotechnological purposes. Nonetheless, recent cultivation efforts from diverse research groups have allowed to bring into axenic culture many strains (including novel species) from underexplored and evolutionarily deep-branching groups such as the Planctomycetota (Almeida, et al. 2022, Dedysh, et al. 2020, Gaurav, et al. 2021, Kaushik, et al. 2020, Kulichevskaya, et al. 2020a, Kulichevskaya, et al. 2022, Kulichevskaya, et al. 2020b, Kumar, et al. 2020a, Kumar, et al. 2020b, Kumar, et al. 2021a, 2021b, Vitorino, et al. 2020, Vitorino et al. 2021b, Vitorino, et al. 2022b, Vitorino, et al. 2022c, Vitorino, et al. 2022d, Wiegand, et al. 2020). The bacterial phylum Planctomycetes (now renamed Planctomycetota (Oren and Garrity 2021)) belongs to the super phylum Planctomycetota-Chlamydiota-Verrucomicrobiota (PVC) (Rivas-Marín and Devos 2018; Wagner and Horn 2006). Planctomycetota have many interesting and distinctive features that makes them relevant bacteria from an ecological and a biological point of view (Lage, et al. 2019; Wiegand, et al. 2018). They can inhabit most of the earth’s biomes and play central roles in the carbon and nitrogen cycles (Lage and Bondoso 2014; Lage, et al. 2019). Their cells have permanently condensed DNA, unusual structures and complex cellular plans with intricate invaginations of the cytoplasmatic membrane, large periplasmic space and an enigmatic cell wall (Boedeker, et al. 2017; Devos 2014; Jeske, et al. 2015; Lage, et al. 2013; Santarella-Mellwig, et al. 2013). Planctomycetota cell division can be by binary fission (Phycisphaerae and Candidatus “Brocadiia”) (Fukunaga, et al. 2009; Jetten, et al. 2010; Lodha, et al. 2021) or by a rare prokaryotic budding (Planctomycetia) (Krieg et al. 2010; Vitorino and Lage 2022). This division is performed without the otherwise universal FtsZ protein, and the exact process is still unknown (Rivas-Marin, et al. 2020). Endocytosis like uptake of molecules and membranous tubulovesicular networks were also proposed for these bacteria (Acehan, et al. 2014; Boedeker, et al. 2017).
It was only in the recent years that Planctomycetota have gained attention as promising organisms for biotechnological purposes due to the lack of isolates (< 200) (Kallscheuer and Jogler 2021; Vitorino and Lage 2022). Furthermore, Planctomycetota do not grow as fast or are not as easy to handle as other microorganisms, so they often require complex media formulations and long cultivation periods. Even so, insights into the biology of these bacteria have highlighted their potential as producers of molecules of pharmacological interest. Most Planctomycetota have large genomes with many genes of unknown function and a comparatively high number of genes coding large proteins (Faria, et al. 2018; Lage and Bondoso 2014; Wiegand, et al. 2018), features that are shared with other well-known producers of bioactive compounds. Genome mining in many strains also demonstrated their rich content of putatively new biosynthetic gene clusters (BGCs) from several structural types (such as nonribosomal peptide synthases (NRPS) polyketide synthases (PKS), ribosomally synthesized and post-translationally modified peptides (RiPPs), lanthipeptides and bacteriocins), which are often linked to the production of bioactive secondary metabolites (Belova, et al. 2020; Graca, et al. 2016; Jeske, et al. 2013; Kallscheuer and Jogler 2021; Vitorino, et al. 2022b, 2022d; Wiegand, et al. 2020). Additionally, the capability of some Planctomycetota to produce antimicrobial and cytotoxic molecules was already confirmed through in vitro bioactivity screenings, including the description of the first (and the only one up to date) antimicrobial compounds from planctomycetal origin (Belova, et al. 2020; Calisto, et al. 2019; Graca, et al. 2016; Jeske, et al. 2016; Kallscheuer, et al. 2020; Sandargo, et al. 2020). These N-acyl-amino acid molecules were named Stieleriacines A-E and were isolated from two species in the genus Stieleria (Kallscheuer, et al. 2020; Sandargo, et al. 2020). Although these studies begin to show the great potential for biosynthesis of natural products in the Planctomycetota, the spectrum of planctomycetal strains and species studied remains limited.
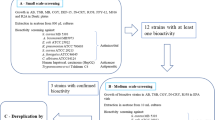
The demand for new chemically distinct molecules, combined with the potential of the promising phylum Planctomycetota, motivated us to study twenty-three marine and brackish water planctomycetal strains from our culture collection. They were screened for various bioactivities following the workflow represented in Fig. 1. Novel recently described species (Godinho et al. 2021; Vitorino, et al. 2020, 2021a, 2022a, 2022b, 2022c, 2023) were also included in this study. Metabolites from Planctomycetota cultured in oligotrophic medium were extracted with organic solvents and followed by antimicrobial and antifungal assays, in a first stage, against two different types of bacteria (the Gram-negative Escherichia coli and the Gram-positive Staphylococcus aureus) and three different fungi (the yeast Candida albicans and the filamentous fungi Trichophyton rubrum and Aspergillus fumigatus). In the second stage, extracts obtained from the selected bioactive strains were additionally screened against five human pathogens (drug resistant and drug sensitive Gram-positive bacteria) and against five human tumoral cell lines (Fig. 1). Planctomycetal crude extracts were also analyzed by liquid chromatography coupled to high-resolution mass spectrometry (LC-HRMS), which allowed the prediction of molecular formulae of the components present in these extracts and their search against natural products databases.
Material and Methods
Biological material
The biological material analyzed in this study included twenty-three planctomycetal strains belonging to our culture collection (Laboratory for Microbial Ecophysiology of the University of Porto, Portugal-LEMUP). These strains were previously isolated from different samples collected in marine and estuarine environments in various regions of Portugal (Bondoso, et al. 2014, 2015; Godinho et al. 2021; Lage and Bondoso 2011; Vitorino, et al. 2020, 2021b, 2022b, 2022c, 2023, 2022d). They are all affiliated to the class Planctomycetia within the phylum Planctomycetota. The diversity screened in this study includes Planctomycetota from three different families (Pirellulaceae/Lacipirellulaceae and Planctomycetaceae) of orders Pirellulales and Planctomycetales in a total of nine genera and thirteen different species, including recently described novel taxa (Vitorino, et al. 2020, 2021b, 2022c, 2023, 2022d).
