Abstract
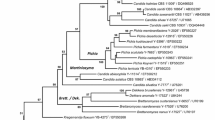
Three new yeast species, Candida kashinagacola (JCM 15019T = CBS 10903T), C. pseudovanderkliftii (JCM 15025T = CBS 10904T), and C. vanderkliftii (JCM 15029T = CBS 10905T) are described on the basis of comparison of nucleotide sequences of large subunit ribosomal DNA D1/D2 region (LSU rDNA D1/D2). The nearest assigned species of the three new species was Candida llanquihuensis. Candida kashinagacola and C. pseudovanderkliftii differed from C. llanquihuensis by 3.8% nucleotide substitution of the region, while C. vanderkliftii did by 4.4%. Three new species differed in a number of physiological and growth characteristics from any previously assigned species and from one another. A phylogenetic tree based on the sequences of LSU rDNA D1/D2 showed that these new species together with Candida sp. ST-246, Candida sp. JW01-7-11-1-4-y2, Candida sp. BG02-7-20-001A-2-1 and C. llanquihuensis form a clade near Ambrosiozyma species. The new species did not assimilate methanol as a sole source of carbon, which supported the monophyly of these non methanol-assimilating species which are closely related to the methylotrophic yeasts. Candida kashinagacola was frequently isolated from the beetle galleries of Platypus quercivorus in three different host trees (Quercus serrata, Q. laurifolia and Castanopsis cuspidata) located in the sourthern part of Kyoto, Japan, thus indicating that this species may be a primary ambrosia fungus of P. quercivorus. On the other hand, C. pseudovanderkliftii and C. vanderkliftii were isolated only from beetle galleries in Q. laurifolia. Candida vanderkliftii was isolated from beetle gallery of Platypus lewisi as well as those of P. quercivorus. Candida pseudovanderkliftii and C. vanderkliftii are assumed to be auxiliary ambrosia fungi of P. quercivorus.



Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. doi:10.1093/nar/25.17.3389
Batra LR (1966) Ambrosia fungi: extent of specificity to ambrosia beetles. Science 153:193–195. doi:10.1126/science.153.3732.193
Batra LR (1967) Ambrosia fungi: a taxonomic revision, and nutritional studies of some species. Mycologia 59:976–1017. doi:10.2307/3757271
Batra LR (1972) Ectosymbiosis between ambrosia fungi and beetles. 1. Indian J Mycol Plant Pathol 2:165–169
Beaver RA (1989) Insect-fungus relationships in the bark and ambrosia beetles. In: Wilding N, Collins NM, Hammond PM, Webber JF (eds) Insect-fungus interactions. 14th symposium of the Royal Entomological Society of London in collaboration with the British Mycological Society. Academic Press, London, UK
Endoh R, Suzuki M, Benno Y (2008) Ambrosiozyma kamigamensis sp. nov. and A. neoplatypodis sp. nov., two new ascomycetous yeasts from ambrosia beetle galleries. Anton Leeuwen. doi:10.1007/s10482-008-9253-z
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution Int J Org Evolution 39:783–791. doi:10.2307/2408678
Francke-Grosmann H (1966) Űber die Ambrosiazucht der holzbrűtenden Ipiden im Hinblick auf das System. Verhandlungen der Deutschen Gesellschaft fűr Angewandte Entomologie 14:139–144
Francke-Grosmann H (1967) Ectosymbiosis in wood-inhabiting insects. In: Henry SM (ed) Symbiosis, vol 2. Academic Press, New York, USA; London, UK
Harrington TC (2005) Ecology and evolution of mycophagous bark beetles and their fungal partners. In: Vega FE, Blackwell M (eds) Insect-fungal associations. Oxford University Press, New York, USA, p 258
Ito S, Yamada T (1998) Distribution and spread of the mass mortality of oak trees. J Jpn For Soc 80:229–232 in Japanese with English abstract
Kajimura H, Hijii N (1992) Dynammics of the fungal symbionts in the gallery system and the mycangia of the ambrosia beetle, Xylosandrus mutilatus (Blandford) (Coleoptera: Scolytidae) in relation to its life history. Ecol Res 7:107–117. doi:10.1007/BF02348489
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. doi:10.1007/BF01731581
Kobayashi M, Ueda A (2005) Wilt disease of Fagaceae trees caused by Platypus quercivorus (Murayama) (Coleoptera: Platypodidae) and the associated fungus: aim is to clarify the damage factor. J Jpn For Soc 87:435–450 in Japanese with English abstract
Kubono T, Ito S (2002) Raffaelea quercivora sp. nov. associated with mass mortality of Japanese oak, and the ambrosia beetle (Platypus quercivorus). Mycoscience 43:255–260. doi:10.1007/s102670200037
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Anton Leeuwen 73:331–371. doi:10.1023/A:1001761008817
Masuya H, Ichihara Y, Kubono T (2005) Fungi isolated from beetle gallery of Platypus quercivorus. In: Proceeding of 116th conference of The Japanese Forest Society. (in Japanese) http://www.jstage.jst.go.jp/article/jfsc/116/0/359/_pdf/-char/ja/. Cited 3 Apr 2008.
