Abstract
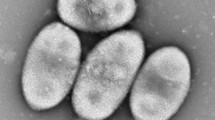
Two Gram-stain-positive, facultatively anaerobic, rod-shaped bacterial strains, S126T and S82T, were isolated from coastal algae of China. Strains S126T and S82T are halotolerant and could grow in the presence of 0–13% NaCl and 0–14% NaCl, respectively. The two strains shared 98.9% 16S rRNA gene sequence similarity with each other and 93.4–99.8% similarity with type strains of Exiguobacterium species. The major fatty acids (> 10%) of strains S126T and S82T were iso-C17:0, iso-C13:0, anteiso-C13:0 and iso-C15:0. The predominant quinones of strains S126T and S82T were MK-7 and MK-8. The polar lipid profiles of strain S126T and S82T contained diphosphatidylglycerol, phosphatidylglycerol and phosphatidylethanolamine. The cell-wall peptidoglycans of both strains S126T and S82T were of the A3α L-Lys-Gly type. The average nucleotide identity (ANI) and average nucleotide index (AAI) between strains S126T and S82T and type strains of Exiguobacterium species were all below the thresholds to discriminate bacterial species, indicating that they constitute two novel species in the genus Exiguobacterium. Based on polyphasic taxonomy characterization and genomic aspects, the names Exiguobacterium algae sp. nov. and Exiguobacterium qingdaonense sp. nov. are proposed for the two novel species, with type strains being S126T (= CGMCC 1.17116T = KCTC 43079 T) and S82T (= CGMCC 1.17115T = KCTC 43078T), respectively.

Similar content being viewed by others
Abbreviations
- Ocs:
-
Orthologous clusters
- ANI:
-
Average nucleotide identity
- AAI:
-
Average nucleotide index
References
Altenburger P, Kämpfer P, Makristathis A, Lubitz W, Busse HJ (1996) Classification of bacteria isolated from a medieval wall painting. J Biotechnol 47:39–52
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using diamond. Nat Methods 12:59–60
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:1–9
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The carbohydrate-active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res 37:D233-238
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) TrimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973
Chaturvedi P, Prabahar V, Manorama R, Pindi PK, Bhadra B, Begum Z, Shivaji S (2008) Exiguobacterium soli sp. nov., a psychrophilic bacterium from the McMurdo Dry Valleys. Antarct Int J Syst Evol MIcrobiol 58:2447–2453
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu X-W, Meye SD, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Collins MD, Lund BM, Farrow JAE, Schleifer KH (1983) Chemotaxonomic study of an alkalophilic bacterium, Exiguobacterium aurantiacum gen. nov., sp. nov. J Gen Microbiol 129:2037–2042
Crapart S, Fardeau ML, Cayol JL, Thomas P, Sery C, Ollivier B, Combet-Blanc Y (2007) Exiguobacterium profundum sp. nov., a moderately thermophilic, lactic acid-producing bacterium isolated from a deep-sea hydrothermal vent. Int J Syst Evol Microbiol 57:287–292
Dong XZ, Cai MY (eds) (2001) Determinative manual for routine bacteriology. Scientific Press, Beijing (English translation)
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fitch WM (1971) Toward defining the course of evolution: minimum changes for a specific tree topology. Syst Zool 20:406–416
Fruhling A, Schumann P, Hippe H, Straubler B, Stackebrandt E (2002) Exiguobacterium undae sp. nov. and Exiguobacterium antarcticum sp. nov. Int J Syst Evol Microbiol 52:1171–1176
Jiang X, Xue Y, Wang L, Yu B, Ma Y (2013) Genome sequence of a novel polymer-grade L-lactate-producing alkaliphile, Exiguobacterium sp. strain 8–11-1. Genome Announc 1:e00616-e713
Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, Mering CV, Bork P (2017) Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol Biol Evol 34:2115–2122
Kasana RC, Pandey CB (2018) Exiguobacterium: an overview of a versatile genus with potential in industry and agriculture. Crit Rev Biotechnol 38:141–156
Kim IG, Lee MH, Jung SY, Song JJ, Oh TK, Yoon JH (2005) Exiguobacterium aestuarii sp. nov. and Exiguobacterium marinum sp. nov., isolated from a tidal flat of the Yellow Sea in Korea. Int J Syst Evol Microbiol 55:885–889
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Konstantinidis KT, Rosselló-Móra R, Amann R (2017) Uncultivated microbesin need of their own taxonomy. ISME J 11:2399–4063
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Letunic I, Bork P (2019) Interactive tree of life (iTOL) V4: recent updates and new developments. Nucleic Acids Res 47:256–259
Liao H, Lin XL, Li YQ, Qu MM, Tian Y (2020) Reclassification of the taxonomic framework of orders Cellvibrionales, Oceanospirillales, Pseudomonadales and Alteromonadales in class Gammaproteobacteria through phylogenomic tree analysis. Msystems 15:e00543-e620
Liu H, Xin B, Zheng J, Zhong H, Yu Y, Peng D, Sun M (2021) Build a bioinformatic analysis platform and apply it to routine analysis of microbial genomics and comparative genomics. Protoc Exch. https://doi.org/10.21203/rs.2.21224/v5