All 16S rRNA gene sequences are deposited in the National Center for Biotechnology Information (NCBI) and the respective GenBank accession numbers presented in Table 1, where a summary of the isolation habitat and taxonomic features of each strain are also specified. Maximum likelihood 16S rRNA gene sequence-based phylogenetic trees were constructed to show their position in the phylum together with the results obtained. Sequences were first aligned with Clustal W (Larkin, et al. 2007) together with publicly sequences from the closest type strains, which were retrieved from NCBI under the following GenBank tags: LR132072 for Gimesia chilikensis JC646T, MK554521 for Alienimonas californiensis CA12T, AJ231190 for Rubinisphaera brasiliensis IFAM 1448 T, MK559971 for Stieleria neptunia Enr13T, MK554549 for Novipirellula galeiformis Pla52aT, NR_136448 for Novipirellula caenicola YM26-125 T, NR_043384 for Rhodopirellula baltica SH1T, EF589351 for Rhodopirellula lusitana UC17T, AP021861 for Lacipirellula parvula PX69T, KF607112 for Bythopirellula goksoeyrii Pr1dT, MK559982 for Aeoliella mucimassa Pan181T. The trees were constructed using the Mega X software with 1000 bootstraps replicates following the general time reversible model and activated gamma distributed with invariant sites (G + I) (Kumar, et al. 2018). iTOL was used for tree visualization and annotation (Letunic and Bork 2021).
Culturing of strains and natural products extraction
Strains were routinely maintained in pure culture at 25 °C in modified M13 medium plates prepared as previously described (Lage and Bondoso 2011). For metabolite extraction, strains were first cultured at 25 °C for 7 days in a liquid pre-inoculum in modified M13 medium prepared as follow: per liter of medium, 920 mL of deionized water, 0.25 g of yeast extract and peptone, 50 mL of HCl-Tris buffer (0.1 mM, pH 7.5) and 33 g of sea salts (Sigma-Aldrich®, St. Louis, MO, USA). After autoclaving, the following solutions were added by sterile filtration (0.22 µm): 10 mL glucose solution (2.5% w/v), 10 mL vitamin solution (following previous formulations (Lage and Bondoso 2011)) and 20 mL Hutner’s basal salts (prepared as described formerly (Cohen-Bazire, et al. 1957). To enhance secondary metabolite production, nutritional stress was induced. The pre-inoculums were diluted 1:5 in 250 mL of the oligotrophic 1:10 M13 medium (Vitorino, et al. 2021b), prepared as follows, per liter of medium: 919 mL of deionized water, 0.025 g of yeast extract and peptone, 50 mL of HCl-Tris buffer (0.1 mM, pH 7.5) and 33 g of sea salts (Sigma-Aldrich®, St. Louis, MO, USA). After autoclaving, the following solutions were added by sterile filtration (0.22 µm): 1 mL glucose solution (2.5% w/v), 10 mL vitamin solution (Lage and Bondoso 2011) and 20 mL Hutner’s basal salts (Cohen-Bazire, et al. 1957). After incubation in glass flasks for 14 days at 25 °C under agitation in the absence of light, cells were harvested by centrifugation (3600 rpm for 15 min in an Eppendorf 5810R Centrifuge) and metabolites extracted using acetone (250 mL) for 1–4 h. Solid residues were achieved by drying the solvent in a rotatory vacuum evaporator (Rotavapor® R-100 equipment from BUCHI). Finally, the crude extracts were suspended in 20% v/v dimethyl sulfoxide (DMSO) in water and stored under refrigerated conditions for subsequent screenings.
Antibacterial assays
All antibacterial assays were performed as previously described, using liquid cultures in a 96-well plate format (Audoin, et al. 2013; Santos, et al. 2020a; Vitorino, et al. 2022d). The two bacterial targets initially tested in this study were methicillin-sensitive Staphylococcus aureus (MSSA) ATCC 29213 and Escherichia coli ATCC 25922 (Gram-positive and Gram-negative representatives, respectively). Briefly, the pathogens were cultured overnight (30 °C under 220 rpm) in nutrient broth (NB) medium (5 g of peptone, 1 g of yeast extract, 1 L of deionized water) and the cultures standardized to an assay inoculum of 5.0 × 105 colony forming units (CFU)/mL. In the antibacterial assay, 10 μL of each planctomycetal extract were diluted 1:10 with the bacterial inoculum for a final volume of 100 μL per well (assay concentration of extract: 1 mg/mL). Internal plate controls were included for validation of the assay: blank control (100 μL of NB medium), growth control (90 μL target inoculum + 10 μL Millipore water), positive control (90 μL target inoculum + 10 μL ampicillin 200 mg/L) and solvent control (90 μL target inoculum + 10 μL DMSO 20% v/v). Three biological independent experiments (n = 3) were conducted. Turbidity (absorbance (Abs) at 600 nm) was measured at the beginning (T0) and after 24 h of incubation (Tf) at 30 °C in a Multiskan GO plate reader from Thermo Scientific™ equipment. The inhibition percentages were calculated as previously described (Vitorino, et al. 2022d).
Additionally, promising bioactive planctomycetal strains were also screened for a panel of pathogenic microorganisms from Fundación MEDINA’s collection (Fig. 1). These included methicillin-resistant Staphylococcus aureus MB 5393 (MRSA), Enterococcus faecalis ATCC 29212, vancomycin-resistant Enterococcus faecium (clinical isolates VanA and VanB) and vancomycin-sensitive Enterococcus faecium (clinical isolate VanS). Briefly, the bacterial pathogens were incubated overnight at 37 °C under 220 rpm in Bacto™ Brain heart infusion (BHI) medium (Becton, Dickinson and company, MD, USA). For the assay, they were diluted to standardized cultures of 1.1 × 106 CFU/mL for MRSA and 5.0 × 105 CFU/mL for all Enterococcus spp. strains. In the assay, 90 µL per well of the corresponding diluted inoculum were mixed with 10 µL per well of the corresponding extract. Turbidity (Abs612 nm) was measured with an Envision plate reader at the beginning (T0) and after 24 h incubation (Tf). Positive and negative internal plate controls were included following the previously described methodologies (Audoin, et al. 2013; Oluwabusola, et al. 2021) and three biological independent experiments (n = 3) were conducted. The Genedata Screener software (Genedata, Inc., Basel, Switzerland) was used for the analysis of the data obtained and to calculate the percentage of inhibition of each extract. The RZ’ factor was used to estimate the robustness of the assays (Zhang, et al. 1999), which was always between 0.87 and 0.95.