Meyer W, Mitchell TG, Freedman EZ, Vilgalys R (1993) Hybridization probes for conventional DNA fingerprinting used as single primers in the polymerase chain reaction to distinguish strains of Cryptococcus neoformans. J Clin Microbiol 31:2274–2280
Meyer W, Latouche GN, Thanos M, Mitchell TG, Yarrow D, Schönian G et al (1997) Identification of pathogenic yeasts of the imperfect genus Candida by polymerase chain reaction fingerprinting. Electrophoresis 18:1548–1559. doi:10.1002/elps.1150180911
Meyer SA, Payne RW, Yarrow D (1998) Candida Berkhout. In: Kurtzman CP, Fell JW (eds) The yeasts, a taxonomic Study, 4th edn. Elsevier Science BV, Amsterdam, The Netherlands, pp 520–521
Nakase T, Suzuki M (1985) Taxonomic studies on Debaryomyces hansenii (Zopf) Lodder et Kreger-van Rij and related species. I. Chemotaxonomic investigations. J Gen Appl Microbiol 31:49–69. doi:10.2323/jgam.31.49
O’Donnel K (1993) Fusarium and its near relatives. In: The fungal holomorph: mitotic, meiotic and pleomorphic speciation in fungal systematics. CAB International, Willingford, UK, pp 225–233
Péter G, Tornai-Lehoczki J, Dlauchy D (2007) Ogataea allantospora sp. nov., an ascomycetous yeast species from phylloplane. Anton Leeuwen 92:443–448. doi:10.1007/s10482-007-9172-4
Ramírez C, González A (1984) Five new filamentous, glucose-fermenting Candida isolated from decayed wood in the evergreen rainy Valdivian forest of southern Chile. Mycopathol 88:83–92. doi:10.1007/BF00436436
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic tree. Mol Biol Evol 4:406–425
Sampaio JP, Gadanho M, Bauer R (2001) Taxonomic studies on the genus Cystofilobasidium: description of Cystofilobasidium ferigula sp. nov. and clarification of the status of Cystofilobasidium lari-marini. Int J Syst Evol Microbiol 51:221–229
Six DL (2003) Bark beetle-fungus symbioses. In: Boutzis K, Miller TA (eds) Insect symbiosis. Contemporary topics in entomology series. CRC Press, Boca Raton, USA; London, UK; New York, USA; Washington D. C., USA
Suzuki M, Nakase T (1999) Cellular neutral sugar compositions and ubiquinone systems of the genus Candida. Microbiol Cult Collect 14:49–62
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. doi:10.1093/nar/22.22.4673
van der Walt JP (1972) The yeast genus Ambrosiozyma gen. nov. (ascomycetes). Mycopathol Mycol Appl 46:305–316. doi:10.1007/BF02052126
van der Walt JP, Scott DB, van der Klift WC (1971) Five new Torulopsis species from South African insect sources. Anton Leeuwen 37:461–471. doi:10.1007/BF02218516
Vassart G, Georges M, Monsieur R, Brocas H, Lequarré AS, Christophe D (1987) A sequence in M13 phage detects hypervariable minisatellites in human and animal DNA. Science 235:683–684. doi:10.1126/science.2880398
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols–a guide to methods and applications. Academic Press, CA, San Diego, Chile, pp 315–322
Yarrow D (1998) Methods for the isolation, maintenance and identification of yeasts. In: Kurtzman CP, Fell JW (eds) The Yeast, A Taxonomic Study, 4th edn edn. Elsevier Science BV, Amsterdam, The Netherlands, pp 77–100
Acknowledgements
We thank Dr. M. Kobayashi, Dr. A. Nozaki and technical staffs working for Kamigamo Experimental Station for their help in collecting isolation sources. We also thank Dr. Zaki A. Siddiqui for his help in checking English sentences. We thank Ms. Y. Omoto for her help in isolating yeasts. This research was partially supported by the Ministry of Education, Science, Sports and Culture, Grant-in-Aid for Scientific Research (A), no. 18208015, 2006–2008.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Endoh, R., Suzuki, M., Benno, Y. et al. Candida kashinagacola sp. nov., C. pseudovanderkliftii sp. nov. and C. vanderkliftii sp. nov., three new yeasts from ambrosia beetle-associated sources. Antonie van Leeuwenhoek 94, 389–402 (2008). https://doi.org/10.1007/s10482-008-9256-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-008-9256-9