Liu SW, Ye JJ, Lu QP, Cheema MT, Abbas M, Huang DL, Sajid I, Sun CH (2020) Motilibacter deserti sp. nov. and Motilibacter aurantiacus sp. nov., two novel actinobacteria isolated from soil of Cholistan Desert and emended description of the genus Motilibacter. Syst Appl Microbiol 43:126150
Lopez-Cortes A, Schumann P, Pukall R, Stackebrandt E (2006) Exiguobacterium mexicanum sp. nov. and Exiguobacterium artemiae sp. nov., isolated from the brine shrimp Artemia franciscana. Syst Appl Microbiol 29:183–190
Margesin R, Gander S, Zacke G, Gounot AM, Schinner F (2003) Hydrocarbon degradation and enzyme activities of cold-adapted bacteria and yeasts. Extremophiles 7:451–458
Meng X, Chang YQ, Zhou LY, Du ZJ (2020) Exiguobacterium flavidum sp. nov., isolated from the Red Maple Lake. Int J Syst Evol Microbiol 70:2359–2365
Mohan Kulshreshtha N, Kumar R, Begum Z, Shivaji S, Kumar A (2013) Exiguobacterium alkaliphilum sp. nov. isolated from alkaline wastewater drained sludge of a beverage factory. Int J Syst Evol Microbiol 63:4374–4379
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274
Raichand R, Pareek S, Singh NK, Mayilraj S (2012) Exiguobacterium aquaticum sp. nov., a member of the genus Exiguobacterium. Int J Syst Evol Microbiol 62:2150–2155
Rodrigues DF, Goris J, Vishnivetskaya T, Gilichinsky D, Thomashow MF, Tiedje JM (2006) Characterization of Exiguobacterium isolates from the Siberian permafrost. Description of Exiguobacterium sibiricum sp. nov. Extremophiles 10:285–294
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. Technical Note 101.Newark, DE: MIDI Inc
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schumann P (2011) Peptidoglycan structure. Methods Microbiol 38:101–129
Schleifer KH (1985) Analysis of the chemical composition and primary structure of murein. Methods Microbiol 18:123–156
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069
Singh NK, Raichand R, Kaur I, Kaur C, Pareek S, Mayilraj S (2013) Exiguobacterium himgiriensis sp. nov. a novel member of the genus Exiguobacterium, isolated from the Indian Himalayas. Antonie Van Leeuwenhoek 103:789–796
Srivastava AK, Srivastava R, Sharma A, Bharati AP, Tiwari PK, Singh AK, Srivastava AK, Chakdar H, Kashyap PL, Saxena AK (2020) Pan-genome analysis of Exiguobacterium reveals species delineation and genomic similarity with Exiguobacterium profundum PHM 11. Environ Microbiol Rep 12:639–650
Stolz A, Busse HJ, Kämpfer P (2007) Pseudomonas knackmussii sp. nov. Int J Syst Evol Microbiol 57:572–576
Tindall BJ (1990a) Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol Lett 66:199–202
Tindall BJ (1990b) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13:128–130
Vishnivetskaya TA, Lucas S, Copeland A, Lapidus A, Glavina del Rio T, Dalin E, Tice H, Bruce DC, Goodwin LA, Pitluck S, Saunders E, Brettin T, Detter C, Han C, Larimer F, Land ML, Hauser LJ, Kyrpides NC, Ovchinnikova G, Kathariou S, Ramaley RF, Rodrigues DF, Hendrix C, Richardson P, Tiedje JM (2011) Complete genome sequence of the thermophilic bacterium Exiguobacterium sp. AT1b. J Bacteriol 193:2880
Vishnivetskaya TA, Kathariou S, Tiedje JM (2009) The Exiguobacterium genus: biodiversity and biogeography. Extremophiles 13:541–555
Wang Q, Liu F, Zhang DC (2020) Pelagihabitans pacificus gen. nov., sp. nov., a member of the family Flavobacteriaceae isolated from a deep-sea seamount. Int J Syst Evol Microbiol 70:4569–4575
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Zhang H, Yohe T, Huang L, Entwistle S, Wu P, Yang Z, Busk PK, Xu Y, Yin Y (2018) dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 46:95–101
Funding
This study was supported by National Natural Science Foundation of China (31670002, awarded to Dechao Zhang; 31970003 and 31770003, awarded to Jinshui Zheng).
Author information
Authors and Affiliations
Contributions
ZDC designed this research, LFM, HWX and WWQ isolated the strain and performed the initial cultivation and strain deposition, LFM, HWX and WWQ performed strain characterization, the electron microscopic and chemotaxonomic analysis, LYJ and ZJS performed the genomic and phylogenetic analysis, LFM and LYJ drafted the manuscript. ZDC supervised the study and contributed to text preparation and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Availability of data and material
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request. All data generated or analysed during this study are included in this published article [and its supplementary information files].
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, F., Li, Y., He, W. et al. Exiguobacterium algae sp. nov. and Exiguobacterium qingdaonense sp. nov., two novel moderately halotolerant bacteria isolated from the coastal algae. Antonie van Leeuwenhoek 114, 1399–1406 (2021). https://doi.org/10.1007/s10482-021-01594-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01594-8