Antifungal assays
The target fungal strains utilized in this study were Candida albicans ATCC 10231, Aspergillus fumigatus ATCC 240305 and a clinical strain of Trichophyton rubrum (FF5), strains that belong to the Mycological Laboratory from the Faculty of Pharmacy, University of Porto (Portugal). The antifungal assays were performed in liquid format using 96-well plates following protocols described previously (Benoutman, et al. 2022; Erbiai, et al. 2021) and according to the Clinical and Laboratory Standard Institute-CLSI guidelines (M38-A2 for filamentous fungi and M27-A3 for yeasts). Briefly, fungal organisms previously cultured in Sabouraud dextrose agar (SDA) were suspended in RPMI-1640 broth buffered with MOPS (pH 7.0 (Sigma-Aldrich®, St. Louis, MO, USA): 1–3 × 103 CFU/mL for T. rubrum, 0.4–5 × 104 CFU/mL for A. fumigatus and 1–5 × 103 CFU/mL for C. albicans. The planctomycetal extracts were then diluted 1:10 with each fungal suspension at an assay concentration of 0.1 mg/mL for a final volume of 200 μL per well. Internal plate controls were added: positive control (1 µg/mL voriconazole for A. fumigatus and 64 µg/mL fluconazole for T. rubrum and C. albicans), blank control (200 μL of RPMI-1640 medium), negative control (180 μL target inoculum + 20 μL of RPMI-1640 medium) and solvent control (180 μL target inoculum + 20 μL of 20% v/v DMSO). The growth of the fungal targets was visually inspected after 2 days at 37 °C for A. fumigatus and C. albicans and after 7 days at 26 °C for T. rubrum. All screenings were done in three biological independent assays (n = 3).
Cytotoxicity screening
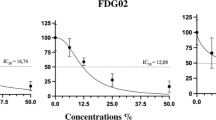
Cytotoxic activities of planctomycetal extracts against five tumoral human cell lines were evaluated using the MTT test for assessing cell viability (Mosmann 1983) in a 96-well-plate format. The assay workflow was made according to MEDINA’s portfolio of high-throughput screenings (Subko, et al. 2021). The cell lines used were human lung carcinoma A549 (ATCC CCL-185), breast adenocarcinoma MCF7 (ATCC HTB-22), human skin melanoma A2058 (ATCC CRL-11147), hepatocyte carcinoma HepG2 (ATCC HB-8065) and pancreas carcinoma MiaPaca-2 (ATCC CRL-1420). Extracts were tested at 0.25 mg/mL per duplicate for 72 h and the data obtained analysed using Genedata Screener Software.
Analysis of extracts by liquid chromatography/high-resolution mass-spectrometry
Bioactive extracts were subjected to a chemical dereplication analysis, the process of identification of already known bioactive compounds in an extract (Perez-Victoria, et al. 2016), in this case achieved by combined liquid chromatography/high-resolution mass-spectrometry (LC-HRMS) techniques. The equipment used was an Agilent 1200 Rapid Resolution HPLC interfaced with a Bruker maXis mass spectrometer. The stationary phase utilized was a Zorbax SB-C8 column (2.1 mm × 30 mm, 3.5 mm particle size) and the mobile phase (10-min run) constituted by two solvents, both composed of water (A) and acetonitrile (B), in a 90:10 ratio for solvent A, and in a 10:90 ratio for B, both additionally supplemented with ammonium formate (13 mM) and 0.01% trifluoracetic acid. The mass spectrometer was operated in positive ESI mode. For putative component identification, the retention time and exact mass of each element were compared against Fundación MEDINA’s high-resolution mass spectrometry database. Additionally, the predicted molecular formulae or the exact masses were searched for in the Chapman and Hall Dictionary of Natural Products (DNP) database (https://dnp.chemnetbase.com/chemical/ChemicalSearch.xhtml?dswid=3475, accessed on June 2023).
Results and Discussion
Antimicrobial activities of marine and estuarine Planctomycetota
In total, twenty-three marine and brackish water planctomycetal strains were screened for antimicrobial activities against different microorganisms following the workflow depicted in Fig. 1. The microorganisms included in this study represent both types of Gram-positive and Gram-negative bacteria as well as other human pathogens (including fungi and drug-resistant bacteria) and human tumoral cell lines. A maximum-likelihood tree was constructed to show the phylogenetic position of each planctomycetal strain together with a heat map to summarize the antimicrobial results obtained (Fig. 2). The distribution of these planctomycetal bioactivities across bacterial and fungal pathogens is also displayed in Supplementary Fig. 1.
Phylogenetic representation of the planctomycetal strains screened for antimicrobial activities in this study (in bold) together with a heat map showing the effects against each of the following microbial targets: SA = methicillin-sensitive Staphylococcus aureus ATCC 29213; EC = Escherichia coli ATCC 25922; TR = Trichophyton rubrum FF5; CA = Candida albicans ATCC 10231 and AF = Aspergillus fumigatus ATCC 240305. Data used for construction of this tree is presented in Supplementary Table 1. Coloured branches were used to differentiate each family: green for Lacipirellulaceae, orange for Pirellulaceae and blue for Planctomycetaceae. *data on the antibacterial effects of strain ICT_E10.1T were taken from previous study (Vitorino, et al. 2022b) and represented here for comparison
In this study, mild to potent antimicrobial activities were observed with Planctomycetota from all the nine genera tested, as represented in Fig. 2 (data available in Supplementary Table 1). Regarding the anti-fungal screenings, both T. rubrum FF5 and C. albicans ATCC 10231 showed susceptibility towards several planctomycetal extracts (Fig. 2). Additionally, extracts from strains Rhodopirellula aestuarii ICT_H3.1 T, Rhodopirellula lusitana LzF4 and Novipirellula caenicola MEMO17_8 induced a clear visual modification on the growth of A. fumigatus ATCC 240305 (such as the inhibition of the production of hyphae or abnormal morphology). Regarding the antibacterial screening, extracts of seven planctomycetal strains consistently inhibited the growth of methicillin-sensitive S. aureus ATCC 29213 in the three assays (growth reduction above 95%): Rubinisphaera margarita ICM_H10T, Rubinisphaera brasiliensis Gr7, Roseimaritima ulvae UC8T, Rhodopirellula aestuarii ICT_H3.1 T, Aeoliella straminimaris ICT_H3.7, Rhodopirellula baltica ICT_H6.1 and Novipirellula caenicola MEMO17_8. Partial activities (> 50% growth inhibition) were also observed with other planctomycetal strains (Fig. 2). On the other hand, only low activities were detected against E. coli ATCC 25922, where the only significant activity (> 50%) was observed with the extract of Roseimaritima ulvae UC8T (52%). These results suggest an overall activity which is more specific towards Gram-positive bacteria. Gram-positive bacteria have a cell wall composed only of peptidoglycan and Gram-negative bacteria have an extra outer membrane. This difference may justify the greatest inhibition against Gram-positive bacteria because the bioactive molecules may interfere with peptidoglycan synthesis and/or may be unable to enter the cell due to the outer membrane barrier. Based on this finding, the six most bioactive strains were additionally screened against five Gram-positive pathogens, including antibiotic resistant strains. As summarized in Table 2 and in Supplementary Table 2, three Planctomycetota (Rubinisphaera margarita ICM_H10T, Rhodopirellula aestuarii ICT_H3.1 T and Rhodopirellula lusitana LzF4) induced moderate to strong inhibitory effects across all the different pathogens tested, which is a strong indication for the production of bioactive molecules by these strains. Particularly, the extract of R. margarita ICM_H10T inhibited the growth of all Enterococcus spp. tested, including vancomycin resistant ones (VanA and VanB, being VanA the most resistant), as it is depicted in Fig. 3. Additionally, Novipirellula caenicola MEMO17_8 showed a strong inhibitory effect on the methicillin-resistant S. aureus (MRSA) while affecting none of the Enterococcus spp. strains (Fig. 3).
Assay wells of the antimicrobial screening of Novipirellula caenicola MEMO17_8 and Rubinisphaera margarita ICM_H10T extracts showing the potent inhibitory effects of these strains against methicillin-resistant Staphylococcus aureus MB 5393 (MRSA) and Enterococcus faecalis ATCC 29212/Enterococcus faecium clinical isolates, respectively (highlighted in red). Positive and solvent controls were also included (to show no bacterial growth and the target growth when exposed only to the extract solvent DMSO 20% v/v, respectively)
In summary, diverse planctomycetal strains from different taxonomic groups within three families (Lacipirellulaceae, Pirellulaceae and Planctomycetaceae) from class Planctomycetia showed promising antimicrobial activities. Highest activities were observed against T. rubrum and methicillin-sensitive S. aureus (Fig. 2 and Supplementary Fig. 1). The strains with the widest range of activities towards the tested targets belong to the genera Rhodopirellula (Rhodopirellula aestuarii ICT_H3.1 T and Rhodopirellula lusitana LzF4) and Novipirellula (Novipirellula caenicola MEMO17_8). Rubinisphaera margarita ICM_H10T also stood out by having a broad effect across all tested Gram-positive bacteria.
With this study, we widen the spectrum of studied planctomycetal species for antimicrobial purposes. Our results reinforce the idea that many Planctomycetota (specifically, those of the class Planctomycetia) can indeed produce antimicrobial metabolites, most likely related to survival in their complex lifestyle and habitats such as biofilms, which has been also evidenced previously by genome mining (Belova, et al. 2020; Graca, et al. 2016; Jeske, et al. 2013; Kallscheuer and Jogler 2021; Wiegand, et al. 2020).
Cytotoxic effects of planctomycetal extracts
Extracts from six planctomycetal strains that showed potent antimicrobial activities were additionally screened for cytotoxic effects against various cancer cell lines (data available at Supplementary Table 3). As represented in the phylogenetic tree combined with the heat map of the results obtained (Fig. 4), all extracts tested exhibited mild to potent activity against, at least, one cancer cell line. Overall, higher activities were observed towards human breast (MCF-7), pancreas (MiaPaca-2) and melanoma (A2058) cell lines (Fig. 4). Extracts from strains of the genus Rhodopirellula demonstrated the strongest effects against most cell lines. Extracts from the other 4 strains exhibited more selective activities to specific cell lines without affecting the most sensible one (HepG2), which is often used as a control for cytotoxicity.
Phylogenetic representation of the planctomycetal strains screened for anti-tumor activities in this study (in bold) together with a heat map showing the effects against each cell line: HepG2 = human hepatocellular carcinoma; MCF7 = human breast cancer; A549 = human lung carcinoma; A2058 = human melanoma and MiaPaca-2 = human pancreatic cancer. Data used for construction of this tree is presented in Supplementary Table 3
Up to date, little is still known about the anti-cancer potential of the bacteria from the phylum Planctomycetota. Only one previous in-vitro screening study has identified promising strains and demonstrated the cytotoxic effects of various planctomycetal species against human prostatic cancer cell line PC3 and human acute myeloid leukemia cell line MOLM-13, as well as normal rat kidney epithelial cell line (Calisto, et al. 2019). MOLM-13 was the most susceptible cell line to planctomycetal extracts. Overall, Rhodopirellula spp. were the most cytotoxic, similarly to what we also observed in the present research. Our study contributes to broadening the number of planctomycetal species screened for anti-tumour activities and has identified promising strains from the species Novipirellula caenicola, Stieleria sedimenti, Rubinisphaera margarita and Aeoliella straminimaris.
Dereplication of bioactive extracts
The chemical composition of some bioactive extracts was analysed through LC-HRMS. After searching for the exact mass or proposed molecular formula and UV/vis spectra in the DNP database of the components obtained, we could not identify any of the peaks with currently recognized bioactive natural products, therefore, the compounds responsible for the bioactivities observed remain unidentified (Fig. 5 and Supplementary Table 4). At this fermentation scale, biomass production by the planctomycetal strains is rather low, which leads to a low apparent complexity in the extracts. Nevertheless, several peaks were detected in the chromatographic runs, some corresponding to putatively non-identified molecules. For example, in the chromatogram run of Novipirellula caenicola MEMO17_8 bioactive extract (Fig. 5a), three major peaks were detected and their respective UV and mass spectra are also displayed (Fig. 5b, c and d). For each of these components, the predicted molecular formulae were the following: C8H8N2S for peak 1, C11H9N3S for peak 2 and C12H11NO2S for peak 3. While the first 2 were not found in the DNP, the molecular formula C12H11NO2S had a match for the compound named chuanghsinmycin, an antibiotic first isolated from a bacterium of the genus Actinoplanes (Shi, et al. 2018). However, the UV/vis spectra of both (peaks for Chuanghsinmycin at UV228, UV285 and UV295, according to literature) do not match, which seems to indicate that they might be distinct molecules.
Chromatogram run (UV210nm) of the bioactive extract of strain MEMO17_8 (a) with 3 possible peaks of interest (P1, P2, P3), corresponding to putatively non-identified molecules. In the following order, the mass spectra and UV/vis of the components of Peak 1 (b), Peak 2 (c) and Peak 3 (d) are displayed, together with the molecular formulas suggested for each element
Due to the phylogenetic distance of Planctomycetota with well-known prokaryotic producers of bioactive compounds (such as Actinomycetota), it can be hypothesized that their metabolomes (which are still largely underexplored) could have very distinctive profiles. This is also supported by genome analyses, which demonstrate that the phylum Planctomycetota currently contains a large number of unknown coding areas (thus coding for still unidentified compounds) (Faria, et al. 2018; Wiegand, et al. 2020, 2018). Additionally, when analysed for their content in BGCs and compared with known databases (for example, using AntiSMASH Blin, et al. 2021, 2019)), planctomycetal genomes rarely present strong identities to known gene clusters. All of this supports the likely production of novel metabolites by Planctomycetota, which is also reinforced by our chemical analysis.
Oligotrophy as a means to obtain bioactive extracts from Planctomycetota
Secondary metabolites of bacterial origin are often bioactive molecules (Donadio, et al. 2007; Seyedsayamdost 2019), which implies that they are not always produced throughout all life cycle nor under all conditions. This implies that changes applied during bacterial fermentation can have a great impact on the molecules produced by each strain through deactivation/activation of metabolic pathways. Besides, it has also been perceived that even one strain can produce different compounds under distinct environmental conditions (the “One Strain, Many Compounds”- OSMAC approach) (Romano, et al. 2018; Santos, et al. 2022; Wei, et al. 2010). The same seems to be applicable for Planctomycetota. In previous planctomycetal studies, modifications in the content of the culturing medium changed the bioactivity outcome (Belova, et al. 2020; Graca, et al. 2016; Jeske, et al. 2016). Furthermore, Jeske and colleagues even obtained different molecule profiles when growing Planctomycetota in a maintenance medium supplemented with two different organic sources, in this case, glucose or N-acetylglucosamine (Jeske, et al. 2016).
Stieleria sedimenti strain ICT_E10.1 T showed promising enhanced antimicrobial activity during oligotrophic stress (Vitorino, et al. 2022b). Thus, in this study, we also tried to apply stress through fermentation under oligotrophic conditions (medium 1:10 M13). The production of biomass under these conditions was greatly reduced when compared to the typical maintenance media such as M13 or M14 (Lage and Bondoso 2011). Yet, this condition seemed to be favourable for production of bioactive compounds, as many strains showed potent bioactivities. Already, a few of these strains have been screened for antimicrobial activities in previous studies against other groups of targets (Graca, et al. 2016; Vitorino, et al. 2022d). This is the case of strains Gr7, UC8T, LF2T, FF15T, Pd1, which were screened for activities against Bacillus subtilis and C. albicans (Graca, et al. 2016) and strains ICT_H3.1 T, ICT_E10.1 T, ICT_H6.1 and ICT_E8.1, which were screened against E. coli/S. aureus (Vitorino, et al. 2022d). In the first study, only strain Gr7 displayed mild inhibitory action towards both targets, while no activity was demonstrated by the other strains (Graca, et al. 2016). Strains ICT_H3.1 T, ICT_E10.1 T, ICT_H6.1 and ICT_E8.1 all demonstrated mild effects against the Gram-positive microorganism (Vitorino, et al. 2022d). However, the media used for fermentation was richer in comparison to the present study which used 1:10 M13 [M13/M14 and M14 supplemented with N-acetylglucosamine, respectively (Graca, et al. 2016; Vitorino, et al. 2022d)]. Since the fermentation of these same strains in oligotrophic medium either yielded new activities or enhanced previously observed ones, it is reinforced the need to test various growth conditions for the obtainment of planctomycetal bioactive extracts and the selection of promising strains.
Conclusions
In this study, the majority of the 23 tested planctomycetal strains exhibited moderate to potent antimicrobial activities against selected pathogens. The six most potent planctomycetal strains also demonstrated mild to high effects against several human cancer cell lines. This study reinforces the relevance of Planctomycetota for biotechnological purposes. We also showed that more research is necessary to find out under which conditions the best bioactive extracts from Planctomycetota can be obtained. We have observed at least 3 putative novel compounds that need further molecular identification and testing.
Abbreviations
- BGCs:
-
Biosynthetic gene clusters
- PKS:
-
Polyketide synthases
- NRPS:
-
Non-ribosomal peptide synthetases
- RiPP:
-
Ribosomally synthesized and post-translationally modified peptide
- rpm:
-
Rotations per minute
- DMSO:
-
Dimethyl sulfoxide
- CFU:
-
Colony forming units
- OD:
-
Optical density
- nm:
-
Nanometers
- LC/HRMS:
-
Liquid chromatography/high-resolution mass spectroscopy
- LEMUP:
-
Laboratory for Microbial Ecophysiology of the University of Porto, Portugal
- MRSA:
-
Methicillin-resistant Staphylococcus aureus
- VRE:
-
Vancomycin-resistant Enterococcus
- SDA:
-
Sabouraud dextrose agar
References
Acehan D, Santarella-Mellwig R, Devos DP (2014) A bacterial tubulovesicular network. J Cell Sci 127:277–280
Almeida E, Carvalho MF, Lage OM (2022) Culturomics remains a highly valuable methodology to obtain rare microbial diversity with putative biotechnological potential from two Portuguese salterns. FBE
Anjum K, Sadiq I, Chen L et al (2018) Novel antifungal janthinopolyenemycins A and B from a co-culture of marine-associated Janthinobacterium spp ZZ145 and ZZ148. Tetrahedron Lett 59:3490–3494
Audoin C, Bonhomme D, Ivanisevic J et al (2013) Balibalosides, an original family of glucosylated sesterterpenes produced by the mediterranean sponge oscarella balibaloi. Mar Drugs 11:1477–1489
Barer MR, Harwood CR (1999) Bacterial viability and culturability. Adv Microb Physiol 41:93–137
Belova SE, Saltykova VA, Dedysh SN (2020) Antimicrobial activity of a novel freshwater planctomycete Lacipirellula parvula PX69T. Microbiol 89:503–509
Benoutman A, Erbiai EH, Edderdaki FZ et al (2022) Phytochemical composition, antioxidant and antifungal activity of Thymus capitatus, a medicinal plant collected from Northern Morocco. Antibiotics 11:681
Blin K, Shaw S, Steinke K et al (2019) antiSMASH 50: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87
Blin K, Shaw S, Kloosterman AM et al (2021) antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res 49:W29–W35
Boedeker C, Schuler M, Reintjes G et al (2017) Determining the bacterial cell biology of Planctomycetes. Nat Commun 8:14853
Bondoso J, Albuquerque L, Lobo-da-Cunha A et al (2014) Rhodopirellula lusitana sp. nov. and Rhodopirellula rubra sp. nov., isolated from the surface of macroalgae. Syst Appl Microbiol 37:157–164
Bondoso J, Albuquerque L, Nobre MF et al (2015) Roseimaritima ulvae gen. nov., sp. nov. and Rubripirellula obstinata gen. nov., sp. nov. two novel planctomycetes isolated from the epiphytic community of macroalgae. Syst Appl Microbiol 38:8–15
Braña AF, Sarmiento-Vizcaino A, Perez-Victoria I et al (2017) Branimycins B and C, antibiotics produced by the abyssal actinobacterium pseudonocardia carboxydivorans M-227. J Nat Prod 80:569–573
Calisto R, Sæbø EF, Storesund JE et al (2019) Anticancer Activity in Planctomycetes. Front Mar Sci 5:499
Cohen-Bazire G, Sistrom WR, Stanier RY (1957) Kinetic studies of pigment synthesis by non-sulfur purple bacteria. J Cell Comp Physiol 49:25–68
Dedysh SN, Kulichevskaya IS, Beletsky AV et al (2020) Lacipirellula parvula gen. nov., sp. nov., representing a lineage of planctomycetes widespread in low-oxygen habitats, description of the family Lacipirellulaceae fam. nov. and proposal of the orders Pirellulales ord. nov., Gemmatales ord. nov. and Isosphaerales ord. nov. Syst Appl Microbiol 43:126050
Devos DP (2014) PVC bacteria: variation of, but not exception to, the Gram-negative cell plan. Trends Microbiol 22:14–20
Donadio S, Monciardini P, Sosio M (2007) Polyketide synthases and nonribosomal peptide synthetases: the emerging view from bacterial genomics. Nat Prod Rep 24:1073–1109
Erbiai EH, Bouchra B, da Silva LP et al (2021) Chemical composition and antioxidant and antimicrobial activities of Lactarius sanguifluus, a wild edible mushroom from northern Morocco. Euro-Mediterr J Environ Integr 6:43
Faria M, Bordin N, Kizina J et al (2018) Planctomycetes attached to algal surfaces: Insight into their genomes. Genomics 110:231–238
Feling RH, Buchanan GO, Mincer TJ et al (2003) Salinosporamide A: a highly cytotoxic proteasome inhibitor from a novel microbial source, a marine bacterium of the new genus salinospora. Angew Chem Int Ed Engl 42:355–357
Fukunaga Y, Kurahashi M, Sakiyama Y et al (2009) Phycisphaera mikurensis gen. nov., sp. nov., isolated from a marine alga, and proposal of Phycisphaeraceae fam nov., Phycisphaerales ord. nov. and Phycisphaerae classis nov. in the phylum Planctomycetes. J Gen Appl Microbiol 55:267–275
Gaurav K, Kumar D, Jagadeeshwari U et al (2021) Phylo-taxogenomics of the genus Tautonia with descriptions of Tautonia marina sp. nov., Tautonia rosea sp. nov., and emended description of the genus. Syst Appl Microbiol 44:126229
Gerwick WH, Moore BS (2012) Lessons from the past and charting the future of marine natural products drug discovery and chemical biology. Chem Biol 19:85–98
Godinho O, Botelho R, Albuquerque L et al (2021) Bremerella alba sp. nov., a novel planctomycete isolated from the surface of the macroalga Fucus spiralis. Syst Appl Microbiol 44
Graca AP, Calisto R, Lage OM (2016) Planctomycetes as novel source of bioactive molecules. Front Microbiol 7:1241
Harinantenaina Rakotondraibe L, Rasolomampianina R, Park HY et al (2015) Antiproliferative and antiplasmodial compounds from selected Streptomyces species. Bioorg Med Chem Lett 25:5646–5649
Jeske O, Jogler M, Petersen J et al (2013) From genome mining to phenotypic microarrays: Planctomycetes as source for novel bioactive molecules. Antonie Van Leeuwenhoek 104:551–567
Jeske O, Schuler M, Schumann P et al (2015) Planctomycetes do possess a peptidoglycan cell wall. Nat Commun 6:7116
Jeske O, Surup F, Ketteniss M et al (2016) Developing techniques for the utilization of planctomycetes as producers of bioactive molecules. Front Microbiol 7:1242
Jetten M, Op den Camp H, Kuenen JG et al (2010) Description of the order brocadiales. The Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes: Springer
Kallscheuer N, Jogler C (2021) The bacterial phylum Planctomycetes as novel source for bioactive small molecules. Biotechnol Adv 53:107818
Kallscheuer N, Jeske O, Sandargo B et al (2020) The planctomycete Stieleria maiorica Mal15(T) employs stieleriacines to alter the species composition in marine biofilms. Commun Biol 3:303
Kaushik R, Sharma M, Gaurav K et al (2020) Paludisphaera soli sp. nov., a new member of the family Isosphaeraceae isolated from high altitude soil in the Western Himalaya. Antonie Van Leeuwenhoek 113:1663–1674
Krieg NR, Staley JT, Brown DR, et al (2010) “Phylum XXV. Planctomycetes Garrity and Holt 2001 137 emend. Ward,” in Bergey’s Manual of Systematic Bacteriology: The Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes volume 4: Springer, New York
Kulichevskaya IS, Ivanova AA, Naumoff DG et al (2020a) Frigoriglobus tundricola gen nov., sp. nov., a psychrotolerant cellulolytic planctomycete of the family Gemmataceae from a littoral tundra wetland. Syst Appl Microbiol 43:126129
Kulichevskaya IS, Naumoff DG, Miroshnikov KK et al (2020b) Limnoglobus roseus gen. nov., sp. nov., a novel freshwater planctomycete with a giant genome from the family Gemmataceae. Int J Syst Evol Microbiol 70:1240–1249
Kulichevskaya IS, Ivanova AA, Suzina NE et al (2022) Anatilimnocola floriformis sp nov, a novel member of the family Pirellulaceae from a boreal lake, and emended description of the genus Anatilimnocola. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-022-01769-x
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kumar D, Gaurav K, Pk S et al (2020a) Gimesia chilikensis sp. nov., a haloalkali-tolerant planctomycete isolated from Chilika lagoon and emended description of the genus Gimesia. Int J Syst Evol Microbiol 70:3647–3655
Kumar D, Gaurav K, Jagadeeshwari U et al (2020b) Roseimaritima sediminicola sp. nov., a new member of Planctomycetaceae isolated from Chilika lagoon. Int J Syst Evol Microbiol 70:2616–2623
Kumar G, Kumar D, Jagadeeshwari U et al (2021a) Crateriforma spongiae sp. Nov., isolated from a marine sponge and emended description of the genus “Crateriforma.” Antonie Van Leeuwenhoek 114:341–353
Kumar G, Lhingjakim KL, Uppada J et al (2021b) Aquisphaera insulae sp. Nov., a new member in the family Isosphaeraceae, isolated from the floating island of Loktak lake and emended description of the genus Aquisphaera. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-021-01615-6
Lage OM, Bondoso J (2011) Planctomycetes diversity associated with macroalgae. FEMS Microbiol Ecol 78:366–375
Lage OM, Bondoso J (2014) Planctomycetes and macroalgae, a striking association. Front Microbiol 5:267
Lage OM, Bondoso J, Lobo-da-Cunha A (2013) Insights into the ultrastructural morphology of novel Planctomycetes. Antonie Van Leeuwenhoek 104:467–476
Lage OM, van Niftrik L, Jogler C et al (2019) Planctomycetes. In: Schmidt TM (ed) Encyclopedia of microbiology, 4th edn. Academic Press, Oxford, pp 614–626. https://doi.org/10.1016/B978-0-12-809633-8.90689-7
Larkin MA, Blackshields G, Brown NP et al (2007) Clustal W and Clustal X version 20. Bioinformatics 23:2947–2948
Leao PN, Costa M, Ramos V et al (2013) Antitumor activity of hierridin B, a cyanobacterial secondary metabolite found in both filamentous and unicellular marine strains. PLoS ONE 8:e69562
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296
Lodha T, Narvekar S, Karodi P (2021) Classification of uncultivated anammox bacteria and Candidatus Uabimicrobium into new classes and provisional nomenclature as Candidatus Brocadiia classis nov. and Candidatus Uabimicrobiia classis nov. of the phylum Planctomycetes and novel family Candidatus Scalinduaceae fam. nov to accommodate the genus Candidatus Scalindua. Syst Appl Microbiol 44:126272
Mosmann T (1983) Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods 65:55–63
Oluwabusola ET, Adebisi OO, Reyes F et al (2021) Isolation and characterization of new phenolic siderophores with antimicrobial properties from Pseudomonas sp. UIAU-6B. Beilstein J Org Chem 17:2390–2398
Oren A, Garrity GM (2021) Valid publication of the names of forty-two phyla of prokaryotes. Int J Syst Evol Microbiol 71.
Perez-Victoria I, Martin J, Reyes F (2016) Combined LC/UV/MS and NMR strategies for the dereplication of marine natural products. Planta Med 82:857–871
Reynolds KA, Luhavaya H, Li J et al (2018) Isolation and structure elucidation of lipopeptide antibiotic taromycin B from the activated taromycin biosynthetic gene cluster. J Antibiot 71:333–338
Rinke C, Schwientek P, Sczyrba A et al (2013) Insights into the phylogeny and coding potential of microbial dark matter. Nature 499:431–437
Rivas-Marin E, Peeters SH, Claret Fernandez L et al (2020) Non-essentiality of canonical cell division genes in the planctomycete Planctopirus limnophila. Sci Rep 10:66
Rivas-Marín E, Devos DP (2018) The Paradigms They Are a-Changin’: past, present and future of PVC bacteria research. Antonie Van Leeuwenhoek 111:785–799
Rodriguez V, Martin J, Sarmiento-Vizcaino A et al (2018) Anthracimycin B, a potent antibiotic against gram-positive bacteria isolated from cultures of the deep-sea actinomycete streptomyces cyaneofuscatus M-169. Mar Drugs 16(11), 406. https://doi.org/10.3390/md16110406
Romano S, Jackson SA, Patry S et al (2018) Extending the “One Strain Many Compounds” (OSMAC) principle to marine microorganisms. Mar Drugs 16:244
Sandargo B, Jeske O, Boedeker C et al (2020) Stieleriacines N-Acyl dehydrotyrosines from the marine planctomycete Stieleria neptunia sp nov. Front Microbiol 11:1408
Santarella-Mellwig R, Pruggnaller S, Roos N et al (2013) Three-dimensional reconstruction of bacteria with a complex endomembrane system. PLoS Biol 11:e1001565
Santos JD, Vitorino I, de la Cruz M et al (2020a) Diketopiperazines and other bioactive compounds from bacterial symbionts of marine sponges. Antonie Van Leeuwenhoek 113:875–887
Santos JDN, João SA, Martín J et al (2022) iChip-inspired isolation, bioactivities and dereplication of actinomycetota from portuguese beach sediments. Microorganisms 10:1471
Santos JD, Vitorino I, Reyes F et al (2020b) From ocean to medicine: pharmaceutical applications of metabolites from marine bacteria. Antibiotics 9(8):455. https://doi.org/10.3390/antibiotics9080455
Schulze CJ, Donia MS, Siqueira-Neto JL et al (2015) Genome-directed lead discovery: biosynthesis, structure elucidation, and biological evaluation of two families of polyene macrolactams against trypanosoma brucei. ACS Chem Biol 10:2373–2381
Seyedsayamdost MR (2019) Toward a global picture of bacterial secondary metabolism. J Ind Microbiol Biotechnol 46:301–311
Shi Y, Jiang Z, Li X et al (2018) Biosynthesis of antibiotic chuangxinmycin from Actinoplanes tsinanensis. Acta Pharm Sin B 8:283–294
Subko K, Wang X, Nielsen FH et al (2021) Mass spectrometry guided discovery and design of novel asperphenamate analogs from penicillium astrolabium reveals an extraordinary NRPS flexibility. Front Microbiol 11. https://doi.org/10.3389/fmicb.2020.618730
Tareq FS, Lee MA, Lee HS et al (2014) Non-cytotoxic antifungal agents: isolation and structures of gageopeptides A-D from a Bacillus strain 109GGC020. J Agric Food Chem 62:5565–5572
Vitorino IR, Lage OM (2022) The Planctomycetia: an overview of the currently largest class within the phylum Planctomycetes. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-021-01699-0
Vitorino I, Albuquerque L, Wiegand S et al (2020) Alienimonas chondri sp. nov., a novel planctomycete isolated from the biofilm of the red alga Chondrus crispus. Syst Appl Microbiol 43 (3):126083. https://doi.org/10.1016/j.syapm.2020.126083
Vitorino I, Albuquerque L, Wiegand S et al (2021a) Corrigendum to “Alienimonas chondri sp. Nov., a novel planctomycete isolated from the biofilm of the red alga Chondrus crispus” [Syst. Appl. Microbiol. 43 (2020) 126083]. Syst Appl Microbiol 44:126219
Vitorino I, Santos JDN, Godinho O et al (2021b) Novel and conventional isolation techniques to obtain planctomycetes from marine environments. Microorganisms 9(10), 2078. https://doi.org/10.3390/microorganisms9102078
Vitorino IR, Klimek D, Calusinska M et al (2022a) Rhodopirellula aestuarii sp. Nov., a novel member of the genus Rhodopirellula isolated from brackish sediments collected in the Tagus River estuary, Portugal. Syst ApplMicrobiol 45(6). https://doi.org/10.1016/j.syapm.2022.126360
Vitorino IR, Klimek D, Calusinska M et al (2022b) Stieleria sedimenti sp. nov., a novel member of the family pirellulaceae with antimicrobial activity isolated in Portugal from Brackish Sediments. Microorganisms 10(11), 2151. https://doi.org/10.3390/microorganisms10112151
Vitorino IR, Lobo-da-Cunha A, Vasconcelos V et al (2022c) Rubinisphaera margarita sp. nov., a novel planctomycete isolated from marine sediments collected in the Portuguese north coast. Int J Syst Evol Microbiol 72(6). https://doi.org/10.1099/ijsem.0.005425
Vitorino IR, Lobo-da-Cunha A, Vasconcelos V et al (2022d) Isolation, diversity and antimicrobial activity of planctomycetes from the Tejo river estuary (Portugal). FEMS Microbiol Ecol 98(7). https://doi.org/10.1093/femsec/fiac066
Vitorino IR, Lobo-da-Cunha A, Vasconcelos V et al (2023) Aeoliella straminimaris sp. nov., a novel member of the phylum Planctomycetota with an unusual filamentous structure. Int J Syst Evol Microbiol 73(4). https://doi.org/10.1099/ijsem.0.005850
Wagner M, Horn M (2006) The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opin Biotechnol 17:241–249
Wei H, Lin Z, Li D et al (2010) OSMAC (one strain many compounds) approach in the research of microbial metabolites—a review. Wei Sheng Wu Xue Bao 50:701–709
WHO (2017) Global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics. Geneva: World Health Organization
WHO (2019a) Assessing national capacity for the prevention and control of noncommunicable diseases: report of the 2019 global survey: World Health Organization
WHO (2019b) Global antimicrobial resistance surveillance system (GLASS) report: early implementation 2017–2018. Geneva: World Health Organization: Geneva, Switzerland
WHO (2022) WHO fungal priority pathogens list to guide research, development and public health action: World Health Organization
Wiegand S, Jogler M, Jogler C (2018) On the maverick planctomycetes. FEMS Microbiol Rev 42:739–760
Wiegand S, Jogler M, Boedeker C et al (2020) Cultivation and functional characterization of 79 planctomycetes uncovers their unique biology. Nat Microbiol 5:126–140
Wiese J, Abdelmohsen UR, Motiei A et al (2018) Bacicyclin, a new antibacterial cyclic hexapeptide from Bacillus sp. strain BC028 isolated from Mytilus edulis. Bioorg Med Chem Lett 28:558–561
Zhang JH, Chung TD, Oldenburg KR (1999) A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J Biomol Screen 4:67–73
Zhang Y, Adnani N, Braun DR et al (2016) Micromonohalimanes A and B: antibacterial halimane-type diterpenoids from a marine micromonospora species. J Nat Prod 79:2968–2972
Zhang B, Wang KB, Wang W et al (2018) Discovery, biosynthesis, and heterologous production of streptoseomycin, an anti-microaerophilic bacteria macrodilactone. Org Lett 20:2967–2971
Funding
Open access funding provided by FCT|FCCN (b-on). Inês Vitorino is supported by a “Fundação para a Ciência e Tecnologia (FCT)” doctoral grant (SFRH/BD/145577/2019). This work was financially co-supported by the strategical funding from FCT UIDB/04423/2020 and UIDP/04423/2020; and by the project ATLANTIDA (ref. NORTE-01–0145-FEDER-000040), supported by the Norte Portugal Regional Operational Program (NORTE 2020), under the PORTUGAL 2020 Partnership Agreement and through the European Regional Development Fund (ERDF). The MEDINA authors disclosed the receipt of financial support from Fundación MEDINA, a public–private partnership of Merck Sharp & Dohme de España S.A./Universidad de Granada/ Junta de Andalucía.
Author information
Authors and Affiliations
Contributions
Scientific work and data analysis were performed by IRV. Experiment design was planned by IRV and OML. IRV, EP, PS and MC were responsible for the antimicrobial assays. Cytotoxic assays were performed by MCR and TAM. IRV, JM and FR performed the dereplication work and analysis. IRV was supervised by OML, FR, FV and VV. IRV wrote the manuscript and all remaining authors revised it. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vitorino, I.R., Pinto, E., Martín, J. et al. Uncovering the biotechnological capacity of marine and brackish water Planctomycetota. Antonie van Leeuwenhoek 117, 26 (2024). https://doi.org/10.1007/s10482-023-01923-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10482-023-01923-z